Back to Journals » The Application of Clinical Genetics » Volume 13
The Importance of Small Non-Coding RNAs in Human Reproduction: A Review Article
Authors Kamalidehghan B, Habibi M, Afjeh SS, Shoai M, Alidoost S , Almasi Ghale R, Eshghifar N , Pouresmaeili F
Received 4 March 2019
Accepted for publication 4 December 2019
Published 8 January 2020 Volume 2020:13 Pages 1—11
DOI https://doi.org/10.2147/TACG.S207491
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Martin Maurer
Behnam Kamalidehghan,1,* Mohsen Habibi,2,* Sara S Afjeh,1 Maryam Shoai,3 Saeideh Alidoost,4 Rouzbeh Almasi Ghale,4 Nahal Eshghifar,5 Farkhondeh Pouresmaeili1,6
1Department of Medical Genetics, Faculty of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran; 2Central Laboratory, Faculty of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran; 3Department of Molecular Neuroscience, UCL Institute of Neurology, London, UK; 4Department of Biology, Science and Research Branch, Islamic Azad University, Tehran, Iran; 5Department of Cellular and Molecular Biology, Faculty of Advanced Science and Technology, Tehran Medical Sciences, Islamic Azad University, Tehran, Iran and Men’s Health and Reproductive Health Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran; 6Men’s Health and Reproductive Health Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran
*These authors contributed equally to this work
Correspondence: Farkhondeh Pouresmaeili
Men’s Health and Reproductive Health Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran
Tel/Fax +98 21-23872572
Email [email protected]
Background: MicroRNAs (miRNA) play a key role in the regulation of gene expression through the translational suppression and control of post-transcriptional modifications.
Aim: Previous studies demonstrated that miRNAs conduct the pathways involved in human reproduction including maintenance of primordial germ cells (PGCs), spermatogenesis, oocyte maturation, folliculogenesis and corpus luteum function. The association of miRNA expression with infertility, polycystic ovary syndrome (PCOS), premature ovarian failure (POF), and repeated implantation failure (RIF) was previously revealed. Furthermore, there are evidences of the importance of miRNAs in embryonic development and implantation. Piwi-interacting RNAs (piRNAs) and miRNAs play an important role in the post-transcriptional regulatory processes of germ cells. Indeed, the investigation of small RNAs including miRNAs and piRNAs increase our understanding of the mechanisms involved in fertility. In this review, the current knowledge of microRNAs in embryogenesis and fertility is discussed.
Conclusion: Further research is necessary to provide new insights into the application of small RNAs in the diagnosis and therapeutic approaches to infertility.
Keywords: miRNA, female fertility, male fertility, piwi-interacting RNAs, piRNAs
Introduction
MicroRNAs (miRNAs) are small fragments made of 20–24 nucleotides in single-stranded noncoding ribonucleic acid structures, influencing the normal function of eukaryotic cells from various organisms, including mammalian cells. Their sequence homology is approximately 90% conserved among human, rat, and mouse.1 The miRNAs are known to be transcribed by RNA polymerase II as primary stem-loop structured miRNA molecules are processed through DROSHA enzyme and DGCR8 proteins cooperation to produce pre-miRNAs with hairpin structures and 60–70-nucleotides length in the nucleus. Finally, the recent molecules are exported to the cytoplasm by exportin 5 (XPO5), and processed by a ribonuclease III enzyme named DICER1 that produces the mature 19–25 nucleotides miRNAs.2 The guide strand of the mature miRNA in accompany with the miRNA-induced silencing complex (miRISC), composed of DICER1 and Argonaute (AGO) proteins, directs the miRISC for a sequence complementary binding to the target mRNAs.3 miRNA plays an important role in almost every biochemical process in the body through post-transcriptional regulation. Many of the miRNA templates are located in intronic regions and are transcribed into a double strand pre-miRNA, often adopting a hairpin conformation. The pre-miRNA is then cleaved by RNase III and unwound to form a single strand mature miRNA in the cytoplasm.4 This strand is eventually unwound at the 5ʹ end, the less stable terminal sequence, and is subsequently packaged into a RNA-induced silencing complex (RISC). RISC has cleaving capacity and most often interacts with the noncoding region of mRNA and is composed of Argonaut proteins.4
The recent studies on miRNAs and their new implied role in the regulation of cellular processes revealed new insight into the regulation of gene expression.5,6
miRNA molecules may induce negative regulation by mRNA degeneration via deadenylation7,8 and/or translational repression by partial complementary mRNA annealing or direct messenger RNA (mRNA) cleavage1,9 in addition to the positive effect on target gene promoters.10 There are up to 50,000 different miRNAs, each with hundreds to thousands of potential target mRNAs, regulating approximately 30% of protein-coding genes as predicted by computational analysis in mammalian cells.11,12 Moreover, there is evidence that miRNAs are associated with several biological processes including cell growth, development, and differentiation.13
MiRNAs are also associated with human diseases including several types of cancers, viral infections, and heart disease.1,13,14 Synthesis and degradation of miRNAs have been proven to play an important role during oogenesis and embryonic development.15,16 Global miRNA expression profiling has shown that miRNAs are inherited maternally and almost 60% of them are being lost during the maternal-zygotic transition from one to two cell stages.17,18
It seems that dynamic degradation and synthesis of miRNAs coexist during the development of the preimplantation mouse embryo with an overall elevation in miRNAs toward the blastocyst stage.19 The animal studies have proved that specific stages of mouse embryonic prenatal development including E9.5, E10.5, and E11.5 indicated different miRNA expression profiles with the identification of new mammalian miRNAs.20
Expressionally, miRNA defects have been identified to be associated with a number of human diseases,21 including benign gynecological conditions, malignancies of female reproductive organs and fertility disorders of the human female reproductive tract.22
It is thought that the new advancements in miRNA technology together with the new insights into the function of miRNAs may enable the development of new biomarkers of female infertility. This can encompass biomarkers of ovarian physiology, the quality of oocyte and embryos, implantation potential of embryos as well as tissue receptivity of the endometrium.
