Back to Journals » OncoTargets and Therapy » Volume 7
Stage-dependent prognostic impact of molecular signatures in clear cell renal cell carcinoma
Authors Weber T, Meinhardt M, Zastrow S, Wienke A, Erdmann K, Hofmann J, Fuessel S, Wirth MP
Received 2 January 2014
Accepted for publication 3 February 2014
Published 2 May 2014 Volume 2014:7 Pages 645—654
DOI https://doi.org/10.2147/OTT.S59983
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 5
Thomas Weber,1,2 Matthias Meinhardt,3 Stefan Zastrow,1 Andreas Wienke,4 Kati Erdmann,1 Jörg Hofmann,1 Susanne Fuessel,1 Manfred P Wirth1
1Department of Urology, Technische Universität Dresden, Dresden, Germany; 2Department of Oncology and Hematology, Martin-Luther-University Halle-Wittenberg, Halle (Saale), Germany; 3Institute of Pathology, Technische Universität Dresden, Dresden, Germany; 4Institute of Medical Epidemiology, Biostatistics, and Informatics, Martin-Luther-University Halle-Wittenberg, Halle (Saale), Germany
Purpose: To enhance prognostic information of protein biomarkers for clear cell renal cell carcinomas (ccRCCs), we analyzed them within prognostic groups of ccRCC harboring different tumor characteristics of this clinically and molecularly heterogeneous tumor entity.
Methods: Tissue microarrays from 145 patients with primary ccRCC were immunohistochemically analyzed for VHL (von Hippel-Lindau tumor suppressor), Ki67 (marker of proliferation 1), p53 (tumor protein p53), p21 (cyclin-dependent kinase inhibitor 1A), survivin (baculoviral IAP repeat containing 5), and UEA-1 (ulex europaeus agglutinin I) to assess microvessel-density.
Results: When analyzing all patients, nuclear staining of Ki67 (hazard ratio [HR] 1.08, 95% confidence interval [CI] 1.04–1.12) and nuclear survivin (nS; HR 1.04, 95% CI 1.01–1.08) were significantly associated with disease-specific survival (DSS). In the cohort of patients with advanced localized or metastasized ccRCC, high staining of Ki67, p53 and nS predicted shorter DSS (Ki67: HR 1.07, 95% CI 1.02–1.11; p53: HR 1.05, 95% CI 1.01–1.09; nS: HR 1.08, 95% CI 1.02–1.14). In organ-confined ccRCC, patients with high p21-staining had a longer DSS (HR 0.96, 95% CI 0.92–0.99). In a multivariate model with stepwise backward elimination, tumor size and p21-staining showed a significant association with DSS in patients with "organ-confined" ccRCCs. The p21-staining increased the concordance index of tumor size from 0.75 to 0.78. In patients with "organ-confined" ccRCC, no disease-related deaths occurred in the group with p21-expression below the threshold of 32.5% p21-positive cells (log rank test: P=0.002).
Conclusion: The prognostic information of the studied protein biomarkers depended on anatomic tumor stages, which displayed different acquired biological tumor characteristics. Analysis of prognostic factors within different clinical ccRCC groups could help to enhance their prognostic power. The p21-staining was an independent prognostic factor and increased prognostic accuracy in a predictive model in "organ-confined" ccRCC.
