Back to Journals » International Journal of Nanomedicine » Volume 15
Study on Adenovirus Infection in vitro with Nanoself-Assembling Peptide as Scaffolds for 3D Culture
Authors Ao DS , Gao LY, Gu JH, Qiao JH, Wang H, Liu YF , Song H
Received 8 April 2020
Accepted for publication 30 July 2020
Published 24 August 2020 Volume 2020:15 Pages 6327—6338
DOI https://doi.org/10.2147/IJN.S239395
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Linlin Sun
Di-Shu Ao,1,* Lu-Yao Gao,1,* Jing-Han Gu,1 Jun-Hua Qiao,1 Huan Wang,1 Yan-Fei Liu,2 Hong Song1
1Department of Microbiology, School of Basic Medical Sciences, Zunyi Medical University, Zunyi 563000, People’s Republic of China; 2Key Laboratory of Cell Engineering of Guizhou Province, Affiliated Hospital of Zunyi Medical University, Zunyi 563000, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Yan-Fei Liu
Key Laboratory of Cell Engineering of Guizhou Province, Affiliated Hospital of Zunyi Medical University, Zunyi, 563000, China
Tel +86-18286276104
Email [email protected]
Hong Song
Department of Microbiology, School of Basic Medical Sciences, Zunyi Medical University, Zunyi 563000, People’s Republic of China
Tel +86-18985259977
Email [email protected]
Purpose: To construct a three-dimensional (3D) culture model of adenovirus in vitro using the nanoself-assembling peptide RADA16-I as a 3D cell culture scaffold combined with virology experimental technology to provide a novel research method for virus isolation and culture, pathogenesis research, antiviral drug screening and vaccine preparation.
Methods: The nanoself-assembling peptide RADA16-I was used as a 3D scaffold material for 293T cell culture, and adenovirus was cultured in the cells. The growth, morphological characteristics and pathological effects of 3D-cultured 293T cells after adenovirus infection were observed with an inverted microscope and MTS. The proliferation of adenovirus in 293T cells was observed by TEM and detected by qPCR. The levels of TNF-α and IL-8 secreted by adenovirus-infected 293T cells in the RADA16-I 3D culture system were detected by ELISA.
Results: The 293T cells grew well in the RADA16-I 3D culture system for a prolonged period of time. The adenovirus infection persisted for a long time with multiple proliferation peaks, which closely resembled those of in vivo infections. The adenovirus virions amplified in the 3D system remained infectious. There were multiple secretion peaks of TNF-α and IL-8 secretion levels in adenovirus-infected 293T cells cultured in 3D culture systems.
Conclusion: The nanoself-assembling peptide RADA16-I can be used as a 3D scaffold for adenovirus isolation, culture and research. The 3D culture system shows more realistic in vivo effects than two-dimensional (2D) culture.
Keywords: nanoself-assembling peptide, 3D culture, adenovirus, 293T cells
Introduction
Infectious diseases pose a great threat to public health. Among the pathogens that cause infectious diseases, viruses from either zoonotic sources or vector-borne sources are the main pathogens that result in the most pandemic threats.1 There are few effective drugs to treat viral diseases, which are highly contagious and widespread. In addition to causing an acute infection, some viruses can cause a persistent infection, and certain viruses are even closely associated with the occurrence of tumours and autoimmune diseases.2,3 Therefore, research on the physical and chemical properties, pathogenic mechanisms, immune mechanisms and antiviral drugs for the treatment of viruses has become one of the hotspots in medical and life science research. Viruses can grow only in living cells because of the lack of genetic information-encoding elements required for energy metabolism or protein synthesis (mitochondria and ribosomes), so it is more difficult to culture viruses than other microorganisms that can grow on inanimate media. Currently, virus culture methods include animal inoculation, chick embryo cultivation and cell culture, wherein cell culture is the most commonly used method for virus isolation, identification and research.4
Viruses are mostly cultured in vitro in two dimensions in glass or plastic dishes. Many cells gradually planarize, differentiate and lose their differentiated phenotypes after being separated from tissues and cultured in two dimensions; in fact, the two-dimensional (2D)-culture method cannot simulate the interaction between cells and the extracellular matrix (ECM) with a spatial structure.5 Over decades, researchers have gradually realized that various signals, such as adhesion between cells and the ECM, cytokines, and neurotransmitters in the extracellular microenvironment, actively participate in regulating cell growth, differentiation, proliferation and apoptosis.6 Therefore, cell culture technology has continuously improved with the progression from 2D to three-dimensional (3D) and static 3D to dynamic 3D cultures. Three-dimensional cell culture technology has been widely used in tissue engineering, regenerative medicine and in vitro studies of tumour cells.7–13 In recent years, some scholars have successfully developed 3D cell models for virus culture,14–20 and the observed infection efficiency of 3D cell culture is higher than that of 2D single-layer cell culture models.18,19 However, there have been few reports on the application of 3D cell culture techniques for virus research.
