Back to Journals » International Journal of General Medicine » Volume 14
A Comprehensive Way to Access Hospital Death Prediction Model for Acute Mesenteric Ischemia: A Combination of Traditional Statistics and Machine Learning
Received 14 January 2021
Accepted for publication 4 February 2021
Published 25 February 2021 Volume 2021:14 Pages 591—602
DOI https://doi.org/10.2147/IJGM.S300492
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Wenhan Wu,1,2 Zongguang Zhou1,2
1Institute of Digestive Surgery of Sichuan University, Chengdu, 610041, Sichuan; 2Department of Gastrointestinal Surgery, West China Hospital, West China School of Medicine, Sichuan University, Chengdu, 610041, Sichuan
Correspondence: Zongguang Zhou
Institute of digestive surgery of Sichuan University, and department of gastrointestinal surgery, West China hospital, west China school of medicine, Sichuan university, Chengdu, 610041, Sichuan
Email [email protected]
Purpose: This study aimed to use traditional statistics and machine learning to develop and validate prediction models for predicting hospital death in patients with AMI and compare these models’ performance.
Patients and Methods: Data were retrieved from the Medical Information Mart for Intensive Care (MIMIC III) electronic clinical database. A total of 338 eligible AMI patients were divided into a training cohort (n = 238) and a validation cohort (n = 100), and all patients were divided into survival groups and nonsurvival groups according to patients’ hospital outcomes. The performance of the traditional statistics prediction model and the optimal machine learning prediction model was evaluated and compared with respect to discrimination, calibration, and clinical utility in the validation cohort.
Results: Univariate and multivariate logistic regression analyses identified the following independent risk factors associated with hospital death for AMI in the training cohort, including diastolic blood pressure, blood lactate, blood creatinine, age, blood pH, and red blood cell distribution width. Both the nomogram (AUC = 77.0%, 67.9– 86.1%) and optimal machine learning model (AUC = 82.9%, 74.9– 91.0%) achieved good discrimination and calibration in the validation cohort. Decision curves analysis showed that the optimal machine learning model has a greater net benefit than that of nomogram in this study.
Conclusion: The nomogram achieved a concise and relatively accurate prediction of hospital death in patients with AMI, the machine learning model also has good discrimination and seems to have better clinical utility. Traditional statistics may help infer the relationship between risk factors and hospital death, while machine learning may contribute to a more accurate prediction. Traditional statistics and machine learning are complementary in developing the prediction model for hospital death of AMI. Therefore, a combination of nomogram–machine learning (Nomo-ML) predictive model may improve care and help clinicians make AMI management-related decisions.
Keywords: acute mesenteric ischemia, hospital mortality, prediction model, nomogram, machine learning
Introduction
Acute mesenteric ischemia (AMI) is a class of disease, usually caused by a sudden lack of intestinal blood supply, including arterial occlusive mesenteric ischemia, mesenteric venous thrombosis, and nonocclusive mesenteric ischemia.1 Because of the non-specific symptoms and immature imaging technology, it led to difficulties in diagnosis, which seriously affected patients’ survival in the last century. Nowadays, the application of contrast-enhanced Computed Tomography (CT) has made it possible for a timely diagnosis of AMI.2 Besides, the development of vascular manipulation and surgical techniques has improved the prognosis of AMI.3 However, there were still many patients who died under current treatment management, with high hospital mortality of about 40%.4 Although AMI is an uncommon surgical emergency, whose incidence is about 0.09–0.2% per year. Unfortunately, in the situation of the aging population and the prevalence of risk factors contributing to AMI, the incidence seems to have increased. Thus, some studies have examined the risk factors related to the hospital death of AMI, aimed to identify high-risk patients early.5–7 Recently, several intestine stroke centers have shown that the multidisciplinary stepwise management strategy for AMI might reduce hospital mortality and improve patient prognosis.8,9 However, factors affecting the hospital mortality and prognosis of AMI are not fully understood, and the accurate prognostic models of early judging the risk of hospital death for AMI are still lacking. Therefore, improving knowledge about the prognosis of AMI patients and creating an accurate mortality risk prediction model would allow clinicians to provide more precise clinical information for patients and their families, optimize current management, and inspire future clinical trial design. This accurate prognostic information is also conducive to realizing AMI’s medical care according to local conditions, reasonably controlling medical expenditures, and maximizing medical resource allocation.
