Back to Archived Journals » Research and Reports in Biology » Volume 6
The instability of the flax element LIS-1 in transgenic Arabidopsis thaliana
Received 27 November 2014
Accepted for publication 3 February 2015
Published 14 May 2015 Volume 2015:6 Pages 89—105
DOI https://doi.org/10.2147/RRB.S78195
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Professor Zvi Kelman
Nasmah K Bastaki, Christopher A Cullis
Department of Biology, Case Western Reserve University, Cleveland, OH, USA
Background: The LIS-1 is an element that appears as a site-specific insertion event in some flax lines in response to certain growth conditions and can be transmitted to subsequent generations. The origin of LIS-1 in the flax genome is uncertain. One possibility is that since LIS-1 does not exist intact in the progenitor line, it is assembled from small sequences found scattered throughout the genome, and that, under stressful growth conditions, induction occurs and these sequences are rearranged and assembled to form the intact LIS-1 element. It is unknown whether the intact LIS-1 element would remain stably integrated in other plant species or if it would be destabilized from their genome.
Results: In this study, Agrobacterium-mediated plant transformation via floral dipping was used to transform different accessions of the Columbia ecotype of Arabidopsis thaliana, with either LIS-1 or the target site into which LIS-1 integrates. The stability and the inheritance patterns of both elements were followed in subsequent generations. Our results indicate that, in the different transformed accessions, the target site of LIS-1 remains stable in the T1 and T2 generations. However, LIS-1 is not found intact in any transformed A. thaliana plants. Instead, it goes through multiple fragmentation events, which seem to be genotype dependent. In the process, the region originally flanking LIS-1 in the T-DNA construct can be converted to the same sequence found at the target site in flax, followed by complete excision of all the flax DNA in the construct.
Conclusion: These results demonstrate that the processes by which LIS-1 is produced in flax are also present in A. thaliana because both plants are capable of destabilizing the intact LIS-1 element.
Keywords: flax (Linum usitatissimum), Arabidopsis thaliana, plant transformation, Linum insertion sequence (LIS-1)
Background
Flax (Linum usitatissimum) is an important crop grown widely for its fibers and oils. Inbred lines of flax have been optimized for either fiber quality, so-called “fiber flax” lines, an example of which is the variety Stormont cirrus, or for oil content, so-called linseed “oil flax” lines such as the variety Bethune. A major difference between fiber flax and linseed varieties appears to be the ability to respond to nutrient stress with heritable genomic alternations; this feature has been observed to date only in some of the fiber flax lines and has not been observed in any linseed lines.1–7
The inbred flax variety, Stormont cirrus (denoted as plastic [Pl]), is the responsive parental line from which other lines (called genotrophs) have been derived by growth in different environments. Unlike the original Pl line, the genotrophs are considered stable and they breed true when grown in a number of different environments. Two extreme genotrophs that were derived for the Pl line, following its growth for one generation under different nutrient regimes, were originally classified as large and small (L and S) genotrophs, based on their weight and stability to further grow under a number of different growth environments.1,2,5,6,8–10
Besides these phenotypic changes, a number of genomic alterations were observed in the genotrophs following this single generation’s growth, which include a change in total DNA content; copy number variation of repetitive regions, such as the ribosomal RNA genes (rDNA) and satellite regions; and the appearance of a novel single copy insertion termed LIS-1.5–7,9
The LIS-1 is a 5.7 kb sequence that is inserted into a specific, single copy target sequence in the progenitor Pl genome (referred to as the “target site”) when grown under the inducing stress condition of low nutrients (Figure S1A). The progenitor target sequence is defined here as a conserved single copy region of 3,792 base pairs within the flax genome that can be present in both fiber and oil-seed varieties. The target sequence following LIS-1 insertion is divided into two sections, separated by the LIS-1 element, and these two regions combined are referred to as the flanking sequence. A comparison between the flanking sequence and the target sequence from the lines with and without the LIS-1 insertion has revealed 129 single nucleotide polymorphisms and indels that differ between these two regions. In addition to these differences, a TCC duplication is present at either end of the insertion site compared with only a single copy of this sequence in the target sequence. This duplication is the only characteristic of LIS-1 and its insertion that is reminiscent of other transposable elements. However, unlike transposable elements, there are neither direct or inverted repeats at the ends of LIS-1, nor any of the genes identified with LTR retrotransposons (such as the gag, pol, and env genes), indicating that LIS-1 is not a typical long terminal repeat transposable element.5–7