Currently, knowledge of viral replication, pathogenicity, and drug screening is mostly obtained through 2D cell cultures and animal experiments, whereas cells that grow in tissues in vivo live beyond all doubt in a 3D microenvironment, with which the occurrence and development of viral infections are closely associated in the body. Traditional 2D cell culture methods cannot reflect the influence of the microenvironment on the occurrence and development of viral infection in vivo. Thus, a 3D culture model that simulates the environment in vitro will facilitate the in-depth exploration of viral infection, replication, and pathogenesis and the relationship between the virus and the host. In our previous research work, some 3D cell culture models were successfully established using a nanoself-assembling peptide.20,21 Herein, a new project based on the 3D cell culture model is proposed. The aim of this project is to construct a 3D virus culture model using a nanoself-assembling peptide RADA16-I by combining virology technology and nanoself-assembling peptide cell culture technology to provide a novel theoretical and experimental basis for the study of viral diseases and the development of antiviral drugs and vaccines.
Materials and Methods
Materials
The 293T cell line was purchased from CCTCC (Wuhan Province, China). Adenovirus (Adenovirus-EGFP) was purchased from Shanghai Genechem Co., Ltd. (Shanghai, China). Dulbecco’s modified Eagle’s medium (DMEM) and foetal bovine serum were purchased from GIBCO (USA). MTS was purchased from Promega (USA). Calcein-AM (Ca-AM), propidium iodide (PI), and DAPI were purchased from Sigma (USA). The nanoself-assembling peptide RADA16-I (Ac-RADARADARADARADA-CONH2, purity 99%) was synthesized by Shanghai Biotech Bioscience & Technology Co., Ltd. (Shanghai, China). Enzyme-linked immunosorbent assay (ELISA) kits for tumour necrosis factor (TNF)-α and interleukin (IL)-8 were purchased from the Cloud Clone Corp (Wuhan Province, China).
Material Characterization
Scanning Electron Microscopy (SEM) Measurements
For SEM, the RADA16-I hydrogel was prepared to 0.5% weight/volume (wt/vol) and washed. The 0.5% RADA16-I hydrogel was fixed with 2.5% (v/v) glutaraldehyde for 2 hours at 37°C and dehydrated in 70%, 80%, 90%, and 100% ethanol. The critical point drying was used to dry the samples. Finally, the samples were mounted on SEM stubs using a double-coated carbon-conductive tape for SEM (Hitachi, S3400N, Japan) observation.
Transmission Electron Microscopy (TEM) Measurements
For TEM, the RADA16-I peptide solution was diluted to 0.05% weight/volume (wt/vol). Before observation, 0.05% RADA16-I peptide solution was dropped on the surface of a copper grid and negatively stained with 3% phosphotungstic acid for 3–5 minutes. After being dried under an incandescent lamp for a few minutes, the grid was observed directly by TEM (JEOL-JEM123, Japan).
3D Culture of 293T Cells
293T cells seeded in 25-cm2 culture flasks were grown at 37°C in a humidified atmosphere containing 5% CO2 in DMEM supplemented with 10% foetal calf serum. When the density reached 90%, the cells were detached using Trypsin-EDTA 0.05%. Subsequently, they were resuspended in 10% sucrose, and 75 µL of a cell suspension containing 5×104 cells was added to 75 µL of 1% RADA16-I, gently mixed, deposited in a 24-well plate with complete medium and incubated at 37°C with 5% CO2 for 30 minutes before the first medium change. Twenty-four hours later, the medium was changed again. Afterwards, the medium was changed every 2 d. For some experiments, 3D cultures were scaled to a 96-well plate using 25 µL of cell suspension containing 1.67×104 cells and 25 µL of 1% RADA16-I.