The nomograms that can provide individualized and accurate risk estimation have been widely used in different clinical settings, mainly to achieve the graphical representation of the traditional statistics-based multivariate predictive models.10 Machine learning, a subfield of artificial intelligence, is a new and developed technology in data mining, which can extract models describing data patterns from experience (existing sample) and predict unseen data results.11
In particular, after variable selection, the classification prediction algorithms could also realize clinical risk estimation, which were also widely used in biomedical research,12,13 and variable selection is pertinent as it is aimed at removing unrelated variables from the clinical predictive models to reduce the complexity without compromising its accuracy.
Our aim was 1) to establish a visual predictive nomogram of hospital death in patients with AMI using traditional statistics. 2) to develop models using variable selection and classification algorithms in machine learning to predict AMI patients’ hospital death risk. 3) to compare and validate the performance of these models in discrimination, calibration, and clinical utility.
Patients and Methods
Data Collection and Study Design
The dataset for this study was derived from MIMIC-III (Medical Information Mart for Intensive Care, https://mimic.physionet.org/).14 All patients admitted to the hospital due to AMI from 2001 to 2012 were retrospectively included in our study. The following were exclusion criteria: patients with AMI induced by other diseases (such as aortic dissection, burn), ischemic colitis, necrotizing enterocolitis or incomplete data. The demographic characteristics, past medical history, laboratory tests on admission, initial interventions, and the outcome of each patient were collected. All work in this study was carried out in accordance with the Declaration of Helsinki, and all data were collected anonymously without affecting medical decisions. The use of MIMIC-III was approved by the institutional review boards of the Massachusetts Institute of Technology and Beth Israel Deaconess Medical Center.
To develop the models for predicting the hospital mortality of patients with AMI, three major experiments were conducted in our study. The dataset was split into two cohorts: 238 records for model construction and 100 records for model evaluation and validation. On the training cohort, we used univariate analysis and multivariate logistic regression to determine independent risk factors for the hospital mortality of patients with AMI, and then the nomogram was constructed based on these independent risk factors. Similarly, we used Lasso and Boruta to select the potential predictors, then based on the selected predictors, SVM, XGBoost, and ELM were adopted to develop the predictive models. Finally, The performance of the nomogram and the optimal machine learning prediction model was evaluated and compared with respect to discrimination, calibration, and clinical utility in the validation cohort. A diagram of this present experiment is illustrated in Figure 1. When performing classification tasks, the grid search strategy was used to determine the hyperparameters. In grid search, we set up a grid of hyperparameters and train/test our predictive model on each of the possible combinations in the training cohort using 5-fold cross-validation, and the hyperparameters of the model with the highest accuracy were considered the best. All analyses in this study were reported according to the Transparent Reporting of a Multivariable Prediction Model for Individual Prognosis or Diagnosis (TRIPOD) guidelines.15
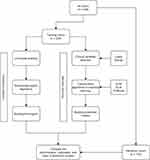 |
Figure 1 Diagram of developing AMI hospital mortality prediction models. |
Outcome Indicator
The outcome indicator was defined as follows: survivors referred to the patients who survived and stable vital signs when discharged from the hospital. Nonsurvivors referred to the patients who died during hospitalization.
Statistical Analysis
Each variable distribution was presented as Mean±SD (normally distributed numerical variable), or median with interquartile range (numerical variables that do not conform to the normal distribution), or frequency (categorical variable). To detect significant differences between nonsurvivors and survivors in AMI, the Student’s t-test or Mann–Whitney U-test for the numerical variable, as well as the Chi-square test for the categorical variables were chosen. All the tests were two-tailed. All statistical analyses were performed using the R software (version 3.5.3). P < 0.05 was considered statistically significant.