Once inserted, LIS-1 can become stabilized and inherited by subsequent generations (eg, in the S genotroph). The absence of the intact LIS-1 from the progenitor Pl and its presence in the S genotrophs has been demonstrated by Southern analysis, polymerase chain reaction (PCR)6,7,9 and whole genome sequencing, (Cullis, in prep). LIS-1 presence or absence can be easily demonstrated by PCR using primers for either the entire inserted sequence, the junctions of LIS-1 and the flanking sequence, or the target sequence without LIS-1 (Figure S1B).6,7
The origin of LIS-1 in the flax genome is uncertain since it is not present intact in the Pl genome, but its appearance has been followed during growth under inducing conditions.9 One possibility is that LIS-1 is assembled from small sequences found scattered throughout the genome. The conditions under which the plant can become induced (stress) result in these small sequences being rearranged and joined together to build the full LIS-1 sequence, which is then inserted into a defined “target site”. Moreover, LIS-1 appears to be the result of programmed DNA rearrangement since it has been reproducibly assembled in independent lines.6,9 Currently, the only comparable system for such a radical, controlled or programmed DNA rearrangement is seen in the ciliate macronucleus and the sea lamprey.11–13
Genomic reorganization has been studied in the ciliate macronucleus, where extensive genomic rearrangements and chromosome amplifications occur when micro nuclear DNA is altered in the formation of the somatic macronucleus. The ciliate macronucleus utilizes an scnRNA mechanism combined with dicer and AGO proteins to process 24–28 nucleotide transcripts and direct activity toward regions of complementarity. The target sequences are eliminated and the genome reassembled.11,12,14 Currently, the scnRNA mechanism is the only related system for the programmed DNA rearrangement consistent with the observations of LIS-1 insertion.
It is unknown whether or not other plants would have the capacity to maintain the intact LIS-1 element stably integrated in their genome, and subsequently transmit it through multiple generations. In an attempt to determine this, and to see if the functions that produce LIS-1 are also present in other plant species, different accessions of Arabidopsis thaliana were transformed with either LIS-1 or the progenitor “target site” of LIS-1. Since only some flax genotypes are Pl and respond to certain growth environments by LIS-1 insertion, it was necessary to screen multiple A. thaliana accessions to determine if the same situation exists in A. thaliana. Therefore, different accessions of A. thaliana Columbia ecotype, plus one additional A. thaliana line “known as the Quick line (from Ward Scientific)”, were used for the transformation process. The stability and inheritance patterns of LIS-1 and the target site were followed in the T1 and T2 generations in these transgenic plants.
The results show that when A. thaliana accessions were transformed with the target site of LIS-1, the target site remained stably integrated in the genome of all transformed plants through both the T1 and T2 generations. However, in every case tested where the Arabidopsis accessions were transformed with the LIS-1 element, LIS-1 was not found intact in either the T1 or T2 generations of any of the transformed A. thaliana plants. Moreover, the fragments of DNA still present within the T-DNA at earlier DNA collections were found to be identical to the progenitor sequence into which LIS-1 was inserted, ie, the “target site”, indicating that on excision of LIS-1 the remaining DNA sequences were converted to the progenitor target site. This was a temporary conversion as it was followed by complete excision of all flax sequences at later DNA collections. Additionally, during this rearrangement process, some sub-fragments of LIS-1 were found to remain stably integrated in the transformed A. thaliana genome, separate from the T-DNA fragment, while other sub-fragments were excised and deleted from the genome. The specific nature of the maintenance and/or deletion of LIS-1 sequences was, however, genotype dependent.
This study is the first to demonstrate that LIS-1 is not a stable element in another plant species. The results demonstrate that A. thaliana and flax have processes for accurately destabilizing an intact LIS-1 element, since the progenitor “target site” is created in A. thaliana accessions transformed with only LIS-1. This study provides an initial indication that an “inducing” growth environment is necessary for maintaining an intact LIS-1 element in the A. thaliana genome, mirroring what has previously been confirmed in flax.
Methods
Plant materials
Flax (L. usitatissimum)
The inbred flax (L. usitatissimum) variety Stormont cirrus (plastic line or Pl) and the genotroph (S) were grown at the greenhouse facilities at the Case Western Reserve University Squire Valleevue Farm. DNAs used as the sources for the LIS-1 and the target site of LIS-1 for the cloning reactions were extracted from leaves.
A. thaliana
Seven natural accessions of A. thaliana, Columbia ecotype (CS28166, CS22681, CS22625, CS76113, CS28167, CS28168, CS70000), seed stocks which were obtained from Arabidopsis Biological Resource Center (ABRC), Columbus, Ohio, plus one additional A. thaliana line “known as the Quick line” from Ward Scientific, were used in this study. Plants were grown to flowering in a growth room under long daylight (18 hour light–6 hour dark) at 22°C–24°C. After about 4 weeks post seed germination, primary inflorescences were clipped off to encourage the emergence of multiple secondary bolts. Plants were dipped when most secondary inflorescences were about 2–10 cm tall (2–4 days after clipping). In each floral dipping, two pots/per seed variety/per treatment were used and each pot contained 6–8 plants.
