Back to Journals » International Journal of Nanomedicine » Volume 10 » Issue 1
Single peptide ligand-functionalized uniform hollow mesoporous silica nanoparticles achieving dual-targeting drug delivery to tumor cells and angiogenic blood vessel cells
Authors Liu Y, Chen Q, Xu M, Guan G, Hu W, Liang Y, Zhao X, Qiao M, Chen D, Liu H
Received 29 September 2014
Accepted for publication 20 November 2014
Published 6 March 2015 Volume 2015:10(1) Pages 1855—1867
DOI https://doi.org/10.2147/IJN.S75098
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Thomas Webster
Yang Liu,1,2 Qing Chen,1 Ming Xu,3 Guannan Guan,1 Wen Hu,3 Ying Liang,2 Xiuli Zhao,1 Mingxi Qiao,1 Dawei Chen,1 Hao Liu2
1School of Pharmacy, Shenyang Pharmaceutical University, Shenyang, 2Department of Pharmacy, Bengbu Medical College, Bengbu, 3College of Pharmaceutical Science, Soochow University, Suzhou, People’s Republic of China
Background: The purpose of this study was to construct hollow mesoporous silica nanoparticles (HMSN) decorated with tLyp-1 peptide (tHMSN) for dual-targeting drug delivery to tumor cells and angiogenic blood vessel cells.
Methods: HMSN were synthesized de novo using a novel cationic surfactant-assisted selective etching strategy and were then modified with tLyp-1. Multiple methods, including transmission electron microscopy, X-ray photoelectron spectroscopy, thermogravimetric analysis, bicinchoninic acid assay, and nitrogen adsorption and desorption isotherms, were used to characterize the tHMSN. Doxorubicin were chosen as the model cargo, and the uptake of doxorubicin-loaded tHMSN into MDA-MB-231 cells and human umbilical vein endothelial cells (HUVECs), as models of tumor cells and tumor neovascular endothelial cells, respectively, were observed and detected by confocal laser scanning microscopy and flow cytometry. An in vitro pharmacodynamic study and a study of the mechanism via which the nanoparticles were endocytosed were also performed.
Results: HMSN with a highly uniform size and well oriented mesopores were synthesized. After tHMSN were characterized, enhanced uptake of the cargo carried by tHMSN into MDA-MB-231 cells and HUVECs compared with that of their unmodified counterparts was validated by confocal laser scanning microscopy and flow cytometry at the qualitative and quantitative levels, respectively. Further, the pharmacodynamic study suggested that, compared with their unmodified counterparts, doxorubicin-loaded tHMSN had an enhanced inhibitory effect on MDA-MB-231 cells and HUVECs in vitro. Finally, a preliminary study on the mechanism by which the nanoparticles were endocytosed indicated that the clathrin-mediated endocytosis pathway has a primary role in the transport of tHMSN into the cytoplasm.
Conclusion: tHMSN might serve as an effective active targeting nanocarrier strategy for anti-mammary cancer drug delivery.
Keywords: hollow mesoporous silica nanoparticles, tLyp-1 peptide, drug delivery, anti-mammary cancer
Introduction
Active tumor-targeted drug delivery systems (ATT-DDS) have opened up a new area for traditional chemotherapy. Typically, there are two paths (targeting to tumor cells or tumor vessels) that can achieve effective enrichment of drugs in tumor tissues.1,2 Unfortunately, current designed ATT-DDS of single-targeting model still have many challenges for the existence of multiple barrier in vivo. For example, drugs targeting to tumor cells may cause drug resistance, which in most cases is caused by high expression of P-glycoprotein.3,4 On the other hand, the more recent therapeutic strategy of targeting to tumor vessels (often referred to as “starving tumors”) may inhibit the growth of tumor locally but increase the probability of tumor invasion and metastasis.5–8 Consequently, construction of a novel ATT-DDS that could be dual-targeting to tumor cells and tumor vessels is expected to make up for the shortcomings of the single-targeting mode and have enhanced antitumor effects. More importantly, there have been reports showing that the combination of two different targeting strategies can generate not only a combined effect, but also a synergistic effect to achieve an unprecedented therapeutic outcome in vivo.9,10 More recently, Pan et al created mesoporous silica nanoparticle (MSN)-based ATT-DDS exploiting RGD and TAT dual peptides (tumor-homing peptides and cell-penetrating peptides, respectively) as ligands to achieve sequential vascular-to-cell nuclear-targeted drug delivery for efficient tumor regression.11
Although implementation of dual-targeting to tumor cells and tumor vessels simultaneously is a promising therapeutic strategy, it is still a dilemma (for example, the preparation process is boring and complicated, and immunogenicity is increased in vivo) of multiple ligand-modified nanomaterials to achieve such an attractive goal.12,13 To address these issues, we hope to be able to construct single ligand-functionalized nanoparticles to achieve dual-targeting to tumor cells and tumor vessels.
