Back to Journals » Journal of Hepatocellular Carcinoma » Volume 11
A Hypoxia-Related miRNA-mRNA Signature for Predicting the Response and Prognosis of Transcatheter Arterial Chemoembolization in Hepatocellular Carcinoma
Authors Zong S, Huang G , Pan B, Zhao S, Ling C, Cheng B
Received 13 December 2023
Accepted for publication 28 February 2024
Published 12 March 2024 Volume 2024:11 Pages 525—542
DOI https://doi.org/10.2147/JHC.S454698
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Imam Waked
Shaoqi Zong,1,2,* Guokai Huang,1,3,* Bo Pan,1,* Shasha Zhao,4 Changquan Ling,1,3 Binbin Cheng1,3
1Oncology Department of Traditional Chinese Medicine, The First Affiliated Hospital of Naval Medical University, Shanghai, 200043, People’s Republic of China; 2Department of Oncology, Yueyang Hospital of Integrated Traditional Chinese and Western Medicine, Shanghai University of Traditional Chinese Medicine, Shanghai, 200437, People’s Republic of China; 3Faculty of Traditional Chinese Medicine, Naval Medical University, Shanghai, 200043, People’s Republic of China; 4Shanghai Municipal Hospital of Traditional Chinese Medicine, Shanghai University of Traditional Chinese Medicine, Shanghai, 200071, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Binbin Cheng, Oncology Department of Traditional Chinese Medicine, The First Affiliated Hospital of Naval Medical University, 168 Changhai Road, Shanghai, 200043, People’s Republic of China, Email [email protected] Changquan Ling, Faculty of Traditional Chinese Medicine, Naval Medical University, 800 Xiangyin Road, Shanghai, 200043, People’s Republic of China, Email [email protected]
Purpose: Transcatheter arterial chemoembolization (TACE) is commonly used in the treatment of hepatocellular carcinoma (HCC). However, not all patients respond to this treatment. TACE typically leads to hypoxia in the tumor microenvironment. Therefore, we aimed to construct a prognostic model based on hypoxia-related differentially expressed microRNA (miRNAs) in hepatocellular carcinoma (HCC) and to investigate the potential target mRNAs for predicting TACE response.
Methods: The hypoxia-related miRNAs (HRMs) were identified in liver cancer cells, then global test was performed to further select the miRNAs which were associated with recurrence and vascular invasion. A prognostic model was constructed based on multivariate Cox regression analysis; qRT-PCR analysis was used to validate the differentially expressed miRNAs in HCC cell lines under hypoxic condition. We further identified the putative target genes of the miRNAs and investigate the relationship between the target genes and TACE response, immune cells infiltration.
Results: We established a HRMs prognostic model for HCC patients, containing two miRNAs (miR-638, miR-501-5p), the patients with high-HRMs score showed worse survival in discovery and validation cohort; qRT-PCR analysis confirmed that these two miRNAs are up-regulated in hepatoma cells under hypoxic condition. Furthermore, four putative target genes of these two miRNAs were identified (ADH1B, CTH, FTCD, RCL1), which were significantly associated with TACE response, immune score, immunosuppressive immune cells infiltration, PDCD1 and CTLA4.
Conclusion: The HCC-HRMs signature may be utilized as a promising prognostic factor and may have implications for guiding TACE and immune therapy.
Keywords: hypoxia, miRNA, signature, hepatocellular carcinoma, prognosis
Introduction
Primary liver cancer (PLC), the incidence ranks sixth and the cancer-related mortality rate is the third leading among all malignant tumors in the world.1 Hepatocellular carcinoma (HCC) is the most common type of PLC, with a 5-year-survival rate of less than 21%.2 Surgical resection is generally considered the mainly radical treatment for HCC.3,4 However, resection is only possible in a small proportion of patients because of the advanced tumor stages at the time of diagnosis and poor hepatic function.
Transarterial chemoembolization (TACE) is recommended as the first-line treatment for intermediate-stage HCC.5 In addition, clinical studies have reported that adjuvant TACE prolongs the overall survival (OS) and recurrence-free survival (RFS) in patients with HCC after surgical resection.6,7 With a treatment-related mortality rate of < 5%, TACE can be expected to result in a median survival of 11–20 months. However, the survival benefit of patients undergoing TACE is variable, and > 40% of patients do not respond to this therapy, highlighting the importance of developing an effective biomarker for predicting the survival benefit of TACE. TACE blocks the blood supply to the tumor and thereby aggravates local hypoxia,8 then promoting cancer growth, metastasis,9–11 and leading to poor prognosis. Rhee et al reported that hypoxia related markers could predict TACE resistance and result in poor outcome in HCC;12 Therefore, it is necessary to develop biomarkers that can predict the response and prognosis based on hypoxia-related markers.
MicroRNAs (miRNAs) are non-coding, single-stranded RNA that are highly stable in fixed tissues and body fluids; miRNAs are attractive biomarkers for assessing clinicopathological conditions and predicting clinical outcome,13,14 which could modulate the translation of one or more mRNAs and regulate several biological processes, including cellular metabolism, differentiation, proliferation and the immune response.15–17 Consistently, a series of circulating miRNAs including miR-221, miR-26a, miR-106b, miR-107, miR-21 and miR-122 have been reported as potential predictive markers for TACE efficacy in HCC patients.18–20 More importantly, it has been confirmed that hypoxia could regulate the expression of several miRNAs to promote angiogenesis, invasion, metastasis and recurrence of HCC.21–24 Notably, miR-125b loss activates the HIF-1α/p-AKT loop, contributing to TACE resistance in HCC patients.25 Therefore, we aimed to construct a hypoxia-related miRNA signature to predict the response and prognosis of HCC patients who underwent TACE.
