Back to Journals » Clinical Interventions in Aging » Volume 15
Using the Systems Biology Approach and Molecular Method to Investigate the Role of the Dopaminergic Pathway in Osteoarthritis: A Case Control Study
Authors Sheikhpour M , Eliaspour D , Arabi I , Raeissadat SA, Lari A , Seif Barghi T
Received 12 November 2019
Accepted for publication 30 January 2020
Published 4 March 2020 Volume 2020:15 Pages 321—327
DOI https://doi.org/10.2147/CIA.S238351
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Richard Walker
Mojgan Sheikhpour,1,2 Darioush Eliaspour,3 Iraj Arabi,3 Seyed Ahmad Raeissadat,3 Arezou Lari,4 Tohid Seif Barghi5
1Department of Mycobacteriology and Pulmonary Research, Pasteur Institute of Iran, Tehran, Iran; 2Microbiology Research Center (MRC), Pasteur Institute of Iran, Tehran, Iran; 3Shahid Beheshti University of Medical Sciences, Department of Physical Medicine and Rehabilitation, Tehran, Iran; 4Biomedicine Unit, Pasteur Institute of Iran, Tehran, Iran; 5Department of Sports and Exercise Medicine, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran
Correspondence: Mojgan Sheikhpour; Darioush Eliaspour
Tel +98 9122969712; +98 9125707896
Fax +98 21 64112313
Email [email protected]; [email protected]
Objective: Osteoarthritis is the most common type of arthritis and one of the leading causes of job loss and motor disabilities. Recently, the involvement of dopaminergic pathways and dopamine receptor genes has been considered in this disease. Therefore, studying and comparing the expression pattern of these receptor genes can lead to a greater understanding of the pathogenesis of this disease.
Methods: In this research, we used the systems biology approach to investigate the role of the dopaminergic pathway in osteoarthritis. Then the gene expression pattern of dopamine receptor genes was examined in an osteoarthritis patientgroup in comparison with healthy individuals by Real-time PCR method.
Results: The analysis of the transcriptome dataset of osteoarthritis identified some genes in the dopaminergic pathway and the six most important genes in this disease are in the network with a significant relationship to dopamine receptors which differentially expressed compared to health groups. Statistical analysis of the case control study showed a significant difference (P-value< 0.05) in DRD1 and DRD2 family in the patients in comparison to healthy individuals.
Discussion: We attained the significant expression pattern of dopamine receptors in the blood of osteoarthritis patients which could be useful to identify new strategies for the diagnosis, management, or treatment of this disease.
Keywords: osteoarthritis, systems biology, dopamine receptor genes, gene expression
Introduction
Osteoarthritis (OA) is the most common type of arthritis. It is one of the leading causes of job loss and motor disabilities. It is usually the target of the joints of the hands, thighs, knees, legs, and lumbar spine. Sometimes the disease attacks the wrist, elbows, and shoulders. Even if the condition does not cause joint malformations such as rheumatoid arthritis, it can significantly effect on daily activities. OA affects the cartilage and it causes wears, and tear, and in most cases, can destroy the cartilage and place the bone on the bone. It is also a multifactorial, aging-related, although not an inflammatory disease. Inflammation is a secondary complication, and this condition can increase the severity of the disease, especially in the tissue surrounding the joint.1
OA is a prevalent disease that can be found in all geographical areas.2 The exact cause of this disorder is unknown and appears to be the result of the combination or interaction of mechanical and other factors such as inflammatory and hereditary.3 Recently, the role of neurological factors, such as the involvement of neurotransmitter pathways, has been considered in the investigation of diagnostic and therapeutic approaches to this disease. Studies, in particular, have shown that the best way to treat and control of this disease in the long term is to use anti-inflammatory drugs. Studies have also shown that increased expression of dopamine receptor genes in diseases such as rheumatoid arthritis and OA to varying degrees can activate inflammatory pathways and increase the secretion of factors such as interleukins.4
On the other hand, the nervous and immune systems are intimately connected. The degree of interconnection between these two systems is such that they are sometimes referred to as the super-body system. The effects of these two systems on each other and the effects of genetic and environmental factors on the immune system cause many autoimmune diseases. The interactions between the nervous system and the immune system in humans involve a variety of message signaling molecules, including cytokines and neurotransmitters.5,6
Dopamine and serotonin are two important agents in the human nervous system. Recent studies have suggested that neurotransmitters, including dopamine and serotonin, act as mediators of the nervous and immune systems. Their receptors are present on lymphocytes and can play a role in causing pathologic changes.7 The dopaminergic system consists of dopamine receptors, which are the type of metabotropic G protein-coupled receptors that are expressed in the central nervous system and are normally stimulated by the dopamine ligand. These receptors mediate neuroendocrine systems. Dopamine binds to receptors attached to G-protein. Dopamine receptors include five subgroups D1, D2, D3, D4, D5, which are D5 and D1 receptors of the D1-like family that bind to Gs receptors and D4, D2, D3 receptors of the D2-like family that is connected to Gi/G0 receptors. There is some evidence suggesting the possibility of D7 and D6 dopamine receptors, but these receptors have not yet explicitly been identified.8 On the other hand, the coordinates of central pain processing pathways that are involved in OA pain processing have also been studied, indicating that the extent of pain in affected individuals may vary based on existing gene polymorphisms.9,10 Therefore, studying and comparing the expression pattern of these receptor genes on human peripheral blood leukocytes can lead to a greater understanding of the pathogenesis of such diseases. Indeed, the results could be provided effective solutions for diagnosis, treatment, management, and even the prognosis of this disease.