Neuropilin (NRP), a modular transmembrane protein identified as a receptor for various forms and isoforms of vascular endothelial growth factor and members of the class 3 semaphorin family, is highly overexpressed on the surface of both mammary cancer cells14,15 and tumor vessels,16–19 where NRP1 is involved in angiogenesis and NRP2 is involved in lymphangiogenesis. Thus, NRP is a potential therapeutic target for dual-targeted anti-mammary drug delivery. tLyp-1 (sequence CGNKRTR),20,21 a newly reported heptapeptide which are screened by phage display library, is a ligand which is selectively targeted both to NRP1 and NRP2. Therefore, tLyp-1-decorated nanoparticles may be expected to have dual-targeting functions, and previous studies22,23 have shown that such polymer nanoparticles have precise dual-targeting efficacy and penetrate extensively into tumor parenchyma.
Newly developed structurally hollow MSN (HMSN)24 not only have the many advantages of traditional MSN, including negligible cytotoxicity, high biocompatibility, and appropriate stability in vivo,25–28 but also have the additional advantages of low density, a large surface area, and high drug-loading capacity.29,30 In previous studies, the most widely used strategies for synthesis of HMSN are hard-template31–33 and soft-template34–36 methods and the procedures are time-consuming, cumbersome, and difficult in terms of maintaining uniformity. Recently, a synthetic cationic surfactant-assisted selective etching strategy was developed37–39 to allow reproducible preparation of large amounts of monodispersed high-quality HMSN. Such HMSN have a uniform shape with narrowly distributed pore structures that can be adapted to allow controlled drug release. At present, encouraging progress have been reported with regard to reversal of multidrug resistance40,41 and synergistic treatment42 using MSN. However, there have been no reports on tLyp-1 peptide-decorated HMSN (tHMSN) being used to achieve dual-targeted drug delivery to tumor cells and tumor vessels.
In this research, we synthesized HMSN using a cationic surfactant-assisted selective etching strategy, and then covalently linked them with polyethylene glycol (PEG) and tLyp-1 successively. Subsequently, the dual-targeting function and relative mechanism of the nanoparticles were studied in human breast cancer (MDA-MB-231) cells and human angiogenic blood vessel cells (human umbilical vein endothelial cells [HUVECs]). All in all, tHMSN, in our opinion, has strongly prospects in active targeted drug delivery for human mammary cancer.
Materials and methods
Materials
Tetraethyl orthosilicate (TEOS), cetyltrimethylammonium bromide (CTAB), and 3-aminopropyltriethoxysilane (APTES) were obtained from Sinopharm Co Ltd (Shanghai, People’s Republic of China). Doxorubicin hydrochloride was obtained from HuaFeng Technology Co Ltd (Beijing, People’s Republic of China). Maleimido-PEG2000-N-hydroxysuccinimide (MAL-PEG2000-NHS) was provided by JenKem Technology Co Ltd (Beijing, People’s Republic of China). tLyp-1 peptide was custom-synthesized by GL Biochem Ltd (Shanghai, People’s Republic of China). Hoechst 33342 were purchased from Molecular Probes (Eugene, OR, USA). Cell Counting Kit-8 (CCK-8) was purchased from Boster Corporation (Wuhan, People’s Republic of China). Chlorpromazine hydrochloride and colchicine were obtained from Genestar Co Ltd (Shanghai, People’s Republic of China). All other chemicals were of analytical or chromatographic grade and were used without further purification.
Synthesis of HMSN
The synthesis of HMSN, PEG-HMSN (pHMSN), and tHMSN is shown in Figure 1. The synthesis process used for HMSN followed that of previous reports37,38 with slight modifications. Broadly, the process consisted of three steps, as follows: first, solid SiO2 nanospheres (SSN) were prepared via a sol-gel process using a modified Stöber method. Briefly, 74 mL of ethanol, 3.15 mL of aqueous ammonia solution (26%–28%), and 10 mL of ultrapure water were mixed and heated to 50°C, at which point 6 mL of tetraethyl orthosilicate was added rapidly. After 3 hours of reaction at 50°C, SSN were obtained by centrifugation and washed repeatedly with ultrapure water and ethanol. Second, 50 mg of SSN were homogeneously dispersed in a 1:2 (v/v) ethanol/water mixture, and 75 mg of CTAB, 300 μL of aqueous ammonia solution (26%–28%), and 125 μL of tetraethyl orthosilicate were added successively to the above mixture. After 12 hours of reaction at room temperature, the formed solid core-shell nanoparticles, denoted SSN@CTAB/SiO2, were centrifugally separated from the suspension and washed with ultrapure water. Third, 100 mg of SSN@CTAB/SiO2 nanomaterials were dispersed in 20 mL of water and heated to 50°C. Next, 464 mg of Na2CO3 was added into the mixture. The reaction was allowed to react for 11 hours at 50°C before the acquired HMSN structures were harvested by centrifugation. To remove the surfactant template in the shell of the HMSN, the products were dispersed in 120 mL of NH4NO3 ethanol solution (10 mg/mL). The mixture was heated to 45°C and stirred for 10 hours. The silica particles were centrifugally separated from the suspension and finally dispersed in dimethyl sulfoxide for further use.