MTS Proliferation Assay of 3D-Cultured 293T Cells
3D-cultivated cells were incubated in 96-well plates and 20 µL of MTS reagent was added directly to the wells at 1, 3, 5, …, and 23 days after cultivation. Then, cell samples were incubated at 37°C for 2 hours in the dark and the absorbance was measured at 490 nm.
Ca-AM, DAPI and PI Staining of 3D-Cultured 293T Cells
Cells grown in 3D conditions in 24-well plates were observed by fluorescence microscopy after staining with Ca-AM, DAPI and PI at 0, 5, 10, 15, and 30 days after cultivation.
Adenovirus Infection of 3D-Precultured 293T Cells
For infection experiments, cells grown in RADA16-I were incubated for 24 hours in a 24-well plate, as previously described. Then, the cells were inoculated with adenovirus (4×106 copies viral nucleotides per well) for 24 hours at 37°C. Afterwards, the solution was removed, and cell-laden 3D cultures were washed three times with PBS and covered with complete medium. Observations were carried out every day after infection.
MTS Proliferation Assay of Adenovirus-Infected 3D-Cultured 293T Cells
Cells seeded in RADA16-I in a 96-well plate were infected with adenovirus (1×106 copies viral nucleotides per well). Afterwards, 20 µL of MTS reagent was added to the wells at 1, 3, 5, …, and 23 dpi and the absorbance was detected as previously mentioned.
Enhanced Green Fluorescent Protein (EGFP) Assay for Adenovirus Proliferation in the 3D Culture System
3D-cultured 293T cells in 24-well plates were infected with adenovirus as previously described. The EGFP reporter gene was used as a label to detect virus proliferation in 293T cells under direct fluorescence microscopy at every second day at 2–22 dpi (3 fields were counted per well with 6 duplicates).
TEM for Adenovirus in 3D and 2D Conditions
Cells cultured in 3D conditions in a 24-well plate were infected with adenovirus as previously described, and the cell cultures (293T cells and hydrogel) were harvested at 9 and 15 dpi. Afterwards, the samples were post-fixed using 1% osmic acid at 4°C for 3 hours, dehydrated through a graded series of alcohol, embedded with Spurr resin, and cut into ultra-thin sections (thickness 70 nm). Specimens were then stained with uranyl acetate and lead citrate, and images were obtained using a JEM1230 TEM (JEOL, Japan) at an acceleration voltage of 80 kV. For 2D samples, the 293T cells were harvested at 3 dpi and observed as previously described.
DNA Extraction and Quantitative qPCR of the 3D-Cultured Adenovirus
3D-cultured 293T cells in 96-well plates were infected with adenovirus (1×106 copies viral nucleotides per well), and the cell cultures (supernatant and hydrogel) were harvested at 1, 3, 5, and 23 dpi and stored at −20°C. Subsequently, viral nucleic acids were extracted with a DNA/RNA extraction kit (Tiangen DP315) according to the manufacturer’s protocol. qPCR systems were prepared using a BIO-RAD qPCR kit in accordance with the manufacturer’s protocol. The primers used for detection were as follows:
EGFP, 5ʹ-TTCAAGATCCGCCACAACA-3ʹ and 5ʹ-CGCTTCTCGTTGGGGTC-3ʹ. The qPCR conditions were as follows: pre-denaturation at 95°C for 2 minutes, 39 cycles of denaturation at 95°C for 5 s, annealing at 60°C for 30 s and a melting curve temperature between 65°C and 95°C. Negative (water) and positive (adenovirus standard samples) controls were established for each amplification. After the reaction was complete, the system automatically generated a standard curve and calculated the copy number of the complete viral genome in the corresponding samples.
Infectivity Detection of Adenovirus Produced in the 3D Cell Cultures
The cell supernatants of 3D cell cultures were collected at 6 dpi (1×107 copies based on the qPCR amplification quantification) and then used to infect 293T cells grown in 2D culture. The supernatants and cells of 2D culture were collected at 2, 4 and 6 dpi. Thereafter, virus DNA was extracted and amplified by qPCR.
Cytokine Detection in Adenovirus-Infected 3D-Cultured 293T Cells
3D-cultured 293T cells seeded in 96-well plates were infected with adenovirus as previously described. The supernatant was collected at 6, 12 hpi and 1–10 dpi and stored at −80°C. The control group was set up without infection. TNF-α and IL-8 secretion levels were detected by ELISA. Cytokine levels in samples were calculated according to a standard curve.