Preparing the constructs and the cells
DNA extraction
Five to six leaves from flax (Pl) and genotroph (S) were used for genomic DNA extraction. Leaves were put in 400 μL buffer AP1 from DNeasy Plant Minikit (Qiagen NV, Venlo, the Netherlands) with sterile sand and ground until no tissue clumps were visible. DNA extraction was then performed according to manufacturer’s instructions, eluted in 100 μL water, and stored at −20°C, until used for the PCR reaction.
PCR reactions to amplify LIS-1 and the target site of LIS-1
The LIS-1 fragment from the S genotroph was amplified using primers 2 and 19′ (Table S1, Figure S1B). The recognition sequence for the restriction enzyme “SalI” was added to the 5′ terminus of primer 2 and the recognition sequence for the restriction enzyme “SacI” was added to the 5′ terminus of primer 19′ (Table S1). PCR was carried out using speedSTAR DNA polymerase (Takara). The cycling conditions involved an initial hold of 24°C, then 2 minutes at 94°C, followed by 30 cycles of 98°C for 5 seconds, 60°C for 15 seconds, and 72°C for 2 minutes. A final extension step was carried out at 72°C for 5 minutes, followed by an indefinite hold at 4°C. The target site of LIS-1 from Stormont cirrus (Pl) was amplified using primers 2 and PL9, with the recognition sequence for the restriction enzyme “SalI” added to the 5′ terminus of 2 primer and the recognition sequence for the restriction enzyme “SacI” added to the 5′ terminus of PL9 primer (Table S1). PCR was carried out using GoTaq DNA polymerase (Promega Corporation, Fitchburg, WI, USA). The cycling conditions involved: an initial hold of 24°C, then 2 minutes at 95°C, followed by 35 cycles at 95°C for 30 seconds, 55°C for 30 seconds, and 72°C for 1 minute. A final extension step was carried out at 72°C for 5 minutes, followed by an indefinite hold at 4°C. PCR products were separated at 1% agarose gel, cut out from the gel and cleaned via Qiagen gel extraction kit in preparation for cloning.
TOPO TA cloning
LIS-1 was cloned into PCR 2.1-TOPO vector (Thermo Fisher Scientific, Waltham, MA, USA), and transformed into one Shot MAX Efficiency DH5α™-T1R Competent Cells according to manufacturer instructions (Thermo Fisher Scientific). Single colonies were purified using Qiagen miniprep, and the plasmid (TOPO+LIS1) was confirmed by PCR and sequencing.
PRI909 cloning and Escherichia coli transformation
Sequential restriction digests with SalI and SacI enzymes were done for the plasmid (TOPO+LIS-1) and for the plant binary vector PRI909 (Takara), ligation between LIS-1 and PRI909 vector was performed using DNA T4 ligase (New England Biolabs), followed by transformation into one Shot MAX Efficiency DH5α™-T1R Competent Cells. Single colonies were purified and the plasmid (PRI909+LIS1) was confirmed by PCR and sequencing. Target site of LIS-1 was cloned directly into the PRI909 vector using New England Biolabs ligation kit and transformed similarly into Shot MAX Efficiency DH5α™-T1R Competent Cells.
Electroporation into Agrobacterium tumefaciens LBA4404
One nanogram of plasmid (PRI909+LIS1 or PRI909+target site) was added to 20 μL Agrobacterium tumefaciens LBA4404 electro-cells (Takara). Electroporation was followed according to manufacturer instructions. The integrity of both constructs (PRI909+target site and PRI909+LIS-1) inside the Agrobacterium cells was confirmed by purifying plasmids from single colonies of Agrobacterium and testing them with different PCR primers (Figure S5) and by sequencing (Eurofins MWG operon).
A. thaliana transformation
Performed as explained in literature15–19 and summarized in the following steps
Culture of A. tumefaciens and floral dipping: A. tumefaciens LBA4404 strain, carrying either the plant binary vector PRI909+LIS1 or PRI909+target site, was used for the plant transformation. Bacteria were grown to stationary phase in LB +50 μg/mL kanamycin + 50 μg/mL streptomycin. Cultures were started from 1:100 dilutions of smaller overnight cultures and grown in a shaking incubator for 24–48 hours at 28°C–30°C at 150 rpm. The starting cultures for the transformation were checked to ensure that the constructs were intact (Figure S5) before the procedure of floral dipping was performed. Once confirmed, cells were collected by centrifuging at 5,000 × g. Cells were resuspended in infiltration medium (half strength Murashige and Skoog basal medium containing 5.0% sucrose, 0.05%–0.003% silwet L-77 and 44 nM benzylaminopurine). In some instances, a revised protocol was used in which the infiltration medium consisted of only 5% sucrose and silwet L-77.
