Back to Journals » Drug Design, Development and Therapy » Volume 15
Temozolomide-Induced Changes in Gut Microbial Composition in a Mouse Model of Brain Glioma
Authors Li XC, Wu BS, Jiang Y, Li J, Wang ZF, Ma C, Li YR, Yao J , Jin XQ, Li ZQ
Received 23 December 2020
Accepted for publication 7 April 2021
Published 21 April 2021 Volume 2021:15 Pages 1641—1652
DOI https://doi.org/10.2147/DDDT.S298261
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Tuo Deng
Xiao-Chong Li,1,* Bang-Sheng Wu,1,2,* Yi Jiang,1,2 Jie Li,3 Ze-Fen Wang,3 Chao Ma,1 Yi-Rong Li,4 Jie Yao,5 Xiao-Qing Jin,6 Zhi-Qiang Li1
1Department of Neurosurgery, Zhongnan Hospital, Wuhan University, Wuhan, 430071, Hubei, People’s Republic of China; 2Second Clinical School, Wuhan University, Wuhan, 430071, Hubei, People’s Republic of China; 3Department of Physiology, Wuhan University School of Basic Medical Sciences, Wuhan, 430071, People’s Republic of China; 4Department of Clinical Laboratory, Zhongnan Hospital, Wuhan University, Wuhan, 430071, Hubei, People’s Republic of China; 5Department of Biological Repositories, Zhongnan Hospital, Wuhan University, Wuhan, 430071, Hubei, People’s Republic of China; 6Emergency Department, Zhongnan Hospital of Wuhan University, Wuhan, 430071, Hubei, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Xiao-Qing Jin; Zhi-Qiang Li Email [email protected]; [email protected]
Background: Gut microbiota is associated with the progression of brain tumors. However, the alterations in gut microbiota observed during glioma growth and temozolomide (TMZ) therapy remain poorly understood.
Methods: C57BL/6 male mice were implanted with GL261 glioma cells. TMZ/sodium carboxymethyl cellulose (SCC) was administered through gavage for five consecutive days (from 8 to 12 days after implantation). Fecal samples were collected before (T0) and on days 7 (T1), 14 (T2), and 28 (T3) after implantation. The gut microbiota was analyzed using 16S ribosomal DNA sequencing followed by absolute and relative quantitation analyses.
Results: Nineteen genera were altered during glioma progression with the most dramatic changes in Firmicutes and Bacteroidetes phyla. During glioma growth, Lactobacillus abundance decreased in the early stage (T1) and then gradually increased (T2, T3); Intestinimonas abundance exhibited a persistent increase; Anaerotruncus showed a transient increase (T2) and then a subsequent decrease (T3). Similar longitudinal changes in Intestinimonas and Anaerotruncus abundance were observed in TMZ-treated mice, but the decrease of Anaerotruncus at T3 in the TMZ-treated group was less than that in the vehicle-treated group. No significant change in Lactobacillus was observed after TMZ treatment. Additionally, compared to vehicle control, TMZ treatment led to an enrichment in Akkermansia and Bifidobacterium.
Conclusion: Glioma development and progression altered the composition of gut microbiota. Induction of Akkermansia and Bifidobacterium as well as the prevention of the reduction in Anaerotruncus may contribute to the anti-tumor effect of TMZ. This study helps to reveal the association between levels of specific microbial species in the gut and the anti-tumor effect of TMZ.