Synthesis of tHMSN and pHMSN
The synthesis of tHMSN consists of three steps (Figure 1) by the Michael addition reaction, and PEG2000 was used as a linker to connect HMSN with the tLyp-1 polypeptide. First, 8 mg of HMSN were dispersed in 50 mL of dimethyl sulfoxide and then 500 μL of 3-aminopropyltriethoxysilane were added. After the mixture was stirred for 20 hours, amine-functionalized HMSN (HMSN-NH2) were obtained by centrifugation and washed with ethanol twice. Next, HMSN-NH2 was redispersed in 5 mL of dimethyl sulfoxide, and 8 mg of NHS-PEG2000-MAL was added. The mixture was then stirred at room temperature for 2 hours under anhydrous conditions before harvesting by centrifugation. The product was then redispersed in dimethyl sulfoxide, and 3 mg of thiol end-capped tLyp-1 peptide was added. After 24 hours of reaction, 10 μL of β-mercaptoethanol was added to the mixture to quench the unreacted maleimide groups. The entire reaction was stopped one hour later by centrifugation, and excess tLyp-1 was removed by repeatedly washing the nanoparticles in distilled water. Finally, tHMSN were dispersed in water and stored at 4°C. For synthesis of pHMSN, the same process was followed as for tHMSN, except that tLyp-1 was not added.
Characterization of nanoparticles
The hydrodynamic size and zeta potential of HMSN, pHMSN, and tHMSN were determined by dynamic laser scattering analysis. Morphological examination of the HMSN was also performed using a transmission electron microscope. Nitrogen adsorption–desorption isotherms were measured using a surface area and porosity analyzer. Specific surface area was calculated by the Brunauer–Emmett–Teller method and the pore size distribution was calculated using the Barrett–Joyner–Halenda model. The bicinchoninic acid assay, X-ray photoelectron spectroscopy, and thermogravimetric analysis were used to verify the surface modification of tHMSN and conjugation of tLyp-1 and PEG on the tHMSN surface. A detailed description of the various methods used is given in the Supplementary materials.
Drug-loading capacity and in vitro release study
Doxorubicin was loaded into tHMSN, pHMSN, and HMSN by mixing doxorubicin (1 mg/mL) with the nanomaterials in dimethyl sulfoxide. The mixture was ultrasonicated for 30 minutes and then stirred at 30°C for 24 hours. The remaining free doxorubicin was removed by centrifugation and the precipitate was washed with ultrapure water twice. To evaluate the doxorubicin-loading efficiency, the supernatant was collected and the residual doxorubicin content was determined using the calibration curve for standard doxorubicin solutions with ultraviolet-visible measurement at 496 nm. The amount of doxorubicin loaded into the nanomaterials was calculated by subtracting the mass of doxorubicin in the supernatant from the total mass of drug in the initial solution. The loading capacity (LC) of doxorubicin in the nanoparticles was calculated using the following equation (n=3): LC (%) = [amount of doxorubicin in the nanoparticles/(nanoparticles weight + amount of doxorubicin in the nanoparticles)] ×100%.
In vitro release characteristics were investigated using the dialysis method. Briefly, the drug-loaded hollow nanoparticles were dispersed in phosphate-buffered saline (PBS, 0.1 M, pH 7.4 or pH 5.0) and placed in a dialysis bag (molecular weight cut-off, 3,500 Da) to simulate the sink condition. The release study was performed at 37°C in a shaking water bath (SHA-C, Jinda Instruments Inc, Ningbo, People’s Republic of China) at 100 rpm. At designated time intervals, 1 mL of solution was removed from outside of the dialysis bag for analysis of fluorescence emission spectra (excitation 467 nm, emission 557 nm) using a fluorescence spectrophotometer (Fluoroma-4, Horiba Scientific, Tokyo, Japan) and replaced with an equal amount of fresh PBS. To assess the drug-release behavior, the cumulated amounts of released drug were calculated, and the percentages of drug released from the nanoparticles were plotted against time.
Cell lines and culture conditions
The MDA-MB-231 cell line was purchased from the Cell Institute of the Chinese Academy of Sciences (Shanghai, People’s Republic of China) and primary HUVECs were obtained from Cell Applications Inc (San Diego, CA, USA). The cells were cultured in Dulbecco’s Modified Eagle’s Medium supplemented with 10% (v/v) fetal bovine serum, 100 U/mL penicillin, and 100 μg/mL streptomycin sulfate at 37°C in 5% CO2.
Cell viability assay
The cytotoxicity of the nanoparticles against MDA-MB-231 cells and HUVECs was evaluated by the CCK-8 assay. The CCK-8 assay is highly sensitive for cell viability, and previous studies43,44 have shown that it may be more suited for MSN than the MTT assay. The cells were seeded in 96-well plates at a density of 5×103 cells/well and allowed to adhere for 24 hours prior to the assay. The cells were then exposed to nanoparticles loaded or not loaded with doxorubicin at the designated doses. Cell viability, based on a CCK-8 assay, was determined after each treatment by measuring absorbance at 450 nm using a microplate reader. Free doxorubicin hydrochloride was used as a positive control and cells that had not been incubated with the test nanoparticles were used as a negative control.
