Back to Journals » International Journal of Nanomedicine » Volume 13
The effects of amine-modified single-walled carbon nanotubes on the mouse microbiota
Authors Mulvey JJ, Littmann ER, Ling L, McDevitt MR, Pamer EG, Scheinberg DA
Received 18 March 2018
Accepted for publication 12 June 2018
Published 10 September 2018 Volume 2018:13 Pages 5275—5286
DOI https://doi.org/10.2147/IJN.S168554
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Thomas Webster
J Justin Mulvey,1,2 Eric R Littmann,3 Lilan Ling,3 Michael R McDevitt,4,5 Eric G Pamer,3,5 David A Scheinberg1,3,5
1Department of Laboratory Medicine, Weill Cornell Medicine, New York, NY, USA; 2Department of Molecular Molecular Pharmacology, Memorial Sloan Kettering Cancer Center, New York, NY, USA; 3Department of Medicine, Memorial Sloan Kettering Cancer Center, New York, NY, USA; 4Department of Radiology, Memorial Sloan Kettering Cancer Center, New York, NY, USA; 5Weill Cornell Medicine, New York, NY, USA
Background: Amine-modified carbon nanotubes are drug delivery platforms with great potential that have not yet been applied in human clinical trials. Although modified nanotube vectors have the ability to carry multiple effectors, targeting agents, and even wrapped RNA, reports on unmodified, insoluble carbon nanotubes have highlighted inflammation in organs, including the intestine, with disruption of its resident microbiota. Disruption of the microbiota may allow for colonization by pathogenic bacteria, such as Clostridoidies difficile, stimulate immunoinfiltrates into the lamina propria or alter the absorption of therapeutics. Most proposed nanotube drugs are soluble, modified structures that are administered parenterally, and the majority of these soluble macromolecules are renally excreted; however, some are released into the bile, gaining access to the gastrointestinal tract.
Methods: Using environmentally isolated BALB/C mice in oral and intraperitoneal dosing models, high dose (3.80 or 4.25 mg/week), we administered amine-modified, soluble carbon nanotubes for 7 or 8 weeks. The general health and weight of the mice were monitored weekly, and upon killing, the diversity and content of their colonic, cecal, and ileal microbiota were assessed using shotgun 16S DNA sequencing.
Results and conclusion: We show that while oral administration at suprapharmacological doses modestly altered the α- and β-diversity of the mouse microbiome, these changes did not result in observed changes in clinical end points. Intraperitoneally-dosed mice exhibited none of the toxicities assessed.
Keywords: SWCNT, toxicity, 16S sequencing, nanopharmaceuticals
Introduction
Despite being excellent antioxidants due to their uniform sp2 bond structure,1 when inhaled in their unmodified state, fibrillar, insoluble, single-walled carbon nanotubes (SWCNTs) have been associated with inflammation and granuloma formation in the lungs.2,3 They have recently been shown to inflame the intestines of mice and disrupt resident microbiota.4 In contrast, highly soluble, SWCNTs achieved through modification can avoid this toxicity, be biocompatible, and be excreted rapidly.5–9 By combining favorable pharmacokinetic properties with the ability to carry numerous, diverse payloads simultaneously, NTs have been proposed as highly effective platforms for drug delivery.10 Better standardization of length, diameter, degree of imperfection, chirality, and adduct density are continued goals of the field.11,12 Formal toxicity studies are also needed. In this paper, we explore the effects of suprapharmacological intraperitoneal (IP) and oral (PO [per os]) dosing of amine-modified, SWCNTs on bacterial populations of rectal, cecal, and ileal contents in adult mice, in addition to general assessments of health. Closely related multiwalled, unmodified CNTs have also shown inflammation-related toxicity,13 but are larger and more heterogeneous, making them less suitable candidates for drug development. Multiwalled NTs are not included in this investigation.