In this review we aim to utilize the current knowledge on miRNAs and their involvement in female reproductive potential from the beginning point in the early developmental stage to the time of embryo formation in order to identify stage-specific biomarkers for the ovarian status in fertile or infertile women, to find causes of impaired ovarian function, the role of miRNAs in infertility and the prediction of Assisted Reproductive Technology (ART) results.
miRNAs and Uterine Function
It has been demonstrated that miRNAs mediate regulation and expression of important genes of uterine during implantation.23 For instance, cyclooxygenase-2 expression, which is pivotal for embryo implantation in the mouse, is regulated following the transcription of two miRNAs including ha-miR-199a, and hsa-miR-101a.24 Additionally, implantation failure has been remarkably considered as a significant contributor to human infertility; therefore, the aberrant expression of critical miRNAs during implantation can also be a factor. To gain insight into miRNA expression amongst human embryos development, quantitative real-time PCR (qPCR) was performed for a set of 12 miRNAs on individual human blastocysts.21 These miRNAs were specifically selected from recent MICE model publications that proved their expression in either mouse embryos4,17 or human embryonic stem cells: RNUU48, hsa-let-7a, hsa-let-7b, hsa-let-7c, hsa-let-7g, hsa-miR-93,hsa-miR-92, hsa-miR-21, and hsa-miR-24.1
miRNAs and Primordial Germ Cell Development
It is assumed that the development of human preimplantation begins with primordial germ cells (PGCs) that singularly emerge outside the genital ridge region and can be first distinguished in the human embryo at about 3 weeks in the yolk sac epithelium beside the base of developing allantois.25
The role of endometrial miRNAs on implantation is beginning to be understood, but, the targeted mRNAs and the regulatory networks induced by miRNAs in PGCs remain unclear. Also, primordial germ cell has a vital role in development/miscarriage. One of the most remarkable functions of miRNAs in PGCs is identified as targeting the epigenetics-related genes and DNA methylation process in developing gonads.26 The related in vivo mouse models have revealed the important role of miR-29b in female gonadal development, targeting epigenetics-related genes including DNMT3A and DNMT3B and modulating methylation of genomic DNA in PGCs.27 Previous studies indicated the importance of miR-290-295 and miR-17-92 clusters during mouse germ cell development, through remarkably abundant miRNAs in PGCs.26 These findings were confirmed similarly in chicken with improved DNMT3B expression in a female germ cell-specific aspect and down-regulation of four miRNAs including miR-15c, miR-29b, miR-383, and miR-222.28 Other researches on the vertebrate species like Carassius auratus (goldfish)29 and Danio rerio (zebrafish)30 have identified some different miRNAs which may play an important role in development and maintenance of PGCs. Although some miRNAs including miR-29b and miR-430 demonstrated overlap between different species, the structure of miRNA expression in PGCs seems to be species-specific.31
miRNAs and Ovarian Development
PGCs migrate and colonize to become oogonia in the forming fetal ovary. The oogonia undergo mitotic divisions and proliferate broadly by up to 5–7 million cells in humans. MiRNAs found in mouse ovarian cells have been recently explored in ovarian cells, revealing their influence on fundamental ovarian aspects like ovulation, steroidogenesis, and corpus luteum development and function.26 Three different miRNAs including miR-224, miR-378, and miR-383 have been identified to play a role in the regulation of aromatase expression during follicle development. Furthermore, miR-21 was implicated in promotion the follicular cell survival during ovulation. Proangiogenic miRNAs including miR-17-5p and let-7b proved to be crucial for normal development of the corpus luteum after ovulation.32 Mouse animal studies showed that ribonuclease III nominated Dicer1 is involved in the process of folliculogenesis. Dicer1 protein is expressed in oocytes and granulosa cells of follicles.33 To determine miRNAs function in the development of mouse ovary, in a new strategy, Dicer1 was deleted specifically in mouse granulosa cells, which resulted in Dicer1 conditional knockout (Cho) mouse. Dicer1 deletion caused significantly miR-503 downregulation.31 This miRNA is more abundant in mouse ovary tissue compared with other tissues. Increasing the pool of primordial follicles hastened recruitment of early follicle and therefore more degenerated follicles can be observed.
