Back to Journals » Cancer Management and Research » Volume 13
The Long Noncoding RNA MAGI1-IT1 Regulates the miR-302d-3p/IGF1 Axis to Control Gastric Cancer Cell Proliferation
Authors Wang Q, Gu M, Zhuang Y, Chen J
Received 3 February 2021
Accepted for publication 17 March 2021
Published 1 April 2021 Volume 2021:13 Pages 2959—2967
DOI https://doi.org/10.2147/CMAR.S305108
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Xueqiong Zhu
Qinge Wang,1 Min Gu,2 Yun Zhuang,1 Jianping Chen1
1Department of Digestive Disease, The First People’s Hospital of Changzhou, The Third Affiliated Hospital of Soochow University, Changzhou, Jiangsu, People’s Republic of China; 2Department of Pediatrics, Changzhou Children’s Hospital, Changzhou, Jiangsu, People’s Republic of China
Correspondence: Jianping Chen
Department of Digestive Disease, The First People’s Hospital of Changzhou, The Third Affiliated Hospital of Soochow University, Juqian Road, Tianning District, Changzhou, Jiangsu, 213003, People’s Republic of China
Email [email protected]
Background: MAGI1-IT1 is a long non-coding RNA (lncRNA) previously reported to regulate several cancer types, but its functional role in gastric cancer (GC) remains to be defined. This study therefore explored the mechanistic role played by MAGI1-IT1 in the regulation of GC cell proliferation.
Methods: 120 pairs of GC patient tumor, paracancerous tissues, human GES-1 control cells and human AGS, MKN-74, MKN-45, and MGC-803 GC cell lines were used to detected MAGI1-IT1, miR-302d-3p, and IGF1 expression by a qPCR approach. An shRNA approach was used to knock down MGI1-IT1 in order to examine the effect of such treatment on GC cell proliferation, and rescue experiments were subsequently conducted. In addition, the functional role of MAGI1-IT1 in GC in vivo was evaluated with a xenograft model system. P < 0.05 was the significance threshold.
Results: Elevated MAGI1-IT1 expression was detected in GC cell lines and tissues, and was linked to poorer patient overall survival. Knocking down this lncRNA disrupted GC cell proliferation in vitro and in vivo, and miR-302d-3p was identified as a MAGI1-IT1 target. Notably, miR-302d-3p inhibition partially reversed the impact of MAGI1-IT1 knockdown on GC cell proliferation. IGF1 was subsequently identified as a miR-302d-3p target gene that was upregulated by MAGI1-IT1 through miR-302d-3p.
Conclusion: Overall, these results indicated that MAGI1-IT1 controlled GC cell proliferation by modulating the miR-302d-3p/IGF1 axis, suggesting that this may be a viable treatment target in those with GC.
Keywords: MAGI1-IT1, cell proliferation, gastric cancer, miR-302d-3p, IGF1
Background
Gastric cancer (GC) is a leading cause of cancer-related death globally.1 While many diagnostic and therapeutic advances for this cancer type have been made in recent years,2,3 those with advanced disease still have a poor prognosis, underscoring the need for the identification of novel therapeutic targets that can be leveraged to better treat this deadly disease.
Long noncoding RNAs (lncRNAs) are RNAs that are > 200 nucleotides long and which exhibit prognostic or diagnostic utility in many oncogenic contexts.4 Several lncRNAs have been shown to be dysregulated in GC, such as FAM230B, which controls the miR-27a-5p/TOP2A axis to drive GC cell metastasis and proliferation.5 Similarly, AC093818.1 is a lncRNA that can induce PDK1 expression via epigenetic mechanisms, thereby enhancing GC metastasis.6 BRD4 can control the expression of the lncRNA MAGI2-AS3, which in turn sequesters miR-141/200a and thereby induces ZEB1 overexpression to drive GC progression.7 MAGI1-IT1 is a lncRNA that has been found to promote ovarian and non-small-cell lung cancer development,8,9 but its role in GC has yet to be clarified.