The mouse digestive tract is the most widely used experimental model for the microbiome, and has led to better understanding of many human pathologies, including obesity,14 inflammatory bowel disease,15 and diabetes.16 However, the mouse digestive tract differs in some significant ways from its human counterpart. Mice have taller villi, larger ceca, decreased rectal goblet cells, and no appendix or haustra, all of which can influence biogeography.17–19 Morphological variation combined with disparate mucosal immunity and diet and frequent coprophagy nurtures a murine microbiota that differs from that of humans in content and diversity.20 Although murine intestines are not perfect models for the human intestine, bacterial communities react similarly to many environmental stressors, allowing for mice as models for microbiota investigations.21
When the microbiota is perturbed by an antibiotic, inflammation, pharmaceutical, or ingested bacteria, the general trends of loss of diversity and the elimination of certain subgroups are often translatable.21,22 For instance, the presence of Clostridium scindens and C. populeti have been shown to be protective factors against C. difficile infections in both mice and humans, while Enterococcus avium encourages the expansion of C. difficile.23 It has generally been observed that diversity is a sign of gut health that limits inflammation and resists domination by pathogenic organisms.24
Most studies on the ability of medications to alter the microbiota have focused on antibiotics, which by their nature can rapidly and drastically alter intestinal populations. However, many compounds make changes to the flora.25,26 The converse is also true: the effects of medications, their uptake, and their metabolism can vary greatly based on the microbiota.27 Here, we explore microbiota changes related to amine-modified SWCNTs, a class of molecules that is under development for a variety of biomedical applications. More specifically, this study will employ lysine-modified SWCNTs (SWCNT-Lys-NH2).7 The effect of these positively charged, primary-amine-coated macromolecules on the microbiota has yet to be examined. This study is more generally an examination of the effect of a large cationic amine load.
While the vast majority of systemically delivered amine-modified NTs are excreted through the kidneys, a small population is released into the bile and thus into the gastrointestinal (GI) tract via a mechanism discovered by this group.28 Distribution, elimination, and toxicity studies have been performed using amine-modified NTs; however, no study detailing the intestinal uptake and bioavailability of PO-administered tubes has been reported in the literature. Though NTs may move from the circulation to the gut, it remains unknown whether the reverse is true.
In this study, PO and systemic routes of NT administration are examined, followed by the collection and analysis of bowel contents from three different bio-geographical regions of the digestive tract. Both routes of administration are important. Most proposed NT-based drugs are administered either intravenously or IP, as bioavailability is unknown and NT-linked adducts may not survive the stomach. Toxicity by systemic administration methods is of greater interest; however, the creation of PO NT-based drugs is a distinct possibility. Furthermore, PO dosing provides far higher amounts of NTs to the GI tract and interacts with the esophagus, stomach, and duodenum, which are avoided in systemic administration. Sequencing three sections of the bowel, each with distinct microbiota populations, allows for more precision in how SWCNT-Lys-NH2 may affect bacterial communities.
Methods
Production of lysine-modified SWCNT solutions
SWCNT-Lys-NH2 were produced using electrocyclic chemistry29 as previously described.7 The NTs had an average length of ~400 nm and an average diameter of 2–3 nm, including adducts (Figure 1), with an average primary amine density of one amine per 121 carbons (~275 amines per tube) or about one amine for every 1,455 Da. All tubes were solvent exchanged by dialysis30 into a 25% methanol–H2O solution and then frozen before being lyophilized to a brown powder and dissolved into either sterile water or sterile saline for use in this study. pH was then adjusted to ~7.4 using Sigma-Aldrich pH-indicator strips (7.2–8.8 range) and HCl or NaOH. Sterile water was reverse osmosis, autoclaved water.31 Solutions were made freshly using lyophilized SWCNT-Lys-NH2 with each set of injections and each cage-water replacement.
  | Figure 1 Model of lysine-modified nanotubes. |
Mice
BALB/c mice were purchased from Jackson Laboratories, mated, and females housed individually once confirmed pregnant in pathogen-isolated cages with irradiated food and distilled water. Selected healthy pups born within 72 hours of one another were harmonized to their median birth date. Adolescents were separated into experimental group cages as soon as they were weaned and after 3 days of cohousing with the other broods for randomization and microbiota harmonization. Physical allocation of same-sex isobiotic BALB/c experimental groups occurred before male sexual maturity. Mouse handling and weekly cage changes were performed by investigators wearing sterile gowns, masks, and gloves in a sterile biosafety hood. All animals were maintained in a specific pathogen-free facility at the Memorial Sloan Kettering Cancer Center Animal Resource Center. All experiments were approved by the facility’s Institutional Animal Care and Use Committee and performed according to their approved protocols.