Keywords: glioma, gut microbiota, temozolomide, tumor microenvironment, epigenetics
Introduction
Glioma is the most common and lethal tumor in the central nervous system in adults.1 Despite significant progress in our understanding of the molecular mechanisms of oncogenesis and tumor progression as well as in clinical therapeutic management, the prognosis of glioma patients remains poor.2,3
It is now well established that gut microbiota may contribute to human health and various diseases. Since the gut microbiota affects inflammation and immunity not only locally at the mucosal level but also systemically, it stands to reason that the dysbiosis of gut microbiota should exist in various diseases, including brain disorders.4 Patients with Parkinson’s disease who suffer from constipation exhibit an altered gut microbiota with increased Enterobacteriaceae and decreased Prevotella, which is related to motor phenotype in patients.5 In individuals with Alzheimer’s disease, the altered microbiota is characterized by increased Bacteroidetes, decreased Firmicutes and Bifidobacterium.6 The bidirectional communication between the central nervous system and the gastrointestinal tract is well recognized, and is referred to as the brain-gut axis. Many studies have shown that gut microbiota modulates brain-gut axis interactions via neuroendocrine, immunological, and direct neural mechanisms, which gives rise to a more inclusive nomenclature – the brain-gut-microbiota axis.7 Therefore, disruption of the gut microbiota may affect the initiation and progress of brain disorders by altering the inflammatory and immune response in the brain. A recent study showed that the variation in gut microbial composition alters gut-immune-brain communication to form a tumor-tolerant microenvironment in the central nervous system, which promotes the development of glioma.8 However, the specific microorganisms related to the initiation and progress of glioma remain to be elucidated.9
Temozolomide (TMZ), an alkylating prodrug, is a commonly used chemotherapeutic agent in the standard Stupp protocol for glioma patients.10 Compared to surgical resection with radiotherapy (RT) alone, a combination of RT and TMZ increases the 2-year survival rate from 10% to 27%.11 Previous studies have indicated that the disruption of gut microbiota influenced the antitumor efficacy of chemotherapy by altering the tumor microenvironment,9,12–14 indicating that sensitive responses to chemotherapy require an optimal microbiota. The antitumor efficacy of cyclophosphamide, another prominent alkylating anticancer agent, is also influenced by intestinal microbiota.12,14 In addition, cyclophosphamide has been shown to alter the composition of intestinal microbiota in healthy mice.12 The result indicates that the microbiota dysbiosis induced by chemotherapeutic agents may also contribute to their antitumor effect. However, TMZ-induced microbiota alterations during glioma therapy have not been investigated.
Next-generation sequencing of bacterial 16S ribosomal DNA is commonly used to analyze differences in the composition of the microbiome. In most studies, relative abundance (the number of different microbial groups divided by the sum of groups in the sample) is used.15–17 However, relative approaches ignore the possibility of altered overall microbiota abundance.18,19 Increased relative abundance of a specific microbial taxa may be attributed to the reduced relative abundance of other microorganisms, rather than the increase in their absolute abundance.19 The relative abundance can be transformed into the absolute abundance using an absolute quantitative approach.20
In this study, we used both relative and absolute abundances to investigate the changes in gut microbiota during glioma growth and following TMZ treatment in a mouse model.
Materials and Methods
This study was carried out in strict accordance with the National Institute of Health Guidelines for the Care and Use of Laboratory Animals. The protocol was approved by the Institutional Animal Care and Ethics Committee of Wuhan University (No.2018117).
Reagents
TMZ and sodium carboxymethyl cellulose (SCC) were purchased from MedChemExpress (New Jersey, USA). TMZ was dissolved in 0.5% SCC at a final concentration of 5 mg/mL. Dulbecco’s modified Eagle’s medium (DMEM) and fetal bovine serum (FBS) were obtained from Gibco (California, USA). Chloral hydrate was from MOLBASE (Shanghai, China).
Animals and GL261 Cell Line
Specific-pathogen-free male C57BL/6 mice (n=12, 8–10 weeks) were purchased from SPF Biotechnology Co. Ltd. (Beijing, China). All mice were housed in the Animal Experimental Center of Zhongnan Hospital at Wuhan University where the temperature was maintained at 25 °C with a 12 h light/dark cycle and free access to food and water. After adaptive feeding for 7 days, mice were randomly allocated to two groups (vehicle- and TMZ-treated groups; n=6/group) and underwent intracranial GL261-cell implantation and subsequent vehicle or TMZ gavage.
The mouse glioblastoma cell line GL261 was obtained from the American Type Culture Collection (Rockville, MD) and cultured in DMEM supplemented with 10% FBS in a humidified 5% CO2 cell incubator at 37 °C.
GL261 Transplantation and TMZ Treatment
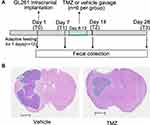
The procedure of intracranial GL261 implantation and TMZ treatment is shown in Figure 1A. After the first collection of fecal samples following a 7-day adaptive feeding regime (T0), the allogeneic orthotopic glioma model was constructed by implanting GL261 cells into the left putamen as described previously.21 Briefly, after anesthesia by intraperitoneal injection of 2% chloral hydrate (0.2 mL/10g) mice were fixed in stereotaxic apparatus (RWD Life Science, Shenzhen, China). A burr hole was made in the left parietal bone 0.1 mm posterior to the bregma and 2.3 mm lateral to the midline. GL261 cells (2.5×104 cells in 2 μL PBS) were administered stereotactically through the burr hole into the left putamen with a depth of 2.3 mm from the brain surface. The cell suspension was delivered slowly for 2–3 min. The injection needle was left in place for an additional 2 min and then slowly withdrawn. The burr hole was sealed with bone wax and the incision sutured. TMZ gavage started at 7 days after implantation (T1) with the dosage of 50 mg/kg/d for five consecutive days (days 8–12 after implantation). The TMZ solution was freshly prepared for gavage. The vehicle-treated mice received gavage of 0.5% SCC.
