Back to Journals » International Journal of Nanomedicine » Volume 10 » Special Issue on diverse applications in Nano-Theranostics
Targeting cancer stem cells by using the nanoparticles
Authors Hong I, Jang G, Lee H, Nam J, An SSA
Received 10 May 2015
Accepted for publication 28 July 2015
Published 10 September 2015 Volume 2015:10(Special Issue on diverse applications in Nano-Theranostics) Pages 251—260
DOI https://doi.org/10.2147/IJN.S88310
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Thomas Webster
In-Sun Hong,1,2,* Gyu-Beom Jang,1,2,* Hwa-Yong Lee,3 Jeong-Seok Nam1,2
1Laboratory of Tumor Suppressor, Lee Gil Ya Cancer and Diabetes Institute, 2Department of Molecular Medicine, School of Medicine, Gachon University, Incheon, 3The Faculty of Liberal Arts, Jungwon University, Chungbuk, Republic of Korea
*These authors contributed equally to this work
Abstract: Cancer stem cells (CSCs) have been shown to be markedly resistant to conventional cancer treatments such as chemotherapy and radiation therapy. Therefore, therapeutic strategies that selectively target CSCs will ultimately lead to better cancer treatments. Currently, accessible conventional therapeutic agents mainly eliminate the bulk tumor but do not eliminate CSCs. Therefore, the discovery and improvement of CSC-targeting therapeutic agents are necessary. Nanoparticles effectively inhibit multiple types of CSCs by targeting specific signaling pathways (Wnt/ß-catenin, Notch, transforming growth factor-ß, and hedgehog signaling) and/or specific markers (aldehyde dehydrogenases, CD44, CD90, and CD133) critically involved in CSC function and maintenance. In this review article, we summarized a number of findings to provide current information about their therapeutic potential of nanoparticles in various cancer cell types and CSCs.
Keywords: ALDH, Wnt/ß-catenin, Hedgehog, Notch, TGF-ß signaling, CD44, CD133
Introduction
Current systemic therapies for cancer such as chemotherapy and radiation are partly effective in inhibiting bulk tumor cell growth and blocking tumor formation. However, a minority of cancer patients with metastases and a quarter of those with early stage disease are at a high risk of relapse due to cancer stem cells (CSCs). The CSCs concept was first suggested to describe a small population of acute myeloid leukemia, which contribute to tumor growth, metastasis, and recurrence.1 The identification of leukemic CSCs prompted further investigation into other solid tumor types. Recently, CSCs have been identified in almost all cancer types, including pancreatic,2 gastric,3 brain,4 colon,5 prostate,6 and lung cancers.7 CSCs are generally defined by a unique set of functional characteristics: 1) CSCs can be purified by specific biomarkers and/or signaling pathways,8–11 2) CSCs are capable of generating colonies in suspension culture conditions,12 and 3) CSCs are resistant to chemotherapeutic agents13–15 and radiation.15,16 These CSC-specific features suggest that the majority of conventional treatments, such as chemotherapy and radiation, can kill the bulk tumor cells but may ultimately fail to induce durable clinical results because conventional approaches are not as effective at eliminating CSCs; thus, the remaining CSCs are able to form new colonies and regenerate tumors in patients. Therefore, new therapeutic strategies that selectively target CSCs will ultimately improve cancer treatments.17 Currently, new treatment modalities in the form of nanoparticles (NPs)-targeting CSC-specific markers or signaling pathways are available or under investigation.18,19 Hirsch et al first introduced the effects of NPs in breast cancer by using silica–gold nanoshells.20 Recently, extensive research has identified various types of NP-targeting CSCs, including NP-mediated hyperthermia,21 curcumin-based NPs,22 and liposomes-based NPs.23 These NP-based therapeutic approaches provide advantages over the small molecule pharmaceutical agents-based therapeutic strategies. However, there is not enough information currently available to make a conclusive statement regarding the therapeutic potential of these NPs. Therefore, in this review article, we provide an overview of the characteristics of CSC and discuss the various NPs-targeting CSC-specific signaling pathways and biomarkers involved in the development and maintenance of CSCs.
Cancer stem cells: implications for tumorigenesis
Identification and isolation of CSCs in various cancers
The majority of cells in bulk tumors have limited self-renewal and tumor-initiating capacity; indeed, only a small subpopulation of cancer cells retains extensive self-renewal and tumorigenic potential. These higher tumorigenic populations are called CSCs or cancer initiating cells. The CSC model of tumor development has been proposed to explain the high degree of phenotypic and functional heterogeneity among cancer cells.24 In the 1960s, Bruce et al found that only 1%–4% of the total number of mouse leukemic cells transplanted in vivo formed colonies and initiated tumor growth in the recipient spleen.25 The identification of leukemic CSCs prompted further investigation into other solid tumor types. CSCs were first identified and isolated from a solid tumor breast cancer. Breast CSCs are typically characterized with a CD44+/CD24−/low phenotype and test positive for the epithelial cell adhesion molecule, also known as the epithelial-specific marker.26,27 As few as 100 cells with these molecular characteristics grew rapidly and extensively in vitro and generated new tumors in vivo.26 Recently, extensive research has identified CSCs in different types of solid tumors, including brain,28 colon,29 head and neck,30 liver,31 lung,32,33 and other cancers.34 CSCs are typically resistant to various chemotherapeutic drugs13–15 and radiation therapies.15,16 These CSC-specific features suggest that the majority of conventional cancer treatments, such as surgery, chemotherapy, and radiation therapy, can kill the bulk tumor cells but may ultimately fail to induce durable clinical responses because they are not as effective at killing CSCs; thus, the remaining CSCs are able to form colonies and initiate new tumors in patients.
CSCs as a selective therapeutic target
Despite promising treatment results, current therapeutic strategies against cancers have many limitations that frequently lead to metastatic failure and a high risk of recurrence and mortality. The most common cause of an unsatisfactory clinical response is resistance to conventional therapeutic strategies. CSC-mediated therapeutic resistance was demonstrated in different tumors, including brain,35 breast,13 colorectal,36 leukemia,37 melanoma,38 and pancreatic2 cancers. Furthermore, CSC-mediated radiation resistance was reported in brain28 and breast39 cancers. Therefore, the development of novel therapeutics and control strategies that selectively target CSCs without unduly affecting normal and healthy cells is urgently required.40–42 A significantly improved therapeutic outcome could be achieved by the selective targeting of subtle differences in surface marker expression or signaling pathways when compared with bulk tumor cells. Since their identification in various cancer types, multiple CSC therapeutic strategies targeting specific stem cell surface markers and unique signaling pathways have been developed for several solid tumors. These alternative therapeutic strategies can successfully eliminate CSCs and thereby prevent tumor recurrence and metastasis.