Keywords: survival, prognostic groups, localized, p21, advanced
Introduction
Clear cell renal cell carcinomas (ccRCCs) have a very heterogeneous course of disease. To date, clinical outcome is not reliably predictable with established clinical and pathological factors. Many attempts were made to enhance the prognostic power of established markers, such as nomograms based on clinical and pathological variables and the evaluation of molecular markers.1 Nonetheless, TNM stage and Fuhrman grade still yield the highest prognostic power.2,3
Based on the TNM staging system, ccRCC can be classified into three major prognostic groups: “organ-confined” (pathological tumor stage, pT1-pT2; lymph node stage, N0; metastases stage, M0), “advanced localized” (pT3-pT4 N0 M0), and “metastasized” (any pT, N1, and/or M1) ccRCC2 with 5-year disease-specific survival (DSS) rates from 82% to 95%,4 24% to 72%,5 and 5% to 30%,6 respectively. These groups display different acquired characteristics of cancers such as tissue invasion and metastasis. Organ-confined ccRCC is nearly always present without invasion of the surrounding renal tissue.7 This is reflected in the low rate of distant metastasis between 4.2% and 7.1% in ccRCC ≤4 cm at initial presentation.8
Protein expression analysis of gene products with important functions in tumorigenesis and with altered expression in cancer initiation and/or progression, respectively, can help to identify prognostic groups more accurately.3,9
Weiss et al showed that the predictive capability of the molecular biomarker p21 (cyclin-dependent kinase inhibitor 1A) is dependent on ccRCC biology, with high nuclear p21-expression being associated with longer DSS in organ-confined and advanced localized disease, but shorter survival in metastasized disease.10 High Ki67 (marker of proliferation 1)-expression, a measure of high proliferation activity, is a negative predictor of DSS in ccRCC patients.3 High p53-staining is associated with TP53-mutations and with shorter survival in ccRCC.1 p21 can arrest the cell cycle as an effector of p53, but also has p53-independent growth promotional, pro-apoptotic, and anti-apoptotic effects in ccRCC.10 The inhibitor of apoptosis, survivin, is highly upregulated in ccRCC and associated with poor outcome.11,12 VHL (von Hippel-Lindau tumor suppressor)-alterations are an initial step in the development of most ccRCC leading to angiogenesis, cell migration, and proliferation.1 In addition, VHL is a regulator of p53 stabilization.13 Angiogenesis was proven to be an important step in development and progression of ccRCC with microvessel density (MVD) as an easily assessable marker of tumor vascularization.14
These molecular markers reflect the complex interplay between apoptosis, cell cycle regulation, growth signaling, angiogenesis, and immortalization in ccRCC. In this study, we analyzed whether the immunohistochemical staining profiles of Ki67, p53, p21, nuclear survivin (nS), cytoplasmic survivin (cS), VHL, and MVD have distinct associations with clinical outcome in the subgroups of organ-confined or advanced localized and metastasized ccRCC. The primary aim of the present study is to identify if these molecular markers add prognostic information to established clinical and pathological markers in biologically distinct subgroups of ccRCC.
Materials and methods
Tissue specimens
Tumor specimens from 145 patients suffering from primary ccRCC with available paraffin blocks and follow-up data were analyzed retrospectively. All patients underwent total or partial nephrectomy at the Department of Urology Dresden between 1993 and 2000. Follow-up of patients was retrieved from in-house medical records and/or from the attending urologist or general practitioner. For all patients, histological classification and tumor grade according to Fuhrman were reevaluated on the histopathological slides in a blinded fashion (by MM and TW). Tumor stage was reclassified according to the Union for International Cancer Control classification 2009.15 Tissue collection and analysis was acknowledged by the internal review board of the Technical University of Dresden (EK59032007 and EK195092004). Patients gave written informed consent.
Tissue microarrays
Tissue microarrays (TMAs) were constructed including two representative tumor cores (one core from the center and one from the margin of the tumor) with a diameter of 0.6 mm from each patient. Selection of representative tissue areas was performed by a trained pathologist (MM) on large sections for each ccRCC. Normal kidney specimens were used as controls. Specimens of 12 metastases were incorporated in the TMA. Sections of 4 μm thickness were mounted on adhesive glass slides. Quality control with regard to type of tissue and grade was performed on hematoxylin and eosin stained TMA sections.