For floral dipping, all aboveground tissues were submerged in the infiltration medium for 5–10 seconds and plants were placed on their sides and covered with plastic wrap to maintain high humidity in the dome. The next day, plants were placed in an upright position and maintained normally for an additional 3–5 weeks until their siliques became dry and their seeds could be collected.
Selection of T1 putative transformants using kanamycin
Seeds from the treated plant were surface sterilized by soaking in 70% ethanol, rinsing in sterile water, soaking in 5% bleach followed by rinsing 3–6 times in sterile water. Sterilized seeds were suspended in 0.05% sterile agarose, plated on kanamycin selection plates at a density of approximately 3,000–5,000 seeds per 150×15 mm2 plate, cold-treated for 2 days and then grown for 10 days in a growth cabinet under long daylight. Selection plates contained half Murashige and Skoog medium, 0.8% agar, and 50 mg/mL kanamycin. Controls include: growing T1 seeds without kanamycin and growing wild-type seeds on kanamycin. As an additional control, carbenicillin was added to kanamycin for plant selection. Carbenicillin inhibits growth of Agrobacterium cells but it does not have any effect on seed germination.
PCR amplifications of genomic DNA
T1 positive transformants were transplanted into moistened potting soil. Genomic DNA was extracted from their leaves and subjected to PCR amplification. Either GoTaq® Green Master Mix (Promega Corporation) or speedSTAR DNA polymerase (Takara) was used for the PCR and cycling conditions were provided as above. For specific details about primers and annealing temperature and extension time for each primer pair, refer to Table S1.
T2 generations were obtained by self-pollination of T1. For the segregation analysis, 50–100 seeds from each T2 transgenic plant were germinated on Murashige and Skoog media supplemented with 50 μg/mL kanamycin and the number of green and white plantlets were scored for each accession. Genomic DNAs were prepared from the leaves of the T2 generation and were subjected for PCR amplification (Table S1). Negative controls for PCR includes: PCR of wild-type (non-dipped) A. thaliana plants and PCR of no DNA template.
Results
The constructs for the transformation, the floral dipping, and selection procedures15–19 are described in the “Materials and methods” section. The results are presented in a logical order starting with the confirmation of transformation based on the presence of T-DNA components, then the fate of the target sequence in Arabidopsis followed by the fate of the LIS-1-containing construct.
Selection of T1 kanamycin resistant transformants
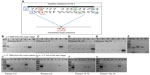
Selection of T1 putative transformants with 50 μg/mL kanamycin is shown in Figure 1. Seeds from wild-type A. thaliana (wild-type was defined as non-dipped control plants) confirming their sensitivity to kanamycin is shown in Figure 1B and b. Positive T1 transformants were selected based on their kanamycin resistance phenotypes: dark green cotyledon, true leaves, and in some cases their long roots that penetrate into the media (Figure 1C, c, D). Positive transformants were transplanted into soil, and the phenotypes of the plants from which leaves were collected at approximately 3 weeks post-transplanting to soil (Figure 1E, early stage), and at approximately 3 months post-transplanting to soil (Figure 1F, late stage) are shown.
Confirmation of T-DNA components in the transformed T1
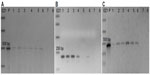
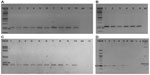
The amplification from genomic DNA extracted from the leaves of the positive T1 transformants was performed using different primers across the T-DNA to confirm its presence in the A. thaliana genome (Table S1 and Figure 2). The amplification with the NPTII (kanamycin) primers, (Figure 2A), and PriC primers, which amplified the region between the vector’s multiple cloning site and right border of T-DNA (Figure 2B), resulted in bands of the expected sizes. The DNAs extracted from transgenic plants transformed with the PRI909+LIS-1 construct were also amplified by PCR using PriMCSF+3′ primers, which amplifies the region between the vector’s multiple cloning site and the 3′ junction fragment of LIS-1 (Figure 2C), and gave rise to a product of expected size. Similarly, amplifications for DNAs from all T1 positive transformants resulting from the floral dip with Agrobacterium containing the construct with PRI909+target site, resulted in the expected sizes and sequence of all of these bands. None of these primers amplified regions from wild-type A. thaliana genome, which was confirmed both by bioinformatic analysis by blastn of the Arabidopsis genome and by PCR (Figure 2, sample 7 in all gels). These results confirm that T-DNA was integrated into the transgenic A. thaliana plants in the T1 generation.