Experimental groups
Experimental groups were designed for 38 healthy pups from 42 littered by 10 mothers (Table 1). Once adolescents, nine groups of pups were divided into nine cages in sets of four to five mice based on sex, planned exposure type, and day of life of first experimental exposure. Female mice were divided into groups 1–6, consisting of three PO-dosing groups and three IP dosing groups. Male mice were divided into groups 7–9 and dosed IP only. Delivery of SWCNT-Lys-NH2 by either PO or IP injection methods were started on day 23 or day 30 of life, depending on the group, to examine the effect of stage of development on the introduction of NTs. All mice received irradiated food, and all mice except those in the OS-dosing experimental groups received sterile water, in keeping with microbiome best research practices.31 All mice were weighed in milligrams rounded to the nearest centigram from day 23 of life until the end of the experiment.
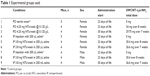  | Table 1 Experimental groups used |
Mice in the PO-dosing experimental groups were housed with sterile water that had been inoculated with SWCNT-Lys-NH2 to a molality of 0.125 g/L. PO solution concentration and dosing were based on a consumption experiment described in Methods section. PO dose was calculated under the assumption that the total volume consumed was divided equally by cagemates, meaning they each drank on average 34 mL tube water per week for a total of 4.25 mg per week.
The mice in IP injection control groups 4 and 7 were injected IP with 200 μL sterile saline once every 7 days from 23 days of life. Mice in IP injection experimental groups 5, 6, 8, and 9 were injected IP with 200 μL saline solution of Lys-modified NTs at a molality of 19 g/L for a total of 3.8 mg per injection from either 23 or 30 days of life. IP dosing was based on an unpublished toxicity dose escalation study and was a third of the morbidity threshold.
Validation of oral-dosing quantity and concentration
In a planning experiment, two sets of four adult female BALB/c mice each were housed in cages where the only source of liquid was a sterile water solution of SWCNT-Lys-NH2 at 0.125 g/L concentration. This was compared with two additional control cages that were given sterile water. All mice were allowed to imbibe ad libitum. The amount of fluid (sterile water or Lys-NT solution) consumed or wasted in the process of drinking was measured every 5 days for 30 days. The difference between cages never exceeded 16%. Stools were assessed qualitatively for gross changes, including liquidity, and no changes were found in any of the four cages. After the allotted 30 days, the mice were weighed. The groups had mean weights within 10% of one another when normalized for starting mean weight.
Sample collection
After 7 or 8 weeks of NT exposure depending on the group (in the 12th week of life), fresh stools were collected in a sterile manner from the three mice with weights closest to the cage mean using a flash-freeze technique that was proven on these mice by pilot sequencing experiments on feces. Subsequently, the mice were killed and the contents of their ceca and ilea extruded in a sterile manner, then flash-frozen. Stool, cecum, and ileum samples were then prepared for sequencing. Liver and kidneys were removed from all female mice, briefly dabbed on gauze, and weighed at time of death.
DNA extraction
Briefly, a frozen aliquot (~100 mg) of each sample was suspended while frozen in a solution containing 500 μL extraction buffer (200 mM Tris, pH 8/200 mM NaCl/20 mM EDTA), 200 μL 20% SDS, 500 μL phenol:chloroform:isoamyl alcohol (25:24:1), and 500 μL 0.1 mm-diameter zirconia/silica beads (BioSpec Products). Microbial cells were lysed by mechanical disruption with a bead beater (BioSpec Products) for 2 minutes, after which two rounds of phenol:chloroform:isoamyl alcohol extraction were performed. DNA was precipitated with ethanol and resuspended in 50 μL TE buffer with 100 μg/mL RNase. Isolated DNA was subjected to additional purification with QiaAmp mini spin columns (Qiagen).
16S rDNA amplification and Illumina sequencing
For each sample, duplicate 50 μL PCRs were performed, each containing 50 ng purified DNA, 0.2 mM dNTPs, 1.5 mM MgCl2, 2.5 U platinum Taq DNA polymerase, 2.5 μL 10× PCR buffer, and 0.5 μM each primer designed to amplify the V4–V5:563F (5′-nnnnnnnn-NNNNNNNNNNNN-AYTGGGYDTAAAGNG-3′) and 926R (5′-nnnnnnnn-NNNNNNNNNNNN-CCGTCAATTYHTTTRAGT-3′). A unique 12-base Golay barcode (Ns) preceded the primers for sample identification,33 and one to eight additional nucleotides were placed in front of the barcode to offset the sequencing of the primers. Cycling conditions were 94°C for 3 minutes, followed by 27 cycles of 94°C for 50 seconds, 51°C for 30 seconds, and 72°C for 1 minute; 72°C for 5 minutes was used for the final elongation step. Replicate PCRs were pooled, and amplicons were purified using the QiaQuick PCR purification kit (Qiagen). PCR products were quantified and pooled at equimolar amounts before Illumina barcodes and adaptors were ligated on using the Illumina TruSeq sample preparation protocol. The completed library was sequenced on an Illumina MiSeq platform following Illumina-recommended procedures with a paired-end 250×250 bp kit.