Many lncRNAs regulate tumor cells by serving as competing endogenous RNAs (ceRNAs) for particular miRNAs.10–12 In cervical squamous cell carcinoma, miR-302d-3p has previously been shown to suppress tumor cell growth,13 while IGF1 is an important glioma progression inhibitor.14 Predictive analyses conducted with the TargetScan database suggested that both MAGI1-IT1 and IGF1 shared sequence complementarity with miR-302d-3p, suggesting that a MAGI1-IT1/miR-302d-3p/IGF1 ceRNA regulatory axis may control tumor cell proliferation. As such, we herein explored MAGI1-IT1 and its interactions with IGF1 and miR-302d-3p as potential regulators of GC cell proliferation through both in vitro and in vivo experiments.
Methods
Clinical Samples and Cell Lines
In total, 120 pairs of GC patient tumor and paracancerous tissues were obtained from The Third Affiliated Hospital of Soochow University following patient diagnosis. Patients participated in this study never received any other therapeutic method but surgery. All patients provided written informed consent, and the Ethics Committee of The Third Affiliated Hospital of Soochow University (No.00019675) approved all human studies detailed herein. The research involving human subjects conform to the principles laid out in the Declaration of Helsinki.
Human GES-1 control cells and human AGS, MKN-74, MKN-45, and MGC-803 GC cell lines were obtained from Bena Culture Collection (Beijing, China) and were grown in DMEM (Sigma, MO, USA) supplemented with 10% fetal bovine serum at 37 °C in a 5% CO2 incubator.
Transfection
A MAGI1‐IT1-specific shRNA was purchased from GenePharma (Shanghai, China), while pcDNA3.1-IGF1 was generated via the sub-cloning of full-length IGF1 cDNA into the pcDNA3.1 vector (Invitrogen), with pcDNA3.1-Ctrl and sh-Ctrl serving as corresponding negative control constructs. Both miR-302d-3p mimics and inhibitors were obtained from RiboBio (Guangzhou, China), along with corresponding scrambled miR-Ctrl constructs. These constructs were then transfected into AGS or MGC-803 cells with Lipofectamine 2000 (Invitrogen) based on provided directions, and cells were used for downstream analyses at 48 h post-transfection.
qPCR
TRIzol (Invitrogen) was used to extract sample RNA, after which a PrimeScript™ strand cDNA synthesis kit (Takara, Dalian, China) was used based on provided directions. All qPCR reactions were performed using an ABI 7900 instrument (Thermo Fisher Scientific, MA, USA) and the Power SYBR Green PCR Master Mix (Thermo Fisher Scientific). GAPDH and U6 served as a normalization control, and primers were as follows: MAGI1‐IT1, F: 5′‐GCAGTCCCCCTTTGTCTGAA‐3′ and R: 5′‐CTGTCCGTGGGTAAAGTGCT‐3′; IGF1, F: 5′‐CATGCCTGCTCAGAAGGGTA‐3′ and R: 5′‐GCCTCTGATCCTTGAGGTGA‐3′; GAPDH, F: 5′-CCATTTGCAGTGGCAAAG-3′ and R: 5′-CACCCCATTTGATGTTAGTG-3′; miR-302d-3p, F: 5’-TCTACTTTAACATGGAGGCACTT-3’ and R: 5’-TCACCAAAACATGGAAGCAC-3’; U6, F: 5’-CGCTTCGGCAGCACATATACTA-3’ and R: 5’-CGCTTCACGAATTTGCGTGTCA-3’.
CCK-8 Assay
Appropriately treated AGS or MGC-803 cells (3×103/well) were seeded in 96-well plates for 24, 48, or 72 h, after which 10µL of CCK-8 solution (KeyGen Biotech, Jiangsu, China) was added per well. Following a 4 h incubation at 37°C, the absorbance was measured by microplate reader (Bio-Rad Laboratories, Inc., Hercules, CA, USA) at a wavelength of 490 nm.
Luciferase Reporter Assay
MAGI1‐IT1 or IGF1 3’UTR sequences harboring putative wild-type (WT) or mutant (Mut) miR-302d-3p binding sites were cloned into the pmirGLO luciferase reporter construct (Promega, WI, USA), after which these reporters were co-transfected into 293T cells along with miR-302d-3p mimics or controls (miR-Ctrl) with Lipofectamine 2000 (Invitrogen) in a 24-well plate format. At 48 h post-transfection, a dual luciferase assay kit (Promega) was used to evaluate luciferase activity.