The ovarian niche is very crucial for the growth and maturation of follicles/oocytes that may cause infertility.31
Oocyte-Cumulus Complex
Each oocyte grows and matures in the practical unit of the ovary, the follicle enveloped and supported by granulosa and theca cells, which demonstrate the important niche for oocyte growth and maturation.34 Normal fertilization and the development of an embryo depend on normal growth and maturation of the oocyte which needs sufficient communication between the oocyte and surrounding follicular cells.34 Recent studies indicate that miRNAs are expressed within the organs of the female reproductive tract where they regulate necessary cellular pathways for proper function of these organs.35 While miRNAs represent a limited and sensitive biological material, which is mostly not available to researchers, their related data in human oocytes are insufficient.36 A few studies on human oocytes resulted from the in vitro fertilization have proved the existence of miRNAs in these germ cells.31 A study showed the dynamic changes of miRNAs from GV- to MII-stages using miR-CURY LNA microarray platform and quantitative RT-PCR, indicating the concordant dynamic changes of miR-15a and miR-20a expression during meiosis.37 Furthermore, increased concentration of follicle-stimulating hormone (FSH) in the in vitro maturation medium, had a reverse effect on the expression of miR-15a and miR-20a, strengthening the evidence for the role of these miRNAs in the oocyte maturation process under FSH induction.37 Another study demonstrated that human mature MII-stage oocytes have an abundance of miR10A, miR100, and miR184 in which they differ from the miRNAs profile in enclosing cumulus oophorus. The cumulus oophorus are a group of cells surrounding the oocytes in the ovarian follicle.38 Moreover, it is shown that ovarian pathologies including polycystic ovary syndrome (PCOS) and premature ovarian failure (POF) or primary ovarian insufficiency affect the miRNA expression in follicular fluid and plasma.39
Polycystic Ovary Syndrome (PCOS)
PCOS is a systematic disease defined by a set of symptoms resulting from a hormonal imbalance in women and is the most common endocrinopathy in women of reproductive age, affecting reproduction performance and metabolic process.40 Patients are diagnosed with anovulation, high androgen levels, and several ovarian cysts detected by ultrasound, low fertility, irregular or no menstrual periods, obesity, heavy periods, pelvic pain, excess body and facial hair (hirsutism), acne and patches of thick and darker skin as typical symptoms.40
Circulating miRNAs are observed in whole blood, serum, plasma and the follicular fluid of PCOS patients which provide potential biomarkers and a new diagnostic tool for the PCOS.41 In PCOS, human follicular fluids microarray profiling demonstrated the upregulation of miR-9, miR18b, miR-32, miR-34c, miR-135a and downregulation of miR-132 and miR-320 between the PCOS patients and normal groups of women.41
Premature Ovarian Failure (POF)
POF is an indication of the early severe ovarian failure which is defined as an ovarian disorder of multifactorial origin characterized by hypergonadotropism (e.g., increased FSH levels) and hypoestrogenism where no follicles/oocytes are available to be fertilized in women under the age of 40 years.31 Due to the stability of miRNAs in body fluids, upregulation of miR-126, miR-202, miR-23a, miR-146a, miR-139-3p, miR-125b-2, miR-654-5p, miR-27a, miR-765, and miR-342-3p and downregulation of miR-22-3p, let-7c, and miR-144 was identified in premature ovarian failure,31 thus giving rise to novel biomarkers for ovarian function.42
Embryo, Development, and Implantation
Human embryos have been found to secrete miRNAs into culture media which were recognized as new biomarkers for embryo development and implantation.43 miRNAs expression from culture media of 55 single-embryo transfer cycles was tested using array-based RT-PCR (or qPCR) analysis. The results demonstrated an association between the expressions of identified miRNA with pregnancy outcomes.44 Two miRNAs including miR-191 and miR-372 were expressed specifically in spent media after embryo culture.44 Other miRNAs were identified in more concentrated status in intracytoplasmic sperm insemination (ICSI) research and day-5 media samples compared with regularly inseminated and day-4 samples, respectively.22
It was found that miRNA levels are associated with some severe conditions. For instance, miR-191 was more highly concentrated in media from aneuploid embryos, while miR-372, miR-645, and miR-191 were more concentrated in media from failed in vitro fertilization cycles without pregnancy.44
The Repeated Implantation Failure (RIF)
RIF is one of the most noteworthy problems that may be experienced in the in vitro fertilization program.45 Studies on implantation miRNAs biomarkers may help clinicians to predict pregnancy outcome and detect occult implantation deficiency. Thirteen different miRNAs have been identified in RIF’s endometrial samples, in comparison to normal ones, which putatively regulate the expression of 3800 genes.41 It was found that ten miRNAs were overexpressed in RIF endometrial samples, including but not limited to miR-23b, miR-99a, and miR-145.46
Endometriosis is out of uterine cavity formation of endometrial tissues including glands and stroma. Endometriomas are Endometrial cysts of the ovary which are known as chocolate cysts,47 in which the most effective treatment for the disease is surgery due to endometriomas resistance to common treatment methods.48 It is reported that approximately 16% of reproductive age women undergoing hysterectomy showed endometriosis,49 while up to 40% of women presenting infertility revealed endometriosis.50 In comparison with eutopic tissues, many mRNA transcripts are differentially regulated in endometriotic.51 A study by Pan et al,52 indicated differential expression of 48 miRNAs in a microarray panel analysis,52 in which most of these miRNAs showed reduced expressions in eutopic and ectopic tissues in affected women with endometriosis in comparison to controls. Additionally, validated microarray panel results of qRT-PCR, confirm dysregulation of two important miRNAs miR-26a and miR-21 in eutopic tissues compared with ectopic tissues.52 Some of infertility associated miRNAs in females is summarized in Table 1.