Sequence analysis
The 16S (V4–V5) paired-end reads were merged and demultiplexed. The Uparse pipeline32 was used to33 perform error filtering, using maximum expected error of 1,32,34 group sequences into operational taxonomic units (OTUs) of 97% distance-based similarity,34 and identify and remove potential chimeric sequences using both de novo and reference-based methods. Prior to clustering, singleton sequences were removed as part of the protocol. In order to gauge the impact of singleton removal on resulting sequences and diversity measures, the analysis was repeated with singletons retained (“Uparse +1” algorithm). Taxonomic assignment to species level was performed for representative sequences from each OTU. This was achieved by using a custom Python script incorporating nucleotide BLAST,35 with NCBI RefSeq36 as the reference training set. We used a minimum E value threshold of 1E–10 for assignments. Sequence designations and identity scores were inspected manually for quality and consistency in terms of taxonomic structure and secondary matches. A phylogenetic tree of OTU-representative sequences was constructed using QIIME. α-Diversity and UniFrac distances (inverse Simpson plots) were calculated using the PhyloSeq R package.37 All statistical analyses were performed using R version 3.2.1. Graphics were made using the GGplot2 package.38 We also made comparisons using linear discriminant analysis effect size (LEfSe): the difference between bacterial groups based on cladogram-based UniFrac distance, operational taxonomic units, quantity, and specific-gene presence.39 One sample from the male IP control group (group 7) did not reach minimum coverage and was excluded from analysis.
Results
Deaths
One mouse from the female sterile saline injection control group (group 4) died rapidly after IP injection on day 58 of life. The mouse was excluded from analysis. This was likely a result of large-vessel trauma during the injection.
Gross toxicity
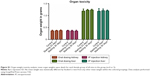
Gross toxicity was assessed by total mouse weight and comparison of liver and kidney weights using all mice in each female group. When results were separated by sex, there were no significant differences between any of the male or female groups by Student’s t-test (Figure 2). Whole-organ liver and kidney weights were compared for the female mice. Organ weights between treatment groups of mice also were not statistically significantly different (Figure S1).
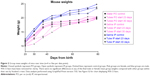  | Figure 2 Group mean weights of mice over time (n=4 or five per data point). |
Microbiota diversity
α-Diversity comparisons were segregated by sex and exposure type. Analyses employed inverse Simpson to determine α-diversity and Kruskal–Wallace to determine significance. IP female groups (4–6) and IP male groups (7–9) had no significant differences in diversity between them for any part of the bowel. IP injection group α-diversity with Kruskal–Wallis-derived P-values may be found in Table S1.
Female PO groups had significant differences between experimental and relevant control groups. There was a significant reduction in diversity in the cecum of mice receiving PO SWCNT-Lys-NH2 P23 (group 2) when compared with PO sterile water (group 1), but no significant change in colon or ileum diversity (Figure 3). There was also a significant reduction in diversity in the colons of mice receiving PO SWCNT-Lys-NH2 P30 (group 3) when compared with PO sterile water (group 1). Female PO group α-diversity with Kruskal–Wallis-derived P-values may be found in Table 2.
  | Table 2 α-Diversity of PO groups |
Differences in bacterial content
Principal coordinate analysis (PCoA) employing UniFrac distances is a way of clustering bacteria by the degree of their many disparate features, including phylogenetic differences. This is a measure of β-diversity. UniFrac PCoA analysis is provided for each sequenced female sample, organized by section of the bowel, and a second PCoA analysis separated by intervention type (ie, SWCNT-Lys-NH2 delivery or lack thereof) is also provided in Figure S3.
When the PCoA colored by bowel section was compared to the PCoA separated by intervention type, there was clustering of bacterial contents taken from the ceca and colons of mice as being separate from contents taken from their ilea, but there was no such clustering between intervention type and controls. If differences in colon contents between mice given PO SWCNT-Lys-NH2 and their controls were great enough, the specimens from these groups would cluster together. However, examination of all samples by biogeography failed to cluster the samples. This means that the introduction of SWCNT-Lys-NH2 did not affect the microbiota drastically enough to make it distinct from controls by either IP or PO administration methods. In other words, the differences between experimental groups and controls were not great enough to overshadow the regional differences.