Segregation analysis of T-DNA components in the T2 generations
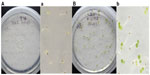
Segregation analysis of kanamycin transgene in T2 generations was performed by screening 50–100 seeds with 50 μg/mL kanamycin (Figure S2). T2s, from all transformed T1s (irrespective of the construct used), segregated in a normal Mendelian fashion (approximately three quarters contained the kanamycin transgene and appeared green and one quarter did not have any copy of kanamycin transgene and appeared white) (Figure S2 B, b). Control, wild-type A. thaliana screened with 50 μg/mL kanamycin, were all white and died within a few days (Figure S2 A, a).
Ten kanamycin resistant T2 plants from single T1 transformants for each accession were examined for the inheritance of the T-DNA components. In every case the T-DNA was inherited in the T2 generation as expected, for a single copy (locus) insertion, all T2 transgenic plants amplified the expected product sizes with primers NPTII and PriC (Figure S3); they all also amplified the small 150 bp fragment of the binary vector region with primers PriMCSF+R, similar to the pattern of PCR amplifications from the DNA from T1 generations at later growth stages (data not shown).
The stability of the target site in T1 and T2 generations
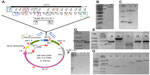
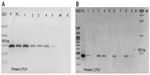
All of the positive T1 A. thaliana plants, which were transformed with PRI909+target site of LIS-1, have amplified the inserted target site of LIS-1 (product size of 478 bp) using primers 2 and PL9, as can be seen in Figure 3A (the location of these primers can be seen in Figure 4A, highlighted in red boxes). The bands were sequenced and confirmed to be an exact match to flax target site for LIS-1. Wild-type A. thaliana failed to amplify any band with PCR using these primers (Figure 3A, wild-type band).
To follow the inheritance of the target site in the T2 generation, PCR with the same primers (2 and PL9) was performed and the results show that the 478 bp product size of the target site was inherited as expected in the T2 generation (Figure 3B).
Therefore, the results of transformation of the target site of LIS-1 into A. thaliana confirm that the target site of LIS-1 was stably transformed and stability inherited through at least two growth generations in the different accessions of the transgenic A. thaliana plants.
The reorganization of LIS-1 in the T1 generations
For the A. thaliana accessions, which were transformed with LIS-1 only, none of the positive T1 A. thaliana plants have amplified the inserted intact LIS-1 fragment using primers 2 and 19′, as can be seen in Figure S4B.
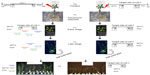
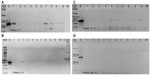
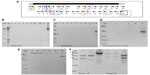
In an attempt to find whether sub-fragments of LIS-1 might still be present in the transformed A. thaliana genome, internal LIS-1 primers were used in PCR across the entire insertion sequence of LIS-1 and only four primer pairs (2 and 3′- 5 and 6′- 14 and 15′- 18a and 19′) were successful in amplifying sub-fragments of LIS-1 in the transformed A. thaliana plants (highlighted in the filled boxes in Figure 4A “top”). These bands were sequenced and confirmed to be an exact match to the sub-fragments of LIS-1. These primers did not amplify with wild-type A. thaliana and their sequences do not exist in wild-type A. thaliana genome database.
We have also observed genotype differences for the stability of these sub-fragments between early DNA collections and late DNA collections. In early DNA collections (Figure 4B–F), primers 2 and 3′ amplified from all T1 accessions. However, primers 5 and 6′ amplified from the following accessions, 22681, 28168, and 76113, yet did not amplify from accessions 28167 and 28168B. Primers 14 and 15′ amplified from all accessions, except 28167, while primers 18a and 19′ amplified from all accessions, except 28168. In Figure 4F, the line “QuickII” amplified with primers 2 and 3′, 18a and 19′, 5 and 6′ but not 14 and 15′.
However, at late DNA collections (Figure 4b–e), primers 2 and 3′, 14 and 15′ and primers 18a and 19′ amplified from 28168, 28167 and 22681 but not from the other accessions, while primers 5 and 6′ amplified from all accessions except 76113. So, at late DNA collections, it appears that the accession 76113 has lost all the sub-fragments of LIS-1. While in other accessions such as 28168 and 22681, these sub-fragments become present in more cells.