So, in this research, the analysis of the transcriptome dataset of OA identified some genes in the dopaminergic pathway, which differentially expressed compared to health groups. A protein-protein interaction (PPI) network of interest DEGs was constructed, and the most important genes, which are dopamine receptor genes was confirmed by Real-Time PCR. Given the rapid and profound advances in researches in disease genetics and the new approach to personalized medicine, in this study, we attempted to attain the expression pattern of dopamine receptors in the blood of two groups of healthy and OA patients.
Materials and Methods
Microarray Data Analysis and Functional Enrichment
The publicly available microarray data set GSE48556 was obtained from GEO. Following data extraction, all analyses were undertaken using the R (version 3.5.2). Differentially expressed genes (DEG) were identified using empirical Bayes methods implemented into the limma package from Bio Conductor.11 A probe set is considered differentially expressed if the FDR-adjusted P-value <5%.12
The Kyoto Encyclopedia of Genes and Genomes (KEGG; http://www.kegg.jp/) is a database resource for functional interpretation and annotation of enriched pathways using large-scale gene datasets.4
PPI Network
The Search Tool for the Retrieval of Interacting Genes (STRING; http://www.string-db.org/) database is an online database containing direct and indirect associations of proteins.13 The DEGs of the dopaminergic pathway and dopaminergic receptors genes were mapped to STRING to assess the interactions. Next, PPI networks were established using Cityscape software (version 3.6.1; http://www.cytoscape.org/), a free software package for visualizing, modeling, and analyzing the integration of bimolecular interaction networks with high-throughput expression data.14
Sampling
This research was done based on a research plan approved by the Medical Ethics Committee of Shahid Beheshti University of Medical Sciences with No. 19084 and with code of ethics: IR.SBMU.MSP.REC.1398. 500. The study population was 30 Osteoarthritis patients with no history of neuropsychiatric, endocrine, malignant, immune and infectious diseases, and 30 healthy individuals. Written informed consent conducted in accordance with the Declaration of Helsinki was provided by patients. 5mL of blood samples were collected from all the subjects and were collected in tubes containing 1.5 mM EDTA.
Molecular Studies and Statistical Analysis
Peripheral blood mononuclear cells (PBMC) were isolated by Ficoll-Hypaque density gradient centrifugation (Pharmacia, Uppsala, Sweden). The RNA was extracted from PBMCs using a Roche company RNA extraction kit (Germany). Then the Fermentase kit and Reverse Transcriptase PCR method were used to synthesis cDNAs. Finally, The Real-time PCR for the studied genes (DRD1-DRD5) and beta-actin as housekeeping genes were done by the designed specific primers, which have been shown in Table 1. The data were analyzed in SPSS software. An independent sample T-test was used, and statistical significance was defined as P-value < 0.05.
 |
Table 1 Information of Dopamine and Beta-Actin Primers |
Results
Microarray Data Analysis and Functional Enrichment
KEGG pathway enrichment analyses were conducted for differentially expressed genes. Eleven genes from the dopaminergic synapse pathway exist in our DEGs (Figure 1 and Table 2). These genes were ATF4, CREB1, GSK3B, GNG11, GNG8, GNAQ, CLOCK, PPP1R1B, GNAL, PRKCG, and GNB1 related to OA [Table 1].