Construction of the Nomogram
Univariate logistic regression was performed to evaluate each statistically significant variables in the training cohort. Then variables with P < 0.05 were included in a final multivariate logistic regression using the backward step-down selection procedure, with a liberal P < 0.05 as the retention criteria to select the independent risk factors for hospital mortality of AMI. The estimate of relative risk was evaluated by the odds ratio (OR) with a 95% confidence interval (CI). Finally, a nomogram was built based on the result of multivariate logistic regression, and the “rms” (version 5.1–4) package was used for creating the nomogram.16
Machine Learning
Variable Selection Method
Lasso is a linear regression method that uses L1 regularization, resulting in a weight of zero for some variables.17 Therefore, this method can handle the tasks of sparseness and variable selection. In order to find the optimal hyperparameter penalty coefficient, we calculated the mean-squared error using ten-fold cross-validation. The ratio at a standard deviation from the minimum mean-squared error model was considered as the optimal hyperparameter in this method. The “glmnet” (version 4.0) package was used for fitting Lasso regression.18
Boruta is a variable selection wrapper algorithm that can output the importance of variables. By default, Boruta uses random forest. This method compares the importance of the original variables and randomly available (shadow) variables, and gradually eliminates irrelevant variables to stabilize that procedure, thereby achieving a top-down selection of variables. The “Boruta” (version 7.0.0) package was used for Boruta variable selection.19
Classification Algorithm
SVM is a typical kernel-based supervised learning algorithm. The basic idea is to create a hyperplane among data points to maximize the classification interval.20 The probability measurement theory and the law of large numbers were not involved, which undoubtedly simplifies the classification tasks. The “kernlab” (version 0.9–29) package was used for carrying out SVM.21
XGBoost is an efficient system implementation of Gradient Boosting. This method provides a parallel tree boosting, and it can explicitly regularize the tree model. So it can solve many classification tasks in a fast and accurate way. The “xgboost” (version 1.0.0.2) package was used for carrying out XGBoost.22
ELM is a learning algorithm for solving a single hidden layer feedforward neural network. The key innovation of this method is that the connection weights of the input layers and the hidden layers are randomly assigned, and the connection weights between the hidden layers and output layers are not required iterative adjustment but are determined by solving equations.23 The “elmNNRcpp” (version 1.0.2) package was used for carrying out ELM.24
Evaluation Techniques
In this study, three essential measures were adopted to evaluate the performance of the prediction model on the validation cohort. The receiver operating characteristic (ROC) was used to assess the nomogram and machine learning model discrimination. The discriminative power of the prediction models was determined by the area under the curve (AUC) with 95% confidence interval (CI). The calibration curve and Hosmer-Lemeshow test (the non-statistical significance of the test indicates a good agreement) were used to assess the nomogram and optimal machine learning model with the highest discriminative power. Finally, we analyzed the net benefit (proportion of true-positive results minus the proportion of weighted false-positive results) of the nomogram and the optimal machine learning model. We plotted each model’s DCA (Decision Curve Analysis) curve according to different weights (threshold probability).25 The prediction models with higher net benefits were considered to have higher clinical utility.
Results
Patients Characteristics and Survival
A total of 338 eligible patients with AMI were involved in this study, including 162 men (47.9%) and 176 women (52.1%), with a median age of 67.9 years old. The overall hospital mortality rate was 34.6% (n = 117). Among them, 238 patients were included in the training cohort, and 100 patients were included in the validation cohort. There were no differences in the clinical characteristics between the two datasets in most of the comparisons. The comparison of demographic characteristics, clinical and laboratory examinations upon admission, and outcome in survivors and nonsurvivors were listed in Table 1. The nonsurvivors were more likely to have lower systolic (SBP) and diastolic blood pressure (DBP), lower hematocrit, lower mean corpuscular hemoglobin concentration (MCHC), lower platelet, lower blood pH, higher mean corpuscular volume (MCV), higher red cell distribution width (RDW), higher lactate, higher anion gap, higher blood creatinine, higher blood urea nitrogen, and higher age. Besides, the nonsurvivors experienced congestive heart failure, chronic kidney disease more often.