Materials and Methods
Data Collection and Preprocessing
A series of datasets were downloaded from the NCBI GEO database (https://www.ncbi.nlm.nih.gov/geo/) and analyzed. The accession numbers include GSE68593, GSE67140, GSE30297, GSE31384, GSE76903, GSE21362, GSE116182, and GSE104580. Of these microarray data, GSE68593,26 GSE31384,27 GSE21362, GSE116182 were based on GPL20157, GPL14140, GPL10312, GPL14613; GSE6714028 and GSE3029729 were based on the GPL8786; GSE104580 was based on the GPL570; Additionally, TCGA Xena database (https://xenabrowser.net/datapages/) was used to further select the putative target genes of miRNAs. All the data were obtained from the public database and therefore this study was waived the requirement of ethical approval. This study was conducted according to the ethical guidelines of the 1975 Declaration of Helsinki. We confirmed that all clinical data from the database contained in this study are anonymous or confidential.
Identification of Hypoxia-Related miRNAs
miRNA expression profiles of GSE68593 were downloaded from the GEO data repository. In this dataset, the SK-Hep-1 cells were exposed in normoxia and hypoxia condition, each group contain 3 independent replications. The original annotation files were downloaded, and the hypoxia-associated miRNAs of the SK-Hep-1 cells were identified using “limma” R package.30 According to the Benjamini and Hochberg (BH) procedure,31 the cut-off criteria were set as P<0.05, and the absolute value of fold change (FC) > 1.5.
Development and Validation of a Hypoxia-Related miRNA Signature
As shown in the Figure 1, we identified hypoxia related miRNAs as described above. A global test, a useful tool for the analysis of microarray data, could identify the genes that are significantly associated with clinical parameters of interest,32 then two discovery datasets (GSE67140, GSE30297) were used to identify the hypoxia-related miRNAs that significantly associated with vascular invasion and recurrence using global test, the z-score of each miRNA was calculated, and miRNAs with z-score > 1 were included for further analysis. On the basis, the dataset (GSE31384) containing survival data was further used to construct the HRM signature. First, the expression and clinical datasets were downloaded, 158 samples were retained after removing the NA value. The dataset was then randomly divided into two groups according to a ratio of 7:3, and 7/10 samples (n = 110) were set as the training dataset. Several methods including multivariate Cox regression analysis (forward and backward), Ridge, Lasso, Enet, and random survival forest (RSF) analysis were used to construct a relative prognostic model, the C-index of each model was calculated to select the optimal model.33,34 In this process, a series of R packages including “glment”, “survival”, “survmener”, and “randomForestSRC” were used. The median risk score of the dataset was set as the cut-off value, the OS and disease-free survival (DFS) analyses for each set were confirmed using Log rank tests. The R packages “timeROC”, “pROC” were used to draw the ROC curve and the area under the curve (AUC). Additionally, the entire dataset (GSE31384, n=158) and another dataset (GSE116182, n=64) were used to confirm the prognostic value of the hypoxia-associated miRNA signature.
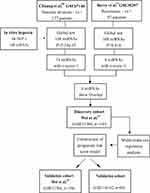 |
Figure 1 The flow-chart of the two hypoxia-related prognostic miRNA signature in HCC. |
Cell Culture
The liver cancer cell lines SK-Hep-1, SMCC-7721, HCCLM3, MHCC-97L, BEL-7402, and Huh-7 and the normal liver cell line L02 were purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, #L110KJ, Shanghai BasalMedia Technologies Co., Ltd., Shanghai, China) containing 10% fetal bovine serum (FBS, Lonsera, #S711-001S, Uruguay).
Quantitative Real-Time PCR (RT-qPCR)
First, L02 and liver cancer cells, including SK-Hep-1, SMCC-7721, HCCLM3, MHCC-97L, BEL-7402, and Huh-7 cells, were cultured in a normoxic chamber (21% O2, 5% CO2) for 24 h. Additionally, SMCC-7721 and SK-Hep-1 cells were cultured in a normoxic chamber (21% O2, 5% CO2) or hypoxic chamber (1% O2, 5% CO2, 94% N2). The total RNA was extracted using RNAiso Plus (Takara #9109, Takara Bio Inc.). To detect the levels of miRNAs and mRNAs, reverse transcription was performed using the RT master mix (Takara, #RR037A, Takara Bio Inc.). SYBR Green Real-time PCR Master Mix (QPK-201; Bio-toyobo) was used for the qPCR analysis. Primers used in this study are listed in Table 1. U6 small nuclear RNA was used as the internal standard.
 |
Table 1 Primers Used in the qRT-PCR Reaction |
Identification of Putative Targets of the miRNAs
Relative targets were downloaded from the TargetScan Human database (Release 7.2). Correlations between miRNAs and their relative targets were explored using GEO datasets (GSE176271 and GSE176288), including the relative expression of miRNAs and mRNAs in 16 patients with HCC. Additionally, differentially expressed genes (DEGs) were analyzed using the GEO dataset (GSE14520: downregulated genes in HCC tissues). Furthermore, the TCGA-LIHC dataset was used to narrow down candidate targets. Genes associated with prognosis were identified using the log-rank survival analysis. Kaplan-Meier survival analysis was used to identify genes that could predict the prognosis of HCC patients who underwent TACE but had no impact on the prognosis of HCC patients who underwent surgical resection.