 |
Table 1 List of miRNAs Expressed in Granulosa Cells, Oocytes and Ovaries |
miRNA, Spermatogenesis and Male Fertility
miRNAs play critical roles in the regulation of gene expression, especially at the post-transcriptional level. They are expressed in a cell- or phase-specific manner during spermatogenesis and have a crucial role in male reproductive health (Figures 1 and 2). Consequently, dysregulation of miRNAs considers as a molecular basis for male infertility and the aberrant expression patterns of these molecules can be transferred to the offspring. Molecular genetic abnormalities are very important in the etiology of male infertility.,53 in which approximately 60-75% of cases have idiopathic oligozoospermia and azoospermia.54 Recent studies on clusters of small RNA demonstrated the different expression patterns of small RNA molecules influence gene expression in the specific process of spermatogenesis.47
Previous studies on miRNAs revealed that they regulate expression through interfering with mRNA translation. This finding was first identified in Caenorhabditis elegans and further studies identified this role in several other organisms including humans.19,55 Currently, miRNA is proved to regulate the expression of approximately 60% of protein-coding genes in the human genome. Since September 2010, 1100 miRNAs have been sequenced on the human genome, and that number is rapidly predicted to grow in the coming years.20 It has been proven that each chromosome, except the Y chromosome, contains miRNA coding genes and X chromosome demonstrated significantly more miRNA coding genes than other chromosomes which are specific to primates and exclusively express in the epididymis with predicted target proteins involved in morphogenesis of the epididymis, direct and indirect regulation of immotile sperm with flagella motility, development of the lumen and maintenance of mature sperm. The SPAG6 protein, sperm-associated antigen 6 that was found as an infertility protein, was also a predicted target.56 These mechanisms of specific clusters of miRNAs are found throughout the genome, and especially on the X chromosome suggesting that small RNA regulate stages of spermatogenesis at different levels. Another study revealed that many miRNAs from X chromosome templates are present in pachytene spermatocyte and the biogenesis of these miRNAs may cause the meiotic sex chromosome inactivation escape (MSCI). This study indicated that miRNA may play a role in suppressing the sex-linked chromosomal transcription, implicating small RNAs at different stages of spermatogenesis like maturation.4 Some of the important small RNAs associated with male reproduction and fertility are summarized in Table 2.
 |
Table 2 A List of the Important Small RNA Molecules Related to Male Reproduction and Fertility |
piRNAs and Infertility
P-element Induced Wimpy testis (PIWI) proteins in complex with PIWI-interacting RNA (piRNA) are a novel member of small non-coding RNAs (ncRNAs) with approximately 24–32 nucleotides.57 piRNAs are expressed in the gonads, suggesting a significant role in germline stem cell maintenance including epigenetic and post-transcriptional gene silencing of retrotransposon and other genetic elements in germline cells, such as reproductive stem cell self-sustainment, differentiation, meiosis, and spermatogenesis. PIWI mutations in Drosophila, C. elegans, mice and Zebrafish caused gametogenic defects such as failure of germline establishment, loss of germline stem cells, meiotic arrest and blockage in spermiogenesis, leading to sterility.58
The PIWI family proteins are expressed predominantly in the germline of various organisms and are highly conserved during the evolutionary process.59,60 PIWI proteins are indispensable for piRNA biogenesis and function.61 However, in humans, the actual involved mechanism is not well understood.61 Recent studies have demonstrated that piRNAs and PIWI proteins play vital roles in developments of germ cells and fertility,62 as mutations that knockout either PIWI protein function or piRNA biogenesis, resulting in germ cell death, gonadal atrophy, and finally sterility in invertebrates and vertebrates.63
PIWIL1/HIWI, PIWIL2/HILI, PIWIL3/HIWI3, and PIWIL4/HIWI2 are four human PIWIs,57 while the mouse genome only encodes three PIWI homologs including Piwil1/Miwi, Piwil2/Mili, and Piwil4/Miwi2.64 The role of piRNAs during spermatogenesis is confirmed by the known functions of their partner, the PIWI proteins.65
Transposable DNA elements make up a fundamental proportion of most eukaryotic genomes.66 As these elements are highly mutagenic, regulatory mechanisms are required for their control. Previous studies on animal germ cells have demonstrated a crucial function of the piRNA pathway in silencing transposable elements (TEs)67,68 through epigenetic and the post-transcriptional level mechanisms, protecting the integrity of the genome and the development of gametes.69
In mice, the de novo DNA methylation of mobile genetic elements is mediated by MILI and MIWI2, which are loaded via transposon-derived piRNAs.70 MILI mutant/deficient models disclose a loss of methylation at L1 (LINE-1) retrotransposon,71 resulting in reactivation of transposable elements that contribute to meiotic arrest and male infertility.66
Previous studies have shown that PIWI/piRNAs complexes methylate DNA to silence repetitive elements, in male germline stem cells. In infertile males with spermatogenic failure, hypermethylation in the promoter regions of PIWIL2 has been consistent with the repression of PIWIL2 due to hypermethylation, hypomethylation of LINE-1 sequences was also observed.66 Consistently, other findings demonstrated the functional result of piRNA depletion is a depression of repetitive elements.72–74
PIWI proteins and piRNAs expression patterns are tightly regulated during spermatogenesis in mouse,75 including successful mitotic, meiotic, and post-meiotic phases.76 MILI and MIWI2 mutant/deficient mice are sterile due to impaired spermatogenesis caused by meiotic arrest at the pachytene spermatocyte and early prophase of meiosis-I,74,77 respectively. Moreover, MIWI-null mice are infertile because of spermatogenic arrest at the round spermatid stage without elongated spermatids or mature spermatozoa.78 In contrast, females with the deficiency in genes that are required for piRNA biogenesis including PIWI proteins family, exhibit normal oogenesis and fertility.79 The presence of piRNA genes that are exclusively expressed in the ovary indicates the probable role of piRNAs in ovarian function.75
In humans, piRNA expression is widely limited to the adult testis and fetal ovary. Unlike chronological age of adult testis and fetal ovary, germ cells in adult testis and fetal ovaries are at similar stages of gametogenesis. The abundance of piRNAs in adult ovary and fetal testis is nearly 25 fold lower compared to the fetal ovary and adult testis. These differences suggest the importance of appropriate levels of piRNAs for fertility.75 However, alterations in piRNAs levels are not the only contributors to male infertility, the supporting proteins for piRNAs including MILI and MIWI2 must also be at appropriate levels to warrant normal spermatogenesis.
