Back to Journals » Drug Design, Development and Therapy » Volume 11
Synergistic effects of antimicrobial peptide DP7 combined with antibiotics against multidrug-resistant bacteria
Authors Wu X, Li Z , Li X, Tian Y, Fan Y, Yu C, Zhou B, Liu Y, Xiang R, Yang L
Received 25 February 2016
Accepted for publication 7 April 2016
Published 22 March 2017 Volume 2017:11 Pages 939—946
DOI https://doi.org/10.2147/DDDT.S107195
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Prof. Dr. Wei Duan
Xiaozhe Wu,1 Zhan Li,1 Xiaolu Li,2,3 Yaomei Tian,1 Yingzi Fan,1 Chaoheng Yu,1 Bailing Zhou,1 Yi Liu,4 Rong Xiang,5 Li Yang1
1State Key Laboratory of Biotherapy/Collaborative Innovation Center of Biotherapy, West China Hospital, Sichuan University, 2International Center for Translational Chinese Medicine, Sichuan Academy of Chinese Medicine Sciences, Chengdu, 3Department of Plastic and Burn Surgery, Affiliated Hospital of Southwest Medical University, Luzhou, 4Department of Microbial Examination, Sichuan Center for Disease Control and Prevention, Chengdu, 5Nankai University School of Medicine, Tianjin, People’s Republic of China
Abstract: Antibiotic-resistant bacteria present a great threat to public health. In this study, the synergistic effects of antimicrobial peptides (AMPs) and antibiotics on several multidrug-resistant bacterial strains were studied, and their synergistic effects on azithromycin (AZT)-resistance genes were analyzed to determine the relationships between antimicrobial resistance and these synergistic effects. A checkerboard method was used to evaluate the synergistic effects of AMPs (DP7 and CLS001) and several antibiotics (gentamicin, vancomycin [VAN], AZT, and amoxicillin) on clinical bacterial strains (Staphylococcus aureus, Pseudomonas aeruginosa, Acinetobacter baumannii, and Escherichia coli). The AZT-resistance genes (ermA, ermB, ermC, mefA, and msrA) were identified in the resistant strains using quantitative polymerase chain reaction. For all the clinical isolates tested that were resistant to different antibiotics, DP7 had high antimicrobial activity (≤32 mg/L). When DP7 was combined with VAN or AZT, the effect was most frequently synergistic. When we studied the resistance genes of the AZT-resistant isolates, the synergistic effect of DP7–AZT occurred most frequently in highly resistant strains or strains carrying more than two AZT-resistance genes. A transmission electron microscopic analysis of the S. aureus strain synergistically affected by DP7–AZT showed no noteworthy morphological changes, suggesting that a molecular-level mechanism plays an important role in the synergistic action of DP7–AZT. AMP DP7 plus the antibiotic AZT or VAN is more effective, especially against highly antibiotic-resistant strains.
Keywords: antimicrobial peptide, drug synergy, resistant bacteria
Introduction
With the ongoing use of antibiotics, increasing numbers of bacteria are becoming resistant to them. Strategies to address this problem include the identification of new antibiotics, the application of the principles of microecology, or the development of antimicrobial peptides (AMPs). AMPs are one of the most promising antibacterial drugs that can be substituted for antibiotics because they have advantages such as high antimicrobial efficacy1–3 and broad-spectrum activity.1,4 Another way to resolve the problem of multidrug-resistant bacteria is by combining different drugs. The effects of AMPs combined with antibiotics often exceed those of the individual drugs, whether in antibacterial activities5–8 or in interruption of biofilm formation.9–13 Drug design research faces a long and difficult challenge to overcome drug resistance, whereas drug combinations require no drug modification and lower doses can be used while enhancing the therapeutic potential.14,15
DP7 is a recently designed AMP derived from Bac2A and has demonstrated good broad-spectrum antimicrobial activity and safety in previous studies.16 CLS001 is an AMP that is currently under investigation in a Phase III clinical trial.17,18 In this study, we evaluated the in vitro antimicrobial activities of gentamicin (GEN), amoxicillin (AMO), azithromycin (AZT), and vancomycin (VAN), alone and in combination with DP7 or CLS001, against antibiotic-resistant Staphylococcus aureus, Pseudomonas aeruginosa, Acinetobacter baumannii, and Escherichia coli strains. The AZT-resistance mechanism was also examined to clarify the synergistic mechanism involved and the traits of multidrug-resistant bacteria in general.