Hematoxylin/Eosin (HE) Staining of Brain Slices
All the mice were sacrificed at 28 days after implantation (T3). The brains were removed and post-fixed overnight in 4% paraformaldehyde at 4 °C, and then transferred into 30% sucrose solution in 0.1 mol/L PBS at 4 °C for 72 h. Coronal section serials were sliced at 15 μm by a Cryostat (Leica Microsystems AG, Germany) and mounted onto glass slides. Sections were stained with hematoxylin and eosin respectively, and then cleared in xylene and mounted.
Fecal Sample Collection and DNA Extraction
As shown in Figure 1A, the stool samples from the mice were collected at four time points: days 1 (the day of glioma cells implantation; T0), 7 (the day before TMZ gavage; T1), 14 (7 days after first TMZ gavage; T2), and 28 (21 days after first TMA gavage; T3). All samples were collected at 8 am. Mice were placed in a sterile plastic box after weighing to collect fecal samples. Samples were transferred into a cell cryopreserved tube using a sterile toothpick, frozen in liquid nitrogen, and then stored at −80 °C until DNA extraction. A total of 48 samples were collected, and bacterial genomic DNA was extracted by the FastDNATM Spin Kit for Soil (MP Biomedicals, USA) according to the manufacturer’s instructions. Extracted DNA was quantified using the Invitrogen Qubit 3.0 Spectrophotometer (Thermo Fisher Scientific, USA). Quality and integrity of the DNA were evaluated using a NanoDropTM 2000 (Thermo Fisher Scientific, USA) followed by 1.5% agarose gel electrophoresis. Isolated DNA was stored at −20 °C until processing.
Polymerase Chain Reaction (PCR)
The V3–V4 region of the bacterial 16S ribosomal DNA was amplified using the following primer pair: 341F (5′-CCTACGGGNGGCWGCAG-3′) and 805R (5′-GACTACHVGGGTATCTAATCC-3′). PCR was performed using the following thermal cycling conditions: 94 °C for 2 min followed by 25 cycles of 94 °C for 20 s, 55 °C for 30 s, and 72 °C for 1 min and a final extension at 72 °C for 5 min.
Spike-In Sequences and Absolute Quantification
A synthetic spike-in plasmid was used as internal reference to quantitatively detect the total number of bacterial 16S rRNA genes. Spike-in plasmids contain randomly arranged base sequences (length and GC content are consistent with the 16S rRNA gene sequencing fragment) and the two ends of the randomly arranged base sequences correspond to the sequences of primers targeting the 16S rRNA gene. Sample (10 ng/μL) and spike-in internal reference DNAs were used as templates for PCR.
Illumina MiSeq Sequencing
PCR amplified products were purified using equal volume of Agencourt AMPure XP PCR Purification Beads (Beckman Coulter, USA). The mixed-sample library was identified using the Agilent 2100 Bioanalyzer (Agilent Technologies, USA) to detect the size of the inserted fragments in the sequencing library (confirming no specific amplification between 120 and 200 bp), and the concentration of these sequencing inserts were quantified. The pooled products were sequenced using the Illumina HiSeq 2500 (Illumina, USA) with a 2×250 bp paired-end protocol according to the manufacturer’s instructions.
Sequencing Analysis
The obtained sequencing reads were quality-filtered using the following steps: (i) the end bases with mass less than 20, possible adapter sequences, and sequences shorter than 100 bp were removed (TrimGalore v0.4.5); (ii) the paired sequences from double-terminal sequencing were spliced, after which the merged low-quality sequences were removed (FLASH2 v2.2.00); (iii) the primer sequences and sequences containing N bases/homopolymer >6 bp were removed (mothur v.1.39.3); (iv) the sequences with a total base error rate of >2 and length <100 bp were removed (usearch v10).