RNA Immunoprecipitation (RIP)
An Imprint RNA Immunoprecipitation Kit (Sigma, USA) was used for RIP assays based upon provided directions using anti-Ago2 (CST, Shanghai, China) and anti-IgG (CST).
ELISAs
ELISAs were conducted using commercial IGF1 ELISA kits from Abcam (USA) based on provided directions.
Xenograft Tumor Model
The Ethics Committee of The Third Affiliated Hospital of Soochow University approved all animal studies, which were consistent with the NIH Guidelines for Animal Use. In total, 12 male BALB/c nude mice (4 weeks old) from the Shanghai Laboratory Animal Center (SLAC, Shanghai, China were subcutaneously implanted with AGS (5x106 cell/mouse) or MGC-803 (5x106 cell/mouse) cells transfected with sh-MAGI1‐IT1 or sh-Ctrl. Tumor growth was monitored once per week, and survival outcomes were monitored.
Statistical Analysis
Date from three independent experiments are shown as mean ± standard deviation (SD). Student’s t-test and ANOVA were respectively conducted for the test of significant difference between two groups or more. The relationship between MAGI1-IT1 expression levels and the overall survival rates of GC patients was assessed by using Kaplan-Meier analysis with log rank test. The correlation significance was determined by Pearson correlation analysis. Statistical analyses were conducted using Prism 7 (GraphPad Software, Inc., CA, USA). P < 0.05 was the significance threshold.
Results
GC Cells and Tissues Exhibit MAGI1‐IT1 Upregulation
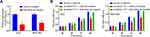
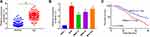
To understand the functional relevance of MAGI1-IT1 in GC, we began by clarifying its expression in GC patient samples and cell lines. GC tumor tissues exhibited significant MAGI1-IT1 upregulation relative to paracancerous tissues (Figure 1A). Similarly, MAGI1-IT1 upregulation was observed in the AGS, MKN-74, MKN-45, and MGC-803 tumor cell lines relative to GES-1 control cells (Figure 1B). Those GC patients with higher MAGI1-IT1 expression also exhibited poorer survival outcomes (Figure 1C), suggesting that MAGI1-IT1 may serve as an important regulator of GC progression.
Knocking Down MAGI1‐IT1 Impairs in vitro and in vivo GC Cell Proliferation
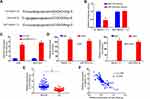
To explore how MAGI1-IT1 influenced GC progression, we next knocked down this lncRNA with a specific shRNA construct (Figure 2A). Subsequent CCK-8 assays indicated that MAGI1-IT1 knockdown impaired the proliferation of these cells (Figure 2B). To test these findings in vivo, we then implanted nude mice with ASG or MGC-803 cells stably transfected with sh-MAGI1‐IT1 or control constructs, and we found tumors in which MAGI1-IT1 had been knocked down grew significantly more slowly in these mice (Figure 2C). Additionally, mice in the sh-MAGI1‐IT1 group survived better than control mice (Figure 2D).
MAGI1‐IT1 Targets miR-302d-3p
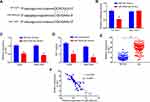
Starbase v2.0 was next employed to identify miR-302d-3p as a putative target for MAGI1-IT1 (Figure 3A). Luciferase reporter assays confirmed the ability of MAGI1-IT1 and miR-302d-3p to interact with one another (Figure 3B), and MAGI1-IT1 knockdown resulted in miR-302d-3p upregulation (Figure 3C). Anti-Ago2 RIP assays additionally confirmed MAGI1‐IT1 and miR-302d-3p enrichment in Ago2 complexes (Figure 3D). When we assessed miR-302d-3p expression in GC tissues, we determined that this miRNA was expressed at lower levels in GC patient tumor tissues relative to paracancerous tissues (Figure 3E). Pearson correlation analyses further confirmed MAGI1-IT1 and miR-302d-3p levels to be negatively correlated with one another in GC tissues (Figure 3F).