Materials and methods
Bacterial strains
The clinical isolates used in this study were isolated and obtained from the Southwest Hospital of China and the Sichuan Center for Disease Control and Prevention; duplicate sample from one patient was excluded. The species were confirmed using the Bacteria Identification Microbial DNA qPCR Multi-Assay Kit (Qiagen NV, Venlo, the Netherlands).
Antimicrobial agents
The peptides were synthesized with Fmoc chemistry by Shanghai Applied Protein Technology Co., Ltd (Shanghai, People’s Republic of China). VAN hydrochloride was purchased from Gentle Pharmaceutical Corporation Kashin Medicines Co., Ltd (Taiwan, Republic of China), AZT powder was purchased from Chen Mei Chemical Co., Ltd (Zhengzhou, People’s Republic of China), gentamicin sulfate was purchased from Hubei Pharmaceutical Co., Ltd (Hubei, People’s Republic of China), and AMO was purchased from Kunming Baker Norton Pharmaceutical Co., Ltd (Kunming, People’s Republic of China).
Susceptibility test
The minimum inhibitory concentrations (MICs) of the antimicrobial drugs were determined using the broth microdilution method, according to Clinical and Laboratory Standards Institute (CLSI) file M7-A7. Serial dilutions of the peptides were prepared in Müller–Hinton broth, and 100 μL of each concentration was added to triplicate wells in a 96-well flat-bottom tissue culture plate. The bacteria were grown on Müller–Hinton agar for 24 hours. A few colonies were diluted to ~1×106 cfu/mL with Müller–Hinton broth, and 100 μL was added in triplicate to each well containing the antimicrobial drugs. The final drug concentrations ranged from 256 mg/L to 0.25 mg/L. The suspensions were then incubated at 37°C for 20 hours. Bacterial concentration was calculated through optical density, and the lowest concentration causing 80% growth inhibition relative to the growth of the control was deemed to be the MIC.
Checkerboard assay
The activities of the AMPs in combination with the antibiotics were analyzed using the checkerboard broth dilution method19 to determine the fractional inhibitory concentration indices (FICIs), calculated as: FICI = MIC of drug A in combination/MIC of drug A alone + MIC of drug B in combination/MIC of drug B alone. Twice the maximum concentration was used in calculation if the MIC was out of measurement in this study. The calculated FICI was interpreted as synergistic (FICI ≤0.5), additive (0.5< FICI <1), indifferent (1≤ FICI <4.0), or antagonistic (FICI ≥4.0).20
Gene identification using quantitative polymerase chain reaction
Total DNA was collected from the bacteria using the DNeasy Blood & Tissue Kit (Qiagen NV). DNA concentration was determined with a Thermo Scientific NanoDrop 2000 spectrophotometer. DNA samples with concentrations (measured at A260) >10 ng/mL and A260/A280 ratios >1.8 were used in the quantitative polymerase chain reaction (qPCR) assay. Bacterial genes, VAN-resistance genes, and AZT-resistance genes were identified with the Microbial DNA qPCR Multi-Assay Kits from Qiagen NV. The level of gene expression was determined as the difference between the quantification cycle (Cq) value of the target sample and that of the nontemplate control (NTC), Cq > NTC −3 was deemed negative for gene expression. Cq < NTC −6 was deemed positive for gene expression; and NTC −6> Cq > NTC −3 was deemed inconclusive.
Transmission electron microscopy
Mid-logarithmic phase S. aureus cells (1×108) were treated with DP7, AZT, or DP7–AZT, and a no-peptide control was included. After incubation for 60 minutes, the bacteria were pelleted by centrifugation at 900× g for 10 minutes and washed twice with 1 M phosphate-buffered saline. The cells were then fixed with 2.5% glutaraldehyde/phosphate-buffered saline at 4°C overnight. The prepared bacterial samples were sent to the Center of Forecasting and Analysis of Sichuan University (Sichuan, People’s Republic of China) for imaging with transmission electron microscopy (TEM, Tecnai G2 F20 S-TWIN, FEI).