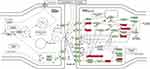 |
Figure 1 Result of microarray data analysis and functional enrichment shows important genes which involve in dopaminergic synapse pathway related to OA. |
 |
Table 2 Description and Adjusted p-value, of Important Genes Which Involve in Dopaminergic Synapse Pathway Related to OA |
PPI Network
It could be observed that the genes linked directly with dopaminergic receptors genes in OA disease [Figure 2, the border node shows fold change]. The expression of the six most important genes in OA disease are in the network with a significant relationship to Dopamine receptors and are shown in Figure 3. In all of them, the gene expression is significantly decreased in OA patients group compared to healthy individuals.
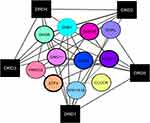 |
Figure 2 The network of the genes which linked directly with dopaminergic receptors genes in OA disease. The borders’ thicknesses of circles nodes indicate absolute LogFc. |
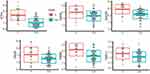 |
Figure 3 The expression of the 6 most interested genes in OA disease (GSK3B, GNAQ, ATF4, GNG11, CREB1 and GNG8) which are in the network with significant relationship to Dopamine receptors. |
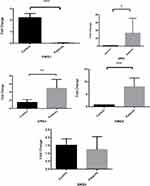
Gene Expression Analysis
Statistical analysis using SPSS software showed a significant difference (P-value<0.05) in DRD1, DRD2 family in OA patients with comparison to healthy individuals. The expression of DRD1 receptor genes (DRD1 and DRD5) expression in the patient group were lower than those in the healthy controls, whiles DRD2 receptor genes (DRD2, DRD3 and DRD4) expression were higher (Figure 4).
Discussion
The existence of common signaling pathways in the cells of the central and peripheral nervous and immune systems, especially through neurotransmitters such as dopamine, serotonin, and related receptors, suggests that these systems are linked to health and disease. Recently, new evidence has emerged of their mediating roles in the immune system in response to infections, inflammation, as well as the control of proliferation and migration regulation of cancer cells and leukocytes.15–17 Research has so far shown that dopamine and serotonin receptors are also present in lymphocytes, macrophages, and neutrophils. These neurotransmitters deliver antigens, activate T lymphocytes, and regulate other cellular activities.18,19 Disruption of any of these actions may lead to misleading immune responses and adverse reactions.7,20,21
Previous studies by researchers have shown that the expression profiles of dopamine receptor genes in people with various diseases, including cancers, diabetes, infectious diseases, and immunity, are different compared to healthy individuals. This pattern of expression is not the same in all of the relevant receptor genes and returns to its original state if treated definitively.22,23 Other researchers reported that generate expression profiles of vulnerable neuronal populations in several neurodegenerative diseases, including Alzheimer’s disease, gene expression profiles of diseased neuronal populations may reveal mechanistic clues to the molecular pathogenesis underlying various diseases such as Parkinson’s disease, schizophrenia, multiple sclerosis, and Creutzfeld–Jakob disease. And aid in identifying potential therapeutic targets.24,25 Recently mediated roles of these receptors in the immune system to respond to infections, control proliferation, and regulate cancer cell migration and diseases such as rheumatoid arthritis (RA) have been identified.25 More recently, studies showed that all dopamine receptors are expressed in synovial fibroblasts isolated from patients with RA).4 Dopamine, by increasing the secretion of pro-inflammatory cytokines such as Il6, has an additive effect on IL17 secretion and subsequently increases inflammation in rheumatoid arthritis. By immunohistochemistry, it has been shown that increased expression of dopamine receptors is an important factor in cartilage injury in this disease.26 Likewise, comparative studies have been performed in OA and RA, and the role of dopamine receptors in the inflammatory pathways leading to inflammation surrounding the osteoarthritic site has been confirmed.4
On the other hand, the coordinates of central pain processing pathways involved in OA pain processing have also been studied, indicating that the degree of pain in affected individuals may vary based on existing gene polymorphisms.27,28
Previous studies by researchers have shown that the expression profiles of dopamine receptor genes in people with various diseases, including cancers, infectious diseases and immunity, are different compared to healthy individuals.29 This expression pattern is not the same in all of the relevant receptor genes and returns to the former state if treated definitively.9,10 Recently mediated roles of these receptors in the immune system to respond to infections, control proliferation, and regulate cancer cell migration and diseases such as RA have been identified. To date, there have been numerous studies on various aspects such as expression pattern determination; different genes have been linked to OA. There have also been scattered studies by using the system biology of this disease, but so far there is no report of a significant association of dopamine receptors with the major genes associated with OA.30,31 Given the rapid and profound advances in research in the field of disease genetics and the new approach to personalized medicine in this study, we attempted to attain the expression pattern of dopamine receptors in the blood of healthy and diseased groups of OA patients using Real Time PCR method. This research may be useful to identify new strategies for the diagnosis, management, or treatment of this disease based on the results. The dopaminergic pathway specifically, dopamine receptors gene expression pattern can lead the clinicians for screening and design of better treatment for success in therapy. Also based on the validation of the bioinformatics studies by the experimental results of this research, dopamine receptors could be used as biomarkers for follow up the therapy of Osteoarthritis before and after treatment.