MAGI1‐IT1 Sequesters miR-302d-3p to Influence GC Cell Proliferation
To assess whether MAGI1-IT1 mediates its effects on GC cells by regulating miR-302d-3p, we next conducted a rescue experiment. We first used a miR-302d-3p inhibitor to knock down this miRNA (Figure 4A), after which we transfected GC cells with sh-MAGI1‐IT1 along with miR-302d-3p inhibitor or control constructs. A subsequent CCK-8 assay revealed that miR-302d-3p inhibition was sufficient to partially reverse the effects of sh-MAGI1‐IT1 on GC cell proliferation (Figure 4B).
IGF1 is a miR-302d-3p Targe Gene
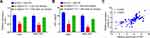
A TargetScan analysis next identified IGF1 as a putative miR-302d-3p target gene (Figure 5A), and this binding interaction was confirmed via luciferase reporter assay in 293T cells (Figure 5B). We then assessed IGF1 expression in GC cells following miR-302d-3p mimic transfection, revealing such transfection to suppress IGF1 levels as measured by qPCR and ELISA (Figure 5C and D). We also assessed IGF1 expression in patient tissues, revealing it to be expressed at significantly higher levels in tumors relative to paracancerous tissues (Figure 5E). Levels of IGF1 in GC tumors were also negatively correlated with miR-302d-3p levels (Figure 5F).
The Impact of miR-302d-3p on GC Cell Proliferation is Mediated by IGF1 Downregulation
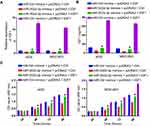
We next explored the influence of the miR-302d-3p/IGF1 axis on GC cell proliferation by transfecting GC cells with IGF1 overexpression and/or miR-302d-3p mimic constructs or corresponding controls, revealing that IGF1 overexpression was sufficient to rescue miR-302d-3p mimic-induced IGF1 downregulation, consistent with successful transfection (Figure 6A and B). Subsequent CCK-8 assays revealed that while miR-302d-3p mimics inhibited GC cell proliferation, the upregulation of IGF1 reversed this effect (Figure 6C), indicating that miR-302d-3p suppresses the proliferation of GC cells by suppressing IGF1 expression.
MAGI1‐IT1 Upregulates IGF1 Through a miR-302d-3p-Dependent Mechanism
Given that MAGI1-IT1 was able to sequester miR-302d-3p, which was in turn able to target IGF1, we next examined the ability of MAGI1-IT1 to indirectly influence GC cell IGF1 expression in a miR-302d-3p-dependent fashion. This analysis revealed that MAGI1-IT1 knockdown resulted in decreased IGF1 expression, while simultaneous miR-302d-3p inhibitor transfection reversed this IGF1 downregulation (Figure 7A and B). Consistent with this, Pearson correlation analyses suggested MAGI1-IT1 and IGF1 expression to be positively correlated in GC tissues (Figure 7C).
Discussion
Many analyses have highlighted the essential regulatory roles of lncRNAs in oncogenic contexts. For example, CASC7 is a lncRNA that suppresses miR-10a and thereby upregulates PTEN to suppress the proliferation of neuroblastoma cells,15 while LUNAR1 is a lncRNA that can modulate the miR4953p/MYCBP axis to promote colorectal cancer progression,16 and TUG1 can sequester miR-1294 to upregulate PLK1 and thereby drive esophageal cancer development.17 MAGI1‐IT1 has been found to play an oncogenic role in non-small-cell lung cancer wherein it serves as a ceRNA to upregulate AKT1,8 and in epithelial ovarian cancer wherein it can control the miR-200a/ZEB axis to promote tumor metastasis.9 Herein, we similarly observed MAGI1‐IT1 upregulation in GC cells and tissues, and found that knocking down this lncRNA was sufficient to suppress GC tumor growth in vitro and in vivo, consistent with its oncogenic role.