Targeting CSC-specific markers
Aldehyde dehydrogenase activity
Aldehyde dehydrogenases (ALDHs) are a superfamily of enzymes that play a key role in the metabolism of various aldehyde derivatives. These enzymes were first described and identified as conferring resistance to cyclophosphamide and other alkylating agents in hematopoietic stem/progenitor cells.43 Recent studies have demonstrated that high levels of ALDH activity are associated with enhanced tumorigenicity and chemoresistance in CSCs.44–46 Indeed, higher ALDH activity has been observed in the highly tumorigenic colon CSC subpopulations with an EpCAMhigh/CD44+ phenotype.47 Moreover, ALDH activity was used to predict a poor clinical outcome in breast cancer patients due to its correlation with tumorigenicity and chemoresistance.45 Ma et al have found that ALDH was preferentially expressed in the CD133-positive subpopulation and could be used to better characterize the tumorigenic CD133-positive CSC population in liver cancers.46 The hierarchical tumorigenic potential was determined as CD133 +ALDH+ > CD133+ALDH− > CD133−ALDH−.46 Therefore, it is reasonable that ALDH can act as a potential prognostic marker and therapeutic target for the treatment of various cancer types. NPs loaded with low-dose decitabine, a DNA hypermethylation inhibitor, that significantly reduced clonogenic growth and ALDH-positive stem-like population in malignant breast cancer by inhibiting cancer cell growth and stem cell phenotypes.48 Chenna et al have also developed polymeric NP from poly(lactic-co-glycolic acid) conjugated with polyethylene glycol encapsulating hedgehog signaling inhibitor (HPI).49 This HPI-incorporated polymeric NP showed a remarkably increased apoptotic effects in secondary mutational pancreatic cells by suppressing the growth of ALDH-positive CSCs in the orthotopic Pa03C xenograft.49
CD44
CD44 is a transmembrane receptor for hyaluronic acid and has recently been identified in CSCs from numerous solid tumors, including breast,26 bladder,50 cervical,51 colon,52 gastric,53 lung,54 ovarian,55 pancreatic,56 and prostate cancers.57 Enhanced CD44 expression in CSCs suggests that CD44 is an attractive new target for the treatment of multiple cancer types. Yang et al demonstrated that CD44+ subpopulations were more tumorigenic than their CD44− counterparts in nude mice.58 Therefore, there is an urgent need for the development of effective CD44-targeted therapeutic strategies. Polymeric micelles have amphiphilic block copolymers with a spherical inner core and outer shell. Hydrophobic inner core serves as a container for hydrophobic drug, while hydrophilic outer core provides structural stability and extends the circulation time.59 Shah et al have reported that paclitaxel-incorporated micelles showed a remarkably increased therapeutic efficacy and specificity in CD44-positive metastatic ovarian cancer cells isolated from patients.60 In addition, liposomal NP can potentially enhance the delivery of suicide gene or chemotherapeutic drugs to the breast and colon cancers.61,62 Wang et al have reported that anti-CD44 antibody-incorporated liposomal NP delivery system loaded with suicide gene or doxorubicin could specifically target the CD44-positive hepatocellular carcinoma cells and effectively induce apoptotic cell death.23 A more recent report suggests that a hyaluronic acid-coated chitosan NPs loaded with 5-fluorouracil (5-FU)/oxaliplatin showed a significantly enhanced cytotoxicity compared with either 5-FU or oxaliplatin alone in human colorectal cancer cells, which overexpress CD44.63,64 Although several NPs for targeting CD44-positive cells are recently developed, their therapeutic effects in vivo have not yet been demonstrated convincingly.
CD90
Various CD cell surface proteins were used as a marker to identify CSCs in human liver cancer cell lines and clinical samples. CD90 is a glycosyl phosphatidylinositol-anchored membrane glycoprotein of the immunoglobulin superfamily expressed mainly on the surface of leukocytes.65 This marker is involved in cell–cell and cell–adhesive matrix interactions. Yang et al found that the CD90-positive subpopulation showed a distinct higher tumorigenicity and metastatic potential in a mouse xenograft model when compared with CD90-negative counterparts.58,66 All clinical tumor specimens and ~90% of blood samples from liver cancer patients contained a CD45−/CD90+ subpopulation capable of initiating and maintaining tumor formation in an immunodeficient mouse model.58,66 Based on the aforementioned findings, CD90 can be used to identify potential hepatic CSCs from tumor specimens and blood samples of liver cancer patients. Thus, CD90 may be an important prognostic marker and effective therapeutic target for the treatment of various hepatic cancer types. Wang et al have reported that anti-CD90 antibody-mediated water-soluble CdSe core nanocrystals loaded with photosensitizers such as trifluoperazine could specifically target the CD90-positive leukemia CSCs and sensitize leukemia CSCs to UV irradiation and promote apoptotic cell death.67
CD133
The stem cell marker CD133, also known as prominin-1, is a transmembrane glycoprotein.68 CD133 is highly expressed in immature hematopoietic stem cells and endothelial progenitor cells.69 In glioblastoma, CD133-positive subpopulations were shown to be considerably more tumorigenic than CD133-negative compartments, which form the bulk tumor. Moreover, a poor clinical outcome is associated with increased CD133 expression in glioblastoma patients.70 In addition, the protein is overexpressed in various cancer types, including metastatic colorectal cancer,71 ovarian cancer,72 glioblastoma,73 and gastric carcinoma.74 Recent evidence suggests that a subpopulation of CD133-positive cancer cells have a significant key molecule that confers resistance to conventional chemotherapeutic agents.75 Moreover, radiation-exposed CD133-positive liver cancer cells induce enhanced radioresistance via MAPK/ERK signaling activation and tumor development in a xenograft model compared with CD133-negative cells, suggesting that the CD133-positive populations confer radioresistance in hepatocellular carcinoma.76 Therefore, the surface marker CD133 should be a potential target to improve the efficiency of treating CSCs. NanoCurc™, a recently developed polymeric NP encapsulating curcumin, significantly reduced clonogenic growth and CD133-positive stem-like population in malignant brain tumors.22 In addition, NPs conjugated with PEGylated poly(lactic-co-glycolic acid) receive extensive attention and are used widely because of their safety in the clinical trials.77 Interestingly, salinomycin-loaded NPs conjugated with CD133 aptamers selectively inhibit CD133-positive osteosarcoma both in vitro and in vivo.78
Targeting CSC-specific signaling pathways
Wnt/β-catenin signaling pathway