Analysis of cellular uptake by flow cytometry
MDA-MB-231 cells or HUVECs were seeded onto six-well plates at 3×105 cells per well and cultured 24 hours for the flow cytometric analysis. After the attachment period, the cells were incubated with tHMSN, pHMSN, or HMSN loaded with doxorubicin for 0.5–2 hours in serum-free medium. After incubation, the cells were washed twice with cold PBS (0.1 M, pH 7.4), detached with 0.05% trypsin, and washed a further three times with cold PBS. Finally, the cells were resuspended in PBS and analyzed using a flow cytometer (BD Biosciences, San Jose, CA, USA). The autofluorescence of the cells was used as the control.
Direct observation of cellular uptake
To directly observe the cellular uptake, MDA-MB-231 cells or HUVECs were seeded onto cover glass in six-well plates at 2×105 cells per well and cultured for 24 hours. After the attachment period, the cells were incubated with the different types of nanoparticles loaded with doxorubicin for 0.5–2 hours in serum-free medium. After removing the nanoparticles and washing the wells twice with cold PBS, the cell nuclei were stained with Hoechst 33342 for 5 minutes and then fixed by 4% polyoxymethylene for 10 minutes. The cells were observed by confocal laser scanning microscopy (LSM-710, Carl Zeiss AG, Germany). Doxorubicin and Hoechst 33342 were excited with an argon laser at 488 nm and 405 nm, respectively.
Investigation of mechanism involved in cellular uptake
In order to study the mechanism of uptake of tHMSN in MDA-MB-231 cells and HUVECs, cellular association of the nanoparticles was determined in the presence of various endocytosis inhibitors, including chlorpromazine (an inhibitor of clathrin-mediated uptake, 10 μg/mL), colchicine (an inhibitor of macropinocytosis, 4 μg/mL), methyl-β-cyclodextrin (a lipid raft inhibitor, 5 mM), genistein (an inhibitor of caveolin-dependent endocytosis, 200 μM), wortmannin (an inhibitor of macropinocytosis and compensatory RhoA-mediated endocytosis, 50 nM), and free tLyp-1 (160 μg/mL), respectively. The cells were pretreated with these inhibitors (all chemical dilutions were prepared in serum-free medium) for 30 minutes followed by immediate incubation with tHMSN in serum-free medium at 37°C for an additional 1 hour. The quantitative analysis was performed by flow cytometry as described above.
Statistical analysis
The data are expressed as the mean ± standard deviation. The resulting values from each experiment were compared by one-way analysis of variance, and multiple comparisons between the groups were performed using the Student–Newman–Keuls method. Statistical significance was set at a level of P<0.05.
Results and discussion
Synthesis and characterization of nanoparticles
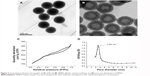
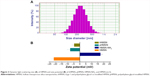
Uniform and monodispersed SSN and HMSN were prepared following the method proposed by Fang et al37,38 with minor modifications. Transmission electron micrographs of the particles showed clearly that both SSN (Figure 2A) and HMSN (Figure 2B) had a spherical morphology and uniform particle size. The mean size of HMSN according to dynamic light scattering was 258.1±9.4 nm (n=3, Figure 3A), which was larger than the value obtained on transmission electron micrographs due to the presence of hydrated layers around the surfaces of the particles. Nitrogen adsorption–desorption isotherm measurements indicated that the HMSN had a relatively high specific surface area of 547.79 m2/g (Figure 2C) and a well-defined mean pore size of 2.54 nm (Figure 2D). Uniform particle sizes with a controllable well-defined pore structure and a high specific surface area indicated that these nanomaterials could be a promising candidate for high-performance drug delivery.45
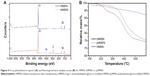
Zeta potential, bicinchoninic acid, X-ray photoelectron spectroscopy, and thermogravimetric analysis assays were carried out to confirm surface modification of the nanoparticles. The particle size of tHMSN and pHMSN showed minor amplification when compared with HMSN, but the zeta potential showed a more marked change (Figure 3B). This change could be attributed to the different chemical groups on the surfaces of the nanoparticles, including a silicon hydroxyl group (Si-OH) on HMSN, a primary amino group (NH2) on HMSN-NH2, and an alkyl hydroxyl group (CH2-OH) originating from PEG (or β-mercaptoethanol) on pHMSN and tHMSN. Further, bicinchoninic acid assay identified peptides on the surfaces of tHMSN but not on pHMSN or HMSN. X-ray photoelectron spectroscopy, a surface chemical analysis technique, was performed. Comparing the X-ray photoelectron spectrum (Figure 4A) of tHMSN with that of HMSN, the peaks seen at 400.2 eV and 286.2 eV indicate that there were additional N and C elements on the surface of tHMSN, which were resourced from tLyp-1 peptide and PEG, respectively. But such elements were not found in HMSN.