By serving as ceRNAs for specific miRNAs, many lncRNAs are able to indirectly influence mRNA expression within cells.18 Several lncRNAs are known to function via this mechanism in GC cells including PTCSC3, MALAT1, AC118344.1, LINC00689, and p4516.19–23 We similarly explore the function of MAGI1‐IT1 as a ceRNA and determined that it was predicted to bind to miR-302d-3p, which is a miRNA that has previously been shown to suppress cervical squamous cell carcinoma tumor growth.13 We confirmed the ability of MAGI1‐IT1 to sequester miR-302d-3p through both luciferase reporter and RIP assays, and we observed negative correlations between these two RNAs in GC patient tumor tissues. Importantly, miR-302d-3p inhibition was sufficient to reverse the impacts of MAGI1‐IT1 knockdown on GC cell proliferation, confirming that MAGI1‐IT1 regulates GC cell malignancy at least in part by serving as a ceRNA for miR-302d-3p.
The ability of miRNAs to suppress gene expression is linked to their binding of complementary 3’-UTR sequences in target mRNAs. We identified IGF1 as a putative miR-302d-3p target gene, which was noteworthy given that miR-422a has previously been shown to suppress the invasion and proliferation of glioma cells by suppressing IGF1 and IGF1R expression,14 and circRAD18 is known to sequester miR-208a/3164 and to thereby regulate the expression of FGF2 and IGF1 to promote the progression of triple-negative breast cancer.24 We confirmed the ability of miR-302d-3p to target IGF1, and observed a negative correlation between IGF1 and miR-302d-3p expression in GC tissues. Importantly, MAGI1‐IT1 was able to upregulate IGF1 in GC cells in a miR-302-3p-dependent manner, indicating that MAGI1‐IT1 controls the miR-302d-3p/IGF1 axis to influence GC progression. However, as lncRNAs can sequester multiple different miRNAs, more research is necessary to identify other MAGI1‐IT1 targets and downstream genes that may impact the role of this lncRNA in GC.
Conclusion
In summary, we determined that MAGI1‐IT1 expression was significantly increased in GC and was linked to poorer outcomes. From a functional perspective, knocking down this lncRNA impaired the proliferation of GC cells by disrupting the miR-302d-3p/IGF1 axis, indicating that this may be a viable target for therapeutic intervention.
Disclosure
The authors declare no conflicts of interest.
References
1. Van Cutsem E, Sagaert X, Topal B, Haustermans K, Prenen H. Gastric cancer. Lancet. 2016;388(10060):2654–2664. doi:10.1016/S0140-6736(16)30354-3
2. Necula L, Matei L, Dragu D, et al. Recent advances in gastric cancer early diagnosis. World J Gastroenterol. 2019;25(17):2029–2044. doi:10.3748/wjg.v25.i17.2029
3. Johnston FM, Beckman M. Updates on management of gastric cancer. Curr Oncol Rep. 2019;21(8):67.
4. Zeng MS. Noncoding RNAs in cancer diagnosis. Adv Exp Med Biol. 2016;927:391–427.
5. Cui Y, Pu R, Ye J, et al. LncRNA FAM230B promotes gastric cancer growth and metastasis by regulating the miR-27a-5p/TOP2A axis. Dig Dis Sci. 2020. doi:10.1007/s10620-020-06581-z
6. Ba MC, Ba Z, Long H, et al. LncRNA AC093818.1 accelerates gastric cancer metastasis by epigenetically promoting PDK1 expression. Cell Death Dis. 2020;11(1):64. doi:10.1038/s41419-020-2245-2
7. Li D, Wang J, Zhang M, et al. LncRNA MAGI2-AS3 is regulated by BRD4 and promotes gastric cancer progression via maintaining ZEB1 overexpression by sponging miR-141/200a. Mol Ther Nucleic Acids. 2020;19:109–123. doi:10.1016/j.omtn.2019.11.003
8. Zhang G, Chen HX, Yang SN, Zhao J. MAGI1-IT1 stimulates proliferation in non-small cell lung cancer by upregulating AKT1 as a ceRNA. Eur Rev Med Pharmacol Sci. 2020;24(2):691–698.