The Wnt/β-catenin signaling pathway is an evolutionarily well-conserved pathway that regulates various physiologic processes, including development, growth, regeneration, and self-renewal.79 The Wnt/β-catenin pathway is activated when a Wnt ligand binds to the transmembrane complex comprising the Frizzled receptor, leading to binding of the low-density lipoprotein-related receptor. This leads to the suppression of glycogen synthase kinase-3β-binding protein, thereby improving the stability of β-catenin, which then accumulates and is translocated to the nucleus. Consequently, β-catenin forms a complex with the transcription factor/lymphocyte enhancer factor and activates the expression of Wnt target gene such as c-myc and cyclin D1.80–82 Altered activation of Wnt/β-catenin signaling is a key feature of various cancer types and is considered to be critical for epithelial–mesenchymal transitions that favor tumor metastasis.83 Aberrant activation of the Wnt/β-catenin signaling has recently been implicated in several types of cancers, including ovarian,84 colon,85 and breast cancer.86 Interestingly, this signaling pathway was initially reported as a key CSC signaling pathway in acute myeloid leukemia87 and was reported to be involved in the maintenance and function of CSCs from breast,88 colon,89 liver,90 and lung91 cancers. Therefore, selective targeting of Wnt/β-catenin signaling may be considered as an effective therapeutic strategy for the treatment of various types of cancer. Yallapu et al have developed curcumin-loaded poly(lactic acid-co-glycolic acid) (PLGA) NP to provide increased bioavailability of curcumin in the blood circulation.92 These curcumin-incorporated PLGA NPs showed a remarkably increased apoptotic effects in cisplatin-resistant ovarian cancer cells by suppressing Wnt/β-catenin signaling component β-catenin.92 Similarly, Tang et al loaded 5-FU to NPs, and demonstrated that these 5-FU NPs can effectively inhibit the peritoneal dissemination of colorectal cancer cells,93 which overexpress Wnt/β-catenin signaling components.94,95
Notch signaling pathway
The Notch signaling pathway is an evolutionarily conserved developmental pathway governing a broad spectrum of events, such as cell differentiation decisions and the formation of precise tissue patterns.96 Notch signaling also plays an important role in regulating stem cell maintenance and differentiation.24,97–99 This signaling pathway is activated through four Notch receptors (Notch 1–4) that can interact with five Jagged family ligands.100,101 It has been suggested that Notch 1 and 2 share the highest degree of sequence and structural similarities and are ubiquitously distributed in a wide variety of tissues. In contrast, Notch 2 and 4 are expressed in a limited range of cell types (eg, vascular endothelial and smooth muscle cells). While the oncogenic effects of Notch signaling in several types of tumors are well-documented, its potential role in CSCs has only recently emerged. A recent study suggested that inhibition of Notch signaling sharply decreased self-renewal, clonogenic, and the tumorigenic potential of glioblastoma CSCs.102 In addition, inhibition of Notch signaling led to a decrease of the CSC-like subpopulation and increased the susceptibility of CSCs to radiation-induced apoptosis in glioblastomas.103 Aberrant activation of Notch signaling has been observed in CD133+ liver CSC subpopulations when compared with CD133− subpopulations.104 Consequentially, targeting Notch signaling pathway may provide an effective therapeutic approach in the treatment of various cancers. Despite the inhibition of Notch signaling can be achieved by γ-secretase inhibitors, inhibitory peptides,105 and antibodies.106 Clinical application of these inhibitory agents are currently restricted by their considerable side effects.107,108 Rosenholm et al have developed mesoporous silica NPs (MSNPs) as drug-delivery systems with high selectivity and demonstrated that these drug-conjugated NPs are suitable for targeted delivery of hydrophobic drugs in vitro.109,110 They demonstrate that these drug-conjugated MSNPs have significantly enhanced cytotoxicity by selective targeting the Notch signaling in various cancer cell types, such as cervical and breast cancer cells.111 In addition, Steg et al have developed Jagged1 (Notch ligand) siRNAs-loaded chitosan NPs, which selectively inhibit ovarian cancer both in vitro and in vivo by selectively targeting Notch ligand Jagged1.112 Despite the number of in vitro studies’ evidences for the therapeutic efficiency and selectivity of drug-conjugated NPs targeting Notch signaling, valid in vivo models are still largely lacking.
Transforming growth factor-β signaling pathway
Transforming growth factor-β (TGF-β) signaling provides important regulatory signals during the initial phase of normal development and regeneration113,114 and exerts promoting effects in the initiation and progression of multiple cancer types including breast,115 colon,116 liver,117 lung,118 and ovary.119 TGF-β ligands bind to a type II receptor, which constitutively recruits and phosphorylates a type I receptor. Next, the type I receptor triggers the phosphorylation of receptor-regulated SMADs and results in ligand-induced transcription.120,121 In addition, cancer patients with enhanced TGF-β levels in urine and serum samples had shorter survival periods when compared with patients with normal TGF-β levels.122–124 Constitutively elevated levels of TGF-β are closely correlated with an advanced disease state and poorer prognosis in cancer patients.125,126 These studies indicate that TGF-β signaling is an important prognostic marker in various types of cancer. While the oncogenic effects of TGF-β in several types of tumors are well-documented, its potential role in CSCs has only recently emerged. Interestingly, this signaling pathway has been reported to be involved in the maintenance and function of CSCs from breast,127 colon,128 liver,129 and lung130 cancers. Thus, selective targeting TGF-β signaling may be considered as an effective therapeutic strategy for the treatment of various types of cancer. In order to improve the drug delivery for breast cancer treatment, a polyethyleneimine/polyethylene glycol-conjugated MSNPs were developed by Meng et al to load a small molecule TGF-β inhibitor, LY364947.131 This approach provided significantly improved therapeutic efficiency in tumor xenograft models compared to the treatment with free LY364947 alone.131 In addition, gold NPs (AuNPs) are widely used as carriers for therapeutic and diagnostic agents because of their great biocompatibility and unique physiochemical properties.132 In this context, Tsai et al found that AuNPs could selectively capture TGF-β1 through S–Au binding between cysteine and disulfides residues resulting in deactivation of TGF-β signaling pathway.133 Interestingly, they also found that the immunosuppressive function of TGF-β was significantly attenuated by AuNPs and resulted in an increased number and frequency of tumor-infiltrating T lymphocytes.133 These results demonstrate that AuNPs may be a promising immune modulator by inhibiting immunosuppressive function of TGF-β1 signaling pathway.