 |
Table 1 Baseline Patient Demographics, Clinical and Laboratory Characteristics, and Outcomes |
Univariate and Multivariate Analysis, Nomogram Development
The results of univariate logistic regression analysis were shown in Table 2. Stepwise multivariate logistic regression indicated that the DBP (OR = 0.955, 95% CI [0.934–0.976]; P < 0.001), blood lactate (OR = 1.407, 95% CI [1.185–1.671]; P < 0.001), blood pH (OR = 0.009, 95% CI [0.001–0.339]; P =0.011), blood creatinine (OR = 1.524, 95% CI [1.210–1.919]; P < 0.001), RDW (OR = 1.431, 95% CI [1.190–1.720]; P < 0.001), and age (OR = 1.048, 95% CI [1.019–1.077]; P = 0.001) were independent predictors for hospital death of AMI (Table 3). These independent predictors were used to construct a AMI hospital death risk estimation nomogram. The nomogram contained a score scale, a total of score scale, and a probality scale, each predictor also corresponded to a scale (Figure 2).
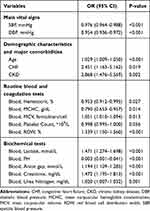 |
Table 2 Univariate Analysis for Potential Risk Factors in the Training Cohort |
 |
Table 3 Multivariate Logistic Regression Model for Hospital Mortality in the Training Cohort |
 |
Figure 2 The hospital death risk-prediction nomogram for AMI. |
Machine Learning
For Lasso variable selection, we used the 10-fold cross-validation to explore the optimal lambda value, and the misclassification error was the target that we wanted to minimize. In order to obtain a model with excellent performance and relatively few features, the penalty coefficient value (lambda lse), whose model is one standard deviation away from the minimum mean-squared model was used as the hyperparameter in the final model (Figure S1). Consequently, a total of seven clinical variables were determined by Lasso, including diastolic blood pressure, blood lactate, blood pH, anion gap, blood creatinine, RDW, age (Table S1). For Boruta variable selection, by default, this method searched for important variables by comparing the importance of original variables with randomly available (shadow) variables (Figure S2). Consequently, a total of nine clinical variables were determined by Boruta, including blood lactate, anion gap, blood creatinine, systolic blood pressure, RDW, diastolic blood pressure, blood pH, age, and blood urea nitrogen (Table S2).
According to the clinical variables selected by Lasso and Boruta, three classification algorithms, including SVM, XGBoost, and ELM, were used to construct the prediction model of AMI hospital death, and the hyperparameters for each classification model are shown in Table S3.
Model Evaluation
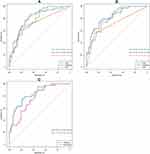
Table 4 summarizes the performance of the different machine learning predictive models tested in the validation cohort. Among the machine learning predictive models with a different combination of variable selection method and classification algorithm, the XGBoost model using clinical variables determined by Boruta achieved the highest accuracy (AUC = 82.9%, 95% CI 74.9–91.0%), which was considered as the optimal machine learning model for predicting hospital death risk of AMI in our study (Figure 3A and B). Figure 3C further presents the ROC curves of the nomogram and optimal machine learning model tested on the validation cohort. The AUC (95% CI) for nomogram and optimal machine learning model was 77.0% (67.9–86.1%) and 82.9% (74.9–91.0%), respectively.
 |
Table 4 The Comparison of Various Machine Learning Classifiers’ Performance Using Different Variable Selection Methods in the Validation Cohort |
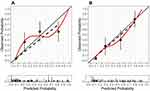
The results of the calibration curve and Hosmer-Lemeshow test statistics (P = 0.076) for nomogram are presented in Figure 4A, which showed the probabilities of AMI hospital death predicted by the nomogram agreed well with the actual probability. The calibration curve and Hosmer-Lemeshow test statistics (P = 0.877) for the optimal machine learning model also showed good calibration in the validation cohort (Figure 4B).
