Back to Journals » Clinical Epidemiology » Volume 7
Validation of International Classification of Diseases coding for bone metastases in electronic health records using technology-enabled abstraction
Authors Liede A, Hernandez RK, Roth M, Calkins G, Larrabee K, Nicacio L
Received 16 July 2015
Accepted for publication 9 September 2015
Published 11 November 2015 Volume 2015:7 Pages 441—448
DOI https://doi.org/10.2147/CLEP.S92209
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Vera Ehrenstein
Alexander Liede,1 Rohini K Hernandez,1 Maayan Roth,2 Geoffrey Calkins,2 Katherine Larrabee,2 Leo Nicacio2
1Center for Observational Research, Amgen Inc., South San Francisco and Thousand Oaks, CA, 2Flatiron Health, New York, NY, USA
Objective: The accuracy of bone metastases diagnostic coding based on International Classification of Diseases, ninth revision (ICD-9) is unknown for most large databases used for epidemiologic research in the US. Electronic health records (EHR) are the preferred source of data, but often clinically relevant data occur only as unstructured free text. We examined the validity of bone metastases ICD-9 coding in structured EHR and administrative claims relative to the complete (structured and unstructured) patient chart obtained through technology-enabled chart abstraction.
Patients and methods: Female patients with breast cancer with ≥1 visit after November 2010 were identified from three community oncology practices in the US. We calculated sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) of bone metastases ICD-9 code 198.5. The technology-enabled abstraction displays portions of the chart to clinically trained abstractors for targeted review, thereby maximizing efficiency. We evaluated effects of misclassification of patients developing skeletal complications or treated with bone-targeting agents (BTAs), and timing of BTA.
Results: Among 8,796 patients with breast cancer, 524 had confirmed bone metastases using chart abstraction. Sensitivity was 0.67 (95% confidence interval [CI] =0.63–0.71) based on structured EHR, and specificity was high at 0.98 (95% CI =0.98–0.99) with corresponding PPV of 0.71 (95% CI =0.67–0.75) and NPV of 0.98 (95% CI =0.98–0.98). From claims, sensitivity was 0.78 (95% CI =0.74–0.81), and specificity was 0.98 (95% CI =0.98–0.98) with PPV of 0.72 (95% CI =0.68–0.76) and NPV of 0.99 (95% CI =0.98–0.99). Structured data and claims missed 17% of bone metastases (89 of 524). False negatives were associated with measurable overestimation of the proportion treated with BTA or with a skeletal complication. Median date of diagnosis was delayed in structured data (32 days) and claims (43 days) compared with technology-assisted EHR.
Conclusion: Technology-enabled chart abstraction of unstructured EHR greatly improves data quality, minimizing false negatives when identifying patients with bone metastases that may lead to inaccurate conclusions that can affect delivery of care.
Keywords: electronic medical records, EHR, US, ICD-9, breast cancer, unstructured data
Introduction
Bone is a common site of metastatic cancer.1,2 Bone metastases can occur in most tumor types but are most common in patients diagnosed with breast, lung, and prostate cancers.3 Bone metastases represent an irreversible condition associated with progressive morbidity. Once diagnosed, patients are vulnerable to bone complications, also referred to as skeletal-related events or SREs (ie, radiation therapy to the bone, pathological or osteoporotic fractures, spinal cord compression, and surgery to the bone) that impact quality of life.4–8 Bone metastases have been associated with higher mortality among women with breast cancer in the US, and the association was stronger for bone metastasis complicated by SREs than for bone metastasis without SREs.9,10 The ability to reliably study patients with bone metastases has potential public health implications.