In an attempt to find the cut off borders from each of the previously amplified sub-fragments, other combinations of primers were used to amplify longer regions inside the LIS-1 (Figure S4). For example, since both primers 2 and 3′ and 5 and 6′ amplified, we tested primer pair 2 and 6′ to amplify the whole region between 2 and 6′; however, it did not amplify from any of the transformed accessions (Figure S4E). A similar strategy was used for other pairs of primers such as 13 and 16′ and 4 and 7′. None of those primer pairs amplified from any of the transformed accessions (Figure S4C, D). Few other primer pairs, such as primers 5 and 19′ and primers 8 and 12′, have amplified from the transformed accessions but gave different product sizes to those expected from the flax (S) positive control (Figure S4F); when they were sequenced, they had no match to LIS-1. These results indicate that the four LIS-1 sub-fragments (2 and 3′- 5 and 6′- 14 and 15′- 18a′ and 19′), present at the T1 transformed accessions, were not contiguous to each other.
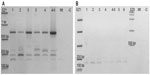
To determine what was left inside the multiple cloning site of the binary vector in these plants transformed with PRI909+LIS1, primers PriMCSF+R were used. This primer pair amplifies the cloned insert region inside the vector’s multiple cloning site. When PCR was performed from the construct PRI909+LIS1, the product size was approximately 6.3 kb, the expected size of the intact LIS-1 fragment (Figure S5f band 3). However, when early DNA stage from transformed T1 accessions was used, three different product sizes, starting with the 700 bp, then approximately 300–250 bp, and the smallest appeared to be at 150 bp (Figure 5A) were amplified. All of these bands were sequenced; the 700 bp bands, surprisingly, aligned perfectly to the target site of LIS-1 in all transformed accessions. The smaller bands aligned to the binary vector sequence. However, when late DNA stage was used, the 700 bp bands disappeared from all accessions, and only the small bands that aligned to the binary vector sequence were present (Figure 5B).
The overall results of LIS-1 transformation in the T1 generation demonstrate that LIS-1 goes through fragmentation events, which seem to be genotype dependent. In the reorganization and deletion of LIS-1 from the T-DNA, some of the DNA sequences remaining within the T-DNA were first converted to the target site, at earlier stages of growth of the T1 plants, followed by complete excision of all flax DNA at later stages. Therefore, the only DNA sequence that is left inside the multiple cloning site of the vector is that pertaining to the binary vector. Furthermore, some smaller sub-fragments of LIS-1 were also present in the transformed plants, and they appeared to be distributed outside the T-DNA region not contiguous to each other.
The inheritance of LIS-1 sub-fragments
The inheritance of LIS-1 sub-fragments in the T2 generation was followed for all the T1s transformed with PRI909+LIS-1. PCRs with the previously amplified LIS-1 sub-fragments’ primers (2 and 3′- 5 and 6′- 14 and 15′- 18a and 19′) were performed. The PCR results show that there are similarities and differences between the Arabidopsis accessions. DNAs from all T2 transgenic plants from all accessions amplified the sub-fragment bounded by 2 and 3′; but all had lost the sub-fragment normally amplified by 18a and 19′. The sub-fragment bounded by 5 and 6′ was amplified from all T2s from transgenic plants of the accessions 28168, 22681, and 76113, but only three T2 transgenic plants from the accessions 28167 and 28168B amplified this fragment and the rest did not. Similarly, for the sub-fragment bounded by 14 and 15′ amplified from DNAs from all T2s from the accessions 28167, 28168 and 28168B, from eight of ten transgenic plants from accession 22681, but from only three of ten transgenic plants from accession 76113 (Table 1). Figure 6A–D shows examples of PCR of DNAs from the T2 generations from selected accessions. Table 1 summarizes the overall results for all PCRs of T2 generations from the accessions transformed with PRI909+LIS1.
The overall results of the T2 generations from the different accessions transformed with LIS-1 confirm that some LIS-1 sub-fragments were stably inherited in the T2 generations, whereas other sub-fragments were lost. Variation in stability was observed between the different accessions, and even between transgenic plants within the same accession.