Thermogravimetric analysis is a thermal method via which changes in the physical and chemical properties of materials can be measured as a function of increasing temperature. The thermogravimetric analysis curve (Figure 4B) shown more obvious weight loss with increasing temperature for tHMSN and pHMSN when compared with HMSN, further demonstrating the difference in chemical constituents of the three types of nanoparticles. Overall, these results confirm successful synthesis of tHMSN from HMSN nanoparticles.
Evaluation of drug-loading and drug-release properties
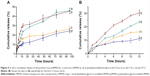
Typically, HMSN had a huge internal cavity and large specific surface area, ensuring a high drug loading capacity (LC). Not surprisingly, the LC of doxorubicin, a model hydrophobic drug, was calculated to be 29.5±1.3 wt%, 24.9±1.1 wt%, and 23.8±2.2 wt% for HMSN, pHMSN, and tHMSN, respectively (n=3). The drug-release profiles for the nanoparticles in neutral or acid PBS buffer were then investigated; similar biphasic patterns were seen, as characterized by fast initial release within the first 12 hours (Figure 5B) and slow continuous release in the subsequent 84 hours (Figure 5A). The rate of release of doxorubicin from the nanoparticles at pH 5.0 was clearly more rapid than that at pH 7.4. These results may be ascribed to the fact that the solubility of doxorubicin is significantly influenced by pH in aqueous medium. It is worth mentioning that such a characteristic (ie, pH-sensitive release of cargo) is very favorable in antitumor treatment given that the microenvironments of extracellular tissues of tumors and intracellular lysosomes and endosomes are acidic.46
Cytotoxicity of nanoparticles
To evaluation the cytotoxicity and targeting capability of tHMSN vector, human mammary cancer MDA-MB-231 cells and HUVECs, both of which NRP were abundantly expressed, were used as the models of tumor cells and tumor neovascular endothelial cells, respectively.20,22,23,47,48
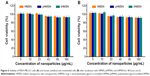
Low cytotoxicity of the carrier itself is a primary concern in the development of a drug delivery system. Figure 6A and B shows the viability of MDA-MB-231 cells and HUVECs incubated with HMSN, pHMSN, or tHMSN for 48 hours (as determined by CCK-8 assay). Obviously, these vehicles showed very little toxicity to cells, even when the nanoparticles reached concentrations up to 160 μg/mL. Importantly, conjugation of the tLyp-1 peptide had a negligible influence on the cytotoxicity of the nanomaterials.
Cellular uptake of nanoparticles
Given that overexpression of NRP has been noted on the surface of endothelial cells in tumor angiogenic blood vessels and mammary cancer cells, tLyp-1 decorated nanoparticles might be taken up in greater amounts by endothelial cells in angiogenic blood vessels and by mammary cancer cells. To test this hypothesis, the model cells were incubated with tHMSN, pHMSN, and HMSN, respectively, and analyzed by flow cytometry or observed by confocal laser scanning microscopy.
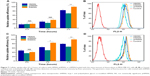
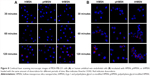
As expected, the flow cytometry assay demonstrated that tHMSN had the highest cellular uptake efficiency of the three types of nanoparticles when loaded with the same amount of doxorubicin (6 μg/mL), although the endocytosis of tHMSN into MDA-MB-231 cells (Figure 7A and B, P<0.01) or HUVECs (Figure 7C and D, P<0.01) was slightly higher than that of HMSN, but was significantly higher than that of their tLyp-1 unmodified counterparts (pHMSN). The lower uptake efficiency of pHMSN compared with that of HMSN can be explained by the sterically hindered effects of PEG chains grafted onto the surface of nanoparticles. However, compared with HMSN and pHMSN, tHMSN, which was tLyp-1 and PEG co-modified HMSN, exhibited higher uptake efficiency into the cytoplasm of tumor cells or angiogenic blood vessels cells and this might ascribed to the specific recognition between tLyp-1 peptide and the overexpressed receptor proteins of NRP on the cell membrane of the target cells. Confocal laser scanning microscopy was then performed to further visualize the behavior of the nanoparticles in cells. After 30 minutes of incubating the cells with nanoparticles loaded with the same amount of doxorubicin (6 μg/mL), no significant difference of the red fluorescence intensity was detected within each group. However, with extension of the incubation time to 2 hours, there was an increasingly discrepancy between the tHMSN group and the pHMSN group (Figure 8A and B).
In vitro pharmacodynamic study
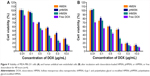
To investigate the pharmacodynamics of the nanoparticles, we compared the cytotoxicity of the different drug-loaded nanoparticles using the CCK-8 assay. The results (Figure 9A and B) show that increased cytotoxicity of doxorubicin-loaded tHMSN compared with doxorubicin-loaded pHMSN, and the IC50 value decreased by 65.06% (from 2.69 μg/mL to 0.94 μg/mL for MDA-MB-231 cells) and by 51.02% (from 0.98 μg/mL to 0.48 μg/mL for HUVECs). Accordingly, tHMSN can not only be regarded as a specific and efficient drug delivery carrier, but also as having huge potential for targeted cancer therapy.