Hedgehog signaling pathway
The Hedgehog (Hh) signaling pathway was first identified as a critical regulator of pattern formation during early development and regeneration, and it also regulates differentiation, growth, and migration in a temporal-, spatial-, and concentration-dependent manner.134–137 The functional significance of this signaling pathway is illustrated by an increase in birth defects and malignancies associated with aberrant activation of this normally quiescent pathway in adults.138,139 Three Hh homologs with different spatial and temporal distribution patterns have been identified in humans: Desert hedgehog (Dhh), Indian hedgehog (Ihh), and Sonic hedgehog (Shh).140 The Hh signaling cascade is initiated by Hh binding to the 12-transmembrane receptor Patched 1, which relieves its inhibition on Smoothened (Smo), culminating in the nuclear localization of DNA-binding Gli transcription factors in target cells.141 Several research groups reported that Hh signaling is aberrantly activated in various cancer types, including colon,142 brain,143 and breast,144 liver,145 lung,146 and ovarian147 cancer. Therefore, targeting Hh signaling pathway may provide an effective therapeutic approach in the treatment of various cancers. Almost currently available Hh small-molecule inhibitors approved for clinical trials for cancer therapy are Smo antagonists.148 However, clinical applications of these inhibitors are currently restricted by their limited binding ability to Smo and poor systemic bioavailability. Therefore, Xu et al developed and characterized Gli antagonist (HPI-1)-conjugated polymeric nanoparticle (NanoHHI).49 NanoHHI significantly inhibited the growth and invasion of CD133-positive cells, which are implicated as CSCs in liver cancers.149 In addition, Verma et al found that Anthothecol (a limonoid derived from plant Khayaanthotheca)-conjugated PLGA NPs effectively inhibited cell proliferation and colony formation and induced apoptosis in pancreatic CSCs by modulating Hh signaling pathway.150 These studies suggest that Hh signaling targeting NPs might be effective therapeutic approaches in patients with recurrence following curative surgical resection.
Conclusion
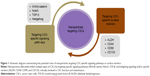
The CSC was first discovered in acute myeloid leukemia >40 years ago. The identification of leukemic CSCs prompted further investigation into other types of solid tumors. Interestingly, CSCs are markedly resistant to conventional cancer treatments, such as chemotherapy and radiation. Therefore, identifying and selectively targeting markers and signaling pathways of CSCs are feasible therapeutic strategies for treating various cancer types, regardless of the underlying cause. Currently, new treatment modalities in the form of NPs-targeting CSC-specific markers or signaling pathways are available or under investigation. Various NPs-targeting CSC-specific surface markers or signaling pathways are listed in Table 1. Despite the number of in vitro and in vivo studies evidences for the therapeutic efficiency and selectivity of chemodrugs- or antibodies-conjugated NPs-targeting CSC-specific markers or signaling pathways, there is not enough information currently available to make a conclusive statement regarding the clinical therapeutic effects of NPs. Therefore, more detailed information about the effects of NPs-targeting CSCs on various cancer types will undoubtedly lead to more effective clinical therapy in the future. Moreover, the majority of the currently available information on CSCs is markedly influenced by the biological characteristics of normal stem/progenitor cells, in terms of their specific surface markers and distinct signaling pathways. Therefore, targeting these markers and/or aberrantly activated signaling pathways to selectively eliminate CSCs may reduce normal stem/progenitor cells and prevent the normal tissue regeneration processes, thus causing tissue or organ damages. Consequently, it remains unclear whether CSCs in various cancers can be selectively eliminated without significantly inhibiting all other normal stem/progenitor cells in the organs or tissues. Therefore, further characteristics related to CSC-specific signaling pathways and surface markers need to be elucidated. These conclusions warrant future studies aimed at providing adequate and unique diagnostic and therapeutic strategies for cancer patients with fewer side effects. Schematic diagram summarizes the potential roles of NPs-targeting CSC-specific markers and signaling pathways in cancer treatments (Figure 1).
Acknowledgments
This research was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI) and the Ministry of Health and Welfare, Republic of Korea (grant number HI11C1512). This work was also supported by a grant (15182MFDS455) from Ministry of Food and Drug Safety, Republic of Korea.
Author contributions
All authors contributed toward data analysis, drafting and critically revising the paper and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
Visvader JE, Lindeman GJ. Cancer stem cells: current status and evolving complexities. Cell Stem Cell. 2012;10(6):717–728. | ||
Hermann PC, Huber SL, Herrler T, et al. Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell. 2007;1(3):313–323. | ||
Fukuda K, Saikawa Y, Ohashi M, et al. Tumor initiating potential of side population cells in human gastric cancer. Int J Oncol. 2009;34(5):1201–1207. | ||
Hemmati HD, Nakano I, Lazareff JA, et al. Cancerous stem cells can arise from pediatric brain tumors. Proc Natl Acad Sci U S A. 2003;100(25):15178–15183. | ||
O’Brien CA, Pollett A, Gallinger S, Dick JE. A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature. 2007;445(7123):106–110. | ||
Collins AT, Berry PA, Hyde C, Stower MJ, Maitland NJ. Prospective identification of tumorigenic prostate cancer stem cells. Cancer Res. 2005;65(23):10946–10951. | ||
Eramo A, Lotti F, Sette G, et al. Identification and expansion of the tumorigenic lung cancer stem cell population. Cell Death Differ. 2008;15(3):504–514. | ||