Discussion
In this study different accessions of A. thaliana Columbia ecotype, plus one additional A. thaliana line “known as the Quick line”, were treated with either PRI909+LIS-1 or PRI909+the target site of LIS-1 (the T0 generation). T1 positive transformants were selected based on their kanamycin resistance and they were all transplanted to soil and grown to maturity. In the T2 generation, the segregation analysis has shown that approximately three quarters of the T2 plants were scored as kanamycin resistant by their dark green phenotype on the plates, confirming that T-DNA is segregating in a normal Mendelian fashion for a single locus. However, these T2 plants did not survive well when they were transplanted to soil. It has previously been noted that epigenetic silencing of the kanamycin transgene could occur in subsequent generations, and confirmation by PCR of kanamycin gene is needed if the transgenic lines are shown to be kanamycin sensitive.20–22 To circumvent any possible bias to the data, another set of plants from the T2 generation was germinated on soil without kanamycin and then those positive for kanamycin using PCR on DNA extracted from their leaves were selected; in other words, T2 generation plants were selected and tested based on kanamycin gene amplification with PCR and not based on selection using kanamycin resistance on plates. All of the T2 transgenic plants that were subjected to further testing, for the presence of either the target site of LIS1 or LIS-1 sub-fragments, were those tested positive by PCR for kanamycin fragment. The inheritance of the four sub-fragments of LIS-1 that had been identified in the T1 generation was followed in the T2 generations since those were the only ones that were amplified in the T1 generations.
The results show that for the A. thaliana accessions transformed with PRI909+target site of LIS1, the target site was stably inherited throughout the T1 and T2 generations. However, for the A. thaliana accessions transformed with PRI909+LIS1 in the T1 generations, LIS-1 goes through fragmentation events, which seem to be genotypic dependent. The loss of LIS-1 from the T-DNA construct during the initial growth of the T1 plants results in the conversion of the remaining DNA sequences (flanking sequences) to the sequence found as the target site in flax. This remaining fragment (identical to the target sequence and now indistinguishable from the transformants with the target sequence) is then completely excised as the T1 continues to grow. None of the transgenic A. thaliana plants maintain an intact LIS-1 element, but four sub-fragments of LIS-1 can be identified in the different accessions. In the T2 generations, three of these sub-fragments were inherited but there were still some variation between the different accessions. Figure S6 summarizes the overall flow of the experiments and results.
In all accessions that were transformed with PRI909+LIS-1, it was observed that LIS-1 was removed from the T-DNA, and sub-fragments of LIS-1 were distributed away from the T-DNA, but possibility linked to it. The latter can be supported by the results in Table 1, in which all T2 transgenic lines that were positive for T-DNA primers, NPTII and PriC, were positive for the sub-fragment 2 and 3′ as well. The results also show that these LIS-1 sub-fragments are not contiguous to each other. For example, transgenic T1 plants that were positive for the sub-fragment 2 and 3′ and 5 and 6′ failed to amplify with 2 and 6′ (Figure S4). So, it seems that LIS-1 has been broken up into smaller sub-fragments that show a complex inheritance pattern. Complete shotgun sequencing by next-generation sequencing of multiple T2 plants from different accessions should identify the sites of insertion of each of these fragments, and also confirm the copy numbers of the fragments. The identification of the sites of these new insertions and the actual ends of the rearranged fragments would also indicate if there was a single mechanism responsible for the dispersal of LIS-1 in Arabidopsis.
Associated with the loss of LIS-1 from the T-DNA construct was the modification of the remaining sequence within the T-DNA (with a reversion back to sequence present in flax prior to LIS-1 insertion) (Figure 5). This sequence is now identical to that used for the treatment of A. thaliana with the target sequence (for early stages of T1 generation, see Figures 5A and S6). However, in lines, which now contain target sequence derived from the original LIS-1 insertion, and fragments of LIS-1, the target sequence was still unstable and was lost from all T1 transformants (for late stages of T1 generation, see Figures 5B and S6). This is in contrast to when the target sequence alone was used in all transformations (PRI909+target site of LIS1) in which the target site was stable in all T1 and T2 generations in all accessions tested (Figures 3 and S6).
To understand the involvement of sub-fragments of LIS-1 in the stability of the target site, a cross between a transgenic line with a stable target site with another transgenic line with stable LIS-1 sub-fragments could be made from either T2 or T3 generations from both of these transgenic lines. This cross would allow the determination of how these sub-fragments may be able to destabilize the target sequence. In this way it should be possible to identify which fragments of LIS-1 were required for the instability of the target site.
It has been noted in previous studies that for the flax variety Stormont cirrus (plastic line or Pl), the inducing growth conditions are required for LIS-1 to be assembled and stably inherited in subsequent generations; when the inducing growth conditions were not maintained, LIS-1 appeared temporarily but was never seen in subsequent generations.9 To understand the effect of the growth conditions on LIS-1 stability in A. thaliana, it might be necessary to germinate the T1 positive transformants under “inducing” growth conditions (although exactly what these might be for A. thaliana has not yet been determined), and examine if changing the growth conditions would have an effect on “intact” LIS-1 stability.