Operational taxonomic units (OTUs) were clustered using a similarity cut-off value of 97%, and chimeric sequences were removed after singleton removal (usearch v10). During relative quantification (RQ) analysis, the spike-in sequence was removed. During absolute quantification (AQ) analysis, the absolute copy number of each OTU was calculated by standard curve of the spike-in sequences in each sample. Sequences were aligned to the RDP 16S rRNA database (trainset 16/release 11.5) with a credibility of >80% using mothur v.1.39.3. Most of the analyses were performed with both AQ and RQ data, but linear discriminant analysis effect size measurements (LEfSe) and Kruskal–Wallis test were performed only with RQ data.
Alpha-diversity was evaluated with Observed species, Chao 1, ACE, Shannon, and Simpson indices. Beta-diversity was evaluated by weighted UniFrac distance matrices and visualized by Principal Coordinate Analysis (PCoA) plots based on weighted UniFrac distance matrices. LEfSe was used to identify the representative microbiome in each group with a logarithmic LDA score threshold of 2 (python v2.7.14). Phylogenetic investigation of the communities by reconstruction of the unobserved states (PICRUSt) was used to predict the functional profiling of the microbiota using the Greengene database as reference (release 13.5). The Kyoto Encyclopedia of Genes and Genomes (KEGG) and Clusters of Orthologous Groups (COG) pathways were used to predict differences in biochemical pathways in the bacteria (PICRUSt v1.0).
Statistical Analysis
The differential genera and α-diversity indices were analyzed by Wilcoxon rank-sum test (R 3.4.3). The Adonis method (Permutational Multivariate Analysis of Variance Using Distance Matrices, PERMANOVA) was used to decompose the total variance and analyze the group-level differences in microbial communities. Alterations in microbial community at phylum and genus levels were analyzed by Kruskal–Wallis test, and changes in functional (COG/KEGG) abundance were evaluated by Welch’s t-test. P<0.05 was considered statistically significant.
Results
Gut Microbial Dysbiosis in Mice with Intracranial Glioma
The experimental procedure in this study was presented in Figure 1A. All mice were sacrificed at the endpoint of the experiment (T3), and the presence of intracranial glioma growth in all mice was confirmed by HE staining (Figure 1B). The stool samples from each mouse were collected longitudinally at each time point, which minimized the impact of inter-individual variations on the dynamic changes in bacterial communities.
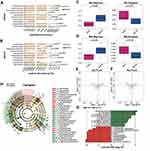
To reveal the changes in microbiota that occurred at the early stage of glioma growth, we performed a comparison of the samples before implantation (T0, n=12) versus 7 days after implantation (T1, n=12). Both relative and absolute abundance demonstrated decreased Lactobacillus (AQ<0.001, RQ<0.001) and increased Intestinimonas (AQ P=0.005; RQ P<0.001) and Anaerotruncus genera (AQ P=0.028; RQ P=0.005) in the glioma mice compared to naive mice (Figure 2A and B). Alpha-diversity analysis showed an increase in the uniformity of bacterial communities in the glioma mice, as demonstrated by the higher Shannon index (AQ P=0.006; RQ P=0.008) and the lower Simpson index (AQ P=0.039; RQ P=0.028, Figure 2C and D). PCoA exhibited differential clustering patterns between naive and glioma mice, indicating a clear separation of microbiome β-diversity (PERMANOVA, AQ pseudo-F=3.59, RQ pseudo-F=4.17; AQ P=0.020, RQ P=0.010, Figure 2E and F). LEfSe analysis confirmed the enrichment of Intestinimonas and Anaerotruncus in the glioma mice (Figure 2G and H). We further analyzed the alterations in microbial function profiles. PICRUSt analysis revealed that lipoic acid metabolism was downregulated in the glioma mice (P<0.05, Supplementary File 1).
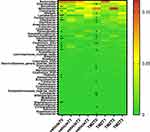
Next, to investigate the dynamic microbiota alterations that occurred during glioma progression, we analyzed the longitudinal changes of microbiota composition in vehicle-treated glioma mice (n=6). At the level of phyla, the abundance of Firmicutes and Bacteroidetes changed most significantly (Figure 3). During the progression of glioma, 19 genera of gut microbiota were altered in vehicle-treated mice (Figure 4, Supplementary File 2). Intestinimonas exhibited a persistent increase during glioma growth (RQ P=0.004, Figure 4). Although Lactobacillus levels were significantly reduced at 7 days after implantation (T1), they subsequently showed a gradual increase, exhibiting their greatest abundance at T3 (RQ P=0.001). Anaerotruncus displayed a transient increase at T2 and then a subsequent decrease at T3 (RQ P=0.009).