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100(1):57–70. | ||
Singh SK, Clarke ID, Terasaki M, et al. Identification of a cancer stem cell in human brain tumors. Cancer Res. 2003;63(18):5821–5828. | ||
Ricci-Vitiani L, Lombardi DG, Pilozzi E, et al. Identification and expansion of human colon-cancer-initiating cells. Nature. 2007;445(7123):111–115. | ||
Zhang M, Behbod F, Atkinson RL, et al. Identification of tumor-initiating cells in a p53-null mouse model of breast cancer. Cancer Res. 2008;68(12):4674–4682. | ||
Dontu G, Abdallah WM, Foley JM, et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 2003;17(10):1253–1270. | ||
Li X, Lewis MT, Huang J, et al. Intrinsic resistance of tumorigenic breast cancer cells to chemotherapy. J Natl Cancer Inst. 2008;100(9):672–679. | ||
Eyler CE, Rich JN. Survival of the fittest: cancer stem cells in therapeutic resistance and angiogenesis. J Clin Oncol. 2008;26(17):2839–2845. | ||
Woodward WA, Chen MS, Behbod F, Alfaro MP, Buchholz TA, Rosen JM. WNT/beta-catenin mediates radiation resistance of mouse mammary progenitor cells. Proc Natl Acad Sci U S A. 2007;104(2):618–623. | ||
Diehn M, Cho RW, Lobo NA, et al. Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature. 2009;458(7239):780–783. | ||
Boman BM, Huang E. Human colon cancer stem cells: a new paradigm in gastrointestinal oncology. J Clin Oncol. 2008;26(17):2828–2838. | ||
Shapira A, Livney YD, Broxterman HJ, Assaraf YG. Nanomedicine for targeted cancer therapy: towards the overcoming of drug resistance. Drug Resist Updat. 2011;14(3):150–163. | ||
Wang D, Lin B, Ai H. Theranostic nanoparticles for cancer and cardiovascular applications. Pharm Res. 2014;31(6):1390–1406. | ||
Hirsch LR, Stafford RJ, Bankson JA, et al. Nanoshell-mediated near-infrared thermal therapy of tumors under magnetic resonance guidance. Proc Natl Acad Sci U S A. 2003;100(23):13549–13554. | ||
Burke AR, Singh RN, Carroll DL, et al. The resistance of breast cancer stem cells to conventional hyperthermia and their sensitivity to nanoparticle-mediated photothermal therapy. Biomaterials. 2012;33(10):2961–2970. | ||
Lim KJ, Bisht S, Bar EE, Maitra A, Eberhart CG. A polymeric nanoparticle formulation of curcumin inhibits growth, clonogenicity and stem-like fraction in malignant brain tumors. Cancer Biol Ther. 2011;11(5):464–473. | ||
Wang L, Su W, Liu Z, et al. CD44 antibody-targeted liposomal nanoparticles for molecular imaging and therapy of hepatocellular carcinoma. Biomaterials. 2012;33(20):5107–5114. | ||
Reya T, Morrison SJ, Clarke MF, Weissman IL. Stem cells, cancer, and cancer stem cells. Nature. 2001;414(6859):105–111. | ||
Bruce WR, Van Der Gaag HA. Quantitative assay for the number of murine lymphoma cells capable of proliferation in vivo. Nature. 1963;199:79–80. | ||
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci U S A. 2003;100(7):3983–3988. | ||
Fillmore CM, Kuperwasser C. Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res. 2008;10(2):R25. | ||
Bao S, Wu Q, McLendon RE, et al. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature. 2006;444(7120):756–760. | ||
Ibrahim EE, Babaei-Jadidi R, Saadeddin A, et al. Embryonic NANOG activity defines colorectal cancer stem cells and modulates through AP1- and TCF-dependent mechanisms. Stem Cells. 2012;30(10):2076–2087. | ||
Prince ME, Sivanandan R, Kaczorowski A, et al. Identification of a subpopulation of cells with cancer stem cell properties in head and neck squamous cell carcinoma. Proc Natl Acad Sci U S A. 2007;104(3):973–978. | ||
Lee TK, Castilho A, Cheung VC, Tang KH, Ma S, Ng IO. CD24(+) liver tumor-initiating cells drive self-renewal and tumor initiation through STAT3-mediated NANOG regulation. Cell Stem Cell. 2011;9(1):50–63. | ||
Jiang F, Qiu Q, Khanna A, et al. Aldehyde dehydrogenase 1 is a tumor stem cell-associated marker in lung cancer. Mol Cancer Res. 2009;7(3):330–338. | ||
Bertolini G, Roz L, Perego P, et al. Highly tumorigenic lung cancer CD133+ cells display stem-like features and are spared by cisplatin treatment. Proc Natl Acad Sci U S A. 2009;106(38):16281–16286. | ||
Bussolati B, Bruno S, Grange C, Ferrando U, Camussi G. Identification of a tumor-initiating stem cell population in human renal carcinomas. FASEB J. 2008;22(10):3696–3705. | ||
Eramo A, Ricci-Vitiani L, Zeuner A, et al. Chemotherapy resistance of glioblastoma stem cells. Cell Death Differ. 2006;13(7):1238–1241. | ||
Dylla SJ, Beviglia L, Park IK, et al. Colorectal cancer stem cells are enriched in xenogeneic tumors following chemotherapy. PLoS One. 2008;3(6):e2428. | ||
Ishikawa F, Yoshida S, Saito Y, et al. Chemotherapy-resistant human AML stem cells home to and engraft within the bone-marrow endosteal region. Nat Biotechnol. 2007;25(11):1315–1321. | ||
Frank NY, Margaryan A, Huang Y, et al. ABCB5-mediated doxorubicin transport and chemoresistance in human malignant melanoma. Cancer Res. 2005;65(10):4320–4333. | ||
Phillips TM, McBride WH, Pajonk F. The response of CD24(-/low)/CD44+ breast cancer-initiating cells to radiation. J Natl Cancer Inst. 2006;98(24):1777–1785. | ||
LaBarge MA. The difficulty of targeting cancer stem cell niches. Clin Cancer Res. 2010;16(12):3121–3129. | ||
Lacerda L, Pusztai L, Woodward WA. The role of tumor initiating cells in drug resistance of breast cancer: Implications for future therapeutic approaches. Drug Resist Updat. 2010;13(4–5):99–108. | ||
Wang Z, Li Y, Ahmad A, et al. Targeting miRNAs involved in cancer stem cell and EMT regulation: an emerging concept in overcoming drug resistance. Drug Resist Updat. 2010;13(4–5):109–118. | ||
Kastan MB, Schlaffer E, Russo JE, Colvin OM, Civin CI, Hilton J. Direct demonstration of elevated aldehyde dehydrogenase in human hematopoietic progenitor cells. Blood. 1990;75(10):1947–1950. | ||
Pearce DJ, Taussig D, Simpson C, et al. Characterization of cells with a high aldehyde dehydrogenase activity from cord blood and acute myeloid leukemia samples. Stem Cells. 2005;23(6):752–760. | ||
Ginestier C, Hur MH, Charafe-Jauffret E, et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 2007;1(5):555–567. | ||