Results
MIC distribution
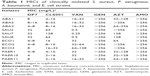
The MIC of each antibiotic and AMP was determined using the broth microdilution method before the synergy test, and the highest concentration obtained was 256 mg/L. Clinical isolates of S. aureus (SAU series), P. aeruginosa (PAER series), A. baumannii (ABA series), and E. coli (ECO series) were examined (Table 1). Most bacterial strains were resistant to the tested antibiotics (GEN, VAN, AMO, and AZT), and GEN and AMO showed negligible activity against these strains. Only strain SAU4 was sensitive to VAN and AZT. Both AMPs CLS001 and DP7 showed strong activity against the clinically isolated strains in this study, while the antimicrobial activity of DP7 was slightly better than that of CLS001.
Synergy assay
The microdilution checkboard method was used to evaluate the synergistic effects of the antibiotics combined with the AMPs, which were determined as FICIs (Table 2). The effects of the DP7–VAN combination were synergistic against 50% of all the isolates tested and additive against 40% of the isolates. The effects of the DP7–AZT combination were synergistic against 60% of the isolates and additive against 20% of the isolates. These two combinations were the most promising of those tested against the antibiotic-resistant bacteria in this study. The percentages of isolates affected synergistically and additively by DP7–AMO were 20% and 10%, respectively, whereas the DP7–GEN combination synergistically affected only 20% of isolates and additively affected no isolates. The synergistic activities of CLS001 combined with these antibiotics were also tested and compared with those of DP7. The synergistic effect rates were 20% for CLS001–VAN, 20% for CLS001–GEN, 10% for CLS001–AZT, and 40% for CLS001–AMO. The effects of CLS001–VAN and CLS001–AZT were additive in 30% and 50% of isolates, respectively, whereas no additive effect was observed for CLS001–GEN or CLS001–AMO. These results indicate that DP7 had a higher synergy rate than CLS001 when combined with VAN or AZT. Another remarkable result was that the combination of the two AMPs, DP7 plus CLS001, showed no synergistic effect on any bacterial isolate tested but had additive effects on 40% of isolates.
Expanded synergy assay
Because DP7–VAN and DP7–AZT had relatively high synergy rates, these two combinations were tested on ten more S. aureus and P. aeruginosa clinical strains to study the synergistic effects on gram-positive and gram-negative bacteria (Table 3). In S. aureus, DP7–VAN had synergistic effects on 40% of isolates and additive effects on another 40% of isolates. DP7–AZT had synergistic effects on 50% of isolates, but an additive on no isolate. In P. aeruginosa, DP7–VAN also showed high synergy rates, exerting a synergistic effect on 50% of isolates and an additive effect on 20% of isolates. However, DP7–AZT had a synergistic effect on only 10% of isolates and an additive effect on 30% of isolates. Although DP7–AZT showed low activity against P. aeruginosa, it showed relatively high activity against AZT-resistant S. aureus strains. DP7–VAN exerted synergistic effects against both S. aureus and P. aeruginosa strains that were either resistant to VAN or not.
Resistance mechanism
Isolates resistant to VAN or AZT were selected to analyze their resistance genes. The VAN-resistance genes vanB and vanC were analyzed with qPCR, but none of the VAN-resistant isolates tested was positive for either gene (data not shown). The occurrence of five AZT-resistance genes (ermA, ermB, ermC, mefA, and msrA) in the AZT-resistant isolates was investigated. Of the 13 AZT-resistant P. aeruginosa isolates, eleven carried ermB and six carried ermC (all of which also carried ermB). Among the eleven S. aureus isolates, nine were resistant to AZT, and of these, four carried ermA, six carried ermB, six carried ermC, two carried both ermA and ermC, three carried both ermB and ermC, and one carried ermA, ermB, and ermC. No ermA gene was found in any P. aeruginosa isolate, and neither the mefA nor msrA gene was detected in any P. aeruginosa or S. aureus isolate in this study (Tables 4 and 5).
  | Table 4 Resistance rates and synergy rates for each type of bacterium |
Relationships between resistance genes, resistance, and synergy
The relationships between the AZT-resistance mechanism, the MICs for AZT, and the FICIs for DP7–AZT were analyzed. As expected, isolates carrying more erm-resistance genes had higher MICs for AZT. Isolate SAU2 carrying the ermA, ermB, and ermC genes had the highest MIC for AZT (>256 mg/L). Of the isolates with two erm genes, 70% (7/10) were highly resistant to AZT (MIC ≥128 mg/L). Of the isolates with one erm gene, 71.4% (5/7) had MICs ≥128 mg/L. Isolates with no erm genes had MICs ≤32 mg/L. The number of resistance genes also had some relationship with the FICI, in that isolates with more resistance genes showed lower FICIs. Of the nine isolates upon which DP7–AZT exerted a synergistic effect, seven (77.8%) carried more than two resistance genes. Of the four isolates upon which DP7–AZT exerted an additive effect, two (50%) carried two resistance genes. Among another nine isolates upon which DP7–AZT exerted an indifferent affect, only three (33.3%) carried two resistance genes. Therefore, bacterial resistance to AZT increased as the number of AZT-resistance genes increased, and the likelihood of a synergistic DP7–AZT effect increased in parallel. The reason why DP7–AZT exerts a greater synergistic effect on more strongly AZT-resistant bacteria warrants further study.