Correlation of Putative Target Genes with TACE Response
In this part, GEO dataset (GSE104580) was used to explore the relationship between the putative genes and the TACE response, this dataset is consisted with 66 TACE responders and 81 TACE non-responders; Wilcoxon test was performed to explore the differences in relative gene expression between the response and non-response groups. In addition, the median level of putative target genes was set as the cut-off value, and the data were arranged as a four-fold contingency table The chi-squared Test was used to evaluate the correlation between putative target genes and TACE responses.
Tumor Microenvironment Analysis
The ESTIMATE algorithm35 was used to evaluate the association between putative target genes and stromal, immune, and estimated scores based on the TCGA-LIHC dataset in tumor tissues.
The Infiltration of Immune Cells in Tumor Microenvironment
TIMER2.0 (http://timer.cistrome.org/)36 was used to further explore the relationship between infiltration of a series of immune cells, PDCD1, CTLA4, and putative target genes. First, the relative data on the correlation between the levels of relative target genes and immune cell infiltration (regulatory T cells, CD4+ T cells, CD8+ T cells, B cells, macrophages, cancer-associated fibroblasts, NK cells, and MDSC) were downloaded. In addition, the relationships between the target genes and levels of PDCD1 and CTLA4 were investigated.
Statistical Analysis
A global test was performed to explore the association between the miRNAs and their clinical features. Multivariate Cox regression analysis (forward and backward) and ridge, lasso, end, and RSF analyses were used to construct a relative hypoxia-associated miRNA prognostic model. The C-index of each model was used to select an optimal model. A two-sided Log rank test was then performed to compare the differences in OS between the low-risk and high-risk groups in both sets using the R (v.3.6.2) software. A paired t-test was conducted to compare differences between liver cancer tissues and paired normal tissues. A t-test was performed to test the difference in miRNA levels between normoxic and hypoxic hepatoma cells, if the data were normally distributed. Otherwise, a non-parametric test (Mann–Whitney) was applied. SPSS 21.0 was used to analysis relative data. P < 0.05 was considered statistically significant.
Results
Identification of Hypoxia-Related Prognostic miRNAs
To investigate the hypoxia-associated miRNAs, the GSE68573 dataset was downloaded from the GEO database. A total of 108 miRNAs (49 upregulated and 59 downregulated) were identified as differentially expressed hypoxia-associated miRNAs. The association of the hypoxia-associated miRNA set with clinicopathological features (tumor recurrence and vascular invasion) was explored using the available datasets submitted to the GEO database. The Goeman global test32 showed that the hypoxia-associated miRNA set was significantly enriched in two discovery cohorts (GSE67140 and GSE30297; P=3.34e-23 for Barry; P=0.014 for Chuang) (Figure 1).
First, 108 hypoxia-related miRNAs were identified from the microarray dataset of SK-Hep-1 cells (GSE68573). Second, the Geoman global test was performed to select miRNAs associated with recurrence (GSE30297) and vascular invasion (GSE67140). Interacting miRNAs were selected for further construction of the hypoxia-related prognostic miRNA signature using a training dataset (GSE31384, n=110). Finally, the entire dataset (GSE31384, n=158) and external dataset (GSE116182, n=64) were used to validate the prognostic value of the two HRM signatures.
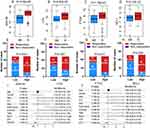
Next, miRNAs with z-scores > 1 were selected as significant miRNAs. Fourteen miRNAs were chosen for the dataset of Barry et al (Figure 2A), and eight miRNAs were chosen by Chuang et al (Figure 2B). Finally, the intersecting miRNAs in the two datasets were identified. Six hypoxia-associated miRNAs (miR-339-3p, miR-501-5p, miR-501-3p, miR-362-5p, and miR-638) were obtained for further analysis.
Construction of a Two-miRNA Prognostic Signature
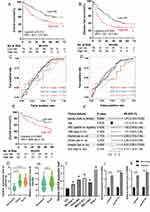
We then constructed a risk score using the six selected miRNAs. First, a variety of methods, including multivariate Cox regression analysis (forward and backward) and Ridge, Lasso and Enet analyses, were performed to screen the miRNAs. The C-index values were calculated for each model. Among these models, the mean C-index value of the multivariate Cox regression analysis (forward) was 0.634, which was the highest (Figure 3A). Therefore, this model was selected as the optimal model and two miRNAs (miR-501-5p and miR-638) were found to be significantly associated with OS (Figure 3B).
Next, we explored whether the two-miRNA signature could predict the OS of HCC patients. The risk score formula was calculated according to the expression of these two miRNAs for OS prediction, as follows: risk score = (1.0225×expression value of miR-501-5p) + (0.7047×expression value of miR-638). The two-miRNA signature risk score was calculated for each patient in the discovery cohort (n=110). The risk score of each patient was ranked, and the median risk score of the dataset was set as the cut-off value. The patients were divided into high-risk (n=55) and low-risk (n=55) groups. As expected, compared with patients with low-risk scores, the patients with high-risk scores showed shorter OS (Log rank test, P=0.034; HR, 1.85 (1.04–3.28); Figure 3C) and DFS (Log rank test P=0.013, HR=1.79 (1.12–2.87), Figure 3D). A time-dependent ROC analysis was performed to assess the prognostic ability of the miRNA signature at different time points. The AUC of the two-miRNA prognostic model for OS was 0.635 at 1 year, 0.68 at 3 years, and 0.605 at 5 years (Figure 3E). The AUC for DFS were 0.684, 0.655, and 0.548 at 1, 3, and 5 years, respectively (Figure 3F).