Ma S, Chan KW, Lee TK, et al. Aldehyde dehydrogenase discriminates the CD133 liver cancer stem cell populations. Mol Cancer Res. 2008;6(7):1146–1153. | ||
Dalerba P, Dylla SJ, Park IK, et al. Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci U S A. 2007;104(24):10158–10163. | ||
Li SY, Sun R, Wang HX, et al. Combination therapy with epigenetic-targeted and chemotherapeutic drugs delivered by nanoparticles to enhance the chemotherapy response and overcome resistance by breast cancer stem cells. J Control Release. 2015;205:7–14. | ||
Chenna V, Hu C, Pramanik D, et al. A polymeric nanoparticle encapsulated small-molecule inhibitor of Hedgehog signaling (NanoHHI) bypasses secondary mutational resistance to Smoothened antagonists. Mol Cancer Ther. 2012;11(1):165–173. | ||
Yang YM, Chang JW. Bladder cancer initiating cells (BCICs) are among EMA-CD44v6+ subset: novel methods for isolating undetermined cancer stem (initiating) cells. Cancer Invest. 2008;26(7):725–733. | ||
Feng D, Peng C, Li C, et al. Identification and characterization of cancer stem-like cells from primary carcinoma of the cervix uteri. Oncol Rep. 2009;22(5):1129–1134. | ||
Haraguchi N, Ohkuma M, Sakashita H, et al. CD133+CD44+ population efficiently enriches colon cancer initiating cells. Ann Surg Oncol. 2008;15(10):2927–2933. | ||
Takaishi S, Okumura T, Tu S, et al. Identification of gastric cancer stem cells using the cell surface marker CD44. Stem Cells. 2009;27(5):1006–1020. | ||
Leung EL, Fiscus RR, Tung JW, et al. Non-small cell lung cancer cells expressing CD44 are enriched for stem cell-like properties. PLoS One. 2010;5(11):e14062. | ||
Zhang S, Balch C, Chan MW, et al. Identification and characterization of ovarian cancer-initiating cells from primary human tumors. Cancer Res. 2008;68(11):4311–4320. | ||
Lee CJ, Dosch J, Simeone DM. Pancreatic cancer stem cells. J Clin Oncol. 2008;26(17):2806–2812. | ||
Dubrovska A, Kim S, Salamone RJ, et al. The role of PTEN/Akt/PI3K signaling in the maintenance and viability of prostate cancer stem-like cell populations. Proc Natl Acad Sci U S A. 2009;106(1):268–273. | ||
Yang ZF, Ho DW, Ng MN, et al. Significance of CD90+ cancer stem cells in human liver cancer. Cancer Cell. 2008;13(2):153–166. | ||
Adams ML, Lavasanifar A, Kwon GS. Amphiphilic block copolymers for drug delivery. J Pharm Sci. 2003;92(7):1343–1355. | ||
Shah V, Taratula O, Garbuzenko OB, Taratula OR, Rodriguez-Rodriguez L, Minko T. Targeted nanomedicine for suppression of CD44 and simultaneous cell death induction in ovarian cancer: an optimal delivery of siRNA and anticancer drug. Clin Cancer Res. 2013;19(22):6193–6204. | ||
Liao D, Liu Z, Wrasidlo WJ, et al. Targeted therapeutic remodeling of the tumor microenvironment improves an HER-2 DNA vaccine and prevents recurrence in a murine breast cancer model. Cancer Res. 2011;71(17):5688–5696. | ||
Liao D, Liu Z, Wrasidlo W, et al. Synthetic enzyme inhibitor: a novel targeting ligand for nanotherapeutic drug delivery inhibiting tumor growth without systemic toxicity. Nanomedicine. 2011;7(6):665–673. | ||
Jain A, Jain SK. In vitro and cell uptake studies for targeting of ligand anchored nanoparticles for colon tumors. Eur J Pharm Sci. 2008;35(5):404–416. | ||
Jain A, Jain SK, Ganesh N, Barve J, Beg AM. Design and development of ligand-appended polysaccharidic nanoparticles for the delivery of oxaliplatin in colorectal cancer. Nanomedicine. 2010;6(1):179–190. | ||
Haeryfar SM, Conrad DM, Musgrave B, Hoskin DW. Antibody blockade of Thy-1 (CD90) impairs mouse cytotoxic T lymphocyte induction by anti-CD3 monoclonal antibody. Immunol Cell Biol. 2005;83(4):352–363. | ||
Yang ZF, Ngai P, Ho DW, et al. Identification of local and circulating cancer stem cells in human liver cancer. Hepatology. 2008;47(3):919–928. | ||
Bakalova R, Ohba H, Zhelev Z, Ishikawa M, Baba Y. Quantum dots as photosensitizers? Nat Biotechnol. 2004;22(11):1360–1361. | ||
Wu Y, Wu PY. CD133 as a marker for cancer stem cells: progresses and concerns. Stem Cells Dev. 2009;18(8):1127–1134. | ||
Corbeil D, Marzesco AM, Wilsch-Brauninger M, Huttner WB. The intriguing links between prominin-1 (CD133), cholesterol-based membrane microdomains, remodeling of apical plasma membrane protrusions, extracellular membrane particles, and (neuro)epithelial cell differentiation. FEBS Lett. 2010;584(9):1659–1664. | ||
Zeppernick F, Ahmadi R, Campos B, et al. Stem cell marker CD133 affects clinical outcome in glioma patients. Clin Cancer Res. 2008;14(1):123–129. | ||
Zhang C, Zhou C, Wu XJ, et al. Human CD133-positive hematopoietic progenitor cells initiate growth and metastasis of colorectal cancer cells. Carcinogenesis. 2014;35(12):2771–2777. | ||
Baba T, Convery PA, Matsumura N, et al. Epigenetic regulation of CD133 and tumorigenicity of CD133+ ovarian cancer cells. Oncogene. 2009;28(2):209–218. | ||
Yi JM, Tsai HC, Glöckner SC, et al. Abnormal DNA methylation of CD133 in colorectal and glioblastoma tumors. Cancer Res. 2008;68(19):8094–8103. | ||
Hibi K, Sakata M, Kitamura YH, et al. Demethylation of the CD133 gene is frequently detected in early gastric carcinoma. Anticancer Res. 2010;30(4):1201–1203. | ||
Zhang Q, Shi S, Yen Y, Brown J, Ta JQ, Le AD. A subpopulation of CD133(+) cancer stem-like cells characterized in human oral squamous cell carcinoma confer resistance to chemotherapy. Cancer Lett. 2010;289(2):151–160. | ||
Piao LS, Hur W, Kim TK, et al. CD133+ liver cancer stem cells modulate radioresistance in human hepatocellular carcinoma. Cancer Lett. 2012;315(2):129–137. | ||
Langer R. Drug delivery and targeting. Nature. 1998;392(6679 suppl):5–10. | ||
Ni M, Xiong M, Zhang X, et al. Poly(lactic-co-glycolic acid) nanoparticles conjugated with CD133 aptamers for targeted salinomycin delivery to CD133+ osteosarcoma cancer stem cells. Int J Nanomedicine. 2015;10:2537–2554. | ||
Branda M, Wands JR. Signal transduction cascades and hepatitis B and C related hepatocellular carcinoma. Hepatology. 2006;43(5):891–902. | ||