Morphological study of S. aureus strain S5375 treated with DP7–AZT
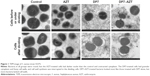
The antimicrobial mechanism of DP7 acts by disrupting the bacterial cell wall and influencing the cell membrane.16 Therefore, the synergistic action of DP7–AZT on the bacterial morphology was investigated. S.aureus strain S5375, which is resistant to AZT but showed a synergistic response to DP7–AZT, was treated with 4× MICs of AZT for 60 minutes, and any damaging effects were observed with TEM (Figure 1). Compared with the bacteria in the control group, the bacteria treated with AZT had darker nuclei and clearer cell walls, which may be attributable to interrupted protein synthesis, with a consequent increased transcription activity in the nucleus and reduction in cell wall proteins. However, in the DP7-treated group, no nuclei were observed, the whole cytoplasm was a mosaic, and the cell edges were fuzzy. Interestingly, the cell walls of dividing cells were detached, indicating that cell wall synthesis was interrupted. However, the shape of the bacteria treated with DP7–AZT did not differ greatly from those treated with DP7 alone, which suggests that DP7 functions earlier than AZT. The only difference between the DP7–AZT-treated and DP7-treated bacteria was that even some nondividing bacteria had no cell walls after treatment with DP7–AZT. This may be attributable to the combined action of the reduced cell wall proteins caused by AZT and the cell wall disruption caused by DP7. These results suggest that the synergistic mechanism of DP7–AZT acts at the molecular level.
Discussion
Many AMPs have been developed in recent years as alternatives to antibiotics to resolve the problem of antimicrobial resistance.21–23 The activity of one of these AMPs, DP7, against antibiotic-resistant clinical isolates was investigated in this study. The broad-spectrum, stable antimicrobial activities of DP7 and CLS001 were confirmed in susceptibility tests against S. aureus, P. aeruginosa, A. baumannii, and E. coli isolates. Although the MICs of the AMPs are usually higher than those of antibiotics,1,4 AMPs are still a promising group of antimicrobial drugs, and AMP–antibiotic combinations have also effectively overcome many drug-resistant bacteria in recent years.5–7
Several studies have shown that antibacterial drugs with similar mechanisms or that influence the same target exert synergistic effects more readily than combinations of very different drugs, and several synergistic mechanisms have been analyzed. However, in this study, DP7 exerted a highly synergistic effect when combined with AZT or VAN. The antibacterial mechanism of AZT involves its combination with 50S ribosomal RNA (rRNA), thus suppressing the transcription process. However, VAN inhibits cell wall synthesis in gram-positive bacteria by binding to D-alanine residues in the glycopeptide chain. Therefore, it is quite strange that DP7 exerts highly synergistic effects with two antibiotics that have totally different mechanisms. The antimicrobial mechanism of GEN involves its binding to 30S rRNA, which is similar to that of AZT, but GEN showed almost no synergy with DP7. It is noteworthy that VAN is only functional against gram-positive bacteria but acted synergistically with DP7 against several gram-negative bacteria in this study. When combined with our TEM results, these data indicate that as well as its cell wall-disrupting function, DP7 has a complex molecular-level antimicrobial mechanism, which requires investigation.