Statistics
All statistical analyses were performed using GraphPad Prism 5.0 unless otherwise noted. Each data point represents the mean of three independent experiments at least. Data are represented as the mean value ± SD of a triplicate experiment.
Results
Characterization of the RADA16-I Peptide Scaffold
The nanoself-assembling peptide RADA16-I is a kind of biomaterial formed by the self-assembly interaction of molecules. It has a unique amino acid sequence (Ac-RADARADARADARADA-CONH2) and can self-assemble into a nanofiber hydrogel with a water content of more than 99% in physiological saline solution;22,23 its fibre mesh aperture is approximately 5–200 nm.24 SEM (Figure 1A) and TEM (Figure 1B) showed that RADA16-I self-assembled into highly crosslinked nanofibers with uniform width and height; the fibres were uniformly long and entangled with each other to form a 3D network. These data suggested that the RADA16-I hydrogel was similar to the ECM structure that makes up the microenvironment in vivo.
 |
Figure 1 RADA16-I was evaluated by SEM and TEM and applied to 3D cell cultures. (A) SEM image of RADA16-I peptide scaffold nanofibers. (B) TEM image of RADA16-I peptide scaffold nanofibers. |
Characterization of 3D Cultures of 293T Cell Lines
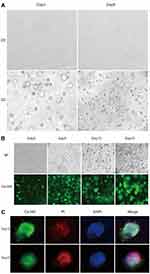
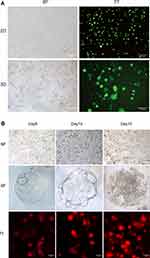
The sequence RAD (arginine-alanine-aspartate) in the RADA16-I and the sequence RGD (arginine-glycine-aspartate) in the integrin ligand-binding site on the cell membrane are similar,25 which can provide a 3D culture environment similar to natural ECM for cell adhesion and growth. In this study, cell proliferation of 293T cells cultured in the RADA16-I hydrogel that began as single cells and gradually formed dense multicellular spheroids and grew well was visible at 3 d, peaked at 11–15 d, and then gradually declined (Figures 2 and 3A).To further check the cell viability in the RADA16-I hydrogel, we stained cells with Ca-AM (green), which is hydrolysed to a green fluorescent product in living cells, and PI (red), which is able to pass through only the compromised membranes of dead cells (Figure 3B). Large green fluorescent areas were observed, and the green fluorescence was still visible in the cell mass after 30 days of culture (Figure 3C), suggesting that cells cultured in the RADA16-I hydrogel maintained good viability. Compared with 3D culture, there is a main limitation associated with a traditional 2D culture that cells were grown on flat dishes optimized for cell attachment and growth (Figure 3A). Even if the serum concentration is reduced to 2%, the culture time can only be appropriately prolonged, but the cells no longer grow. Therefore, 3D culture that can form multicellular spheroids and maintain cell proliferation for a long time is more conducive to study the long-term virus culture in vitro and explore the relationship between viruses and hosts.
 |
Figure 2 Proliferation of 293T cells in 3D culture by MTS. |
Viral Proliferation in 3D Culture
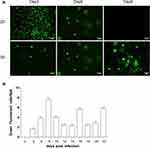
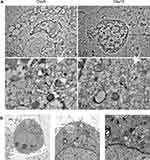
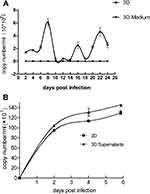
To analyzed whether 3D-cultured 293T cells were susceptible to adenovirus infection, 293T cells in 2D culture and 3D culture were infected with adenovirus-EGFP. As shown in Figures 4A and 5A, the green fluorescence of cells in 2D culture was visible at 24 hpi, and the fluorescence intensity was strongest at 48–72 hpi. In addition, apparent cytopathic effect (CPE) could be observed, and the cells gradually became necrotic and detached at 4–6 dpi. By contrast, green fluorescent cell spheroids in 3D culture were observed at 60–72 hpi, but the fluorescence intensity was very weak. Thereafter, the number of fluorescent cells and fluorescence intensity increased gradually from the edge of the gel, peaking on 8 dpi, then decreasing gradually and peaking on 16 dpi and 22 dpi, respectively (Figures 4A and 5). As shown in Figure 4B vesicles and depressions of multicellular spheroids were visible at 5 dpi in 3D culture, and cytolysis was observed at 9–12 dpi, at which point individual cells were visible. With the extension of the incubation time, the number of dead cells in the RADA16-I hydrogel gradually increased, and fragmentation and cytolysis of cell spheroids were visibly observed at 15 dpi (Figure 4B). To further analyse the difference between the cytopathic effects caused by adenovirus proliferation in the 3D culture and those in the traditional 2D culture, infected-293T cells from these two culture systems were examined by a TEM assay. In the 3D culture, we observed