Immunohistochemistry
As previously described,16 all sections were deparaffinized and antigen retrieval for p21, survivin, and VHL was achieved by heating the sections in 0.01 M citrate buffer pH 6.0 at 120°C for 10 minutes. After blocking endogenous peroxidase, and after incubation with the blocking serum (universal ABC-AP Kit; Vector Laboratories, Inc., Burlingame, CA, USA), sections were incubated overnight at 4°C with the following primary antibodies: mouse monoclonal antihuman antibodies against p21 (clone 4D10; 1:80; Nova Castra, Newcastle upon Tyne, UK) and against VHL (clone immunoglobulin (Ig)32; 1:50; BD Biosciences, San Jose, CA, USA) as well as rabbit polyclonal antihuman survivin antibody (clone AF886; 1:200; R & D Systems, Inc., Abingdon, UK). Detection was achieved by using an anti-mouse/anti-goat ABC-Kit and visualization was accomplished with new fuchsine. For Ki67, p53, and UEA-1 (Ulex europaeus agglutinine-1), antigen retrieval was performed at 95°C for 20 minutes using 0.01 M citrate buffer pH 6.0. Mouse monoclonal antihuman Ki67 (clone MIB-1; 1:300; Dako, Glostrup, Denmark), p53 (clone DO-1; 1:100; Oncogene, Seattle, WA, USA), and UEA-1 (clone B-1065; 1:10; Vector Laboratories, Inc.) antibodies were used. After incubation with ABC-reagent (Vectastain Elite ABC Kit anti-mouse; Vector Laboratories, Inc.), DAB (3,3’-Diaminobenzidin) was used for visualization and Meyer’s Hemalaun for counterstaining. For every TMA-tissue core, the number of positively stained cells per 100 cells was evaluated manually by light microscopy at 400× magnification blinded to the clinical data (performed by TW). The mean of two tumor-cores per case was calculated. Nuclear staining was evaluated for p21, p53, Ki67, and survivin. Cytoplasmic staining was evaluated for VHL and also for survivin. MVD was analyzed by the method of Weidner et al17 using the expression of UEA-1 in endothelial cells. Random samples of all stainings were reevaluated by a second person (MM); no interobserver difference in marker expression of more than 10% was noted. No marked heterogeneity of any staining was observed in parallel analyzed large sections in a subset of patients.
Statistical analysis
All protein markers were analyzed as continuous variables. Associations between protein markers and categorical clinicopathological variables were analyzed by Mann–Whitney U test. DSS was measured as time from nephrectomy to death of disease. Uni- and multivariate survival analyses were performed with Cox proportional hazards regression models. Assumption of proportional hazards was tested for every variable by Schoenfeld’s partial residuals. Multivariate models were performed with stepwise Wald backward exclusion using 0.1, 0.15, and 0.2 as thresholds for exclusion. Hazard ratios (HRs) of the retaining variables were internally validated with bootstrapping analysis with 1,000 samples, and predictive accuracies of the final models were quantified by using the concordance index. For easier assessment of marker expression in clinical routine, p21 expression was classified into two categories: >32.5% (normal) and ≤32.5% (altered) cells with positive p21-staining as used by Weiss et al on renal cell carcinoma TMA sections.10 Survival functions of categorized p21 staining were visualized with Kaplan–Meier plots, and differences between groups were assessed with log rank tests. Generally, P-values <0.05 were considered significant. Due to the explorative character of the study, we did not correct alpha-error due to multiple testing. Statistical analysis was accomplished with PASW 18.0 (IBM Corporation, Armonk, NY, USA) and R 2.15.1 (http://www.r-project.org).
Results
Twenty-eight of the 145 patients (19.3%) died of disease and 27 (18.6%) died of other causes. Mean and median follow-up time of the surviving patients was 9.2 (standard error 4.1) and 9.7 years (interquartile range [IQR] 8.8–10.5), respectively. Patient characteristics are given in Table 1.
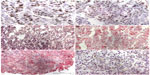
Immunohistochemistry was evaluable in all cases for Ki67, in 144 cases for p21 and UEA-1, in 143 cases for p53 and VHL, and in 141 cases for survivin. Representative stainings are shown in Figure 1. The median percentage of positive-stained ccRCC cells for Ki67, p53, p21, cS, nS, and VHL were 5.5% (IQR 3–8.7), 15.5% (IQR 7–24.5), 26% (IQR 17–37.5), 34.5% (IQR 17–50.5), 6.5% (IQR 2.8–12.8), and 18% (IQR 4.5–28.5), respectively. Median vessel count according to Weidner et al17 was 19.5 (IQR 7.8–33.9). High Fuhrman grade (grade 3–4) was significantly associated with higher median Ki67-staining, lymph node involvement with lower cS-staining, and MVD (Table S1). No other significant associations to clinicopathological parameters were observed.
In univariate Cox proportional hazards model analyses of the whole cohort, high pT stage, TNM stage, Fuhrman grade, larger tumor size, as well as positive N stage and M stage were predictors for a shorter DSS (Table 2).