Most observational studies of the incidence, costs, morbidity, and mortality of bone metastases have relied on International Classification of Diseases, ninth revision (ICD-9) codes in claims databases to identify cases. However, claims-based data are administrative in nature and used for billing purposes; therefore, the validity for epidemiological investigations using these data has been questioned.11 Accordingly, claims-based studies of bone metastasis are inherently limited in their generalizability and capture of patients with metastatic bone disease who are treated with bone-targeting agents (BTAs) whereby ICD-9 coding is populated in the patient record for billing purposes, resulting in underestimation of the burden of bone metastases in the population and potential bias in interpreting treatment trends.12
The at times nonspecific nature of the ICD-9 or ICD-10 coding usually requires various proxy codes and code combinations to improve sensitivity and increase complete capture of potential bone metastasis cases. For example, a claims-based study aimed to estimate the prevalence of bone metastases in the US population on December 31, 2008, relied on either ICD-9 coding for secondary malignant neoplasm of bone or bone marrow (198.5) to identify cases or a claim carrying Healthcare Common Procedure Coding System codes for antiresorptive therapies used to prevent SREs among patients with bone metastases: intravenous bisphosphonates zoledronic acid (Zometa®) or pamidronate (Aredia®).3 In a study of patients with breast cancer in the United Kingdom, data from three databases (General Practice Research Database, National Cancer Registry, and Hospital Episodes Statistics) were linked to create an algorithm for identifying bone metastases.13 Even with the aid of such algorithms, results were thought to underestimate the prevalence of bone metastases.3 Similar attempts using algorithms to improve capture of patients with bone metastases have been attempted in other population-based data sources, such as in the Danish National Registry of Patients (DNRP),14,15 demonstrating limited sensitivity but high specificity.
In the US, the Surveillance, Epidemiology, and End Results (SEER) cancer registries represent a high-quality source for population-based cancer incidence data that is linkable to Medicare data to improve data capture for the Medicare-insurance eligible population aged ≥66 years. SEER registries collect standardized information that meet stringent quality standards, but there are limitations in the number of clinical variables collected and availability of information on disease progression and health utilization. These limitations are significant for breast and other cancers where multigene testing is routine for the diagnostic and prognostic characterization of tumors or for understanding disease progression such as development of metastases.16 A recent validation study of SEER-Medicare linked data in prostate cancer found 59% sensitivity, 54% specificity, and 68% positive predictive value (PPV) for identifying bone metastasis at diagnosis, concluding that claims-based measures using ICD-9 coding may be insufficient to identify patients with incident bone metastases and should be validated against chart data to maximize their potential for population-based analyses.12
Electronic health records (EHR) have been adopted widely by oncology clinics in the US and represent the primary source of information for clinicians tracking the care of their patients. Information fed into these systems may be found in structured fields for which values are sometimes selected from a drop-down menu (eg, sex) or inputted electronically (eg, laboratory test orders and results). More commonly, however, information is found in an unstructured manner, including clinically relevant information such as tumor-specific histopathology, physician notes, or in a scanned PDF-formatted document. Until recently, most research that included information from the unstructured EHR fields utilized manual abstraction, a time-consuming and error-fraught process. Electronic data capture methods have the potential to speed up data collection and improve data quality for this type of research.
In this study, we examined the validity of ICD-9 code 198.5 to identify patients diagnosed with bone metastases in the oncology clinic. To do this, we identified bone metastases among patients with breast cancer using structured EHR and administrative claims data and compared the results with data generated from a technology-enabled review of the source EHR, used as the reference standard for comparison and for calculations of the diagnostic accuracy measures. We calculated sensitivity, specificity, PPV, and negative predictive value (NPV). We evaluated effects of misclassification by studying the proportion of patients treated with BTAs (zoledronic acid, pamidronate, and denosumab), BTA timing relative to diagnosis, and SREs.
Methods
Data sources and study population
For this validation study, cohort assembly began by identifying patients treated at oncology practices in the US as compiled in a large longitudinal EHR database (Flatiron Health, Inc. New York, NY, USA; April 2014). The study cohort included patients with a clinic visit or treatment after November 2010. November 2010 relates to the FDA approval date of denosumab (XGEVA®), a fully human monoclonal antibody against receptor activator of nuclear factor kappa-B ligand (RANKL), for use in adults diagnosed with bone metastases from solid tumors for the prevention of SREs.17 This criterion ensured that women in the cohort were eligible to receive at least one of the BTA treatments approved in the US for bone metastasis secondary to breast cancer. Included were women diagnosed with invasive breast cancer, identified using ICD-9 174 and confirmed during chart abstraction. Women with breast cancer represent a large and suitable population for the study of bone metastasis as bone is the most common site of distant metastasis,18 and survival rates allow for sufficient follow-up in the EHR.19,20
We examined the validity of ICD-9 code 198.5 for diagnosis of bone metastasis from two oncology clinic-specific sources: codes used for diagnoses in the structured portion of the EHR (“structured EHR”) and codes used for billing medications (“claims”). Both sources were compared with the full medical chart. Medical chart review using technology-enabled abstraction of the EHR was used as the reference standard to establish the presence or absence of metastases.