significant cytopathic effects such as swelling of the mitochondria and expansion of the endoplasmic reticulum at the 9th and 15th dpi. Furthermore, a small number of virions with diameters of 60–90 nm were significantly observed in the cytoplasm of 3D-cultured 293T cells (Figure 6A). In addition to swelling of the mitochondria and expansion of the endoplasmic reticulum, the formation of autophagosomes and a large number of autophagic lysosomes were observed at 3 dpi in the 2D culture system. Furthermore, we found that most of the virions with diameters of 60–90 nm were located in autophagic lysosomes, and the number of virions was more than that in 3D culture (Figure 6B). The viability of 3D-cultured 293T cells was affected by adenovirus infection. Therefore, an MTS assay was used to evaluate the viability of 293T cells after infection. As shown in Figure 2, the cells’ viability appeared to be decreased at 3 dpi, reaching its lowest point at 7–9 dpi, then increasing gradually to the 13th dpi and decreasing gradually, reaching its lowest point at 15 and 22 dpi. Furthermore, we next monitored viral DNA at different time points post-infection by qPCR. The results showed that multiple peaks appeared in adenovirus DNA levels, and their proliferation characteristics were consistent with the results of the EGFP assay (Figure 7A). We also included controls without cells in 3D conditions. Also shown in Figure 7A, adenovirus DNA was detected only when 293T cells were present, demonstrating that the signal obtained was not due to the amplification of material-bound input adenovirus DNA. These results indicated that adenovirus could replicate and proliferate continuously in the 3D culture system with the nanoself-assembling peptide RADA16-I as scaffolds and could cause cytopathic effects that were not exactly the same as those cultured in the 2D system.
Characterization of Adenovirus Produced by 3D-Cultured Cells
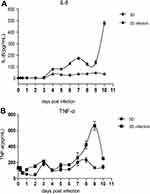
We then investigated the infectivity of virus particles released from adenovirus-infected spheroids.We found that supernatants derived from adenovirus-infected 293T multicellular spheroids were infective, and their infectivity was not significantly different from that of 2D culture (Figure 7B). When cells are infected by viruses, to resist viral invasion, the interaction with viruses can stimulate cells to secrete cytokines such as TNF-α, IP-10, IL-6, and IL-8 to activate an immune response.26,27 To further confirm whether 3D cultures were infected by adenovirus and whether virus proliferation has effects on host cells, TNF-α and IL-8 in the media of the 3D cultures were examined by ELISA at the indicated time points after infection with adenovirus. As shown in Figure 8, the cytokine increases appeared as multiple peaks in the 3D cultures after infection with adenovirus, but the expression levels of IL-8 and TNF-α as well as the secretion trends were different in the cell cultures. Compared to the uninfected cell cultures, expression of IL-8 increased significantly at 3 dpi, reaching three peaks at 4 dpi, 7 dpi and 10 dpi (Figure 8A). The expression of TNF-α increased less in the early stage of infection than in the uninfected cell cultures, the first peak appeared at 3 dpi, then the level of TNF-α increased significantly at 7 dpi and reached its another peak at 9 dpi (Figure 8B). The above data suggested that the virions produced by the 3D cultures were still infectious, and virus proliferation had effects on 3D-cultured 293T cells. In aword, this 3D culture model can be applied to isolation and cultivation of adenovirus in vitro.
Discussion
The nanoself-assembling peptide RADA16-I as a good 3D scaffold has been successfully used in 3D culture for different cells20,21 because it can be chemically synthesized and has several advantages over other traditional 3D culture materials (such as Collagen and Matrigel) including that it is a single-ingredient with high purity, is non-immunogenic, and is pathogen-free. As an ionic hydrogel, RADA16-I can be self-assembled to form a 3D-nanofibre mesh structure under ionic conditions with porosity that is similar to that of natural ECM28 with high moisture, which is beneficial for cell adhesion, growth, diffusion and absorption of various nutrients and signal molecules in the microenvironment.29,30 Moreover, it is easier to operate than Collagen or Matrigel. Our study found that 293T cells in RADA16-I hydrogel can maintain cell proliferation for a long time, which provides a possibility to study long-term virus culture in vitro and explore the relationship between viruses and hosts.