The most frequent VAN-resistance genes are the van genes,24,25 and the most frequent AZT-resistance genes are the erm genes.26,27 In this study, neither vanB nor vanC was detected, while several AZT-resistance genes were detected. Genes ermA, ermB, and ermC encode methyl transferases that can change the structure of rRNA, and the altered AZT-binding site is responsible for the bacterium’s resistance to AZT. Resistance genes msrA and mefA encode a macrolide efflux pump system.26,28,29 Our analysis showed that the ermB and ermC genes were most common in the AZT-resistant isolates and two or more erm genes appeared together quite frequently (12/22). However, neither msrA nor mefA was related to the AZT resistance of the clinical isolates tested. This result is consistent with other studies of the relative resistance conferred by different genes.29 In this study, the synergistic activity of AZT showed that the synergistic effect of AZT–DP7 was greatest in the most highly resistant bacteria, but among the eight S. aureus isolates sensitive to VAN, VAN–DP7 exerted synergistic effects on three and additive effects on four. Therefore, an antibiotic–DP7 combination also exerted highly synergistic effects against antibiotic-sensitive bacteria. Although the target strain was not drug resistant, the sub-MIC concentration used in the drug combination also reduces the toxicity or adverse effects of the antibiotic and prevents the development of drug resistance.2
Conclusion
Several drug combination studies published in recent years were strain or clone specific,30,31 but a drug or drug combination that is therapeutic against many types of antibiotic-resistant bacteria is ideal. However, further studies of DP7 must be undertaken. This study not only confirmed that AMPs are broad-spectrum antimicrobial drugs effective against multidrug-resistant bacteria but also suggests that when antibiotics are combined with the AMP DP7, some antibiotic-resistant bacteria can be eradicated. However, several antibiotic-sensitive bacteria were most sensitive to the combination, and several antibiotics that only act against gram-positive bacteria acted across a broader spectrum when administered in combination with an AMP.
Acknowledgments
The study was mainly supported by the National Major Scientific and Technological Special Project for “Significant New Drugs Development” (grant no 2013ZX09102030) and the National Natural Science Foundation of China (grant no 81171780).
Disclosure
The authors report no conflicts of interest in this work.
References
Fuchs PC, Barry AL, Brown SD. In vitro antimicrobial activity of MSI-78, a magainin analog. Antimicrob Agents Chemother. 1998;42(5):1213–1216. | ||
Andersson DI, Hughes D. Microbiological effects of sublethal levels of antibiotics. Nat Rev Microbiol. 2014;12(7):465–478. | ||
Makobongo MO, Gancz H, Carpenter BM, McDaniel DP, Merrell DS. The oligo-acyl lysyl antimicrobial peptide C(1)(2)K-2beta(1)(2) exhibits a dual mechanism of action and demonstrates strong in vivo efficacy against Helicobacter pylori. Antimicrob Agents Chemother. 2012;56(1):378–390. | ||
Ge Y, MacDonald DL, Holroyd KJ, Thornsberry C, Wexler H, Zasloff M. In vitro antibacterial properties of pexiganan, an analog of magainin. Antimicrob Agents Chemother. 1999;43(4):782–788. | ||
Livermore DM, Warner M, Mushtaq S. Activity of MK-7655 combined with imipenem against Enterobacteriaceae and Pseudomonas aeruginosa. J Antimicrob Chemother. 2013;68(10):2286–2290. | ||
Park Y, Kim HJ, Hahm KS. Antibacterial synergism of novel antibiotic peptides with chloramphenicol. Biochem Biophys Res Commun. 2004;321(1):109–115. | ||
Giacometti A, Cirioni O, Kamysz W, et al. In vitro activity of MSI-78 alone and in combination with antibiotics against bacteria responsible for bloodstream infections in neutropenic patients. Int J Antimicrob Agents. 2005;26(3):235–240. | ||
Cirioni O, Silvestri C, Ghiselli R, et al. Protective effects of the combination of alpha-helical antimicrobial peptides and rifampicin in three rat models of Pseudomonas aeruginosa infection. J Antimicrob Chemother. 2008;62(6):1332–1338. | ||
Dosler S, Karaaslan E. Inhibition and destruction of Pseudomonas aeruginosa biofilms by antibiotics and antimicrobial peptides. Peptides. 2014;62:32–37. | ||