On the basis of the results for cellular uptake and cytotoxicity of the drug-loaded nanoparticles, it can be concluded that tHMSN could enhance endocytosis of cargoes and thereby mediating more robust cytotoxicity compared with pHMSN. Meanwhile, given the overexpression of NRP in many tumor cells and angiogenic blood vessel cells,18–20 such an ability should have a degree of selectivity and dual-targeting performance.
Cellular uptake mechanism of nanoparticles
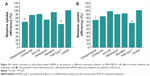
To explore the endocytic pathway of tHMSN further, the influence of different endocytosis inhibitors (competitive and noncompetitive) on the rate of uptake of the doxorubicin-loaded nanomaterials by cells was investigated by flow cytometry. The results shown that free tLyp-1, as the competitive inhibitors, were effective in inhibiting the uptake of tHMSN into cytoplasma of MDA-MB-231 (Figure 10A, P<0.01) or HUVECs (Figure 10B, P<0.01). This finding indicates that the tLyp-1 peptide, when covalently linked to tHMSN, has an important role in the uptake of these nanoparticles. Meanwhile, investigation of the inhibitory effect of five noncompetitive endocytosis inhibitors (chlorpromazine, colchicine, genistein, wortmannin, and methyl-β-cyclodextrin) on the uptake of tHMSN into the cytoplasm of MDA-MB-231 (Figure 10A, P<0.01) and HUVECs (Figure 10B, P<0.05) showed that chlorpromazine both had the strongest inhibitory effect. These results demonstrate (or at least to some extent indicate) that clathrin-mediated endocytosis might play a primary role in transport of tHMSN into the cytoplasm.49
Conclusion
In this study, we developed a novel drug delivery system based on incorporation of HMSN, PEG, and target peptides (tLyp-1). The tHMSN vectors obtained had excellent biocompatibility and a high endocytosis rate, thereby mediating a remarkable dual-targeting cytotoxic effect on both tumor cells and angiogenic blood vessels cells after drug loaded. Therefore, we believe that this novel, high-capacity, and efficient HMSN-based drug delivery system has considerable potential for translation into a treatment for cancer.
Acknowledgment
We acknowledge the National Natural Science Foundation of China (No 81341134) for financial support.
Disclosure
The authors report no conflicts of interest in this work.
References
Arap W, Pasqualini R, Ruoslahti E. Cancer treatment by targeted drug delivery to tumor vasculature in a mouse model. Science. 1998;279(5349):377–380. | ||
Quail DF, Joyce JA. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19(11):1423–1437. | ||
Yan Y, Björnmalm M, Caruso F. Particle carriers for combating multidrug-resistant cancer. ACS Nano. 2013;7(11):9512–9517. | ||
Wu CP, Hsieh CH, Wu YS. The emergence of drug transporter-mediated multidrug resistance to cancer chemotherapy. Mol Pharm. 2011;8(6):1996–2011. | ||
Ebos JM, Kerbel RS. Antiangiogenic therapy: impact on invasion, disease progression, and metastasis. Nat Rev Clin Oncol. 2011;8(4):210–221. | ||
Pàez-Ribes M, Allen E, Hudock J, et al. Antiangiogenic therapy elicits malignant progression of tumors to increased local invasion and distant metastasis. Cancer Cell. 2009;15(3):220–231. | ||
Ebos JM, Lee CR, Cruz-Munoz W, Bjarnason GA, Christensen JG, Kerbel RS. Accelerated metastasis after short-term treatment with a potent inhibitor of tumor angiogenesis. Cancer Cell. 2009;15(3):232–239. | ||
Valastyan S, Weinberg RA. Tumor metastasis: molecular insights and evolving paradigms. Cell. 2011;147(2):275–292. | ||
Pastorino F, Brignole C, Di Paolo D, et al. Targeting liposomal chemotherapy via both tumor cell-specific and tumor vasculature-specific ligands potentiates therapeutic efficacy. Cancer Res. 2006;66(20):10073–10082. | ||
Gao H, Yang Z, Cao S, et al. Tumor cells and neovasculature dual targeting delivery for glioblastoma treatment. Biomaterials. 2014;35(7):2374–2382. | ||
Pan L, Liu J, He Q, Shi J. MSN – mediated sequential vascular to cell nuclear-targeted drug delivery for efficient tumor regression. Adv Mater. 2014;39(26):6742–6748. | ||
Gao H, Xiong Y, Zhang S, Yang Z, Cao S, Jiang X. RGD and interleukin-13 peptide functionalized nanoparticles for enhanced glioblastoma cells and neovasculature dual targeting delivery and elevated tumor penetration. Mol Pharm. 2014;11(3):1042–1052. | ||