Patients included in this study were treated at three independent community oncology practices. Practices varied in size from five oncologists to >30 oncologists. Each practice is located in a different state within the US. All practices use Elekta’s MOSAIQ® EHR, and no coding or documentation differences were observed between the practices. The Institutional Review Board of each oncology practice approved collaboration and contribution of data to a large longitudinal EHR database (Flatiron Health), and individual patient-level EHR data were encrypted at rest and in transit so that all analyses were de-identified to protect patient privacy consistent with the final Health Insurance Portability and Accountability Act (HIPAA) Security Rule from the US Department of Health and Human Services.
EHR technology-enabled abstraction
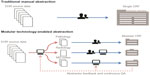
We developed a technology-enabled, modular approach to optimize both the accuracy and efficiency of data abstraction from unstructured sources in the EHR (Figure 1). This approach uses a software tool developed for the identification and targeted display of selected portions of the patient chart for abstraction. A team of clinically trained professionals (Flatiron Health oncology nurses and certified tumor registrars) access each chart within this software to collect only specific data elements across many patients’ charts, rather than review the entirety of any individual patient chart. This modular approach allows chart abstractors to focus their attention on specific data elements (eg, dates of bone metastasis) and to collect these elements uniformly. The approach also enables iterative assessment and improvement of the quality of each data element separately. Quality is continuously assessed by computing inter-abstractor agreement (inter-rater reliability) after subsets of charts were shown to multiple abstractors. An assessment of the inter-rater and test–retest reliability achieved via technology-enabled abstraction is the subject of a manuscript in preparation.
Additional information generated using technology-enabled chart abstraction of the unstructured EHR data included date of bone metastases diagnosis, SREs (defined by ie, radiation therapy to the bone, pathological or osteoporotic fractures, spinal cord compression and surgery to the bone), phenotype (estrogen receptor [ER], progesterone receptor [PR], and human epidermal growth factor receptor 2 [HER2] status), pain status, and Eastern Cooperative Oncology Group status.
Statistical analysis
For the patients with breast cancer with bone metastases, we described distributions of patients’ age at diagnosis, date of bone metastasis diagnosis, and treatment with BTAs.
We defined sensitivity as the proportion of women with breast cancer for whom a diagnosis of bone metastasis was recorded in the chart who also had the bone metastasis diagnosis recorded in structured EHR or claims data. Specificity was the proportion of individuals for whom the diagnosis of bone metastasis was not recorded in the chart and who did not have the outcome of interest recorded in structured EHR or claims data.
We computed the PPV as the proportion of positive results that are true positives, or the proportion of patients for whom the diagnosis of a bone metastasis was recorded in the structured EHR or claims data who also had the diagnosis confirmed by chart review including the unstructured EHR. The NPV represents the proportion of negative results that are true negatives, calculated as the proportion of patients for whom no bone metastasis was recorded in the structured EHR or claims data who had this negative result confirmed by chart review.
We estimated the impact of misclassification in sensitivity analyses with the inclusion/exclusion of cases without ICD-9 coding for bone metastases that would have been missed in the structured EHR or claims.
All estimates are presented with 95% confidence intervals (CIs). Statistical analyses were performed with R software (Version 3.1.0, Vienna, Austria).
Results
Across three large oncology practices, we identified 39,722 patients with cancer who received treatment with any drug therapy, 28,419 of whom were treated after November 2010 (after commercial availability of denosumab [XGEVA®]). Among these, we identified all women with a first-time breast cancer diagnosis confirmed in the medical chart physician note (n=8,796; Figure 2). Of these, 524 women had a bone metastasis recorded in their charts. The median age of breast cancer diagnosis was 53.3 years (mean =54.2 years; range =21.3–92.2 years; Table 1). Patients had a median follow-up of 21.1 months.