The diameter of the fibre mesh formed by the nanoself-assembling peptide RADA16-I is approximately 5–200 nm.24 Therefore, adenovirus with a diameter of approximately 60–90 nm could infect 293T cells through the scaffold pores of RADA16-I; this was confirmed in our study that the virus particles in 293T cells cultured in RADA16-I were observed under TEM at 9 and 15 dpi. In vivo, the virus infects susceptible cells and releases a large number of progeny virus that will infect the surrounding cells again, causing the CPEs and corresponding clinical symptoms in the body. However, in vitro, the cells cultured in the 2D system were a monolayer in-plane. After inoculation of virus, all the cells in plane growth could contact the virus in a short time, CPEs appeared soon, and the cells gradually became round and necrotic and fell off. Therefore, 2D culture is not suitable for the study of long-term infection. In the 3D cell culture system, by contrast, the virus adsorbs on the surface of the multicellular spheroids and first infects the outermost cells. After replication and proliferation in these cells, a large number of progeny viruses are released to infect the central cells or adjacent multicellular spheroids, while the uninfected cells continue to divide and grow. Therefore, our study found that the period of virus culture in RADA16-I was much longer than that in the 2D culture, and there were multiple proliferation peaks, which was similar to the reports of Berto et al.18,31 This characteristic of proliferation may be related to the interaction between the virus and the host (ie, infection, release, reinfection), which may also be the cause of multiple secretion peaks of the cytokines TNF-α and IL-8 after virus infection in the 3D culture, which is similar to Chen et al’s18 findings in the study of a 3D culture of HBoV and HRV-C. Taken together, compared with a 2D cell culture, a 3D cell culture can simulate the process of virus infection in vivo.
In this study, RADA16-I was used as a scaffold to construct a 3D cell culture model of adenovirus in vitro, but there are still some limitations. First, adenovirus can proliferate in the RADA16-I 3D culture system for a long time, but there is no further study on the infection mechanism because the virus is an empty vector. Moreover, whether other viruses can also be cultured in the 3D culture system for long-term culture remains to be verified. Second, with the continuous development of 3D cell culture technology, researchers clearly recognized that the mechanical properties (such as material hardness and pore structure) and biochemical properties (such as biocompatibility and degradability) of scaffold materials may have an important impact on cell viability, adhesion, differentiation and migration.32 Although the nanoself-assembling peptide RADA16-I is a type of good 3D culture scaffold material, it still has problems such as insufficient mechanical strength and low shear resistance.33 It is uncertain whether the nanoself-assembling peptide RADA16-I is the best 3D scaffold material at present, but it is clear that any in vitro 3D culture system cannot completely and accurately simulate the human microenvironment because the organism is a complex environment in which many cells, organs and tissues coexist. Moreover, the process of the virus infecting the organism, replicating and proliferating in cells, and causing a series of pathological changes is a complex pathophysiological process in which many cells and tissues participate. Therefore, if RADA16-I can be used to construct a 3D co-culture model of various cells, it will be important to study the pathogenic mechanism of viruses and explore the interactions between viruses and hosts.
Conclusion
To date, 3D cell culture is considered to be a better cell model in vitro. However, 3D culture scaffolds have different materials, which may have some problems such as being immunogens or pathogens, having too large pores, and being difficult to obtain and shape. Most importantly, it is difficult to standardize and obtain accurate results. Our research showed that the nanoself-assembling peptide RADA16-I can be used as scaffolds for 3D cell culture of adenovirus. Combined with the characteristics of adenovirus vectors such as high transgenic efficiency, high titres regardless of the state of cell division, and easy evaluation for biological viability, we hope that this 3D culture method of adenovirus can be used not only in the study of viral pathogenesis but also in the study of antiviral drugs, vaccines, viral gene therapy and other fields in the future.