Short-Term Effects of TMZ on Gut Microbiota in Glioma Mice
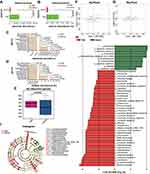
TMZ is one of the most commonly recommended chemotherapeutic regimens for glioma. Conventionally, TMZ is administered orally for five days followed by an interval of 22 days. Short-term effects of TMZ on gut microbiota were examined at 7 days after the first TMZ gavage (T2). The abundance of Verrucomicrobia phylum in TMZ-treated mice was significantly higher than that in the vehicle group (AQ, RQ P=0.002, Figure 5A and B). At the genus level, TMZ-treated mice exhibited a higher abundance of Akkermansia (AQ P=0.002; RQ P=0.002), Bifidobacterium (AQ P=0.005; RQ P=0.005), Coprobacillus (AQ P=0.009; RQ P=0.02), and Clostridium_XVIII (AQ P=0.004; RQ P=0.015), and a lower abundance of Coprobacter (AQ P=0.041; RQ P=0.004) than mice in the vehicle group (Figure 5C and D). However, α-diversity analysis showed no significant differences in microbiomes between the groups. PCoA analysis revealed that the microbiome in TMZ-treated mice differed from that in vehicle-treated mice (PERMANOVA, AQ pseudo-F=3.49, RQ pseudo-F=3.70; AQ P=0.002, RQ P=0.002, Figure 5E and F). LEfSe analysis showed that Bifidobacterium, Verrucomicrobia, and especially Akkermansia were highly enriched in the TMZ group (Figure 5G and H). PICRUSt showed that pathways involved in glutathione metabolism and fatty acid biosynthesis and metabolism were upregulated in TMZ-treated mice (P<0.05, Supplementary File 3).
Long-Term Effects of TMZ on Gut Microbiota in Glioma Mice
Next, we studied the long-term effects of TMZ on gut microbiota at 21 days after the first TMZ gavage (T3). The abundance of the Anaerotruncus genus increased in the TMZ group (AQ=0.041, RQ P=0.026, Figure 6A–D). Community diversity was higher in the TMZ group than that in the vehicle group using observed species index, but this difference was only found in AQ analysis (P=0.030, Figure 6E). Weighted UniFrac-based PCoA of β-diversity demonstrated significant differences in the overall microbial composition between TMZ and vehicle groups (PERMANOVA, AQ pseudo-F=2.67, RQ pseudo-F=2.77; AQ P=0.004, RQ P=0.004, Figure 6F and G). LEfSe analysis further confirmed this change in gut microbiota (Figure 6H and I). PICRUSt showed that the mineral absorption pathway was upregulated in the TMZ group compared to the vehicle group (P<0.05, Supplementary File 4).
Trend of Changes in Gut Microbiota After TMZ Treatment
To investigate the trend of microbiota alterations in glioma mice after TMZ treatment, we used the Kruskal–Wallis test to identify the differential genera in the TMZ group over the course of the entire experiment. Twenty genera were significantly different between the groups with major alterations in the Firmicutes and Bacteroidetes phyla (Figures 3 and 4, Supplementary File 5). The changes of Intestinimonas and Anaerotruncus abundance in TMZ-treated mice were similar to those in the vehicle group. Intestinimonas exhibited a constant increase during TMZ treatment (RQ P=0.01, Figure 4). The Anaerotruncus abundance peaked at T2 before a slight decrease at T3 (RQ P=0.031, Figure 4). However, it was noted that Anaerotruncus abundance in the TMZ group was still higher than that in the vehicle group at T3 (Figure 6C and D, Supplementary Files 2 and 5). The abundance of Bifidobacterium remained steady at T1 but increased at T2 and T3 (RQ P=0.031, Supplementary File 5). No significant change in Lactobacillus was observed in TMZ-treated mice.
Discussion
Gut microbiota is extremely important for the homeostasis of the gastrointestinal tract and proper functioning of the immune system. The dysbiosis of gut microbiota contributes to inflammation and the activation of the immune system in several pathologies.22 This study showed that both glioma growth and TMZ treatment altered the composition of gut microbiota, especially Firmicutes and Bacteroidetes.