Conclusion
The miRNAs represent a great challenge in reproductive and regenerative medicine for better understanding the preimplantation development in humans including PGCs, gametes, female reproductive tissues, embryos, and embryonic stem cells. Upcoming new knowledge can provide new biomarkers and therapeutic targets and approaches for fertility and degenerative disorders in the future.
The miRNAs have indirect DNA methylation function. Therefore, they can control transcriptional activity. This indicates their role in genetic imprinting that may demonstrate epigenetic patterns for embryonic development. Recent surveys have revealed a strong role of epigenetic changes in the regulation of the process of fertility and spermatogenesis. It has been revealed that changes in the epigenetic system could change the performance of important genes/pathways in spermatogenesis or fertility. Thus, future studies can explore the effect of small RNA molecules on the process of epigenetic modifications which not only influences the individuals having the modifications but are also transmitted to the next generation. Therefore, the small RNAs contribution and regulation displayed in epigenetic modifications can help further knowledge of the spermatogenesis and associated infertility processes. Infertility has been combated with Assisted Reproductive Techniques (ART), in particular, thanks to extracting abnormal functional sperm and fertilizing an oocyte via ICSI. Despite ICSI which permits sperm to enter the oocytes for fertilization, genetic factors that may be compromised and are still being propagated to progeny through ART must be considered. Many sperm maturation steps directly correlate with expression levels of certain small RNAs. Many of the specific roles of small RNAs are yet unknown, but the genes and proteins that interact are new windows through which their involvement can be understood.
Further research is required to comprehend the function of lncRNAs in reproductive disorders and this field is likely to be a remarkable point in the coming years.
With the aid of novel technologies and searchable databases, which includes ontology and bioinformatics databases, lncRNAs can additionally give us an exquisite outlook to analysis, prevent and manage some reproductive disorders.
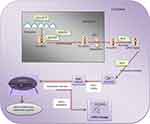 |
Figure 1 Mirnas Biogenesis Pathway. Data from Harchegani et al.53 |
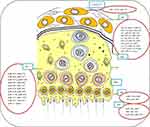 |
Figure 2 The expression of relevant miRNAs in testicular cells. A large number of miRNAs are responsible for the normal function of reproductive organ and spermatogenesis. They control all levels of spermatogenesis including 1-Leydig and Sertoli cells development, 2-spermatogonial stem cells formation and their differentiation to 3-spermatogonia, 4-primary spermatocytes, 5-secondary spermatocytes, 6-round spermatids, and 7-mature sperm cells. Data from Harchegani et al.53 Abbreviations: SSCs indicates spermatogonial stem cells; SPG, spermatogonia; PSPC, primary spermatocytes; SSPC, secondary spermatocytes; RS, round spermatids; SPs, sperm cells; SCs, sertoli cells; miRNAs, microRNAs. |
Disclosure
The authors report no conflicts of interest in this work.
References
1. McCallie B, Schoolcraft WB, Katz-Jaffe MG. Aberration of blastocyst microRNA expression is associated with human infertility. Fertil Steril. 2010;93(7):2374–2382. doi:10.1016/j.fertnstert.2009.01.069
2. Anfossi S, Fu X, Nagvekar R, Calin GA. MicroRNAs, regulatory messengers inside and outside cancer cells. Adv Exp Med Biol. 2018;1056:87–108.