Abbreviations
3D, three-dimensional; 2D, two-dimensional; ECM, extracellular matrix; TEM, transmission electron microscope; SEM, scanning electron microscope; DMEM, dulbecco’s modified eagle medium; Ca-Am, calcein-AM; dpi, days post infection; EGFP, enhanced green fluorescent protein; qPCR, real-time fluorescent quantitative PCR; ELISA, enzyme-linked immunosorbent assay; hpi, hours post infection; CPE, cytopathic effect.
Disclosure
Di-Shu Ao and Lu-Yao Gao should be regarded as co-first authors. The authors declare no conflict of interest.
References
1. Graham BS, Sullivan NJ. Emerging viral diseases from a vaccinology perspective: preparing for the next pandemic. Nat Immunol. 2018;19(1):20–28. doi:10.1038/s41590-017-0007-9
2. Lunn RM, Jahnke GD, Rabkin CS. Tumour virus epidemiology. Philos Trans R Soc Lond B Biol Sci. 2017;372(1732):20160266. doi:10.1098/rstb.2016.0266
3. Getts DR, Chastain EM, Terry RL, Miller SD. Virus infection, antiviral immunity, and autoimmunity. Immunol Rev. 2013;255(1):197–209. doi:10.1111/imr.12091
4. Hematian A, Sadeghifard N, Mohebi R, et al. Traditional and modern cell culture in virus diagnosis. Osong Public Health Res Perspect. 2016;7(2):77–82. doi:10.1016/j.phrp.2015.11.011
5. Antoni D, Burckel H, Josset E, Noel G. Three-dimensional cell culture: a breakthrough in vivo. Int J Mol Sci. 2015;16(12):5517–5527. doi:10.3390/ijms16035517
6. Alhaque S, Themis M, Rashidi H. Three-dimensional cell culture: from evolution to revolution. Philos Trans R Soc Lond B Biol Sci. 2018;373(1750):20170216. doi:10.1098/rstb.2017.0216
7. Park J, Lee SJ, Chung S, et al. Cell-laden 3D bioprinting hydrogel matrix depending on different compositions for soft tissue engineering: characterization and evaluation. Mater Sci Eng C. 2017;71:678–684. doi:10.1016/j.msec.2016.10.069
8. Sato T, Clevers H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science. 2013;340(6137):1190–1194. doi:10.1126/science.1234852
9. Seow WY, Kandasamy K, Purnamawati K, Sun W, Hauser CAE. Thin peptide hydrogel membranes suitable as scaffolds for engineering layered biostructures. Acta Biomater. 2019;88:293–300. doi:10.1016/j.actbio.2019.02.001
10. Tong C, Liu T, Saez Talens V, et al. Squaramide-based supramolecular materials for three-dimensional cell culture of human induced pluripotent stem cells and their derivatives. Biomacromolecules. 2018;19(4):1091–1099. doi:10.1021/acs.biomac.7b01614
11. Ortega-Prieto AM, Skelton JK, Wai SN, et al. 3D microfluidic liver cultures as a physiological preclinical tool for hepatitis B virus infection. Nat Commun. 2018;9(1):682. doi:10.1038/s41467-018-02969-8
12. Verjans ET, Doijen J, Luyten W, Landuyt B, Schoofs L. Three-dimensional cell culture models for anticancer drug screening: worth the effort? J Cell Physiol. 2018;233(4):2993–3003. doi:10.1002/jcp.26052
13. Ishiguro T, Ohata H, Sato A, Yamawaki K, Enomoto T, Okamoto K. Tumor-derived spheroids: relevance to cancer stem cells and clinical applications. Cancer Sci. 2017;108(3):283–289. doi:10.1111/cas.13155
14. Molina-Jimenez F, Benedicto I, Dao Thi VL, et al. Matrigel-embedded 3D culture of Huh-7 cells as a hepatocyte-like polarized system to study hepatitis C virus cycle. Virology. 2012;425(1):31–39. doi:10.1016/j.virol.2011.12.021
15. Gardner JK, Herbst-Kralovetz MM. Three-dimensional rotating wall vessel-derived cell culture models for studying virus-host interactions. Viruses. 2016;8(11):304–320. doi:10.3390/v8110304
16. Bramley JC, Drummond CG, Lennemann NJ, Good CA, Kim KS, Coyne CB. A three-dimensional cell culture system to model RNA virus infections at the blood-brain barrier. mSphere. 2017;2(3):e00206–00217. doi:10.1128/mSphere.00206-17