For all analyzed protein marker variables, assumption of linearity could be made and Cox proportional hazard models were constructed. Higher Ki67- and nS-staining were associated with an 8% and 4% increased risk of death caused by ccRCC for every 1% increase in stained cells, respectively (Ki67: HR 1.08, 95% confidence interval [CI] 1.04–1.12, P<0.001; nS: HR 1.04, 95% CI 1.01–1.08, P=0.016). Patients with higher p53-staining or lower MVD tended to have a higher risk of death due to ccRCC (p53: HR 1.03, 95% CI 0.96–1.06, P=0.096; MVD: HR 0.98, 95% CI 0.95–1, P=0.071). The other biomarkers were not predictive for DSS (Table 2).
When analyzing the 36 patients with advanced localized or metastasized ccRCC (pT3-4 N0 M0 or any pT, N1, and/or M1, respectively) together, the presence of lymph node or distant metastasis were the strongest predictors for shorter DSS. Higher Fuhrman grade tended to be associated with DSS (Table 2). In these patients, Ki67-, p53-, and nS-stainings were significantly predictive for DSS (Ki67: HR 1.07, 95% CI 1.02–1.11, P=0.002; p53: HR 1.05, 95% CI 1.01–1.09, P=0.025; nS: HR 1.08, 95% CI 1.02–1.14, P=0.006; Table 2). These associations of Ki67-, p53-, and nS-stainings with DSS were also noticed in the 24 patients with advanced localized disease (Ki67: HR 1.08, 95% CI 1–1.16, P=0.047; p53: HR 1.09, 95% CI 1.02–1.16, P=0.009; nS: HR 1.18, 95% CI 1.06–1.32, P=0.004; Table 2). We did not perform Cox proportional hazards model analysis for the subgroup of the 12 patients with metastasized ccRCC due to the small number.
For the 109 patients with organ-confined ccRCC (pT1-2 N0 M0), larger tumor size as well as pT2 were associated with shorter DSS in univariate survival analyses (Table 2). Patients with higher p21-staining had a decreased risk of death due to ccRCC (HR 0.96, 95% CI 0.92–0.99, P=0.031). Hence, every 1% decrease of p21-staining increases risk of death due to ccRCC by 4%. No significant association to DSS was seen for the other biomarkers (Table 2).
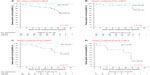
After classification of the p21-staining in patients with normal (>32.5%) and altered (≤32.5%) staining according to Weiss et al,10 68 patients with organ-confined ccRCC (63%) had an altered p21-staining. These patients had a significantly increased risk of death due to ccRCC (log rank test: P=0.002; Figure 2). This effect could be observed in patients with pT1a, pT1b, and pT2 tumors (log rank test for trend: P=0.009; Figure 2). Cox regression analysis could not be performed as no ccRCC-specific deaths occurred in the subgroup of patients with altered p21-staining.
These results indicate a differential prognostic impact of immunohistochemical staining signatures in organ-confined, advanced localized, and metastasized ccRCC. Therefore, we performed multivariate analysis separately for organ-confined ccRCC. The multivariate model comprised Fuhrman grade (grade 1 versus grade 2), tumor size, pT stage (pT1 versus pT2), and continuous p21-staining. After stepwise backward deletion with different deletion thresholds, continuous p21-staining and tumor size remained in the model. Consecutively, bootstrapping analysis was performed for the remaining variables. Continuous p21-staining and tumor size were independent prognostic factors for DSS in the group of organ-confined ccRCC (p21: HR 0.96, 95% CI 0.92–0.99, P=0.027; tumor size: HR 1.02, 95% CI 1.01–1.05, P=0.011). Compared to tumor size alone, inclusion of continuous p21-staining increased the concordance index from 0.75 to 0.78. However, this difference was not statistically significant. Due to the small numbers of patients with advanced localized or metastasized disease, we did not construct multivariate models for these cohorts.