Dosler S, Karaaslan E, Alev Gerceker A. Antibacterial and anti-biofilm activities of melittin and colistin, alone and in combination with antibiotics against Gram-negative bacteria. J Chemother. Epub 2015 Mar 24. | ||
Reffuveille F, de la Fuente-Nunez C, Mansour S, Hancock RE. A broad-spectrum antibiofilm peptide enhances antibiotic action against bacterial biofilms. Antimicrob Agents Chemother. 2014;58(9):5363–5371. | ||
de la Fuente-Nunez C, Reffuveille F, Mansour SC, et al. D-enantiomeric peptides that eradicate wild-type and multidrug-resistant biofilms and protect against lethal Pseudomonas aeruginosa infections. Chem Biol. 2015;22(2):196–205. | ||
de la Fuente-Nunez C, Cardoso MH, de Souza Candido E, Franco OL, Hancock RE. Synthetic antibiofilm peptides. Biochim Biophys Acta. 2016;1858(5):1061–1069. | ||
Anantharaman A, Rizvi MS, Sahal D. Synergy with rifampin and kanamycin enhances potency, kill kinetics, and selectivity of de novo-designed antimicrobial peptides. Antimicrob Agents Chemother. 2010;54(5):1693–1699. | ||
Desbois AP, Coote PJ. Bactericidal synergy of lysostaphin in combination with antimicrobial peptides. Eur J Clin Microbiol Infect Dis. 2011;30(8):1015–1021. | ||
Wu X, Wang Z, Li X, et al. In vitro and in vivo activities of antimicrobial peptides developed using an amino acid-based activity prediction method. Antimicrob Agents Chemother. 2014;58(9):5342–5349. | ||
Roscia G, Falciani C, Bracci L, Pini A. The development of antimicrobial peptides as new antibacterial drugs. Curr Protein Pept Sci. 2013;14(8):641–649. | ||
Hilchie AL, Wuerth K, Hancock RE. Immune modulation by multifaceted cationic host defense (antimicrobial) peptides. Nat Chem Biol. 2013;9(12):761–768. | ||
JA M. Synergism testing: broth microdilution checkerboard and broth macrodilution methods. In: Garcia LS, editor. Clinical Microbiology Procedures Handbook. 2nd ed. Washington, DC: ASM Press; 2007:1–23. | ||
Hauser C, Hirzberger L, Unemo M, Furrer H, Endimiani A. In vitro activity of fosfomycin alone and in combination with ceftriaxone or azithromycin against clinical Neisseria gonorrhoeae isolates. Antimicrob Agents Chemother. 2015;59(3):1605–1611. | ||
Deslouches B, Gonzalez IA, DeAlmeida D, et al. De novo-derived cationic antimicrobial peptide activity in a murine model of Pseudomonas aeruginosa bacteraemia. J Antimicrob Chemother. 2007;60(3):669–672. | ||
Kim H, Jang JH, Kim SC, Cho JH. De novo generation of short antimicrobial peptides with enhanced stability and cell specificity. J Antimicrob Chemother. 2014;69(1):121–132. | ||
Scott MG, Dullaghan E, Mookherjee N, et al. An anti-infective peptide that selectively modulates the innate immune response. Nat Biotechnol. 2007;25(4):465–472. | ||
Gold HS. Vancomycin-resistant enterococci: mechanisms and clinical observations. Clin Infect Dis. 2001;33(2):210–219. | ||
Courvalin P. Vancomycin resistance in gram-positive cocci. Clin Infect Dis. 2006;42(suppl 1):S25–S34. | ||
Zmantar T, Bekir K, Elgarsadi SI, Hadad O, Bakhrouf A. Molecular investigation of antibiotic resistance genes in methicillin resistant Staphylococcus aureus isolated from nasal cavity in pediatric service. Afr J Microbiol Res. 2013;7(34):4414–4421. | ||
Schmitz FJ, Sadurski R, Kray A, et al. Prevalence of macrolide-resistance genes in Staphylococcus aureus and Enterococcus faecium isolates from 24 European university hospitals. J Antimicrob Chemother. 2000;45(6):891–894. | ||
Yin J, Yu S, Liu X, et al. Prevalence and mechanism of resistance to antimicrobial agents in group G streptococcal isolates from China. Antimicrob Agents Chemother. 2011;55(1):402–404. | ||
Ding ZF, Zhang H, Tang W, et al. Methylase genes-mediated erythromycin resistance in Staphylococcus aureus from bovine mastitis in China. Isr J Vet Med. 2012;67(3):170–179. | ||
Furuya R, Koga Y, Irie S, et al. In vitro activities of antimicrobial combinations against clinical isolates of Neisseria gonorrhoeae. J Infect Chemother. 2013;19(6):1218–1220. | ||
Barbee LA, Soge OO, Holmes KK, Golden MR. In vitro synergy testing of novel antimicrobial combination therapies against Neisseria gonorrhoeae. J Antimicrob Chemother. 2014;69(6):1572–1578. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.