9. Gao H, Li X, Zhan G, et al. Long noncoding RNA MAGI1-IT1 promoted invasion and metastasis of epithelial ovarian cancer via the miR-200a/ZEB axis. Cell Cycle. 2019;18(12):1393–1406.
10. Yang B, Gu B, Zhang J, Xu L, Sun Y. CASC8 lncRNA promotes the proliferation of retinoblastoma cells through downregulating miR34a methylation. Cancer Manag Res. 2020;12:13461–13467. doi:10.2147/CMAR.S268380
11. Wang K, Zhao Y, Wang YM. LncRNA MALAT1 promotes survival of epithelial ovarian cancer cells by downregulating miR-145-5p. Cancer Manag Res. 2020;12:11359–11369. doi:10.2147/CMAR.S267355
12. Gong Z, Chen X, Zhang Y, et al. LncRNA GATA6-AS1 inhibits the progression of non-small cell lung cancer via repressing microRNA-543 to up-regulating RKIP. Cancer Manag Res. 2020;12:9327–9338. doi:10.2147/CMAR.S254184
13. Wang J, Chen S. RACK1 promotes miR-302b/c/d-3p expression and inhibits CCNO expression to induce cell apoptosis in cervical squamous cell carcinoma. Cancer Cell Int. 2020;20:385. doi:10.1186/s12935-020-01435-0
14. Wang H, Tang C, Na M, et al. miR-422a inhibits glioma proliferation and invasion by targeting IGF1 and IGF1R. Oncol Res. 2017;25(2):187–194. doi:10.3727/096504016X14732772150389
15. Zhou X, Lu H, Li F, et al. LncRNA cancer susceptibility candidate (CASC7) upregulates phosphatase and tensin homolog by downregulating miR-10a to inhibit neuroblastoma cell proliferation. Neuroreport. 2020;31(5):381–386. doi:10.1097/WNR.0000000000001411
16. Qian J, Garg A, Li F, Shen Q, Xiao K. lncRNA LUNAR1 accelerates colorectal cancer progression by targeting the miR4953p/MYCBP axis. Int J Oncol. 2020. doi:10.3892/ijo.2020.5128
17. Zong M, Feng W, Wan L, Yu X, Yu W. LncRNA TUG1 promotes esophageal cancer development through regulating PLK1 expression by sponging miR-1294. Biotechnol Lett. 2020;42(12):2537–2549. doi:10.1007/s10529-020-02984-0
18. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146(3):353–358. doi:10.1016/j.cell.2011.07.014
19. Dai Q, Zhang T, Li C. LncRNA MALAT1 regulates the cell proliferation and cisplatin resistance in gastric cancer via PI3K/AKT pathway. Cancer Manag Res. 2020;12:1929–1939.
20. Yin G, Tian P, BuHe A, Yan W, Li T, Sun Z. LncRNA LINC00689 promotes the progression of gastric cancer through upregulation of ADAM9 by sponging miR-526b-3p. Cancer Manag Res. 2020;12:4227–4239. doi:10.2147/CMAR.S231042
21. Cai L, Xue Y, Ding J, Zheng B. Long non-coding RNA AC118344.1 promotes gastric cancer cell proliferation, invasion, and metastasis via AKT2 and its downstream molecules HK2 and MMP2. Cancer Manag Res. 2020;12:12613–12621. doi:10.2147/CMAR.S281608
22. Hong L, Wang H, Wang J, et al. LncRNA PTCSC3 inhibits tumor growth and cancer cell stemness in gastric cancer by interacting with lncRNA linc-pint. Cancer Manag Res. 2019;11:10393–10399. doi:10.2147/CMAR.S231369
23. Nie ML, Han J, Huang HC, et al. The novel lncRNA p4516 acts as a prognostic biomarker promoting gastric cancer cell proliferation and metastasis. Cancer Manag Res. 2019;11:5375–5391. doi:10.2147/CMAR.S201793
24. Zou Y, Zheng S, Xiao W, et al. circRAD18 sponges miR-208a/3164 to promote triple-negative breast cancer progression through regulating IGF1 and FGF2 expression. Carcinogenesis. 2019;40(12):1469–1479. doi:10.1093/carcin/bgz071
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.