Discussion
In the last decades, substantial achievements in predicting disease-specific mortality of patients with ccRCC were made (eg, by further improvement of the TNM-staging system or by introduction of algorithms and nomograms incorporating clinical and pathological data).1,2 These scores, such as the SSIGN (Mayo Clinic “stage, size, grade, and necrosis” score) or Leibovich score have already been externally validated.18,19 However, prediction of biology of ccRCC remains difficult and TNM-staging is currently the most relevant predictor of ccRCC.20 With increased understanding of tumor biology, molecular biomarkers are becoming an attractive target for improvement of prognosis prediction.6 Many studies analyzing the prognostic value of biomarkers were conducted, but results were sometimes conflicting, potentially reflecting the nonuniform molecular changes in ccRCC during its evolution.21 For example, carbonic anhydrase IX is one of the most important predictors of survival in metastasized ccRCC but has no prognostic value in localized disease.22 Taking this into account, we stratified our analyses of the prognostic biomarkers in organ-confined ccRCC (noninvasive) as well as advanced localized and metastasized (invasive) ccRCC as organ-confined ccRCC do not usually invade the surrounding kidney tissue,7 which is a characteristic of more advanced tumors. In this study, immunohistochemical staining patterns of VHL, Ki67, p53, p21, nS, and cS as well as MVD were analyzed to investigate whether they displayed the heterogeneity of ccRCC.
The proliferation marker Ki67 is one of the best established prognostic biomarkers for disease-specific recurrence in non-metastasized and metastasized ccRCC.3,23,24 In our patient cohort, high Ki67-staining was related to shorter DSS when analyzing all patients as well as the subgroup with advanced localized or metastasized ccRCC. In accordance with two studies analyzing Ki67-expression in pT1 renal cell carcinomas, we could not identify a relation to DSS in organ-confined ccRCC.25,26
The role of p53 in ccRCC is ambiguous.1 Whole genome sequencing of ccRCC revealed that p53 mutations are not frequent in ccRCC.21 However, high p53-expression is associated with poor survival.3,24,27 In our cohort and the cohort of Kramer et al,28 p53-staining only tended to be associated with DSS, when evaluating all patients. In our study, it was not associated with survival in the subgroup of organ-confined ccRCC. However, in patients with advanced localized and metastasized ccRCC, p53-staining was significantly linked to DSS in our cohort, regardless of whether studied separately or combined. This indicates that indirect p53-alterations might be a later step in ccRCC progression.
The cell cycle control, microtubule stabilization, and antiapoptotic functions of survivin seem to be associated with intracellular protein localization.29 Similar to Parker et al,11 we observed a significant relation between the expression of the nuclear survivin pool and prognosis when analyzing all patients. High nS-staining was associated with shorter DSS in patients with advanced localized and metastasized ccRCC or with locally advanced ccRCC only, but not in patients with organ-confined ccRCC. To our knowledge, no other study analyzed the value of nS-expression in organ-confined ccRCC. Parker et al,11 also observed a cS-staining in ccRCC but did not pursue it further. In our analysis, the cS-staining was not a predictor of prognosis for ccRCC, in contrast to another study on 85 ccRCC patients.12 One reason for the different results might be the antibodies used. We applied the commonly used polyclonal antibody AF886, whereas the monoclonal anti-survivin antibody used by Byun et al12 might not detect all survivin splice variants.
Low MVD is strongly associated with advanced disease.14 This might be a reason why MVD tended to be related with DSS in the whole cohort but lost its predictive ability when studied in the ccRCC subsets.
In a large cohort and in our study, VHL-staining was not associated with DSS.9 Possibly, VHL loss-of-function mutations lead to decreased immunohistochemical staining, but patients with lack of VHL mutations and, thus, with high VHL-staining display a shorter DSS.30,31
Interestingly, in our study, high nuclear p21-staining was associated with longer DSS in organ-confined but not in advanced localized or metastasized ccRCC. This association was seen when analyzing p21-staining as a continuous variable. Most notably, no disease-related death occurred in patients with altered p21-staining when p21-staining results were categorized according to Weiss et al.10 This group10 and Klatte et al22 also observed an association between high p21-staining and favorable prognosis, but they included patients with organ-confined and advanced localized ccRCC. As both studies did not stratify the analysis of p21-staining for these two subgroups, it is unknown if this association would be more pronounced if the subgroup of organ-confined ccRCC was analyzed separately. In addition, p21-staining lost predictive ability when analyzed in patients with advanced localized ccRCC only.
In contrast to localized ccRCC, high nuclear p21-staining was associated with poor prognosis in metastasized disease in the study of Weiss et al.10 Analyzing the 12 patients with metastases, we observed a trend for a shorter DSS in patients with high nuclear p21-staining (data not shown). Studies evaluating p21-staining in localized and metastasized ccRCC together could not uncover its prognostic value.32,33 Function of p21 depends on its subcellular localization. After phosphorylation, p21 is localized in the cytoplasm and possesses p53-independent, growth promotional, and anti-apoptotic functions.10 In the study of Weiss et al, high cytoplasmic p21-staining was associated with shorter survival in metastasized ccRCC.10 Future studies on larger patient cohorts should elucidate the prognostic potential of subcellular p21 localization.