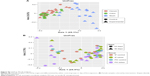
In order to examine smaller effects, when the female PO groups (1–3), female IP groups (4–6), and male IP groups (7–9) were analyzed by region of the bowel (Figure 4), subtle microbiota differences became evident. Mirroring the α-diversity differences, there was tight clustering of PO control cecal samples compared with experimental groups 2 and 3. There were no other tight sample clustering that separated groups. There was a regional association of IP control cecal samples compared with experimental groups 5 and 6, but identification of this as a notable difference in β-diversity would require a higher power study. This regional association was not true of IP-injected male counterparts, also shown in Figure 4.
  | Figure 4 PCoA plots of all samples divided into sets of groups – A (1–3), B (4–6), and C (7–9) – and separated by biogeography. |
UniFrac PCoA and inverse Simpson analysis measure group differences as a whole and may overlook clinically significant individual differences, such as increases in pathogenic species. For this reason, genus- and species-level bacterial content percentage plots along with LEfSe analyses were compared. LEfSe analyses in part reflect manyfold differences in the abundance of bacterial phylogenetic subgroups, and do not equate to gross content. No LEfSe differences in known pathogenic bacteria were identified between any of the groups.
Discussion
This is a first-reported attempt to understand the effect of amine-modified CNTs on the resident bacteria of the intestines. Three different intestinal regions, differences in sex of experimental mice, and time of first exposure to SWCNT-Lys-NH2 were additional variables in this analysis. SWCNT-Lys-NH2 are highly stable, large, polycationic amine structures unlike any naturally ingested material in terms of size and charge capacity. There was reason to suspect these structures might affect the microbiota. Ammonia cations are excellent antimicrobials, and so are polycationic polymers used as surface materials.40 Cationic antimicrobial peptides, such as neutrophil-generated indolicidin, have the potential to inhibit calmodulin,41 of which bacteria have analogs.42
The effects of prolonged high-dose PO administration of SWCNT-Lys-NH2 on microbiota were nontoxic by our metrics, with the exception of decreased diversity of bacteria in the ceca and colons of some female mice receiving PO tubes compared with controls. IP-dosed females did not display reportable differences in α- or β-diversity by our measures. PCoA UniFrac analysis showed that changes in microbiota of treated mice were dwarfed by the natural difference in microbiota in different biogeographical regions of the intestine. In mice with significantly perturbed microbiota, the level of change in α- and β-diversity was more prominent than regional differences. When small group-content differences were examined, the clustering of PO female control cecal samples reinforced that experimental groups were different from controls, if only in subtle ways. Specific increases and decreases in bacterial content as assessed by LEfSe analysis were not revealing for known pathogenic bacteria in any case.
The features of the SWCNT-Lys-NH2 that caused the changes in diversity were not investigated, but could have been size, charge, electron-scavenging potential, or an interaction with bacterial surface membranes or receptors. Importantly, there were no gross changes in the weight or observed health of the mice in any treated group when compared with untreated controls, including the PO groups. Mice injected with SWCNT-Lys-NH2 IP did not demonstrate significantly altered microbiota compared to controls by inverse Simpson, LEfSe, or PCoA UniFrac analysis. Secondary forms of toxicity related to the microbiome, such as susceptibility to infection, were not assessed. Our results stand as a counterpoint to resent research by Chen et al4 that employed unmodified SWCNTs and showed intestinal inflammation and significant microbiota alterations. Our study used a weekly total NT dose that was more than 10-fold the weekly dose used in Chen et al’s highest dosed mice, and a total dose close to 100-fold. Another recent study43 examined the microbiota toxicity of PO-dosed, unmodified multiwalled CNTs. This study did not show toxicity to the microbiome by PCoA analysis, but was conducted on a different variety of NTs at weekly NT doses that were a tenth of the doses used in Chen et al, and conducted with half the total dose.
Although this manuscript used small data sets, mice in experimental groups were exposed to suprapharmacological levels of NTs, >1.5 g/kg, over the course of the experiment. These doses are substantially beyond the proposed doses of modified CNTs as drug delivery vehicles, and should increase the sensitivity and specificity of detecting microbiota alterations or gross toxicity in experimental groups. Importantly, mice in these experimental groups showed no effect on their gross weight, organ weights, or symptom-assessed well-being.