Kibria G, Hatakeyama H, Ohga N, Hida K, Harashima H. Dual-ligand modification of PEGylated liposomes shows better cell selectivity and efficient gene delivery. J Control Release. 2011;153(2):141–148. | ||
Soker S, Takashima S, Miao HQ, Neufeld G, Klagsbrun M. Neuropilin-1 is expressed by endothelial and tumor cells as an isoform-specific receptor for vascular endothelial growth factor. Cell. 1998;92(6):735–745. | ||
Bachelder RE, Crago A, Chung J, et al. Vascular endothelial growth factor is an autocrine survival factor for neuropilin-expressing breast carcinoma cells. Cancer Res. 2001;61(15):5736–5740. | ||
Murga M, Fernandez-Capetillo O, Tosato G. Neuropilin-1 regulates attachment in human endothelial cells independently of vascular endothelial growth factor receptor-2. Blood. 2005;105(5):1992–1999. | ||
Wang L, Zeng H, Wang P, Soker S, Mukhopadhyay D. Neuropilin-1-mediated vascular permeability factor/vascular endothelial growth factor-dependent endothelial cell migration. J Biol Chem. 2003;278(49):48848–48860. | ||
Pan Q, Chanthery Y, Liang WC, et al. Blocking neuropilin-1 function has an additive effect with anti-VEGF to inhibit tumor growth. Cancer Cell. 2007;11(1):53–67. | ||
Dallas NA, Gray MJ, Xia L, et al. Neuropilin-2-mediated tumor growth and angiogenesis in pancreatic adenocarcinoma. Clin Cancer Res. 2008;14(24):8052–8060. | ||
Roth L, Agemy L, Kotamraju VR, et al. Transtumoral targeting enabled by a novel neuropilin-binding peptide. Oncogene. 2011;31(33):3754–3763. | ||
Alberici L, Roth L, Sugahara KN, et al. De novo design of a tumor-penetrating peptide. Cancer Res. 2013;73(2):804–812. | ||
Hu Q, Gu G, Liu Z, et al. F3 peptide-functionalized PEG-PLA nanoparticles co-administrated with tLyp-1 peptide for anti-glioma drug delivery. Biomaterials. 2013;34(4):1135–1145. | ||
Hu Q, Gao X, Gu G, et al. Glioma therapy using tumor homing and penetrating peptide-functionalized PEG–PLA nanoparticles loaded with paclitaxel. Biomaterials. 2013;34(22):5640–5650. | ||
He Q, Shi J. Mesoporous silica nanoparticle based nano drug delivery systems: synthesis, controlled drug release and delivery, pharmacokinetics and biocompatibility. J Mater Chem. 2011;21(16):5845–5855. | ||
He Q, Shi J. MSN anti-cancer nanomedicines: chemotherapy enhancement, overcoming of drug resistance, and metastasis inhibition. Adv Mater. 2014;26(3):391–411. | ||
He Q, Zhang Z, Gao Y, Shi J, Li Y. Intracellular localization and cytotoxicity of spherical mesoporous silica nano-and microparticles. Small. 2009;5(23):2722–2729. | ||
He Q, Zhang Z, Gao F, Li Y, Shi J. In vivo biodistribution and urinary excretion of mesoporous silica nanoparticles: effects of particle size and PEGylation. Small. 2011;7(2):271–280. | ||
Wu SH, Mou CY, Lin HP. Synthesis of mesoporous silica nanoparticles. Chem Soc Rev. 2013;42(9):3862–3875. | ||
Mamaeva V, Sahlgren C, Lindén M. Mesoporous silica nanoparticles in medicine – recent advances. Adv Drug Deliv Rev. 2013;65(5):689–702. | ||
Shen J, Song G, An M, et al. The use of hollow mesoporous silica nanospheres to encapsulate bortezomib and improve efficacy for non-small cell lung cancer therapy. Biomaterials. 2014;35(1):316–326. | ||
Zhang Q, Zhang T, Ge J, Yin Y. Permeable silica shell through surface-protected etching. Nano Lett. 2008;8(9):2867–2871. | ||
Zhang T, Ge J, Hu Y, Zhang Q, Aloni S, Yin Y. Formation of hollow silica colloids through a spontaneous dissolution-regrowth process. Angew Chem Int Ed Engl. 2008;120(31):5890–5895. | ||
Chen Y, Chen H, Guo L, et al. Hollow/rattle-type mesoporous nanostructures by a structural difference-based selective etching strategy. ACS Nano. 2009;4(1):529–539. | ||
Wu X J, Xu D. Soft template synthesis of yolk/silica shell particles. Adv Mater. 2010;22(13):1516–1520. | ||
Wang J, Xiao Q, Zhou H, et al. Budded, mesoporous silica hollow spheres: hierarchical structure controlled by kinetic self-assembly. Adv Mater. 2006;18(24):3284–3288. | ||
Liu J, Hartono SB, Jin YG, et al. A facile vesicle template route to multi-shelled mesoporous silica hollow nanospheres. J Mater Chem. 2010;20(22):4595–4601. | ||
Fang X, Chen C, Liu Z, Liu P, Zheng N. A cationic surfactant assisted selective etching strategy to hollow mesoporous silica spheres. Nanoscale. 2011;3(4):1632–1639. | ||