In summary, these data suggest that p21 might play an ambiguous role in the biological behavior of ccRCC. Consistent with that, p21 is involved in various opposing processes. Depending on conditions, it can act as inhibitor or promoter of cell cycle progression.34 Besides the antiapoptotic effects of the cytoplasmic phosphorylated form, it has many oncogenic transcriptional activities.34 In breast cancer, p21 is a downstream target of transforming growth factor (TGF)-β and is essential for tumor cell invasion.35 Interestingly, TGF-β is regulated via VHL in ccRCC.36,37 p21 possibly acts as cell cycle regulator in organ-confined ccRCC, but its function switches during ccRCC progression to promote cell migration and invasion.
In our multivariate analyses, p21-staining was an independent predictor of DSS in organ-confined ccRCC and it enhanced the predictive ability of tumor size, whereas it was not an independent prognostic factor in the studies of Weiss et al10 and Klatte et al.22 Due to the low number of patients with advanced localized and metastasized disease, we only performed multivariate analyses for patients with organ-confined ccRCC.
Kim et al3 first described that incorporation of the independent survival predictors p53, vimentin, and carbonic anhydrase IX in a system of clinical markers improved predictive ability in nonselected ccRCC from 0.75 to 0.79. To our knowledge, the present analysis is the first study evaluating prognostic ability of biomarkers in the subset of organ-confined ccRCC. However, due to the high predictive accuracy (up to 88%) of prognostic models based only on clinical and radiological parameters, to date, none of the prognostic models incorporating biomarkers are recommended for routine use.1,20
Our analysis is mainly limited by the retrospective design, a relatively limited number of cases, and the small number of patients with metastases. The multiple testing procedures could have led to significant associations by chance, even if we evaluated results not only by statistical but also clinically immanent association. Due to the explorative design of the study, we did not use procedures to reduce alpha-error besides bootstrapping analysis in the multivariate model. Therefore, independent external validation of these data is needed to support our findings. After testing for nonlinearity, we analyzed all biomarkers as continuous variables to avoid insignificant associations by arbitrarily chosen cutoff values.
The study is further limited by the lack of standards for staining technique, storage, and fixation of immunohistochemical procedures. We used normal renal tissue as negative control in our study. Recent studies using techniques such as exome gene sequencing revealed an underestimated heterogeneity by analyzing only a few parts of the tumor.38 To address this, we analyzed two TMA cores per tumor. Immunohistochemical studies revealed that, even in tumors with higher intratumor heterogeneity than ccRCC, the evaluation of two TMA cores leads to accordance of staining results with whole section slides in 90%–95% of cases.39 For all analyzed proteins, the percentage of positive-stained cells between tumor center and periphery correlated significantly (P<0.001, data not shown).
The evaluation of prognostic factors stratified to ccRCC subgroups helped to enhance their prognostic ability in the context of the varying tumor biology and the molecular evolution of this disease. Embedded in the literature, these data will help to improve the prediction of disease-specific mortality and disclose targets for targeted therapy. For a robust conclusion regarding the prognostic value of p21-staining in organ-confined ccRCC shown in the present study, it should be validated in larger independent cohorts.
Conclusion
The analyzed molecular markers had distinct associations with prognosis when stratified to clinical and tumor biological subgroups. High expression of Ki67, nS, and p53 were significantly associated with worse disease-specific mortality in localized advanced or metastasized ccRCC. In contrast, p21-staining was only associated with prognosis in organ-confined disease. p21-expression remained a significant survival predictor in multivariate analyses. Further analyses are needed to confirm this association.
Acknowledgments
We acknowledge support by the German Research Foundation and the Open Access Publication Funds of the Technische Universität Dresden.
Disclosure
The authors declare no conflicts of interest in this work.