Conclusion
Clearly, these results have shown that LIS-1 is unstable in A. thaliana. However, it appears that the excision/deletion of LIS-1 from T-DNA introduced into A. thaliana has similarity to the reversal of the assembly of LIS-1 in flax, since the target sequence is re-created during this excision. In other words, it appears that the processes by which LIS-1 is produced in flax are also present in A. thaliana. The future experiments outlined here will provide better understanding of these processes in both flax and A. thaliana.
Acknowledgment
This work was supported by the Ogelbay fund.
Author contributions
NKB performed the experiments and wrote the manuscript. CAC contributed to the design and supervision of the study and did the drafting and critical reviewing of the manuscript.
Disclosure
The authors declare that they have no conflict of interest to disclose.
References
Durrant A. The environmental induction of heritable change in Linum. Heredity. 1962;17(1):27–61. | |
Durant A. Induction and growth of flax genotrophs. Heredity. 1971; 27:277–298. | |
Cullis CA. Molecular aspects of the environmental induction of heritable changes in flax. Heredity. 1977;38(2):129–154. | |
Cullis CA. Phenotypic consequences of environmentally induced changes in plant DNA. Trends in Genetics. 1986;2:307–309. | |
Cullis CA. Mechanisms and control of rapid genomic changes in flax. Annals of Botany. 2005;95(1):201–206. | |
Chen Y, Schneeberger RG, Cullis CA. A site-specific insertion sequence in flax genotrophs induced by environment. New Phytol. 2005;167(1):171–180. | |
Bickel CL, Lukacs M, Cullis CA. The loci controlling plasticity in flax. Res and Reports in Biol. 2012;3:1–11. | |
Evans GM. Nuclear changes in flax. Heredity. 1968;23(1):25–38. | |
Chen Y, Lowenfeld R, Cullis CA. An environmentally induced adaptive (?) insertion event in flax. Int J Genet Mol Biol. 2009;1(3):38–47. | |
Johnson C, Moss T, Cullis C. Environmentally induced heritable changes in flax. J Vis Exp. 2011;(47):2332. | |
Mochizuki K, Gorovsky MA. Small RNAs in genome rearrangement in Tetrahymena. Current Opinion in Genetics and Development. 2004; 14(2):181–187. | |
Deak JC, Doerder FP. High frequency intragenic recombination during macronuclear development in Tetrahymena thermophila restores the wild-type SerH1 gene. Genetics. 1998;148(3):1109–1115. | |
Smith JJ, Baker C, Eichler EE, Amemiya CT. Genetic consequences of programmed genome rearrangement. Curr Biol. 2012;22(16):1524–1529. | |
Nowacki M, Haye JE, Fang W, Vijayan V, Landweber LF. RNA-mediated epigenetic regulation of DNA copy number. Proc Natl Acad Sci U S A. 2010;107(51):22140–22144. | |
Bent A. Arabidopsis thaliana floral dip transformation method. In: Wang K, editor. Agrobacterium Protocols: Springer; 2006:87–104. | |
Clough SJ, Bent AF. Floral dip: a simplified method for Agrobacterium-mediated transformation ofArabidopsis thaliana. Plant J. 1998;16(6):735–743. | |
Davis AM, Hall A, Millar AJ, Darrah C, Davis SJ. Protocol: Streamlined sub-protocols for floral-dip transformation and selection of transformants in Arabidopsis thaliana. Plant Methods. 2009;5:3. | |
Mara C, Grigorova B, Liu Z. Floral-dip transformation of Arabidopsis thaliana to examine pTSO2::beta-glucuronidase reporter gene expression. J Vis Exp. 2010;(40):1952. | |
Zhang X, Henriques R, Lin SS, Niu QW, Chua NH. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc. 2006;1(2):641–646. | |
Yenofsky RL, Fine M, Pellow JW. A mutant neomycin phosphotransferase II gene reduces the resistance of transformants to antibiotic selection pressure. Proc Natl Acad Sci U S A. 1990;87(9):3435–3439. | |
Zhan XC, Jones DA, Kerr A. Regeneration of flax plants transformed by Agrobacterium rhizogenes. Plant Mol Biol. 1988;11(5):551–559. | |
Meza TJ, Kamfjord D, Håkelien AM, et al. The frequency of silencing in Arabidopsis thaliana varies highly between progeny of siblings and can be influenced by environmental factors. Transgenic Res. 2001;10(1):53–67. |
Supplementary materials
  | Figure S1 A diagram to show the target site of LIS-1 and the detailed primers along LIS-1. |
References
Bickel CL, Lukacs M, Cullis CA. The loci controlling plasticity in flax. Res and Reports in Biol. 2012;3:1–11. | |
Chen Y, Schneeberger RG, Cullis CA. A site-specific insertion sequence in flax genotrophs induced by environment. The New phytologist. 2005;167(1):171–180. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.