  | Figure 2 Study cohort selection. |
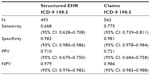
Of 524 patients with confirmed bone metastases in their chart, 350 (67%) also had a diagnosis recorded in the structured EHR. Thus, the sensitivity of the structured EHR was 0.67 (95% CI =0.63–0.71). The specificity, as expected, was high, at 0.98 (95% CI =0.98–0.99). The corresponding PPV was 0.71 (95% CI =0.67–0.75) and NPV was 0.98 (95% CI =0.98–0.98; Table 2). We had the opportunity to examine clinic-based reimbursement claims. This approach identified 406 of the 524 patients with bone metastases, resulting in a sensitivity of 0.78 (95% CI =0.74–0.81) with a specificity of 0.98 (95% CI =0.98–0.98), PPV of 0.72 (95% CI =0.68–0.76), and NPV of 0.99 (95% CI =0.98–0.99).
Basing bone metastases diagnosis on ICD-9 codes in either the structured EHR data or the associated claims missed 17% of bone metastases cases (89 of 524; Figure 3). The structured EHR and claims would have identified additional 183 patients as having bone metastases who did not have confirmation of this diagnosis upon review of the unstructured patient data: a false positive rate of 0.26 (95% CI =0.23–0.29).
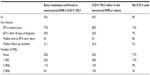
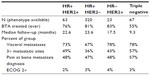
We conducted sensitivity analyses to compare the effects of excluding the 89 cases without an ICD-9 code for bone metastases with results generated using the 435 cases that were identified by ICD-9 code in EHR or claims data (Table 3). Among the 89 women without an ICD-9 code, there was shorter follow-up (9.2 months vs 23.4 months) compared with the 435 women who had a bone metastases coded in either the EHR or claims. In addition, more patients who were not coded for bone metastases were triple-negative phenotype (ER, PR, and HER2 negative; 21% vs 11%), fewer patients experienced an SRE (29% vs 55%), and fewer patients received a BTA (15% vs 89%) compared with those who were coded for bone metastases. The exclusion of these cases would introduce bias across key metrics related to BTA penetration, disease aggressiveness, outcomes such as SREs, or risk-adjustment formulas.
The inclusion of cases (n=618) identified using ICD-9 coding from structured EHR and claims data would result in an overestimation of the BTA penetration (89%) vs the BTA treatment among the 524 confirmed cases (77%). Similarly, the median time from bone metastasis diagnosis to BTA initiation was 1 day in the structured EHR or 0 day in claims data, whereas the confirmed 524 cases indicate the median time to BTA start is 43 days. Utilization of codes only would have conveyed an inaccurate representation of BTA initiation patterns. In the subpopulation with bone metastasis that was not identified by ICD-9 codes, the median time to BTA initiation was 86 days. Interestingly, even when an ICD-9 code is present, the median time from a confirmed physician note to structured EHR diagnosis and medical claim diagnosis were 32 days and 43 days, respectively.
Discussion
To our knowledge, this is the first study using observational data from routine clinical care in the US to evaluate the validity of ICD-9 code 198.5 for identifying a cohort of patients with breast cancer with bone metastases as recorded in structured EHR fields or in claims-based data. Validity of ICD-9 coding was assessed against the reference standard – the complete patient medical chart using technology-enabled abstraction of the source EHR. Studies based in EHR and claims are inherently dependent on the accuracy and completeness of ICD-9 coding; however, as a by-product of billing and administrative procedures, these codes may not accurately represent the clinical picture and may introduce misclassification bias when used for identifying a cohort of individuals with bone metastases.
Using chart review with a technology-enabled abstraction approach, we were able to demonstrate that ICD-9 codes in both structured EHR and claims-based data exhibit good specificity but missed approximately 17% of bone metastasis cases. We also found that the missed cases differed from the ICD-9 identified cases in ways that could bias treatment characterization and outcomes research (eg, lower BTA treatment and higher SRE rates for missed cases). Furthermore, the two data sources had limited sensitivity and low PPV (ie, never exceeding 80%). The validity of studies utilizing codes in structured EHR fields and claims-based data is limited, at best, for inferring bone metastasis or studying its incidence, costs, morbidity, or mortality.