Acknowledgments
We thank all of the patients and healthy individuals who contributed in this research and helped us by donating their blood samples. Also, we thank our colleagues from the Department of Mycobacteriology and Pulmonary Research, Microbiology Research Center, Pasteur Institute of Iran. The authors acknowledge the directors of the Imam Khomeini and Shohaha Tajrish hospitals for the arrangement of the sampling process.
Author Contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors declare that they have no conflicts of interest regarding this article and no funding sources.
References
1. Dyer J, Davison G, Marcora SM, et al. Effect of a Mediterranean type diet on inflammatory and cartilage degradation biomarkers in patients with Osteoarthritis. J Nutr Health Aging. 2017;21(5):562–566. doi:10.1007/s12603-016-0806-y
2. Bruyere O, Collette J, Kothari M, et al. Osteoarthritis, magnetic resonance imaging, and biochemical markers: a one year prospective study. Ann Rheum Dis. 2006;65(8):1050–1054. doi:10.1136/ard.2005.045914
3. Rousseau JC, Garnero P. Biological markers in Osteoarthritis. Bone. 2012;51(2):265–277. doi:10.1016/j.bone.2012.04.001
4. Capellino S, Cosentino M, Luini A, et al. Increased expression of dopamine receptors in synovial fibroblasts from patients with rheumatoid arthritis: inhibitory effects of dopamine on Interleukin‐8 and Interleukin‐6. Arthritis Rheumatol. 2014;66(10):2685–2693. doi:10.1002/art.v66.10
5. Kordon C, Bihoreau C. Integrated communication between the nervous, endocrine and immune systems. Horm Res Paediatr. 1989;31(1–2):100–104. doi:10.1159/000181096
6. Ziemssen T, Kern S. Psychoneuroimmunology–cross-talk between the immune and nervous systems. J Neurol. 2007;254(2):II8–II11. doi:10.1007/s00415-007-2003-8
7. Entschladen F, Lang K, Drell T, et al. Neurotransmitters are regulators for the migration of tumor cells and leukocytes. Cancer Immunol Immunother. 2002;51(9):467–482. doi:10.1007/s00262-002-0300-8
8. Vallone D, Picetti R, Borrelli E. Structure and function of dopamine receptors. Neurosc Biobehav Rev. 2000;24(1):125–132. doi:10.1016/S0149-7634(99)00063-9
9. Sofat N, Ejindu V, Kiely P. What makes Osteoarthritis painful? The evidence for local and central pain processing. Rheumatology. 2011;50(12):2157–2165. doi:10.1093/rheumatology/ker283
10. Comings DE, MacMurray JP. Molecular heterosis: a review. Mol Genet Metab. 2000;71(1–2):19–31. doi:10.1006/mgme.2000.3015
11. Fernandez SV, Huang Y, Snider KE, et al. Expression and DNA methylation changes in human breast epithelial cells after bisphenol an exposure. Int J Oncol. 2012;41(1):369–377. doi:10.3892/ijo.2012.1444
12. Turcan S, Rohle D, Goenka A, et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature. 2012;483(7390):479. doi:10.1038/nature10866
13. Bai B, Xie B, Pan Z, et al. Identification of candidate genes and long non-coding RNAs associated with the effect of ATP5J in colorectal cancer. Int J Oncol. 2018;52(4):1129–1138. doi:10.3892/ijo.2018.4281
14. Zhang L, Guo XQ, Chu JF, et al. Potential hippocampal genes and pathways involved in Alzheimer’s disease: a bioinformatic analysis. Genet Mol Res. 2015;14(2):7218–7232. doi:10.4238/2015.June.29.15
15. Cloëz-Tayarani I, Changeux J-P. Nicotine and serotonin in immune regulation and inflammatory processes: a perspective. J Leukoc Biol. 2007;81(3):599–606. doi:10.1189/jlb.0906544
16. Tóth BE, Vecsernyés M, Zelles T, et al. Role of peripheral and brain-derived dopamine (DA) in immune regulation. Adv Neuroimmune Biol. 2012;3(2):111–155. doi:10.3233/NIB-2012-012044