Validation of the Two-miRNA Signature for Survival Prediction
To confirm our findings, the prognostic power of the two-miRNA signature was further evaluated in the entire cohort (158 samples) and in external cohort (64 samples). According to the same risk formula, the patients in the entire cohort were divided into high- and low-risk groups. Patients with HCC with high-risk scores for the miRNA signature showed shorter OS and DFS (Figure 4A and B). The AUC of the two-miRNA prognostic model for OS at 1, 3, and 5 years were 0.662, 0.659, and 0.614, respectively (Figure 4C). The AUC for DFS were 0.675, 0.651, and 0.565 at 1, 3, and 5 years, respectively (Figure 4D), similar to those observed in the discovery cohort.
In the external cohort (GSE116182, 64 samples), we found that patients with high-risk scores for the miRNA signature tended to have shorter OS (log-rank P=0.086, HR=1.95 (0.92–4.15), Figure 4E). Multivariate Cox analysis also showed that the high-risk score of the miRNA signature was associated with poor OS (P=0.016, HR=3.13(1.24–7.89), Table 2, Figure 4F). The levels of miR-501-5p and miR-638 were higher in the tumor tissues than in the normal hepatic tissues (Figure 4G and H). The expression of miR-638 was significantly upregulated in liver cancer cells compared with that in normal liver cells (Figure 4I). Additionally, miR-501-5p and miR-638 were significantly upregulated under hypoxic condition in liver cancer cells (SK-Hep-1, Figure 4J; SMCC-7721, Figure 4K).
 |
Table 2 Multivariate Analysis of Features Associated with Overall Survival |
Identification of Potential Targets of miR-638 and miR-501-5p in HCC Patients
To identify the potential target genes of miR-638 and miR-501-5p, a total of 1877 and 3892 genes were identified as predictive targets, respectively (Figure 5). Because the upregulation of these two miRNAs predicts poor prognosis in HCC, the target genes should be downregulated. GEO dataset (GSE14520) was analyzed to identify the down-regulated genes in tumor tissues. A total of 581 genes were significantly downregulated in the tumor tissues compared to the adjacent normal tissues (log2 fold change≤-1.0, P-value <0.05). The intersecting genes of these two datasets were further analyzed using the TCGA-LIHC dataset. The results showed that 11 target genes of miR-638 and 25 target genes of miR-501-5p were significantly downregulated in cancer tissues compared to those in normal tissues (log2 fold change≤-2.0) (Figure s1). We found that 18 genes were significantly downregulated in tumor tissues compared with paired normal tissues. Based on this, the Log rank test results revealed that low gene expression was significantly associated with poor prognosis (Tables 3 and 4, Figure s2).
 |
Table 3 Eleven Candidate Target Genes Regulated by miR-638 |
 |
Table 4 Twenty-Five Candidate Target Genes Regulated by miR-501-5p |
 |
Figure 5 The flow-chart for selecting the putative tumor-suppressive targets of miR-501-5p and miR-638. |
Clinical Significance of Potential Targets of miR-501-5p and miR-638
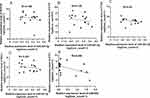
Next, we analyzed whether these genes could be used to predict the prognosis of patients who underwent TACE. We found that low expression of alcohol dehydrogenase 1B (ADH1B), cystathionine γ-lyase (CTH), RNA terminal phosphate cyclase like 1 (RCL1) and formimidoyltransferase cyclodeaminase (FTCD) may predict poor prognosis of HCC patients underwent TACE (Tables 3, 4, Figure 6). To analyze the relationship between the four identified genes and the TACE response, the GEO dataset (GSE104580) was further analyzed. The gene levels of ADH1B (P=3.06e-08), CTH (P=9.92e-05), FTCD (P=1.78e-03), and RCL1 (P=3.33e-02) were significantly lower in the TACE non-responsive group than in the responsive group (Figure 7A–D). Additionally, high levels of ADH1B (P<0.001), CTH (P=0.002), FTCD (P=0.017), and RCL1 (P=0.029) were positively associated with TACE response (Figure 7E–H). In addition, multivariate Cox regression analysis revealed that ADH1B (P=2.18e-04, HR=0.71(0.59–0.85)) and CTH (P=3.80e-02, HR=0.77 (0.61–0.99)) were independent prognostic factors in patients with HCC who received TACE (Figure 7I and J). We further explored the correlation between the two miRNAs and their targets. The results showed that miR-501-5p was negatively associated with ADH1B, CTH, FTCD, and RCL1 expression levels. In addition, miR-638 expression was negatively associated with CTH (Figure 8A–E). Moreover, the identified protein expression (ADH1B, FTCD, and CTH) in normal and HCC patient tissues was confirmed using the HPA database (https://www.proteinatlas.org/). As shown in Figure 9, compared to normal tissues, the identified protein expression was significantly downregulated in patients with HCC.