3. Galagali H, Kim JK. miRISC composition determines target fates in time and space. Dev Cell. 2018;47(2):142–143. doi:10.1016/j.devcel.2018.10.009
4. Mineno J, Okamoto S, Ando T, et al. The expression profile of microRNAs in mouse embryos. Nucleic Acids Res. 2006;34(6):1765–1771. doi:10.1093/nar/gkl096
5. Kloosterman WP, Plasterk RH. The diverse functions of microRNAs in animal development and disease. Dev Cell. 2006;11(4):441–450. doi:10.1016/j.devcel.2006.09.009
6. Perez-Cremades D, Mompeon A, Vidal-Gomez X, Hermenegildo C, Novella S. Role of miRNA in the regulatory mechanisms of estrogens in cardiovascular ageing. Oxid Med Cell Longev. 2018;2018:6082387. doi:10.1155/2018/6082387
7. Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc Natl Acad Sci U S A. 2008;105(5):1608–1613. doi:10.1073/pnas.0707594105
8. Zhang X, Xiao S, Rameau RD, et al. Nucleolin phosphorylation regulates PARN deadenylase activity during cellular stress response. RNA Biol. 2018;15(2):251–260. doi:10.1080/15476286.2017.1408764
9. Bastet L, Turcotte P, Wade JT, Lafontaine DA. Maestro of regulation: riboswitches orchestrate gene expression at the levels of translation, transcription and mRNA decay. RNA Biol. 2018;15(6):679–682. doi:10.1080/15476286.2018.1451721
10. Tang F, Kaneda M, O’Carroll D, et al. Maternal microRNAs are essential for mouse zygotic development. Genes Dev. 2007;21(6):644–648. doi:10.1101/gad.418707
11. Park K, Kim KB. miRTar Hunter: a prediction system for identifying human microRNA target sites. Mol Cells. 2013;35(3):195–201. doi:10.1007/s10059-013-2165-4
12. Schwarzenbacher D, Balic M, Pichler M. The role of microRNAs in breast cancer stem cells. Int J Mol Sci. 2013;14(7):14712–14723. doi:10.3390/ijms140714712
13. Ha T-Y. MicroRNAs in human diseases: from cancer to cardiovascular disease. Immune Netw. 2011;11(3):135–154. doi:10.4110/in.2011.11.3.135
14. Khan F, Momtaz S, Abdollahi M. The relationship between mercury exposure and epigenetic alterations regarding human health, risk assessment and diagnostic strategies. J Trace Elem Med Biol. 2019;52:37–47. doi:10.1016/j.jtemb.2018.11.006
15. Minogue AL, Tackett MR, Atabakhsh E, Tejada G, Arur S. Functional genomic analysis identifies miRNA repertoire regulating C. elegans oocyte development. Nat Commun. 2018;9(1):5318. doi:10.1038/s41467-018-07791-w
16. Real FM, Sekido R, Lupianez DG, Lovell-Badge R, Jimenez R, Burgos M. A microRNA (mmu-miR-124) prevents Sox9 expression in developing mouse ovarian cells. Biol Reprod. 2013;89(4):78. doi:10.1095/biolreprod.113.110957
17. Lakshmipathy U, Love B, Goff LA, et al. MicroRNA expression pattern of undifferentiated and differentiated human embryonic stem cells. Stem Cells Dev. 2007;16(6):1003–1016. doi:10.1089/scd.2007.0026
18. Yang Q, Lin J, Liu M, et al. Highly sensitive sequencing reveals dynamic modifications and activities of small RNAs in mouse oocytes and early embryos. Sci Adv. 2016;2(6):e1501482. doi:10.1126/sciadv.1501482
19. Tatsuguchi M, Seok HY, Callis TE, et al. Expression of microRNAs is dynamically regulated during cardiomyocyte hypertrophy. J Mol Cell Cardiol. 2007;42(6):1137–1141. doi:10.1016/j.yjmcc.2007.04.004
20. Yang Y, Bai W, Zhang L, et al. Determination of microRNAs in mouse preimplantation embryos by microarray. Dev Dyn. 2008;237(9):2315–2327. doi:10.1002/dvdy.21666
21. Singh SK, Pal Bhadra M, Girschick HJ, Bhadra U. MicroRNAs–micro in size but macro in function. FEBS J. 2008;275(20):4929–4944. doi:10.1111/j.1742-4658.2008.06624.x
22. Carletti M, Christenson L. MicroRNA in the ovary and female reproductive tract. J Anim Sci. 2009;87(14_suppl):E29–E38. doi:10.2527/jas.2008-1331
23. Krawczynski K, Bauersachs S, Reliszko ZP, Graf A, Kaczmarek MM. Expression of microRNAs and isomiRs in the porcine endometrium: implications for gene regulation at the maternal-conceptus interface. BMC Genomics. 2015;16:906. doi:10.1186/s12864-015-2172-2
24. Chakrabarty A, Tranguch S, Daikoku T, Jensen K, Furneaux H, Dey SK. MicroRNA regulation of cyclooxygenase-2 during embryo implantation. Proc Natl Acad Sci U S A. 2007;104(38):15144–15149. doi:10.1073/pnas.0705917104