Decision curve analysis was used to assess the clinical utility of the nomogram and the optimal machine learning model. The decision curve graphically presented the clinical utility of the model using a continuous probability threshold (X-axis) and the model’s net benefit (Y-axis). The decision curve indicated that the net benefit of the optimal machine learning model is greater than that of the nomogram, when the threshold probability for a doctor is within a range from 0.22–0.85 (Figure 5).
 |
Figure 5 Decision curves for the nomogram and the optimal machine learning classifier (XGBoost using clinical variables determined by Boruta) in the validation cohort. |
Discussion
In this retrospective analysis, we investigated the clinical characteristics, admission status, and initial interventions of AMI. Then stepwise multivariate logistic regression was used to recognize the independent risk factors related to hospital mortality of AMI and predictors including diastolic blood pressure, blood lactate, blood creatinine, age, blood pH, and RDW. A nomogram was generated based on six variables at admission to predict hospital death of AMI. Moreover, we also used machine learning techniques to develop the prediction models for hospital mortality of AMI, and compared the performance of the nomogram and the optimal machine learning model on a separate validation set. To the best of our knowledge, this study is the first study to relatively comprehensively use traditional statistical methods and machine learning techniques in the context of hospital mortality prediction of AMI. This combinatorial analysis is necessary to ensure that a full understanding of risk factors and the best model is selected for the prediction of AMI hospital death.
In medical research, age is associated with high mortality in patients with AMI.4,26 This was similar to the finding of our research. Our study also indicated that low blood pressure upon admission was also closely related to the hospital death of patients with AMI, which was compatible with previous medical literature.27 Acidosis has been associated with high mortality in many reports.6,26 We found a similar relationship, low pH was a significant predictor in nomogram, which was considered to be associated with many adverse prognostic factors, such as renal failure, symptom duration, and range of intestine necrosis. Lactate is usually a key parameter closely related to necrosis, inflammation, and hypoxia. Our results showed that high lactate levels were significantly related to the occurrence of AMI hospital death. Moreover, previous studies have shown that high blood lactate was often significantly associated with transmural necrosis of the intestine, and removal of the necrotic intestine could reduce blood lactate.28 In previous studies, high serum creatinine was reported as an essential risk factor for hospital mortality of AMI.4,26 Our study also indicated similar results, which highlight the importance of normal kidney function in AMI management. Besides, our study also found that elevated RDW was also an independent risk factor. Although few previous studies have reported the association between RDW and AMI prognosis, studies have shown that elevated RDW is closely related to sepsis.29 The damaged intestinal mucosa loses its resistance to bacteria, which leads to bacteria or even sepsis in some AMI patients might explain why AMI patients with high RDW had a poor prognosis.
Several studies have shown the significance of machine learning techniques in predicting disease prognosis.12,30 XGBoost using clinical variables determined by Boruta models achieve the highest accuracy in the machine learning models, which also outperformed the nomogram in discrimination and clinical utility. The predictors used in this machine learning model included six predictors in the nomogram, as well as SBP, anion gap, and blood urea nitrogen. The three additional variables were also confirmed to be related to hospital deaths of AMI in univariate analysis. However, the relatively strong correlation of variables (Pearson’s r creatinine-urea nitrogen = 0.537, Pearson’s r DBP-SBP = 0.569, Pearson’s r lactate-anion gap = 0.543) might cause the regression coefficients of the three variables were not statistically significant in stepwise logistic regression. While XGBoost uses the gradient boosting method to fit the residuals of the last prediction to create new classification trees until the last tree continuously, and the prediction of the model is the integration of the results of all trees.22 So, this method might better handle the large coverage of the correlation between the variables to improve the accuracy in this study.