The glioma-induced microbiota dysbiosis observed in this study included a persistent increase in Intestinimonas, a transient increase in Anaerotruncus, and a late-onset increase in Lactobacillus during glioma growth and progression. Lactobacillus is a probiotic that modulates the degradation of lipid and lactose. Previous studies have shown that Lactobacillus supplementation improves intestinal immune function,23 and promotes the recovery of patients with colorectal cancer after surgery.24 The increase of Lactobacillus abundance during the late stage of glioma growth (T2 and T3) may be attributed to an automatic anti-tumor response following tumor growth. The increase in abundance of Intestinimonas correlates with the biosynthesis of sesquiterpenoid and triterpenoid,25 compounds that have anti-tumor and antioxidant activities.26 Studies have shown the correlation between Anaerotruncus and the biosynthesis of steroids and terpenoids.25 Anaerotruncus also produces short-chain fatty acids to elicit a strong anti-tumor response by downregulating cell proliferation-related genes, stimulating pro-apoptotic pathways, and reducing intestinal inflammation.27,28 Therefore, the altered levels in gut bacterial communities may themselves exert anti-tumor activity through a variety of pathways. There is an increase in the uniformity of the microbial community in glioma mice, and this increase has also been observed in children with cerebral palsy and epilepsy.17 We found that lipoic acid metabolism was downregulated in glioma mice, which suppresses oxidative stress in astroglial cells.29
TMZ treatment also altered the gut microbiota in glioma mice. TMZ-treated mice exhibited similar changes in Intestinimonas and Anaerotruncus abundance as those seen in the vehicle group. However, no significant alteration in Lactobacillus was observed after TMZ treatment. The similar increase of Intestinimonas and Anaerotruncus in both TMZ and vehicle groups may be an innate anti-tumor response that occurs during the development of tumors. Anaerotruncus eventually decreased at T3 in both the TMZ and vehicle groups, but its abundance in the TMZ group was nevertheless higher than that in the vehicle group at T3, indicating that TMZ treatment could prevent the reduction of Anaerotruncus.
We noted that the microbiota alterations varied at different stages following TMZ treatment in comparison to the vehicle-treated group. At 7 days after the initial TMZ treatment (T2), the abundance of Akkermansia, Bifidobacterium, and Verrucomicrobia increased, and at 21 days after the treatment (T3), the abundance of Anaerotruncus increased. Increased Akkermansia has been observed in patients who respond actively to immunotherapeutic PD-1 blockade, pointing to a role in mediating tumor response to immunotherapy.30 The authors of that study also showed that transplantation of the fecal microbiome from patients who responded positively to PD-1 blockade into germ-free or antibiotic-treated mice ameliorated the anti-tumor response to the PD-1 antibody, whereas transplantation from nonresponding patients failed to do so.30 These results indicate a key role of gut microbiota in the tumor response to chemotherapy and immunotherapy.
Studies in mice have reported that Akkermansia is associated with glucose and lipid metabolism, thereby improving metabolic disorders.31 Administration of Bifidobacterium improved some symptoms in a mouse model of autoimmune disease by inducing regulatory T cells and regulating the release of inflammatory cytokines.32 The result indicates that Bifidobacterium has anti-inflammatory and immunomodulatory activities. Moreover, Bifidobacterium has the capability to produce folate,33 which is closely associated with the DNA methylation of O6-methylguanine DNA methyltransferase (MGMT).34 In this study, pathways involved in cofactor- and vitamin-metabolism were significantly upregulated after TMZ treatment. The status of MGMT promoter methylation is associated with the suppression of tumor proliferation and tumor response to TMZ therapy. Therefore, increased Bifidobacterium may ameliorate the therapeutic effect of TMZ by producing folate to induce MGMT promoter methylation. Additionally, we found that the pathways of glutathione and fatty acid metabolism were upregulated after TMZ treatment, indicating a correlation between TMZ, oxidative stress, and fatty acid levels.
There were several limitations in this study. First, the study was performed only with male mice. Whether the alterations seen in gut microbiota are sex-dependent remains unknown. Second, PICRUSt analysis is associated with specific drawbacks as compared to those that use metabolomics. Further experiments should use metabolomics to reveal the link among gut microbiota composition, functional pathway, and glioma growth. Third, there are differences in bacterial composition in intestinal mucosa and feces, but only fecal samples were collected for analysis here.