The time points for first NT administration were chosen in order to interrogate whether SWCNT-Lys-NH2 affected development. A time point of 23 days was as close to possible to the date of weaning when the mouse was still truly an adolescent. Differences between mice that received first administration at 23 days and those that received first administration at 30 days of life were found to be insignificant in these experiments. The number of male and female groups were determined by the availability of pups, of which many more were female.
Conclusion
Parenterally delivered amine-modified NTs were nontoxic at the dosage and time frames investigated and did not have a major effect on the microbiota. PO-administered NTs may perturb the α- and β-diversity of the microbiota in the lower-GI tract but were also nontoxic at the levels and time frames investigated.
Disclosure
The authors report no conflicts of interest in this work.
References
Lucente-Schultz RM, Moore VC, Leonard AD, et al. Antioxidant single-walled carbon nanotubes. J Am Chem Soc. 2009;131(11):3934–3941. | ||
Brown DM, Kinloch IA, Bangert U, et al. An in vitro study of the potential of carbon nanotubes and nanofibres to induce inflammatory mediators and frustrated phagocytosis. Carbon N Y. 2007;45(9):1743–1756. | ||
Bhattacharya K, Andón FT, el-Sayed R, Fadeel B. Mechanisms of carbon nanotube-induced toxicity: focus on pulmonary inflammation. Adv Drug Deliv Rev. 2013;65(15):2087–2097. | ||
Chen H, Zhao R, Wang B, et al. Acute oral administration of single-walled carbon nanotubes increases intestinal permeability and inflammatory responses: association with the changes in gut microbiota in mice. Adv Healthc Mater. 2018;7(13):e1701313. | ||
Mulvey JJ, Villa CH, Mcdevitt MR, Escorcia FE, Casey E, Scheinberg DA. Self-assembly of carbon nanotubes and antibodies on tumours for targeted amplified delivery. Nat Nanotechnol. 2013;8(10):763–771. | ||
Ruggiero A, Villa CH, Bander E, et al. Paradoxical glomerular filtration of carbon nanotubes. Proc Natl Acad Sci U S A. 2010;107(27):12369–12374. | ||
Mulvey JJ, Feinberg EN, Alidori S, Mcdevitt MR, Heller DA, Scheinberg DA. Synthesis, pharmacokinetics, and biological use of lysine-modified single-walled carbon nanotubes. Int J Nanomedicine. 2014;9:4245–4255. | ||
Ali-Boucetta H, Nunes A, Sainz R. Asbestos-like pathogenicity of long carbon nanotubes alleviated by chemical functionalization. Angew Chemie Int Ed Engl. 2013;52(8):2274–2278. | ||
Battigelli A, Ménard-Moyon C, da Ros T, Prato M, Bianco A. Endowing carbon nanotubes with biological and biomedical properties by chemical modifications. Adv Drug Deliv Rev. 2013;65(15):1899–1920. | ||
Tibbitt MW, Dahlman JE, Langer R. Emerging frontiers in drug delivery. J Am Chem Soc. 2016;138(3):704–717. | ||
Liu H, Nishide D, Tanaka T, Kataura H. Large-scale single-chirality separation of single-wall carbon nanotubes by simple gel chromatography. Nat Commun. 2011;2:309. | ||
Sanchez-Valencia JR, Dienel T, Gröning O, et al. Controlled synthesis of single-chirality carbon nanotubes. Nature. 2014;512(7512):61–64. | ||
Zhou L, Forman HJ, Ge Y, Lunec J. Multi-walled carbon nanotubes: a cytotoxicity study in relation to functionalization, dose and dispersion. Toxicol In Vitro. 2017;42:292–298. | ||
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444(7122):1027–1131. | ||
Wlodarska M, Kostic AD, Xavier RJ. An integrative view of microbiome-host interactions in inflammatory bowel diseases. Cell Host Microbe. 2015;17(5):577–591. | ||
Livanos AE, Greiner TU, Vangay P, et al. Antibiotic-mediated gut microbiome perturbation accelerates development of type 1 diabetes in mice. Nat Microbiol. 2016;1(11):16140. | ||
Linnenbrink M, Wang J, Hardouin EA, Künzel S, Metzler D, Baines JF. The role of biogeography in shaping diversity of the intestinal microbiota in house mice. Mol Ecol. 2013;22(7):1904–1916. | ||