Generally, the experience of the clinician plays an essential role in the patient’s risk estimation and decision-making, but it may have a considerable risk of deviation and is relatively subjective.31 The nomogram has been widely used in the field of disease risk prediction, and machine learning techniques have also shown encouraging results. In this study, traditional statistical techniques (univariate, multivariate, and nomogram) allow us to identify independent risk factors related to hospital death of AMI, and construct a transparent and concise risk prediction model that could estimate the death risk without the need for the internet or computers. The machine learning model seemed to have higher accuracy and higher clinical utility because it can identify and understand the indistinguishable relationship between variables. However, the lack of explicit models made it difficult for machine learning models to associate with existing biological knowledge directly. Therefore, the combination of nomogram-machine learning (Nomo-ML) may provide a more transparent and accurate method for assessing the risk of hospital death in patients with AMI and help to optimize the management of AMI.
As with any study, this work had limitations. Firstly, this study was a retrospective study, some bias inevitably existed and might affect the nomogram and machine learning models. Therefore, it is still necessary to further compare the performance of these tools through a prospective cohort. Secondly, due to data limitations, we cannot construct a hospital death prediction model for each subtype of AMI, which is still worth exploring in the future.
Conclusion
In conclusion, we have used traditional statistical methods to identify potential risk factors related to hospital death of AMI, and constructed a concise and accurate nomogram for risk prediction. Also, machine learning models achieved high accuracy and seemed to have higher clinical utility.
Traditional statistics may help infer the relationship between risk factors and hospital death of AMI, while machine learning may contribute to a more accurate prediction. The combination of nomogram and machine learning techniques may help provide a transparent and accurate disease risk prediction model.
Acknowledgments
This study was supported by Sichuan University West China Hospital Disciplinary Excellence Development 1.3.5 Project (ZY2016105). Besides, we thank M.S. Jia He for her consultation on data collection.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Clair DG, Beach JM. Mesenteric ischemia. New Engl J Med. 2016;374(10):959–968. doi:10.1056/NEJMra1503884
2. Menke J. Diagnostic accuracy of multidetector CT in acute mesenteric ischemia: systematic review and meta-analysis. Radiology. 2010;256(1):93–101. doi:10.1148/radiol.10091938
3. Duran M, Pohl E, Grabitz K, Schelzig H, Sagban TA, Simon F. The importance of open emergency surgery in the treatment of acute mesenteric ischemia. WJES. 2015;10:45. doi:10.1186/s13017-015-0041-6
4. Wu W, Liu J, Zhou Z. Preoperative risk factors for short-term postoperative mortality of acute mesenteric ischemia after laparotomy: a systematic review and meta-analysis. Emerg Med Int. 2020;2020:1382475. doi:10.1155/2020/1382475
5. Huang HH, Chang YC, Yen DH, et al. Clinical factors and outcomes in patients with acute mesenteric ischemia in the emergency department. J Chinese Med Assoc. 2005;68(7):299–306. doi:10.1016/S1726-4901(09)70165-0
6. Acosta-Merida MA, Marchena-Gomez J, Hemmersbach-Miller M, Roque-Castellano C, Hernandez-Romero JM. Identification of risk factors for perioperative mortality in acute mesenteric ischemia. World J Surg. 2006;30(8):1579–1585. doi:10.1007/s00268-005-0560-5
7. Wu W, Yang L, Zhou Z. Clinical features and factors affecting postoperative mortality for obstructive acute mesenteric ischemia in China: a hospital-based survey. Vasc Health Risk Manag. 2020;16:479–487. doi:10.2147/VHRM.S261167
8. Corcos O, Castier Y, Sibert A, et al. Effects of a multimodal management strategy for acute mesenteric ischemia on survival and intestinal failure. Clin Gastroenterol Hepatol. 2013;11(2):158–165. e152. doi:10.1016/j.cgh.2012.10.027
9. Yang S, Fan X, Ding W, et al. Multidisciplinary stepwise management strategy for acute superior mesenteric venous thrombosis: an intestinal stroke center experience. Thromb Res. 2015;135(1):36–45. doi:10.1016/j.thromres.2014.10.018
10. Iasonos A, Schrag D, Raj GV, Panageas KS. How to build and interpret a nomogram for cancer prognosis. J clin oncol. 2008;26(8):1364–1370. doi:10.1200/JCO.2007.12.9791