Conclusions
Glioma development and progression altered the composition of gut microbiota in both TMZ- and vehicle-treated mice. However, treatment with TMZ further altered the levels of several key bacterial populations. Induction of Akkermansia and Bifidobacterium as well as the prevention of the reduction in Anaerotruncus may contribute to the anti-tumor effect of TMZ. Our study helps to establish the association between certain micromaterial communities and the anti-tumor effect of TMZ. Future metabolomics, animal (eg microbiota transplantation), and human studies are necessary to confirm the role of specific gut microbiota in glioma growth and the anti-tumor properties of TMZ.
Data Sharing Statement
The sequencing data was deposited and made publicly available on Gene Expression Ominous database with accession number GSE171041.
Funding
This research was supported by grants from National Health Commission of China (2018ZX-07S-011), Medical Science Advancement Program of Wuhan University (No.TFJC2018003), and Translational Medicine and Interdisciplinary Research Joint Fund of Zhongnan Hospital of Wuhan University (No. ZLYNXM202011, ZNLH201901).
Disclosure
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Louis DN, Ohgaki H, Wiestler OD, et al. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007;114(2):97–109. doi:10.1007/s00401-007-0243-4
2. Peñaranda-Fajardo NM, Meijer C, Liang Y, et al. ER stress and UPR activation in glioblastoma: identification of a noncanonical PERK mechanism regulating GBM stem cells through SOX2 modulation. Cell Death Dis. 2019;10(10):690. doi:10.1038/s41419-019-1934-1
3. Unger MM, Spiegel J, Dillmann KU, et al. Short chain fatty acids and gut microbiota differ between patients with Parkinson’s disease and age-matched controls. Parkinsonism Relat Disord. 2016;32:66–72. doi:10.1016/j.parkreldis.2016.08.019
4. Nie P, Li Z, Wang Y, et al. Gut microbiome interventions in human health and diseases. Med Res Rev. 2019;39(6):2286–2313. doi:10.1002/med.21584
5. Scheperjans F, Aho V, Pereira PA, et al. Gut microbiota are related to Parkinson’s disease and clinical phenotype. Mov Disord. 2015;30(3):350–358. doi:10.1002/mds.26069
6. Vogt NM, Kerby RL, Dill-McFarland KA, et al. Gut microbiome alterations in Alzheimer’s disease. Sci Rep. 2017;7(1):13537. doi:10.1038/s41598-017-13601-y
7. Mulak A, Bonaz B. Brain-gut-microbiota axis in Parkinson’s disease. World J Gastroenterol. 2015;21(37):10609–10620. doi:10.3748/wjg.v21.i37.10609
8. D’Alessandro G, Antonangeli F, Marrocco F, et al. Gut microbiota alterations affect glioma growth and innate immune cells involved in tumor immunosurveillance in mice. Eur J Immunol. 2020;50(5):705–711. doi:10.1002/eji.201948354
9. Mehrian-Shai R, Reichardt JKV, Harris CC, et al. The gut-brain axis, paving the way to brain cancer. Trends Cancer. 2019;5(4):200–207. doi:10.1016/j.trecan.2019.02.008
10. Margison GP, Santibáñez Koref MF, Povey AC. Mechanisms of carcinogenicity/chemotherapy by O6-methylguanine. Mutagenesis. 2002;17(6):483–487. doi:10.1093/mutage/17.6.483
11. Stupp R, Mason WP, van den Bent MJ, et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 2005;352(10):987–996. doi:10.1056/NEJMoa043330
12. Viaud S, Saccheri F, Mignot G, et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science. 2013;342(6161):971–976. doi:10.1126/science.1240537
13. Iida N, Dzutsev A, Stewart CA, et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342(6161):967–970. doi:10.1126/science.1240527
14. Daillère R, Vétizou M, Waldschmitt N, et al. Enterococcus hirae and Barnesiella intestinihominis facilitate cyclophosphamide-induced therapeutic immunomodulatory effects. Immunity. 2016;45(4):931–943. doi:10.1016/j.immuni.2016.09.009
15. Li F, Wang P, Chen Z, et al. Alteration of the fecal microbiota in North-Eastern Han Chinese population with sporadic Parkinson’s disease. Neurosci Lett. 2019;707:134297. doi:10.1016/j.neulet.2019.134297
16. Peng A, Qiu X, Lai W, et al. Altered composition of the gut microbiome in patients with drug-resistant epilepsy. Epilepsy Res. 2018;147:102–107. doi:10.1016/j.eplepsyres.2018.09.013
17. Huang C, Li Y, Feng X, et al. Distinct gut microbiota composition and functional category in children with cerebral palsy and epilepsy. Front Pediatr. 2019;7:394. doi:10.3389/fped.2019.00394
18. Vandeputte D, Kathagen G, D’Hoe K, et al. Quantitative microbiome profiling links gut community variation to microbial load. Nature. 2017;551(7681):507–511. doi:10.1038/nature24460
19. Props R, Kerckhof FM, Rubbens P, et al. Absolute quantification of microbial taxon abundances. Isme j. 2017;11(2):584–587. doi:10.1038/ismej.2016.117
20. Smets W, Leff JW, Bradford MA, et al. A method for simultaneous measurement of soil bacterial abundances and community composition via 16S rRNA gene sequencing. Soil Biol Biochem. 2016;96:145–151.