There are several potential explanations for the degree of under-coding for bone metastases in the EHR. The numeric ICD-9 coding system is used to characterize observed medical events, and the presence of a bone metastasis may not be clinically obvious. The clinical decision to order diagnostic procedures used to detect bone metastases may depend on the patient’s expected prognosis. For instance, if a patient’s overall status is deemed inappropriate for BTA intervention to prevent subsequent bone complications (ie, poor prognosis), then there may be little incentive to code for bone metastases. Indeed, we found that cohort patients with poor prognostic indicators (eg, triple-negative breast cancer) were less likely to receive a BTA and more likely to have visceral metastases, three or more metastatic sites, and to report pain (Table 4).
It should be noted that the reporting of bone metastases is not mandatory. As our data suggest, bone metastases are more likely to be present in the patient record if a clinical decision is made to intervene on the condition, specifically the initiation of a BTA or referral to a specialist such as an orthopedic surgeon or radiation oncologist. This conclusion is substantiated by the short time interval between the date of the diagnostic ICD-9 code for bone metastasis and the initiation date of a BTA that is evident from the structured EHR or claims data.
This validation study demonstrated that access to the full oncology EHR history has advantages over structured EHR fields and claims data for the identification and characterization of patients with bone metastases. Due to limitations of claims and incompleteness of structured EHR data, chart abstraction should be seen as the preferred methodology to understand diseases and treatment patterns in women with bone metastasis from breast cancer.
Our findings are consistent with a study in Denmark evaluating ICD-10 coding for bone metastases in the DNRP.14 The validation study by Jensen et al compared the DNRP EHR data against manual review of patients’ medical records, resulting in a sensitivity for bone metastases ICD-10 coding of 0.58 (95% CI =0.34–0.80) among 100 patients with breast cancer with a high specificity of 0.95 (95% CI =0.88–0.99),14 meaning that the code failed to capture 42% of the patients with bone metastases from breast cancer. Depending on the role of the bone metastases capture in a breast cancer observational research study, low sensitivity (with underlying differences between identified and missed cases) will lead to underestimation of absolute risks, dilution of associations, or residual confounding.21 Although we recognize that, depending on the research question, the specificity and sensitivity of the ICD-9 coding for bone metastases in the structured EHR may be sufficient for epidemiologic investigations, we demonstrated in sensitivity analyses the impact of misclassification bias on metrics related to treatment patterns and outcomes of patients with bone metastases. Additionally, one key outcome of interest in this population, SREs, was only identified from physician note abstraction.
The findings from this study suggest that breast cancer researchers using EHR data should use caution when designing and interpreting results from studies in which patient cohorts are identified using ICD-9 codes on structured EHR data only. From the sensitivity, specificity, and predictive value measures reported here, under-ascertainment appears to be appreciable and related to patient characteristics portending bias in related analyses.21 Future investigations in the bone metastasis setting should incorporate chart abstraction, technology assisted or not, or adjust findings to account for under-ascertainment thereby avoiding conclusions based on overrepresentation of treated patients or those who experience SREs that could affect delivery of proper care and treatment of patients.
Conclusion
In conclusion, researchers should be cautious when interpreting results from studies in subpopulations with bone metastases that are anchored on ICD-9 codes from structured EHR or claims databases. Technology-enabled abstraction of EHR improved data completeness and accuracy, and it should be seen as a valid reference standard, until further investigations validate the utilization of structured data as a valid surrogate. By using unstructured data to validate structured data (eg, biomarker data), we demonstrated improved accuracy of insights from EHR analyses.
Acknowledgments
Results submitted to ASCO 2015 (publication only), Abstract Number e12652, Title: Abstract Title: Improving misclassification of ICD-9 coding for bone metastases in electronic medical records (EMR) using technology-enabled abstraction. This study represents unfunded research in 2014. However, as of 2015, Amgen and Flatiron are in partnership to conduct similar research.
Disclosure
AL, RKH are employees of Amgen Inc. MR, GC, KL are employees of Flatiron Health. LN was an employee of Flatiron Health at the time of this research.