References
Sun M, Shariat SF, Cheng C, et al. Prognostic factors and predictive models in renal cell carcinoma: a contemporary review. Eur Urol. 2011;60(4):644–661. | |
Moch H, Artibani W, Delahunt B, et al. Reassessing the current UICC/AJCC TNM staging for renal cell carcinoma. Eur Urol. 2009;56(4):636–643. | |
Kim HL, Seligson D, Liu X, et al. Using protein expressions to predict survival in clear cell renal carcinoma. Clin Cancer Res. 2004;10(16):5464–5471. | |
Ficarra V, Schips L, Guillè F, et al. Multiinstitutional European validation of the 2002 TNM staging system in conventional and papillary localized renal cell carcinoma. Cancer. 2005;104(5):968–974. | |
Ficarra V, Galfano A, Guille F, et al. A new staging system for locally advanced (pT3-4) renal cell carcinoma: a multicenter European study including 2,000 patients. J Urol. 2007;178(2):418–424; discussion 423–424. | |
Lane BR, Kattan MW. Predicting outcomes in renal cell carcinoma. Curr Opin Urol. 2005;15(5):289–297. | |
Eble JN, Sauter G, Epstein JI, Sesterhenn IA, editors. World Health Organization Classification of Tumours. Pathology and Genetics of Tumours of the Urinary System and Male Genital Organs. Lyon: IARC Press; 2004;7. | |
Lughezzani G, Jeldres C, Isbarn H, et al. Tumor size is a determinant of the rate of stage T1 renal cell cancer synchronous metastasis. J Urol. 2009;182(4):1287–1293. | |
Dahinden C, Ingold B, Wild P, et al. Mining tissue microarray data to uncover combinations of biomarker expression patterns that improve intermediate staging and grading of clear cell renal cell cancer. Clin Cancer Res. 2010;16(1):88–98. | |
Weiss RH, Borowsky AD, Seligson D, et al. p21 is a prognostic marker for renal cell carcinoma: implications for novel therapeutic approaches. J Urol. 2007;177(1):63–68; discussion 68–69. | |
Parker AS, Kosari F, Lohse CM, et al. High expression levels of survivin protein independently predict a poor outcome for patients who undergo surgery for clear cell renal cell carcinoma. Cancer. 2006;107(1):37–45. | |
Byun SS, Yeo WG, Lee SE, Lee E. Expression of survivin in renal cell carcinomas: association with pathologic features and clinical outcome. Urology. 2007;69(1):34–37. | |
Roe JS, Kim H, Lee SM, Kim ST, Cho EJ, Youn HD. p53 stabilization and transactivation by a von Hippel-Lindau protein. Mol Cell. 2006;22(3):395–405. | |
Sandlund J, Hedberg Y, Bergh A, Grankvist K, Ljungberg B, Rasmuson T. Evaluation of CD31 (PECAM-1) expression using tissue microarray in patients with renal cell carcinoma. Tumour Biol. 2007;28(3):158–164. | |
Sobin L, Gospodarowicz M, Wittekind C, editors. TNM Classification of Malignant Tumours. 7th ed. New York: UICC International Union Against Cancer, Wiley-Blackwell; 2009. | |
Weber T, Meinhardt M, Zastrow S, Wienke A, Fuessel S, Wirth MP. Immunohistochemical analysis of prognostic protein markers for primary localized clear cell renal cell carcinoma. Cancer Invest. 2013;31(1):51–59. | |
Weidner N, Semple JP, Welch WR, Folkman J. Tumor angiogenesis and metastasis – correlation in invasive breast carcinoma. N Engl J Med. 1991;324(1):1–8. | |
Pichler M, Hutterer GC, Chromecki TF, et al. External validation of the Leibovich prognosis score for nonmetastatic clear cell renal cell carcinoma at a single European center applying routine pathology. J Urol. 2011;186(5):1773–1777. | |
Zigeuner R, Hutterer G, Chromecki T, et al. External validation of the Mayo Clinic stage, size, grade, and necrosis (SSIGN) score for clear-cell renal cell carcinoma in a single European centre applying routine pathology. Eur Urol. 2010;57(1):102–109. | |
Ljungberg B, Bensalah K, Bex A, et al. Guidelines on Renal Cell Carcinoma. European Association of Urology; 2013. Available from: http://www.uroweb.org/gls/pdf/10_Renal_Cell_Carcinoma_LRV2.pdf. Accessed December 30, 2013. | |