21. Feldman LA, Fabre MS, Grasso C, et al. Perfluorocarbon emulsions radiosensitise brain tumors in carbogen breathing mice with orthotopic GL261 gliomas. PLoS One. 2017;12(9):e0184250. doi:10.1371/journal.pone.0184250
22. Round JL, Mazmanian SK. The gut microbiota shapes intestinal immune responses during health and disease. Nat Rev Immunol. 2009;9(5):313–323. doi:10.1038/nri2515
23. Wu S, Liu Y, Duan Y, et al. Intestinal toxicity of deoxynivalenol is limited by supplementation with Lactobacillus plantarum JM113 and consequentially altered gut microbiota in broiler chickens. J Anim Sci Biotechnol. 2018;9:74. doi:10.1186/s40104-018-0286-5
24. Sivamaruthi BS, Kesika P, Chaiyasut C. The role of probiotics in colorectal cancer management. Evid Based Complement Alternat Med. 2020;2020:3535982. doi:10.1155/2020/3535982
25. Loke MF, Chua EG, Gan HM, et al. Metabolomics and 16S rRNA sequencing of human colorectal cancers and adjacent mucosa. PLoS One. 2018;13(12):e0208584. doi:10.1371/journal.pone.0208584
26. Wang Z, Wang H, Kang Z, et al. Antioxidant and anti-tumour activity of triterpenoid compounds isolated from Morchella mycelium. Arch Microbiol. 2020;202(7):1677–1685. doi:10.1007/s00203-020-01876-1
27. Li M, van Esch B, Wagenaar GTM, et al. Pro- and anti-inflammatory effects of short chain fatty acids on immune and endothelial cells. Eur J Pharmacol. 2018;831:52–59. doi:10.1016/j.ejphar.2018.05.003
28. Fernández J, Redondo-Blanco S, Gutiérrez-Del-Río I, et al. Colon microbiota fermentation of dietary prebiotics towards short-chain fatty acids and their roles as anti-inflammatory and antitumour agents: a review. J Funct Foods. 2016;25:511–522.
29. Park E, Gim J, Kim DK, et al. Protective effects of alpha-lipoic acid on glutamate-induced cytotoxicity in C6 glioma cells. Biol Pharm Bull. 2019;42(1):94–102. doi:10.1248/bpb.b18-00603
30. Routy B, Le Chatelier E, Derosa L, et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359(6371):91–97. doi:10.1126/science.aan3706
31. Cani PD, de Vos WM. Next-generation beneficial microbes: the case of Akkermansia muciniphila. Front Microbiol. 2017;8:1765. doi:10.3389/fmicb.2017.01765
32. Consonni A, Cordiglieri C, Rinaldi E, et al. Administration of bifidobacterium and lactobacillus strains modulates experimental myasthenia gravis and experimental encephalomyelitis in Lewis rats. Oncotarget. 2018;9(32):22269–22287. doi:10.18632/oncotarget.25170
33. Rossi M, Amaretti A, Raimondi S. Folate production by probiotic bacteria. Nutrients. 2011;3(1):118–134. doi:10.3390/nu3010118
34. Sanchez H, Hossain MB, Lera L, et al. High levels of circulating folate concentrations are associated with DNA methylation of tumor suppressor and repair genes p16, MLH1, and MGMT in elderly Chileans. Clin Epigenetics. 2017;9:74. doi:10.1186/s13148-017-0374-y
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.