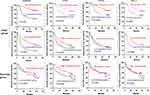 |
Figure 6 Kaplan-Meier curves of 5-year OS and RFS rates in the TACE group according to the expression of ADH1B, CTH, FTCD and RCL1. |
 |
Figure 9 The identified proteins expression in normal and HCC patient tissues. Expression of FTCD (A), CTH (B), and ADH1B (C) in patients with HCC and healthy controls in the HPA database. |
Patients with low ADH1B and RCL1 levels had significantly worse OS and RFS in the TACE group. Low levels of CTH and FTCD predicted worse prognosis in the TACE group. Low levels of ADH1B, CTH, FTCD, and RCL1 showed no significant difference in the OS rate compared with the high expression level group in the resection dataset.
Correlation Analysis Between ADH1B, CTH, FTCD, RCL1 and Immune Score
Hypoxia may lead to immunosuppression in HCC.37,38 Therefore, we further explored the role of these genes in the immune microenvironment of HCC patients. The results showed that ADH1B (R=−0.15, P=0.0027) and CTH (R=−0.2, P=8.8e-05) were negatively associated with the immune score using TCGA dataset (Figure 10A and B). However, there was no significant difference in the immune score between FTCD and RCL1 (Figure s3).
Next, the relationship between ADH1B, CTH, and infiltration of various immune cells was studied. We found that ADH1B expression was negatively correlated with regulatory T cells (Tregs), macrophages, and B cells (Figure 10C). CTH negatively correlated with macrophages and myeloid dendritic cells (Figure 10D). Moreover, the relationship between these two genes (ADH1B and CTH) and immune checkpoint genes CTLA-4 and PDCD1 was further investigated. The results showed that ADH1B and CTH levels were negatively associated with PDCD1 (R=−0.33, P=7.07e-11; R=−0.312, P=7.91e-10) and CTLA4 (R=−0.335, P=3.68e-11; R=−0.289, P=1.42e-08) (Figure 10E–H).
Discussion
Despite its widespread use in clinical practice for HCC treatment, TACE still has many limitations. First, it is difficult to identify patients who would benefit from this treatment because of heterogeneity in the cellular composition and metabolism of HCCs. In addition, although TACE can destroy the tumor blood supply, it also induces severe hypoxia in tumors, which may enhance communication between tumor cells and the microenvironment and aggravate immunosuppression. Therefore, there is an urgent need to improve TACE efficacy and optimize methods for patient selection to identify patients who will benefit from treatment.
TACE is a procedure involving treatment with chemotherapeutic drugs and embolization, exposing cells to high concentrations of chemotherapeutic drugs such as doxorubicin and fluorouracil. In this study, we gathered a hypoxia-related dataset and identified a two-miRNA signature that could stratify patients with HCC into two risk groups to evaluate their prognosis. The signature was validated using both entire and external cohort datasets. Additionally, ROC curve analysis confirmed that the hypoxia-related miRNA signature is highly accurate and sensitive, indicating that this signature may be an effective biomarker for HCC prognosis. We then reviewed comprehensive literature to clarify the biological functions of these miRNAs. Huang et.al39 has confirmed that miR-501-5p was significantly upregulated in HCC cell lines comparing with normal liver cells. Consistently, miR-501-5p is also upregulated in gastric cancer and can promote cancer stem cell properties by regulating the Wnt/β-catenin pathway.40 Additionally, Ma et.al41 demonstrated that miR-501-5p accelerated gastric cancer cell migration and proliferation. MiR-638, another risk factor identified in this study, has been reported to be an oncogenic factor in breast cancer and esophageal squamous cell carcinoma (ESCC), which could induce autophagy and tumorigenesis by suppressing DACT3.42 These results indicated the potential role of hypoxia-related miRNAs (miR-638 and miR-501-5p) as prognostic biomarkers in HCC.
To clarify the molecular mechanisms of miRNAs, their target genes were identified in HCC. ADH1B, CTH, RCL1, and FTCD were significantly associated with the prognosis of patients with HCC treated with TACE. Importantly, two of these (ADH1B and CTH) were independent prognostic factors in patients with HCC who received TACE treatment. ADH1B, an alcohol metabolism-related gene, was significantly downregulated in HCC tissues.43–45 ADH1B mediates alcohol consumption and significantly decreases the risk of HCC.46 Additionally, univariate and multivariate Cox regression analyses showed that lower ADH1B expression was an independent factor for poor prognosis in HCC patients.47 Our results also suggest that ADH1B is an independent prognostic factor in HCC patients who have undergone TACE. CTH, a cysteine metabolism-related gene, was markedly downregulated in the HCC tissues. According to multivariate Cox regression analysis, low expression of CTH is an independent risk factor for OS and recurrence in patients according to multivariate Cox regression analysis. In addition, silencing of CTH induces proliferation, epithelial-mesenchymal transition (EMT), and metastasis in HCC.48,49 Moreover, since hypoxia also influences the tumor immune microenvironment by promoting the recruitment of innate immune cells and interfering with the differentiation and function of adaptive immune cells,50,51 we found that ADH1B and CTH were negatively associated with the immune score and infiltration of various immune cells. ADH1B expression is negatively associated with the infiltration of Tregs, macrophages, and B cells. Tregs are considered immunosuppressive cells.52 Additionally, He et al53 found that B cells exacerbate type-2 inflammation by increasing the production IL-13 in ICOS+ ICL2a cells, thereby inducing an immunosuppressive microenvironment in HCC. Infiltrated macrophages can be polarized into M2 macrophages by hypoxia, thus contributing to an immunosuppressive microenvironment.54,55 CTH is negatively associated with infiltration of myeloid dendritic cells and macrophages. Previous studies have shown that Tumor-associated dendritic cells (tDCs) are known as one type of immunosuppressive immune cells that suppress the generation of cytotoxic T cells and induce the activation of immunosuppressive Tregs, thereby promoting the formation of an immunosuppressive microenvironment.56,57 Therefore, these immunosuppressive immune cells can promote tumor growth and metastasis, thus inducing a poor prognosis in HCC. However, the roles of ADH1B and CTH in immunosuppression of the TME should be further investigated.