Donaldson GP, Lee SM, Mazmanian SK. Gut biogeography of the bacterial microbiota. Nat Rev Microbiol. 2016;14(1):20–32. | ||
Stearns JC, Lynch MD, Senadheera DB, et al. Bacterial biogeography of the human digestive tract. Sci Rep. 2011;1:170. | ||
Xiao L, Feng Q, Liang S, et al. A catalog of the mouse gut metagenome. Nat Biotechnol. 2015;33(10):1103–1108. | ||
Becattini S, Taur Y, Pamer EG. Antibiotic-induced changes in the intestinal microbiota and disease. Trends Mol Med. 2016;22(6):458–478. | ||
Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R. Diversity, stability and resilience of the human gut microbiota. Nature. 2012;489(7415):220–230. | ||
Buffie CG, Bucci V, Stein RR, et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature. 2015;517(7533):205–208. | ||
Buffie CG, Pamer EG. Microbiota-mediated colonization resistance against intestinal pathogens. Nat Rev Immunol. 2013;13(11):790–801. | ||
Devkota S. Prescription drugs obscure microbiome analyses. Science. 2016;351351(6272):452–453. | ||
Maier L, Pruteanu M, Kuhn M, et al. Extensive impact of non-antibiotic drugs on human gut bacteria. Nature. 2018;555(7698):623–628. | ||
Servick K. Mouse microbes may make scientific studies harder to replicate. 2016. Available from: http://www.sciencemag.org/news/2016/08/mouse-microbes-may-make-scientific-studies-harder-replicate. Accessed July 25, 2018. | ||
Alidori S, Bowman RL, Yarilin D, et al. Deconvoluting hepatic processing of carbon nanotubes. Nat Commun. 2016;7:12343. | ||
Tsuge O, Kanemasa S. Recent advances in azomethine ylide chemistry. In: Katritzky AR, editor. Advances in Heterocyclic Chemistry. Vol 45. San Diego: Academic Press; 1989:231–349. | ||
Mulvey J, Feinberg E, Mcdevitt M, Scheinberg D. Dialytic separation of bundled, functionalized carbon nanotubes from carbonaceous impurities. Crystals. 2014;4(4):450–465. | ||
Dubé P. 2017 Microbiome in Mouse Models Workshop review – best practices. 2017. Available from: https://www.taconic.com/taconic-insights/microbiome-and-germ-free/2017-microbiome-workshop-review-part-three.html. Accessed July 25, 2018. | ||
Edgar RC. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods. 2013;10(10):996–998. | ||
Caporaso JG, Lauber CL, Walters WA, et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012;6(8):1621–1624. | ||
Edgar RC, Flyvbjerg H. Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics. 2015;31(21):3476–3482. | ||
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–410. | ||
Tatusova T, Ciufo S, Fedorov B, O’Neill K, Tolstoy I. RefSeq microbial genomes database: new representation and annotation strategy. Nucleic Acids Res. 2014;42(D1):D553–D559. | ||
Mcmurdie PJ, Holmes S. Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One. 2013;8(4):e61217. | ||
Wickham H. Ggplot2: Elegant Graphics for Data Analysis. Heidelberg: Springer; 2009. | ||
Segata N, Izard J, Waldron L, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12(6):R60. | ||
Carmona-Ribeiro AM, Carrasco LD. Cationic antimicrobial polymers and their assemblies. Int J Mol Sci. 2013;14(5):9906–9946. | ||
Sitaram N, Subbalakshmi C, Nagaraj R. Indolicidin, a 13-residue basic antimicrobial peptide rich in tryptophan and proline, interacts with Ca2+-calmodulin. Biochem Biophys Res Commun. 2003;309(4):879–884. | ||
Swan DG, Hale RS, Dhillon N, Leadlay PF. A bacterial calcium-binding protein homologous to calmodulin. Nature. 1987;329(6134):84–85. | ||
Christophersen DV, Jacobsen NR, Andersen MH, et al. Cardiovascular health effects of oral and pulmonary exposure to multi-walled carbon nanotubes in ApoE-deficient mice. Toxicology. 2016;371:29–40. |
Supplementary materials
  | Table S1 α-Diversity of IP injection groups |
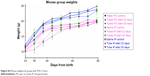  | Figure S2 Mouse weights by group with 95% CI bars. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.