References
Roodman GD. Mechanisms of bone metastasis. N Engl J Med. 2004; 350(16):1655–1664. | |
Coleman RE. Metastatic bone disease: clinical features, pathophysiology and treatment strategies. Cancer Treat Rev. 2001;27(3):165–176. | |
Li S, Peng Y, Weinhandl ED, et al. Estimated number of prevalent cases of metastatic bone disease in the US adult population. Clin Epidemiol. 2012;4:87–93. | |
Clare C, Royle D, Saharia K, et al. Painful bone metastases: a prospective observational cohort study. Palliat Med. 2005;19(7):521–525. | |
Diel IJ. Effectiveness of bisphosphonates on bone pain and quality of life in breast cancer patients with metastatic bone disease: a review. Support Care Cancer. 2007;15(11):1243–1249. | |
Martin M, Bell R, Bourgeois H, et al. Bone-related complications and quality of life in advanced breast cancer: results from a randomized phase III trial of denosumab versus zoledronic acid. Clin Cancer Res. 2012;18(17):4841–4849. | |
Solomayer EF, Diel IJ, Meyberg GC, Gollan C, Bastert G. Metastatic breast cancer: clinical course, prognosis and therapy related to the first site of metastasis. Breast Cancer Res Treat. 2000;59(3):271–278. | |
von Moos R, Body JJ, Egerdie B, et al. Pain and health-related quality of life in patients with advanced solid tumours and bone metastases: integrated results from three randomized, double-blind studies of denosumab and zoledronic acid. Support Care Cancer. 2013;21(12):3497–3507. | |
Sathiakumar N, Delzell E, Morrisey MA, et al. Mortality following bone metastasis and skeletal-related events among women with breast cancer: a population-based analysis of US. Medicare beneficiaries, 1999–2006. Breast Cancer Res Treat. 2012;131(1):231–238. | |
Saad F, Lipton A, Cook R, Chen YM, Smith M, Coleman R. Pathologic fractures correlate with reduced survival in patients with malignant bone disease. Cancer. 2007;110(8):1860–1867. | |
Chawla N, Yabroff KR, Mariotto A, McNeel TS, Schrag D, Warren JL. Limited validity of diagnosis codes in Medicare claims for identifying cancer metastases and inferring stage. Ann Epidemiol. 2014;24(9):666–672, 672. e1–e2. | |
Onukwugha E, Yong C, Hussain A, Seal B, Mullins CD. Concordance between administrative claims and registry data for identifying metastasis to the bone: an exploratory analysis in prostate cancer. BMC Med Res Methodol. 2014;14:1. | |
Hagberg KW, Taylor A, Hernandez RK, Jick S. Incidence of bone metastases in breast cancer patients in the United Kingdom: results of a multi-database linkage study using the general practice research database. Cancer Epidemiol. 2013;37(3):240–246. | |
Jensen AO, Norgaard M, Yong M, Fryzek JP, Sorensen HT. Validity of the recorded International Classification of Diseases, 10th edition diagnoses codes of bone metastases and skeletal-related events in breast and prostate cancer patients in the Danish National Registry of Patients. Clin Epidemiol. 2009;1:101–108. | |
Ehrenstein V, Hernandez RK, Maegbaek ML, et al. Validation of algorithms to detect distant metastases in men with prostate cancer using routine registry data in Denmark. Clin Epidemiol. 2015;7:259–265. | |
Howlader N, Chen VW, Ries LA, et al. Overview of breast cancer collaborative stage data items – their definitions, quality, usage, and clinical implications: a review of SEER data for 2004–2010. Cancer. 2014;120(Suppl 23):3771–3780. | |
XGEVA® (denosumab) [package insert]. Thousand Oaks, CA: Amgen Inc.; 2013. | |
Scheid V, Buzdar AU, Smith TL, Hortobagyi GN. Clinical course of breast cancer patients with osseous metastasis treated with combination chemotherapy. Cancer. 1986;58(12):2589–2593. | |
Early Breast Cancer Trialists’ Collaborative Group. Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: an overview of the randomised trials. Lancet. 2005;365(9472):1687–1717. | |
Harries M, Taylor A, Holmberg L, et al. Incidence of bone metastases and survival after a diagnosis of bone metastases in breast cancer patients. Cancer Epidemiol. 2014;38(4):427–434. | |
Rothman K, Greenland S, Lash T. Modern Epidemiology. 3rd ed. Philadelphia, PA: Lippincott Williams & Wilkins; 2008. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.