Our study has some limitations. First, the results of our study were based on GEO datasets containing retrospective studies. Therefore, multicenter large-sample studies are required to validate the prognostic value of this signature. Second, the biological processes of these two miRNAs require further in vitro and vivo.
In summary, we constructed a hypoxia-associated miRNA signature for HCC patients. Four target genes, miR-501-5p and miR-638, ADH1B, CTH, FTCD, and RCL1, were positively associated with TACE response, and two (ADH1B and CTH) were confirmed to be independent prognostic factors for HCC patients treated with TACE. However, the mechanism of action of these target genes (ADH1B, CTH, FTCD, and RCL1) under hypoxic conditions remains unclear. Further studies are needed to confirm this hypothesis.
Data Sharing Statement
The data supporting the findings of this study are available from the GEO database (https://www.ncbi.nlm.nih.gov/geo/).
Acknowledgments
The authors declare no potential conflicts of interest, and thank the GEO database for data availability.
Supplemental Information
Supplemental information for this article can be found online.
Funding
This study was supported by National Natural Science Foundation of China (No. 82030117 and 82074203).
Disclosure
The authors declare no conflict of interest.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Miller KD, Fidler-Benaoudia M, Keegan TH, Hipp HS, Jemal A, Siegel RL. Cancer statistics for adolescents and young adults, 2020. CA Cancer J Clin. 2020;70(6):443–459. doi:10.3322/caac.21637
3. Pei Y, Chen X, Zhang W. Liver resection for BCLC 0-A stage hepatocellular carcinoma: does the time to surgery not impact the prognosis? J Hepatol. 2018;68(5):1101. doi:10.1016/j.jhep.2017.12.032
4. Li C, Wen TF. Does surgical resection provide better outcomes than radiofrequency ablation in patients with BCLC very early-stage hepatocellular carcinoma? Ann Surg. 2017;266(6):e54–e55. doi:10.1097/SLA.0000000000001357
5. Liang L, Xing H, Zhang H, et al. Surgical resection versus transarterial chemoembolization for BCLC intermediate stage hepatocellular carcinoma: a systematic review and meta-analysis. HPB. 2018;20(2):110–119. doi:10.1016/j.hpb.2017.10.004
6. Ren ZG, Lin ZY, Xia JL, et al. Postoperative adjuvant arterial chemoembolization improves survival of hepatocellular carcinoma patients with risk factors for residual tumor: a retrospective control study. World J Gastroenterol. 2004;10(19):2791–2794. doi:10.3748/wjg.v10.i19.2791
7. Chen X, Zhang B, Yin X, Ren Z, Qiu S, Zhou J. Lipiodolized transarterial chemoembolization in hepatocellular carcinoma patients after curative resection. J Cancer Res Clin Oncol. 2013;139(5):773–781.
8. Kluger MD, Halazun KJ, Barroso RT, et al. Bland embolization versus chemoembolization of hepatocellular carcinoma before transplantation. Liver Transpl. 2014;20(5):536–543. doi:10.1002/lt.23846
9. Liu Z, Wang Y, Dou C, et al. Hypoxia-induced up-regulation of VASP promotes invasiveness and metastasis of hepatocellular carcinoma. Theranostics. 2018;8(17):4649–4663. doi:10.7150/thno.26789
10. Dou C, Zhou Z, Xu Q, et al. Hypoxia-induced TUFT1 promotes the growth and metastasis of hepatocellular carcinoma by activating the Ca(2+)/PI3K/AKT pathway. Oncogene. 2019;38(8):1239–1255. doi:10.1038/s41388-018-0505-8
11. Guo BJ, Ruan Y, Wang YJ, et al. Jiedu Recipe, a compound Chinese herbal medicine, inhibits cancer stemness in hepatocellular carcinoma via Wnt/beta-catenin pathway under hypoxia. J Integr Med. 2023;21(5):474–486. doi:10.1016/j.joim.2023.06.008
12. Rhee H, Nahm JH, Kim H, et al. Poor outcome of hepatocellular carcinoma with stemness marker under hypoxia: resistance to transarterial chemoembolization. Mod Pathol. 2016;29(9):1038–1049. doi:10.1038/modpathol.2016.111
13. Ambros V, Bartel B, Bartel DP, et al. A uniform system for microRNA annotation. Rna. 2003;9(3):277–279. doi:10.1261/rna.2183803
14. Lan H, Lu H, Wang X, Jin H. MicroRNAs as potential biomarkers in cancer: opportunities and challenges. Biomed Res. Int. 2015;2015:125094. doi:10.1155/2015/125094
15. Chen JF, Mandel EM, Thomson JM, et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nature Genet. 2006;38(2):228–233. doi:10.1038/ng1725
16. Suksangrat T, Phannasil P, Jitrapakdee S. miRNA regulation of glucose and lipid metabolism in relation to diabetes and non-alcoholic fatty liver disease. Adv Exp Med Biol. 2019;1134:129–148.