Fleming HE, Janzen V, Lo Celso C, et al. Wnt signaling in the niche enforces hematopoietic stem cell quiescence and is necessary to preserve self-renewal in vivo. Cell Stem Cell. 2008;2(3):274–283. | ||
Choi H, Chun YS, Kim TY, Park JW. HIF-2alpha enhances beta-catenin/TCF-driven transcription by interacting with beta-catenin. Cancer Res. 2010;70(24):10101–10111. | ||
MacDonald BT, Tamai K, He X. Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell. 2009;17(1):9–26. | ||
Zhao C, Blum J, Chen A, et al. Loss of beta-catenin impairs the renewal of normal and CML stem cells in vivo. Cancer Cell. 2007;12(6):528–541. | ||
Rask K, Nilsson A, Brännström M, et al. Wnt-signalling pathway in ovarian epithelial tumours: increased expression of beta-catenin and GSK3beta. Br J Cancer. 2003;89(7):1298–1304. | ||
Kaler P, Godasi BN, Augenlicht L, Klampfer L. The NF-kappaB/AKT-dependent Induction of Wnt Signaling in Colon Cancer Cells by Macrophages and IL-1beta. Cancer Microenviron. 2009;2(1):69–80. | ||
Zhang J, Li Y, Liu Q, Lu W, Bu G. Wnt signaling activation and mammary gland hyperplasia in MMTV-LRP6 transgenic mice: implication for breast cancer tumorigenesis. Oncogene. 2010;29(4):539–549. | ||
Jamieson CH, Weissman IL, Passegue E. Chronic versus acute myelogenous leukemia: a question of self-renewal. Cancer Cell. 2004;6(6):531–533. | ||
Reya T, Clevers H. Wnt signalling in stem cells and cancer. Nature. 2005;434(7035):843–850. | ||
de Sousa EM, Vermeulen L, Richel D, Medema JP. Targeting Wnt signaling in colon cancer stem cells. Clin Cancer Res. 2011;17(4):647–653. | ||
Gedaly R, Galuppo R, Daily MF, et al. Targeting the Wnt/beta-catenin signaling pathway in liver cancer stem cells and hepatocellular carcinoma cell lines with FH535. PLoS One. 2014;9(6):e99272. | ||
Teng Y, Wang X, Wang Y, Ma D. Wnt/beta-catenin signaling regulates cancer stem cells in lung cancer A549 cells. Biochem Biophys Res Commun. 2010;392(3):373–379. | ||
Yallapu MM, Maher DM, Sundram V, Bell MC, Jaggi M, Chauhan SC. Curcumin induces chemo/radio-sensitization in ovarian cancer cells and curcumin nanoparticles inhibit ovarian cancer cell growth. J Ovarian Res. 2010;3:11. | ||
Tang Q, Wang Y, Huang R, et al. Preparation of anti-tumor nanoparticle and its inhibition to peritoneal dissemination of colon cancer. PLoS One. 2014;9(6):e98455. | ||
Segditsas S, Tomlinson I. Colorectal cancer and genetic alterations in the Wnt pathway. Oncogene. 2006;25(57):7531–7537. | ||
Najdi R, Holcombe RF, Waterman ML. Wnt signaling and colon carcinogenesis: beyond APC. J Carcinog. 2011;10:5. | ||
Yin L, Velazquez OC, Liu ZJ. Notch signaling: emerging molecular targets for cancer therapy. Biochem Pharmacol. 2010;80(5):690–701. | ||
Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: cell fate control and signal integration in development. Science. 1999;284(5415):770–776. | ||
Androutsellis-Theotokis A, Leker RR, Soldner F, et al. Notch signalling regulates stem cell numbers in vitro and in vivo. Nature. 2006;442(7104):823–826. | ||
Tien AC, Rajan A, Bellen HJ. A Notch updated. J Cell Biol. 2009;184(5):621–629. | ||
Cerdan C, Bhatia M. Novel roles for Notch, Wnt and Hedgehog in hematopoesis derived from human pluripotent stem cells. Int J Dev Biol. 2010;54(6–7):955–963. | ||
Bray SJ. Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol. 2006;7(9):678–689. | ||
Hovinga KE, Shimizu F, Wang R, et al. Inhibition of notch signaling in glioblastoma targets cancer stem cells via an endothelial cell intermediate. Stem Cells. 2010;28(6):1019–1029. | ||
Fan X, Khaki L, Zhu TS, et al. NOTCH pathway blockade depletes CD133-positive glioblastoma cells and inhibits growth of tumor neurospheres and xenografts. Stem Cells. 2010;28(1):5–16. | ||
Liu J, Mao Z, Huang J, Xie S, Liu T, Mao Z. Blocking the NOTCH pathway can inhibit the growth of CD133-positive A549 cells and sensitize to chemotherapy. Biochem Biophys Res Commun. 2014;444(4):670–675. | ||
Moellering RE, Cornejo M, Davis TN, et al. Direct inhibition of the NOTCH transcription factor complex. Nature. 2009;462(7270):182–188. | ||
Yan M, Callahan CA, Beyer JC, et al. Chronic DLL4 blockade induces vascular neoplasms. Nature. 2010;463(7282):E6–E7. | ||
van Es JH, van Gijn ME, Riccio O, et al. Notch/gamma-secretase inhibition turns proliferative cells in intestinal crypts and adenomas into goblet cells. Nature. 2005;435(7044):959–963. | ||
Wong GT, Manfra D, Poulet FM, et al. Chronic treatment with the gamma-secretase inhibitor LY-411,575 inhibits beta-amyloid peptide production and alters lymphopoiesis and intestinal cell differentiation. J Biol Chem. 2004;279(13):12876–12882. | ||
Rosenholm JM, Peuhu E, Bate-Eya LT, Eriksson JE, Sahlgren C, Linden M. Cancer-cell-specific induction of apoptosis using mesoporous silica nanoparticles as drug-delivery vectors. Small. 2010;6(11):1234–1241. | ||
Rosenholm JM, Meinander A, Peuhu E, et al. Targeting of porous hybrid silica nanoparticles to cancer cells. ACS Nano. 2009;3(1):197–206. | ||
Mamaeva V, Rosenholm JM, Bate-Eya LT, et al. Mesoporous silica nanoparticles as drug delivery systems for targeted inhibition of Notch signaling in cancer. Mol Ther. 2011;19(8):1538–1546. | ||
Steg AD, Katre AA, Goodman B, et al. Targeting the notch ligand JAGGED1 in both tumor cells and stroma in ovarian cancer. Clin Cancer Res. 2011;17(17):5674–5685. | ||
Greenow K, Clarke AR. Controlling the stem cell compartment and regeneration in vivo: the role of pluripotency pathways. Physiol Rev. 2012;92(1):75–99. | ||
Wang W, Rigueur D, Lyons KM. TGFbeta signaling in cartilage development and maintenance. Birth Defects Res C Embryo Today. 2014;102(1):37–51. | ||
Moses H, Barcellos-Hoff MH. TGF-beta biology in mammary development and breast cancer. Cold Spring Harb Perspect Biol. 2011;3(1):a003277. | ||
Bellam N, Pasche B. Tgf-beta signaling alterations and colon cancer. Cancer Treat Res. 2010;155:85–103. | ||
Wu K, Ding J, Chen C, et al. Hepatic transforming growth factor beta gives rise to tumor-initiating cells and promotes liver cancer development. Hepatology. 2012;56(6):2255–2267. | ||