17. Koban R, Neumann M, Daugs A, et al. A novel three-dimensional cell culture method enhances antiviral drug screening in primary human cells. Antiviral Res. 2018;150:20–29. doi:10.1016/j.antiviral.2017.12.005
18. Chen YX, Xie GC, Pan D, et al. Three-dimensional culture of human airway epithelium in matrigel for evaluation of human rhinovirus c and bocavirus infections. Biomed Environ Sci. 2018;31(2):136–145. doi:10.3967/bes2018.016
19. Rosellini A, Freer G, Quaranta P, et al. Enhanced in vitro virus expression using 3-dimensional cell culture spheroids for infection. J Virol Methods. 2019;265:99–104. doi:10.1016/j.jviromet.2018.12.017
20. Yang Z, Zhuang H, Song H, Liu J, Zhao X, Lin W. A miniature cell pattern formation of ovarian cancer cell lines on self-assembling peptide nanofiber-coated coverslip and in vitro chemosensitivity assay. J Nanosci Nanotechnol. 2018;18(4):2370–2378. doi:10.1166/jnn.2018.14384
21. Liu J, Song H, Zhang L, Xu H, Zhao X. Self-assembly-peptide hydrogels as tissue-engineering scaffolds for three-dimensional culture of chondrocytes in vitro. Macromol Biosci. 2010;10(10):1164–1170. doi:10.1002/mabi.200900450
22. Zhao X, Zhang S. Fabrication of molecular materials using peptide construction motifs. Trends Biotechnol. 2004;22(9):470–476. doi:10.1016/j.tibtech.2004.07.011
23. Cormier AR, Pang X, Zimmerman MI, Zhou HX, Paravastu AK. Molecular structure of RADA16-I designer self-assembling peptide nanofibers. ACS Nano. 2013;7(9):7562–7572. doi:10.1021/nn401562f
24. Koutsopoulos S, Unsworth LD, Nagai Y, Zhang S. Controlled release of functional proteins through designer self-assembling peptide nanofiber hydrogel scaffold. Proc Natl Acad Sci U S A. 2009;106(12):4623–4628. doi:10.1073/pnas.0807506106
25. Masuko T, Iwasaki N, Yamane S, et al. Chitosan–RGDSGGC conjugate as a scaffold material for musculoskeletal tissue engineering. Biomaterials. 2005;26(26):5339–5347. doi:10.1007/s00281-017-0640-2
26. Teijaro JR. Cytokine storms in infectious diseases. Semin Immunopathol. 2017;39(5):501–503. doi:10.1007/s00281-017-0640-2
27. Walsh KB, Teijaro JR, Rosen H, Oldstone MB. Quelling the storm: utilization of sphingosine-1-phosphate receptor signaling to ameliorate influenza virus-induced cytokine storm. Immunol Res. 2011;51(1):15–25. doi:10.1007/s12026-011-8240-z
28. Kisiday J, Jin M, Kurz B, et al. Self-assembling peptide hydrogel fosters chondrocyte extracellular matrix production and cell division: implications for cartilage tissue repair. Proc Natl Acad Sci U S A. 2002;99(15):9996–10001. doi:10.1073/pnas.142309999
29. Yokoi H, Kinoshita T, Zhang S. Dynamic reassembly of peptide RADA16 nanofiber scaffold. Proc Natl Acad Sci U S A. 2005;102(24):8414–8419. doi:10.1073/pnas.0407843102
30. Wang X, Horii A, Zhang S. Designer functionalized self-assembling peptide nanofiber scaffolds for growth, migration, and tubulogenesis of human umbilical vein endothelial cells. Soft Matter. 2008;4:2388–2395. doi:10.1039/b807155a
31. Berto A, van der Poel WH, Hakze-van der Honing R, et al. Replication of hepatitis E virus in three-dimensional cell culture. J Virol Methods. 2013;187(2):327–332. doi:10.1016/j.jviromet.2012.10.017
32. Zhang W, Zhuang A, Gu P, Zhou H, Fan X. A review of the three-dimensional cell culture technique: approaches, advantages and applications. Curr Stem Cell Res Ther. 2016;11(4):370–380. doi:10.2174/1574888x11666160217154007
33. Koutsopoulos S. Self-assembling peptide nanofiber hydrogels in tissue engineering and regenerative medicine: progress, design guidelines, and applications. J Biomed Mater Res A. 2016;104(4):1002–1016. doi:10.1002/jbm.a.35638
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.