17. Zhao C, Li XY, Li ZY, Li M, Liu ZD. Moxibustion regulates T-regulatory/T-helper 17 cell balance by modulating the microRNA-221/suppressor of cytokine signaling 3 axis in a mouse model of rheumatoid arthritis. J Integr Med. 2022;20(5):453–462. doi:10.1016/j.joim.2022.06.002
18. Ali HEA, Emam AA, Zeeneldin AA, et al. Circulating miR-26a, miR-106b, miR-107 and miR-133b stratify hepatocellular carcinoma patients according to their response to transarterial chemoembolization. Clin Biochem. 2019;65:45–52. doi:10.1016/j.clinbiochem.2019.01.002
19. Pinjaroen N, Chailapakul P, Sriphoosanaphan S, Chuaypen N, Tangkijvanich P. Predictive role of pretreatment circulating miR-221 in patients with hepatocellular carcinoma undergoing transarterial chemoembolization. Diagnostics. 2023;13(17):2794. doi:10.3390/diagnostics13172794
20. Pelizzaro F, Cardin R, Sartori A, et al. Circulating MicroRNA-21 and MicroRNA-122 as prognostic biomarkers in hepatocellular carcinoma patients treated with transarterial chemoembolization. Biomedicines. 2021;9(8):890. doi:10.3390/biomedicines9080890
21. Liu T, Tang J, Li X, et al. The key network of mRNAs and miRNAs regulated by HIF1A in hypoxic hepatocellular carcinoma cells. Front Genet. 2022;13:857507. doi:10.3389/fgene.2022.857507
22. Zhao B, Ke K, Wang Y, et al. HIF-1alpha and HDAC1 mediated regulation of FAM99A-miR92a signaling contributes to hypoxia induced HCC metastasis. Signal Transduct Target Ther. 2020;5(1):118. doi:10.1038/s41392-020-00223-6
23. Jia W, Liang S, Lin W, et al. Hypoxia-induced exosomes facilitate lung pre-metastatic niche formation in hepatocellular carcinoma through the miR-4508-RFX1-IL17A-p38 MAPK-NF-κB pathway. Int J Biol Sci. 2023;19(15):4744–4762. doi:10.7150/ijbs.86767
24. Yao M, Liang S, Cheng B. Role of exosomes in hepatocellular carcinoma and the regulation of traditional Chinese medicine. Front Pharmacol. 2023;14:1110922. doi:10.3389/fphar.2023.1110922
25. Wei X, Zhao L, Ren R, et al. MiR-125b loss activated HIF1alpha/pAKT Loop, leading to transarterial chemoembolization resistance in hepatocellular carcinoma. Hepatology. 2021;73(4):1381–1398. doi:10.1002/hep.31448
26. Du C, Weng X, Hu W, et al. Hypoxia-inducible MiR-182 promotes angiogenesis by targeting RASA1 in hepatocellular carcinoma. J Exp Clin Cancer Res. 2015;34(1):67. doi:10.1186/s13046-015-0182-1
27. Wei R, Huang GL, Zhang MY, et al. Clinical significance and prognostic value of microRNA expression signatures in hepatocellular carcinoma. Clin Cancer Res. 2013;19(17):4780–4791. doi:10.1158/1078-0432.CCR-12-2728
28. Xie QY, Almudevar A, Whitney-Miller CL, Barry CT, McCall MN. A microRNA biomarker of hepatocellular carcinoma recurrence following liver transplantation accounting for within-patient heterogeneity. BMC Med Genomics. 2016;9(1):18. doi:10.1186/s12920-016-0179-4
29. Barry CT, D’Souza M, McCall M, et al. Micro RNA expression profiles as adjunctive data to assess the risk of hepatocellular carcinoma recurrence after liver transplantation. Am J Transplant. 2012;12(2):428–437. doi:10.1111/j.1600-6143.2011.03788.x
30. Ritchie ME, Phipson B, Wu D, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47–e47. doi:10.1093/nar/gkv007
31. Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Royal Stat Soc Series B. 1995;57(1):289–300.