17. Arreola R, et al. Immunomodulatory effects mediated by serotonin. J Immunol Res. 2015;2015.
18. Levite M. Nervous immunity: neurotransmitters, extracellular K+ and T-cell function. Trends Immunol. 2001;22(1):2–5. doi:10.1016/S1471-4906(00)01799-3
19. Levite M. Neurotransmitters activate T-cells and elicit crucial functions via neurotransmitter receptors. Curr Opin Pharmacol. 2008;8(4):460–471. doi:10.1016/j.coph.2008.05.001
20. Bianchi G, Molfetta L, Saggini R. Italian Survey on the Use of Anti-Inflammatory Drugs in Osteoarthritis. London, England: SAGE Publications Sage UK; 2014.
21. Herrington C, Hall P. Molecular and cellular themes in inflammation and immunology. J Pathol. 2008;214(2):123–125. doi:10.1002/path.v214:2
22. Shaikhpoor M, Ahangari G, Sadeghizadeh M, et al. Significant changes in D2-like dopamine gene receptors expression associated with non-small-cell lung cancer: could it be of potential use in the design of future therapeutic strategies? Curr Cancer Ther Rev. 2012;8(4):304–310.
23. Sheikhpour M, Ahangari G, Sadeghizadeh M, et al. A novel report of apoptosis in human lung carcinoma cells using selective agonist of D2-like dopamine receptors: a new approach for the treatment of human non-small cell lung cancer. Int J Immunopathol Pharmacol. 2013;26(2):393–402. doi:10.1177/039463201302600212
24. Manoel-Caetano FS, Xavier DJ, Evangelista AF, et al. Gene expression profiles displayed by peripheral blood mononuclear cells from patients with type 2 diabetes mellitus focusing on biological processes implicated on the pathogenesis of the disease. Gene. 2012;511(2):151–160. doi:10.1016/j.gene.2012.09.090
25. Mufson EJ, Counts SE, Che S, Ginsberg SD. Neuronal gene expression profiling: uncovering the molecular biology of neurodegenerative disease. Prog Brain Res. 2006;158:197–222.
26. Nakano K, Yamaoka K, Hanami K, et al. Dopamine induces IL-6–Dependent IL-17 production via D1-like receptor on CD4 Naive T cells and D1-like receptor antagonist SCH-23390 inhibits cartilage destruction in a human rheumatoid arthritis/SCID mouse chimera model. J Immunol. 2011;186(6):3745–3752. doi:10.4049/jimmunol.1002475
27. Phillips K, Clauw DJ. Central pain mechanisms in the rheumatic diseases: future directions. Arthritis Rheumatism. 2013;65(2):291–302. doi:10.1002/art.37739
28. Gracely RH, Petzke F, Wolf JM, et al. Functional magnetic resonance imaging evidence of augmented pain processing in fibromyalgia. Arthritis Rheumatism. 2002;46(5):1333–1343. doi:10.1002/(ISSN)1529-0131
29. Li S, Huang S, Peng S-B. Overexpression of G protein-coupled receptors in cancer cells: involvement in tumor progression. Int J Oncol. 2005;27(5):1329–1338.
30. Rao Z, Wang S, Wang J. Exploring the osteoarthritis-related genes by gene expression analysis. Eur Rev Med Pharmacol Sci. 2014;18(20):3056–3062.
31. Ma C, Lv Q, Cao Y, et al. Genes relevant with Osteoarthritis by comparison gene expression profiles of synovial membrane of osteoarthritis patients at different stages. Eur Rev Med Pharmacol Sci. 2014;18(3):431–439.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