11. Alpaydin E. Introduction to Machine Learning. MIT press; 2020.
12. Vieira SM, Mendonça LF, Farinha GJ, Sousa JM. Modified binary PSO for feature selection using SVM applied to mortality prediction of septic patients. Appl Soft Comput. 2013;13(8):3494–3504. doi:10.1016/j.asoc.2013.03.021
13. Shen Y, Wu C, Liu C, Wu Y, Xiong N. Oriented feature selection SVM applied to cancer prediction in precision medicine. IEEE Access. 2018;6:48510–48521. doi:10.1109/ACCESS.2018.2868098
14. Johnson AE, Pollard TJ, Shen L, et al. MIMIC-III, a freely accessible critical care database. Scientific Data. 2016;3:160035. doi:10.1038/sdata.2016.35
15. Moons KG, Altman DG, Reitsma JB, et al. Transparent Reporting of a multivariable prediction model for Individual Prognosis or Diagnosis (TRIPOD): explanation and elaboration. Ann Intern Med. 2015;162(1):W1–W73. doi:10.7326/M14-0698
16. Harrell FE. Regression modeling strategies. BIOS. 2017;330:2018.
17. Tibshirani R. Regression shrinkage and selection via the lasso. J Royal Stat Soc Series B. 1996;58(1):267–288.
18. Friedman J, Hastie T, Tibshirani R. Regularization paths for generalized linear models via coordinate descent. J Stat Softw. 2010;33(1):1. doi:10.18637/jss.v033.i01
19. Kursa MB, Rudnicki WR. Feature selection with the Boruta package. J Stat Softw. 2010;36(11):1–13. doi:10.18637/jss.v036.i11
20. Hastie T, Tibshirani R, Friedman J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction. Springer Science & Business Media; 2009.
21. Karatzoglou A, Smola A, Hornik K, Zeileis A. kernlab-an S4 package for kernel methods in R. J Stat Softw. 2004;11(9):1–20. doi:10.18637/jss.v011.i09
22. Chen T, He T, Benesty M, Khotilovich V, Tang Y. Xgboost: extreme gradient boosting. R package version 04-2. 2015;1–4.
23. Huang G-B, Zhu Q-Y, Siew C-K. Extreme learning machine: theory and applications. Neurocomputing. 2006;70(1–3):489–501. doi:10.1016/j.neucom.2005.12.126
24. Mouselimis L, Gosso A. elmNNRcpp: the extreme learning machine Algorithm, R Package Version 1.0. 1.
25. Vickers AJ, Elkin EB. Decision curve analysis: a novel method for evaluating prediction models. Med Decision Making. 2006;26(6):565–574. doi:10.1177/0272989X06295361
26. Akyildiz HY, Sozuer E, Uzer H, Baykan M, Oz B. The length of necrosis and renal insufficiency predict the outcome of acute mesenteric ischemia. Asian J Surg. 2015;38(1):28–32. doi:10.1016/j.asjsur.2014.06.001
27. Edwards MS, Cherr GS, Craven TE, et al. Acute occlusive mesenteric ischemia: surgical management and outcomes. Ann Vasc Surg. 2003;17(1):72–79. doi:10.1007/s10016-001-0329-8
28. Nuzzo A, Maggiori L, Ronot M, et al. Predictive factors of intestinal necrosis in acute mesenteric ischemia: prospective study from an intestinal stroke center. Am J Gastroenterol. 2017;112(4):597–605. doi:10.1038/ajg.2017.38
29. Jandial A, Kumar S, Bhalla A, Sharma N, Varma N, Varma S. Elevated red cell distribution width as a prognostic marker in severe sepsis: a prospective observational study. Indian J Critical Care Med. 2017;21(9):552. doi:10.4103/ijccm.IJCCM_208_17
30. Rajkomar A, Dean J, Kohane I. Machine learning in medicine. New Engl J Med. 2019;380(14):1347–1358. doi:10.1056/NEJMra1814259
31. Elstein AS. Heuristics and biases: selected errors in clinical reasoning. Acad Med. 1999;74:791–794. doi:10.1097/00001888-199907000-00012
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