32. Goeman JJ, van de Geer SA, de Kort F, van Houwelingen HC. A global test for groups of genes: testing association with a clinical outcome. Bioinformatics. 2004;20(1):93–99. doi:10.1093/bioinformatics/btg382
33. Liu Z, Guo C, Dang Q, et al. Integrative analysis from multi-center studies identities a consensus machine learning-derived lncRNA signature for stage II/III colorectal cancer. EBioMedicine. 2022;75:103750. doi:10.1016/j.ebiom.2021.103750
34. Liu Z, Liu L, Weng S, et al. Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in colorectal cancer. Nat Commun. 2022;13(1):816. doi:10.1038/s41467-022-28421-6
35. Yoshihara K, Shahmoradgoli M, Martínez E, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4(1):2612. doi:10.1038/ncomms3612
36. Li T, Fu J, Zeng Z, et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020;48(W1):W509–W514. doi:10.1093/nar/gkaa407
37. Suthen S, Lim CJ, Nguyen PHD, et al. Hypoxia-driven immunosuppression by Treg and type-2 conventional dendritic cells in HCC. Hepatology. 2022;76(5):1329–1344. doi:10.1002/hep.32419
38. Sceneay J, Parker BS, Smyth MJ, Moller A. Hypoxia-driven immunosuppression contributes to the pre-metastatic niche. Oncoimmunology. 2013;2(1):e22355. doi:10.4161/onci.22355
39. Huang DH, Wang GY, Zhang JW, Li Y, Zeng XC, Jiang N. MiR-501-5p regulates CYLD expression and promotes cell proliferation in human hepatocellular carcinoma. Jpn J Clin Oncol. 2015;45(8):738–744. doi:10.1093/jjco/hyv063
40. Fan D, Ren B, Yang X, Liu J, Zhang Z. Upregulation of miR-501-5p activates the wnt/beta-catenin signaling pathway and enhances stem cell-like phenotype in gastric cancer. J Exp Clin Cancer Res. 2016;35(1):177. doi:10.1186/s13046-016-0432-x
41. Ma X, Feng J, Lu M, et al. microRNA-501-5p promotes cell proliferation and migration in gastric cancer by downregulating LPAR1. J Cell Biochem. 2020;121(2):1911–1922. doi:10.1002/jcb.29426
42. Ren Y, Chen Y, Liang X, Lu Y, Pan W, Yang M. MiRNA-638 promotes autophagy and malignant phenotypes of cancer cells via directly suppressing DACT3. Cancer Lett. 2017;390:126–136. doi:10.1016/j.canlet.2017.01.009
43. Teng L, Li Z, Shi Y, et al. Development and validation of a microenvironment-related prognostic model for hepatocellular carcinoma patients based on histone deacetylase family. Transl Oncol. 2022;26:101547. doi:10.1016/j.tranon.2022.101547
44. Strommer J. The plant ADH gene family. Plant J. 2011;66(1):128–142. doi:10.1111/j.1365-313X.2010.04458.x
45. Wei R, Li P, He F, et al. Comprehensive analysis reveals distinct mutational signature and its mechanistic insights of alcohol consumption in human cancers. Brief Bioinform. 2021;22(3). doi:10.1093/bib/bbaa066
46. Liu J, Yang HI, Lee MH, et al. Alcohol drinking mediates the association between polymorphisms of ADH1B and ALDH2 and Hepatitis B-related hepatocellular carcinoma. Cancer Epidemiol Biomarkers Prev. 2016;25(4):693–699. doi:10.1158/1055-9965.EPI-15-0961
47. Liu X, Li T, Kong D, You H, Kong F, Tang R. Prognostic implications of alcohol dehydrogenases in hepatocellular carcinoma. BMC Cancer. 2020;20(1):1204. doi:10.1186/s12885-020-07689-1
48. Lin Z, Huang W, He Q, et al. FOXC1 promotes HCC proliferation and metastasis by Upregulating DNMT3B to induce DNA Hypermethylation of CTH promoter. J Exp Clin Cancer Res. 2021;40(1):50. doi:10.1186/s13046-021-01829-6
49. Zhang H, Song Y, Zhou C, et al. Blocking Endogenous H(2)S signaling attenuated radiation-induced long-term metastasis of residual HepG2 Cells through inhibition of EMT. Radiat Res. 2018;190(4):374–384. doi:10.1667/RR15074.1
50. Terry S, Buart S, Chouaib S. Hypoxic Stress-induced tumor and immune plasticity, suppression, and impact on tumor heterogeneity. Front Immunol. 2017;8:1625. doi:10.3389/fimmu.2017.01625
51. Guo W, Qiao T, Dong B, Li T, Liu Q, Xu X. The effect of hypoxia-induced exosomes on anti-tumor immunity and its implication for immunotherapy. Front Immunol. 2022;13:915985. doi:10.3389/fimmu.2022.915985
52. Genova C, Dellepiane C, Carrega P, et al. Therapeutic implications of tumor microenvironment in lung cancer: focus on immune checkpoint blockade. Front Immunol. 2021;12:799455. doi:10.3389/fimmu.2021.799455
53. He Y, Luo J, Zhang G, et al. Single-cell profiling of human CD127(+) innate lymphoid cells reveals diverse immune phenotypes in hepatocellular carcinoma. Hepatology. 2022;76(4):1013–1029. doi:10.1002/hep.32444
54. You Y, Wen D, Zeng L, et al. ALKBH5/MAP3K8 axis regulates PD-L1+ macrophage infiltration and promotes hepatocellular carcinoma progression. Int J Biol Sci. 2022;18(13):5001–5018. doi:10.7150/ijbs.70149
55. Chen J, Ji K, Gu L, Fang Y, Pan M, Tian S. HMGA1 promotes macrophage recruitment via activation of NF-κB-CCL2 signaling in hepatocellular carcinoma. J Immunol Res. 2022;2022:4727198. doi:10.1155/2022/4727198
56. Laoui D, Keirsse J, Morias Y, et al. The tumour microenvironment harbours ontogenically distinct dendritic cell populations with opposing effects on tumour immunity. Nat Commun. 2016;7(1):13720. doi:10.1038/ncomms13720
57. Scarlett UK, Rutkowski MR, Rauwerdink AM, et al. Ovarian cancer progression is controlled by phenotypic changes in dendritic cells. J Exp Med. 2012;209(3):495–506. doi:10.1084/jem.20111413
 © 2024 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2024 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.