Ischenko I, Liu J, Petrenko O, Hayman MJ. Transforming growth factor-beta signaling network regulates plasticity and lineage commitment of lung cancer cells. Cell Death Differ. 2014;21(8):1218–1228. | ||
Gao J, Zhu Y, Nilsson M, Sundfeldt K. TGF-beta isoforms induce EMT independent migration of ovarian cancer cells. Cancer Cell Int. 2014;14(1):72. | ||
Massague J. How cells read TGF-beta signals. Nat Rev Mol Cell Biol. 2000;1(3):169–178. | ||
Itoh S, Itoh F, Goumans MJ, Ten Dijke P. Signaling of transforming growth factor-beta family members through Smad proteins. Eur J Biochem. 2000;267(24):6954–6967. | ||
Kim HG, Chung YH, Song BC, et al. Expression of transforming growth factor beta-1 in chronic hepatitis and hepatocellular carcinoma associated with hepatitis C virus infection. Korean J Int Med. 2000;15(3):165–170. | ||
Tsai JF, Jeng JE, Chuang LY, et al. Elevated urinary transforming growth factor-beta1 level as a tumour marker and predictor of poor survival in cirrhotic hepatocellular carcinoma. Br J Cancer. 1997;76(2):244–250. | ||
Tsai JF, Chuang LY, Jeng JE, et al. Clinical relevance of transforming growth factor-beta 1 in the urine of patients with hepatocellular carcinoma. Medicine. 1997;76(3):213–226. | ||
Lu Y, Wu LQ, Li CS, Wang SG, Han B. Expression of transforming growth factors in hepatocellular carcinoma and its relations with clinicopathological parameters and prognosis. Hepatobiliary Pancreat Dis Int. 2008;7(2):174–178. | ||
Ikeguchi M, Iwamoto A, Taniguchi K, Katano K, Hirooka Y. The gene expression level of transforming growth factor-beta (TGF-beta) as a biological prognostic marker of hepatocellular carcinoma. J Exp Clin Cancer Res. 2005;24(3):415–421. | ||
Liu Z, Bandyopadhyay A, Nichols RW, et al. Blockade of Autocrine TGF-beta Signaling Inhibits Stem Cell Phenotype, Survival, and Metastasis of Murine Breast Cancer Cells. J Stem Cell Res Ther. 2012;2(1):1–8. | ||
Mishra L, Shetty K, Tang Y, Stuart A, Byers SW. The role of TGF-beta and Wnt signaling in gastrointestinal stem cells and cancer. Oncogene. 2005;24(37):5775–5789. | ||
Amin R, Mishra L. Liver stem cells and tgf-Beta in hepatic carcinogenesis. Gastrointest Cancer Res. 2008;2(4 suppl):S27–S30. | ||
Gomez-Casal R, Bhattacharya C, Ganesh N, et al. Non-small cell lung cancer cells survived ionizing radiation treatment display cancer stem cell and epithelial-mesenchymal transition phenotypes. Mol Cancer. 2013;12(1):94. | ||
Meng H, Zhao Y, Dong J, et al. Two-wave nanotherapy to target the stroma and optimize gemcitabine delivery to a human pancreatic cancer model in mice. ACS Nano. 2013;7(11):10048–10065. | ||
Shukla R, Bansal V, Chaudhary M, Basu A, Bhonde RR, Sastry M. Biocompatibility of gold nanoparticles and their endocytotic fate inside the cellular compartment: a microscopic overview. Langmuir. 2005;21(23):10644–10654. | ||
Tsai YS, Chen YH, Cheng PC, et al. TGF-beta1 conjugated to gold nanoparticles results in protein conformational changes and attenuates the biological function. Small. 2013;9(12):2119–2128. | ||
Nusslein-Volhard C, Wieschaus E. Mutations affecting segment number and polarity in Drosophila. Nature. 1980;287(5785):795–801. | ||
Lee JJ, von Kessler DP, Parks S, Beachy PA. Secretion and localized transcription suggest a role in positional signaling for products of the segmentation gene hedgehog. Cell. 1992;71(1):33–50. | ||
Mohler J, Vani K. Molecular organization and embryonic expression of the hedgehog gene involved in cell-cell communication in segmental patterning of Drosophila. Development. 1992;115(4):957–971. | ||
Tabata T, Eaton S, Kornberg TB. The Drosophila hedgehog gene is expressed specifically in posterior compartment cells and is a target of engrailed regulation. Genes Dev. 1992;6(12B):2635–2645. | ||
Bai LY, Chiu CF, Lin CW, et al. Differential expression of Sonic hedgehog and Gli1 in hematological malignancies. Leukemia. 2008;22(1):226–228. | ||
Villavicencio EH, Walterhouse DO, Iannaccone PM. The sonic hedgehog-patched-gli pathway in human development and disease. Am J Hum Genet. 2000;67(5):1047–1054. | ||
Rubin LL, de Sauvage FJ. Targeting the Hedgehog pathway in cancer. Nat Rev Drug Discov. 2006;5(12):1026–1033. | ||
Ingham PW, McMahon AP. Hedgehog signaling in animal development: paradigms and principles. Genes Dev. 2001;15(23):3059–3087. | ||
Gulino A, Ferretti E, De Smaele E. Hedgehog signalling in colon cancer and stem cells. EMBO Mol Med. 2009;1(6–7):300–302. | ||
Xu Q, Yuan X, Liu G, Black KL, Yu JS. Hedgehog signaling regulates brain tumor-initiating cell proliferation and portends shorter survival for patients with PTEN-coexpressing glioblastomas. Stem Cells. 2008;26(12):3018–3026. | ||
Ramaswamy B, Lu Y, Teng KY, et al. Hedgehog signaling is a novel therapeutic target in tamoxifen-resistant breast cancer aberrantly activated by PI3K/AKT pathway. Cancer Res. 2012;72(19):5048–5059. | ||
Kappler R, von Schweinitz D. A better way forward: targeting hedgehog signaling in liver cancer. Front Biosci. 2012;4:277–286. | ||
Park KS, Martelotto LG, Peifer M, et al. A crucial requirement for Hedgehog signaling in small cell lung cancer. Nat Med. 2011;17(11):1504–1508. | ||
Bhattacharya R, Kwon J, Ali B, et al. Role of hedgehog signaling in ovarian cancer. Clin Cancer Res. 2008;14(23):7659–7666. | ||
Bisht S, Brossart P, Maitra A, Feldmann G. Agents targeting the Hedgehog pathway for pancreatic cancer treatment. Curr Opin Investig Drugs. 2010;11(12):1387–1398. | ||
Xu Y, Chenna V, Hu C, et al. Polymeric nanoparticle-encapsulated hedgehog pathway inhibitor HPI-1 (NanoHHI) inhibits systemic metastases in an orthotopic model of human hepatocellular carcinoma. Clin Cancer Res. 2012;18(5):1291–1302. | ||
Verma RK, Yu W, Singh SP, Shankar S, Srivastava RK. Anthothecol-encapsulated PLGA nanoparticles inhibit pancreatic cancer stem cell growth by modulating sonic hedgehog pathway. Nanomedicine. 2015; S1549-9634(15)00144-6. [Epub ahead of print]. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.