25. Johnson M, Everitt B. Adult ovarian function. In: Essential Reproduction. Vol. 5. 2000.
26. Bhin J, Jeong HS, Kim JS, et al. PGC-Enriched miRNAs Control Germ Cell Development. Mol Cells. 2015;38(10):895–903.
27. Takada S, Berezikov E, Choi YL, Yamashita Y, Mano H. Potential role of miR-29b in modulation of Dnmt3a and Dnmt3b expression in primordial germ cells of female mouse embryos. RNA. 2009;15(8):1507–1514. doi:10.1261/rna.1418309
28. Rengaraj D, Lee BR, Lee SI, Seo HW, Han JY. Expression patterns and miRNA regulation of DNA methyltransferases in chicken primordial germ cells. PLoS One. 2011;6(5):e19524. doi:10.1371/journal.pone.0019524
29. Mei J, Yue H-M, Li Z, et al. C1q-like factor, a target of miR-430, regulates primordial germ cell development in early embryos of Carassius auratus. Int J Biol Sci. 2014;10(1):15–24. doi:10.7150/ijbs.7490
30. Takeda Y, Mishima Y, Fujiwara T, Sakamoto H, Inoue K. DAZL relieves miRNA-mediated repression of germline mRNAs by controlling poly (A) tail length in zebrafish. PLoS One. 2009;4(10):e7513. doi:10.1371/journal.pone.0007513
31. Virant-Klun I, Ståhlberg A, Kubista M, Skutella T. MicroRNAs: from female fertility, germ cells, and stem cells to cancer in humans. Stem Cells Int. 2016a;2016:3984937. doi:10.1155/2016/3984937
32. Donadeu FX, Schauer S, Sontakke S. Involvement of miRNAs in ovarian follicular and luteal development. J Endocrinol. 2012;215(3):323–334. doi:10.1530/JOE-12-0252
33. Lei L, Jin S, Gonzalez G, Behringer RR, Woodruff TK. The regulatory role of Dicer in folliculogenesis in mice. Mol Cell Endocrinol. 2010;315(1–2):63–73. doi:10.1016/j.mce.2009.09.021
34. Telfer EE, Zelinski MB. Ovarian follicle culture: advances and challenges for human and nonhuman primates. Fertil Steril. 2013;99(6):1523–1533. doi:10.1016/j.fertnstert.2013.03.043
35. Nothnick WB. The role of micro-RNAs in the female reproductive tract. Reproduction. 2012;143(5):559–576. doi:10.1530/REP-11-0240
36. Robert C. Microarray analysis of gene expression during early development: a cautionary overview. Reproduction. 2010;140(6):787–801. doi:10.1530/REP-10-0191
37. Xu Y-W, Wang B, Ding C-H, Li T, Gu F, Zhou C. Differentially expressed micoRNAs in human oocytes. J Assist Reprod Genet. 2011;28(6):559–566. doi:10.1007/s10815-011-9590-0
38. Assou S, Al-Edani T, Haouzi D, et al. MicroRNAs: new candidates for the regulation of the human cumulus–oocyte complex. Hum Reprod. 2013;28(11):3038–3049. doi:10.1093/humrep/det321
39. Imbar T, Eisenberg I. Regulatory role of microRNAs in ovarian function. Fertil Steril. 2014;101(6):1524–1530. doi:10.1016/j.fertnstert.2014.04.024
40. Ecklund LC, Usadi RS. Endocrine and reproductive effects of polycystic ovarian syndrome. Obstet Gynecol Clin North Am. 2015;42(1):55–65. doi:10.1016/j.ogc.2014.09.003
41. Sørensen AE, Wissing ML, Salö S, Englund ALM, Dalgaard LT. MicroRNAs related to polycystic ovary syndrome (PCOS). Genes (Basel). 2014a;5(3):684–708. doi:10.3390/genes5030684
42. Ilie IR, Georgescu CE. Polycystic ovary syndrome-epigenetic mechanisms and aberrant MicroRNA. Adv Clin Chem. 2015;71:25–45.
43. Kropp J, Salih SM, Khatib H. Expression of microRNAs in bovine and human pre-implantation embryo culture media. Front Genet. 2014;5:91.
44. Rosenbluth EM, Shelton DN, Wells LM, Sparks AE, Van Voorhis BJ. Human embryos secrete microRNAs into culture media—a potential biomarker for implantation. Fertil Steril. 2014;101(5):1493–1500. doi:10.1016/j.fertnstert.2014.01.058
45. Simon A, Laufer N. Assessment and treatment of repeated implantation failure (RIF). J Assist Reprod Genet. 2012;29(11):1227–1239. doi:10.1007/s10815-012-9861-4
46. Revel A, Achache H, Stevens J, Smith Y, Reich R. MicroRNAs are associated with human embryo implantation defects. Hum Reprod. 2011;26(10):2830–2840. doi:10.1093/humrep/der255
47. Hawkins SM, Creighton CJ, Han DY, et al. Functional microRNA involved in endometriosis. Mol Endocrinol. 2011;25(5):821–832. doi:10.1210/me.2010-0371
48. Aznaurova YB, Zhumataev MB, Roberts TK, Aliper AM, Zhavoronkov AA. Molecular aspects of development and regulation of endometriosis. Reprod Biol Endocrinol. 2014;12:50. doi:10.1186/1477-7827-12-50
49. Merrill RM, Layman AB, Oderda G, Asche C. Risk estimates of hysterectomy and selected conditions commonly treated with hysterectomy. Ann Epidemiol. 2008;18(3):253–260. doi:10.1016/j.annepidem.2007.10.011
50. Verkauf B. Incidence, symptoms, and signs of endometriosis in fertile and infertile women. J Fla Med Assoc. 1987;74(9):671.