Fang W, Tang S, Liu P, Fang X, Gong J, Zheng N. Pd nanosheet-covered hollow mesoporous silica nanoparticles as a platform for the chemo-photothermal treatment of cancer cells. Small. 2012;8(24):3816–3822. | ||
Fang X, Zhao X, Fang W, Chen C, Zheng N. Self-templating synthesis of hollow mesoporous silica and their applications in catalysis and drug delivery. Nanoscale. 2013;5(6):2205–2218. | ||
Pan L, Liu J, He Q, Wang L, Shi J. Overcoming multidrug resistance of cancer cells by direct intranuclear drug delivery using TAT-conjugated mesoporous silica nanoparticles. Biomaterials. 2013;34(11):2719–2730. | ||
Meng H, Mai WX, Zhang H, et al. Codelivery of an optimal drug/siRNA combination using mesoporous silica nanoparticles to overcome drug resistance in breast cancer in vitro and in vivo. ACS Nano. 2013;7(2):994–1005. | ||
Shen S, Tang H, Zhang X, et al. Targeting mesoporous silica-encapsulated gold nanorods for chemo-photothermal therapy with near-infrared radiation. Biomaterials. 2013;34(12):3150–3158. | ||
Fisichella M, Dabboue H, Bhattacharyya S, et al. Mesoporous silica nanoparticles enhance MTT formazan exocytosis in HeLa cells and astrocytes. Toxicol In Vitro. 2009;23(4):697–703. | ||
Laaksonen T, Santos H, Vihola H, et al. Failure of MTT as a toxicity testing agent for mesoporous silicon microparticles. Chem Res Toxicol. 2007;20(12):1913–1918. | ||
Argyo C, Weiss V, Bräuchle C, Bein T. Multifunctional mesoporous silica nanoparticles as a universal platform for drug delivery. Chem Mater. 2013;26(1):435–451. | ||
Quail DF, Joyce JA. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19(11):1423–1437. | ||
Kang T, Gao X, Hu Q, et al. iNGR-modified PEG-PLGA nanoparticles that recognize tumor vasculature and penetrate gliomas. Biomaterials. 2014;35(14):4319–4332. | ||
Zhu S, Qian L, Hong M, Zhang L, Pei Y, Jiang Y. RGD-Modified PEG-PAMAM-DOX conjugate: in vitro and in vivo targeting to both tumor neovascular endothelial cells and tumor cells. Adv Mater. 2011;23(12):H84–H89. | ||
Kaksonen M, Toret CP, Drubin DG. Harnessing actin dynamics for clathrin-mediated endocytosis. Nat Rev Mol Cell Biol. 2006;7(6):404–414. |
Supplementary material
Detailed description of methods used to characterize the nanoparticles
The particle hydrodynamic size and zeta potential of hollow mesoporous silica nanoparticles (HMSN), tLyp-1 and polyethylene glycol co-modified HMSN (tHMSN), and polyethylene glycol-modified HMSN (pHMSN) were determined by dynamic light scattering analysis using a zeta potential/particle sizer (Nicomp 380, Particle Sizing Systems, Horiba Scientific, Edison, NJ, USA). Morphological examination of HMSN was performed using a transmission electron microscope (Tecnai G-20, FEI, Hillsboro, NJ, USA). Nitrogen adsorption–desorption isotherms were measured at 77 K using a surface area and porosity analyzer (ASAP2020, Micromeritics Instrument Corporation, Norcross, GA, USA). The specific surface area was calculated by the Brunauer–Emmett–Teller method and the pore size distribution was calculated using the Barrett–Joyner–Halenda model.
Bicinchoninic acid assay (BCA), X-ray photoelectron spectroscopy, and thermogravimetric analysis were used to confirm conjugation of tLyp-1 and polyethylene glycol on the surface of the tHMSN. Qualitative assessment of tLyp-1 bound on tHMSN was determined by BCA. Briefly, the BCA working solution was prepared by mixing reagent A with reagent B (reagent A:B 50:1, v/v) for further use. Next, 20 μL of tHMSN (HMSN and pHMSN acted as control) were added into a 96-well plate, followed by incubation with 200 μL of BCA working solution. After 30 minutes at 60°C, the assay was carried out at 570 nm using a microplate reader (Synergy TM2, Bio-Tek Instruments Inc, Winooski, VT, USA). The nanoparticle samples were also lyophilized (0.180 mbar vacuum, −50°C) using a freeze dryer (Alpha 2–4, Labconco, Kansas City, MO, USA) and then analyzed by X-ray photoelectron spectroscopy (Escalab 250Xi, Thermo Fisher Scientific, Waltham, MA, USA) to determine the surface composition. Further, the samples were subjected to thermogravimetric analysis (SDT-2960, TA Instruments, New Castle, DE, USA) in a controlled atmosphere of nitrogen from 90°C up to 590°C at a constant heating rate of 10°C per minute.
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.