Arai E, Kanai Y. Genetic and epigenetic alterations during renal carcinogenesis. Int J Clin Exp Pathol. 2010;4(1):58–73. | |
Klatte T, Seligson DB, LaRochelle J, et al. Molecular signatures of localized clear cell renal cell carcinoma to predict disease-free survival after nephrectomy. Cancer Epidemiol Biomarkers Prev. 2009;18(3):894–900. | |
Bui MH, Visapaa H, Seligson D, et al. Prognostic value of carbonic anhydrase IX and KI67 as predictors of survival for renal clear cell carcinoma. J Urol. 2004;171(6 Pt 1):2461–2466. | |
Kroeze SG, Bijenhof AM, Bosch JL, Jans JJ. Diagnostic and prognostic tissuemarkers in clear cell and papillary renal cell carcinoma. Cancer Biomark. 2010;7(6):261–268. | |
Cheville JC, Zincke H, Lohse CM, et al. pT1 clear cell renal cell carcinoma: a study of the association between MIB-1 proliferative activity and pathologic features and cancer specific survival. Cancer. 2002;94(8):2180–2184. | |
Gelb AB, Sudilovsky D, Wu CD, Weiss LM, Medeiros LJ. Appraisal of intratumoral microvessel density, MIB-1 score, DNA content, and p53 protein expression as prognostic indicators in patients with locally confined renal cell carcinoma. Cancer. 1997;80(9):1768–1775. | |
Phuoc NB, Ehara H, Gotoh T, et al. Immunohistochemical analysis with multiple antibodies in search of prognostic markers for clear cell renal cell carcinoma. Urology. 2007;69(5):843–848. | |
Kramer BA, Gao X, Davis M, Hall M, Holzbeierlein J, Tawfik O. Prognostic significance of ploidy, MIB-1 proliferation marker, and p53 in renal cell carcinoma. J Am Coll Surg. 2005;201(4):565–570. | |
Mahotka C, Krieg T, Krieg A, et al. Distinct in vivo expression patterns of survivin splice variants in renal cell carcinomas. Int J Cancer. 2002;100(1):30–36. | |
Di Cristofano C, Minervini A, Menicagli M, et al. Nuclear expression of hypoxia-inducible factor-1alpha in clear cell renal cell carcinoma is involved in tumor progression. Am J Surg Pathol. 2007;31(12):1875–1881. | |
Patard JJ, Rioux-Leclercq N, Masson D, et al. Absence of VHL gene alteration and high VEGF expression are associated with tumour aggressiveness and poor survival of renal-cell carcinoma. Br J Cancer. 2009;101(8):1417–1424. | |
Aaltomaa S, Lipponen P, Ala-Opas M, Eskelinen M, Syrjanen K, Kosma VM. Expression of cyclins A and D and p21(waf1/cip1) proteins in renal cell cancer and their relation to clinicopathological variables and patient survival. Br J Cancer. 1999;80(12):2001–2007. | |
Haitel A, Wiener HG, Neudert B, Marberger M, Susani M. Expression of the cell cycle proteins p21, p27, and pRb in clear cell renal cell carcinoma and their prognostic significance. Urology. 2001;58(3):477–481. | |
Abbas T, Dutta A. p21 in cancer: intricate networks and multiple activities. Nat Rev Cancer. 2009;9(6):400–414. | |
Dai M, Al-Odaini AA, Arakelian A, Rabbani S, Ali S, Lebrun JJ. A novel function for p21Cip1 and acetyltransferase p/CAF as critical transcriptional regulators of TGFbeta-mediated breast cancer cell migration and invasion. Breast Cancer Res. 2012;14(5):R127. | |
Ananth S, Knebelmann B, Gruning W, et al. Transforming growth factor beta1 is a target for the von Hippel-Lindau tumor suppressor and a critical growth factor for clear cell renal carcinoma. Cancer Res. 1999;59(9):2210–2216. | |
Shang D, Liu Y, Yang P, Chen Y, Tian Y. TGFBI-promoted adhesion, migration and invasion of human renal cell carcinoma depends on inactivation of von Hippel-Lindau tumor suppressor. Urology. 2012;79(4):966. e961–e967. | |
Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012;366(10):883–892. | |
Axelrod DE, Miller N, Chapman JA. Avoiding pitfalls in the statistical analysis of heterogeneous tumors. Biomed Inform Insights. 2009;2:11–18. |
Supplementary material
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.