51. Ohlsson Teague EMC, Print CG, Hull ML. The role of microRNAs in endometriosis and associated reproductive conditions. Hum Reprod Update. 2009;16(2):142–165. doi:10.1093/humupd/dmp034
52. Pan Q, Luo X, Toloubeydokhti T, Chegini N. The expression profile of micro-RNA in endometrium and endometriosis and the influence of ovarian steroids on their expression. Mol Hum Reprod. 2007;13(11):797–806. doi:10.1093/molehr/gam063
53. Harchegani AB, Shafaghatian H, Tahmasbpour E, Shahriary A. Regulatory functions of MicroRNAs in male reproductive health: a new approach to understanding male infertility. Reprod Sci. 2018. doi:10.1177/1933719118765972
54. Rajender S, Meador C, Agarwal A. Small RNA in spermatogenesis and male infertility. Front Biosci (Schol Ed). 2012;4:1266–1274. doi:10.2741/s330
55. Hever A, Roth RB, Hevezi P, et al. Human endometriosis is associated with plasma cells and overexpression of B lymphocyte stimulator. Proc Natl Acad Sci. 2007;104(30):12451–12456. doi:10.1073/pnas.0703451104
56. Eyster KM, Klinkova O, Kennedy V, Hansen KA. Whole genome deoxyribonucleic acid microarray analysis of gene expression in ectopic versus eutopic endometrium. Fertil Steril. 2007;88(6):1505–1533. doi:10.1016/j.fertnstert.2007.01.056
57. Litwin M, Szczepańska‑Buda A, Piotrowska A, Dzięgiel P, Witkiewicz W. The meaning of PIWI proteins in cancer development. Oncol Lett. 2017;13(5):3354–3362. doi:10.3892/ol.2017.5932
58. Gong J, Zhang Q, Wang Q, et al. Identification and verification of potential piRNAs from domesticated yak testis. Reproduction. 2018;155(2):115–125. doi:10.1530/REP-17-0592
59. Juliano C, Wang J, Lin H. Uniting germline and stem cells: the function of piwi proteins and the piRNA pathway in diverse organisms. Annu Rev Genet. 2011;45:447–469. doi:10.1146/annurev-genet-110410-132541
60. Tan Y, Liu L, Liao M, et al. Emerging roles for PIWI proteins in cancer. Acta Biochim Biophys Sin. 2015;47(5):315–324. doi:10.1093/abbs/gmv018
61. Thomson T, Lin H. The biogenesis and function PIWI proteins and piRNAs: progress and prospect. Annu Rev Cell Dev Biol. 2009;25:355. doi:10.1146/annurev.cellbio.24.110707.175327
62. Klattenhoff C, Theurkauf W. Biogenesis and germline functions of piRNAs. Development. 2008;135(1):3–9. doi:10.1242/dev.006486
63. Clark JP, Lau NC. Piwi proteins and piRNAs step onto the systems biology stage. In: Systems Biology of RNA Binding Proteins. Springer; 2014:159–197.
64. Qiu L, Xu L, Chang G, et al. DNA methylation-mediated transcription factors regulate Piwil1 expression during chicken spermatogenesis. J Reprod Dev. 2016;62(4):367–372. doi:10.1262/jrd.2016-003
65. Bak CW, Yoon T-K, Choi Y. Functions of PIWI proteins in spermatogenesis. Clin Exp Reprod Med. 2011;38(2):61–67. doi:10.5653/cerm.2011.38.2.61
66. Heyn H, Ferreira HJ, Bassas L, et al. Epigenetic disruption of the PIWI pathway in human spermatogenic disorders. PLoS One. 2012;7(10):e47892. doi:10.1371/journal.pone.0047892
67. Lim SL, Qu ZP, Kortschak RD, et al. Correction: HENMT1 and piRNA stability are required for adult male germ cell transposon repression and to define the spermatogenic program in the mouse. PLoS Genet. 2015;11(12):e1005782.
68. Ma X, Wang S, Do T, et al. Piwi is required in multiple cell types to control germline stem cell lineage development in the Drosophila ovary. PLoS One. 2014;9(3):e90267. doi:10.1371/journal.pone.0090267
69. Assumpção CB, Calcagno DQ, Araújo TMT, et al. The role of piRNA and its potential clinical implications in cancer. Epigenomics. 2015;7(6):975–984. doi:10.2217/epi.15.37
70. Aravin AA, Sachidanandam R, Bourc’his D, et al. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell. 2008;31(6):785–799. doi:10.1016/j.molcel.2008.09.003
71. Aravin AA, Sachidanandam R, Girard A, Fejes-Toth K, Hannon GJ. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science. 2007;316(5825):744–747. doi:10.1126/science.1142612
72. Aravin AA, Hannon GJ, Brennecke J. The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science. 2007;318(5851):761–764. doi:10.1126/science.1146484
73. Brennecke J, Aravin AA, Stark A, et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell. 2007;128(6):1089–1103. doi:10.1016/j.cell.2007.01.043
74. Carmell MA, Girard A, van de Kant HJ, et al. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev Cell. 2007;12(4):503–514. doi:10.1016/j.devcel.2007.03.001
75. Williams Z, Morozov P, Mihailovic A, et al. Discovery and characterization of piRNAs in the human fetal ovary. Cell Rep. 2015;13(4):854–863. doi:10.1016/j.celrep.2015.09.030
76. Guo R, Yu Z, Guan J, et al. Stage‐specific and tissue‐specific expression characteristics of differentially expressed genes during mouse spermatogenesis. Mol Reprod Dev. 2004;67(3):264–272. doi:10.1002/(ISSN)1098-2795
77. Kuramochi-Miyagawa S, Kimura T, Ijiri TW, et al. Mili, a mammalian member of piwi family gene, is essential for spermatogenesis. Development. 2004;131(4):839–849. doi:10.1242/dev.00973
78. Deng W, Lin H. Miwi, a murine homolog of piwi, encodes a cytoplasmic protein essential for spermatogenesis. Dev Cell. 2002;2(6):819–830. doi:10.1016/S1534-5807(02)00165-X
79. Malki S, van der Heijden GW, O’Donnell KA, Martin SL, Bortvin A. A role for retrotransposon LINE-1 in fetal oocyte attrition in mice. Dev Cell. 2014;29(5):521–533. doi:10.1016/j.devcel.2014.04.027
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
