Back to Journals » Therapeutics and Clinical Risk Management » Volume 10
Sexual dimorphism of miRNA expression: a new perspective in understanding the sex bias of autoimmune diseases
Received 23 October 2013
Accepted for publication 3 December 2013
Published 3 March 2014 Volume 2014:10 Pages 151—163
DOI https://doi.org/10.2147/TCRM.S33517
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 6
Rujuan Dai, S Ansar Ahmed
Department of Biomedical Sciences and Pathobiology, Virginia-Maryland Regional College of Veterinary Medicine, Virginia Tech, Blacksburg, VA, USA
Abstract: Autoimmune diseases encompass a diverse group of diseases which emanate from a dysregulated immune system that launches a damaging attack on its own tissues. Autoimmune attacks on self tissues can occur in any organ or body system. A notable feature of autoimmune disease is that a majority of these disorders occur predominantly in females. The precise basis of sex bias in autoimmune diseases is complex and potentially involves sex chromosomes, sex hormones, and sex-specific gene regulation in response to internal and external stimuli. Epigenetic regulation of genes, especially by microRNAs (miRNAs), is now attracting significant attention. miRNAs are small, non-protein-coding RNAs that are predicted to regulate a majority of human genes, including those involved in immune regulation. Therefore, it is not surprising that dysregulated miRNAs are evident in many diseases, including autoimmune diseases. Because there are marked sex differences in the incidence of autoimmune diseases, this review focuses on the role of sex factors on miRNA expression in the context of autoimmune diseases, an aspect not addressed thus far. Here, we initially review miRNA biogenesis and miRNA regulation of immunity and autoimmunity. We then summarize the recent findings of sexual dimorphism of miRNA expression in diverse tissues, which imply a critical role of miRNA in sex differentiation and in sex-specific regulation of tissue development and/or function. We also discuss the important contribution of the X chromosome and sex hormones to the sexual dimorphism of miRNA expression. Understanding sexually dimorphic miRNA expression in sex-biased autoimmune diseases not only offers us new insight into the mechanism of sex bias of the disease but will also aid us in developing new sex-based therapeutic strategies for the efficient treatment of these diseases with a sex bias.
Keywords: sex differences, microRNA, immune regulation, autoimmune diseases, X chromosome, sex hormones
Introduction
It is now well-established that males and females have different immune capabilities, consequently displaying differing degrees of susceptibility to and severity of various diseases, including autoimmune diseases.1–4 Autoimmune diseases arise from a devastating attack on self antigens and self tissues by a malfunctioning immune system. Thus far, more than 80 different human diseases with an autoimmune component have been recognized. These autoimmune-related disorders collectively affect approximately 5%–10% of the population of the Western countries.5 Autoimmune disorders are often lifelong and seriously compromise quality of life, resulting in an enormous health care cost burden globally.6
Strikingly, about 80% of autoimmune patients are women. Autoimmune diseases are among the top ten causes of all deaths and are the fourth largest cause of disability among women.6,7 The female-to-male patient ratio varies from 2:1 to 25:1 for those autoimmune diseases with female prevalence.1,3 It must be noted that not all autoimmune diseases occur exclusively or preferentially in females. In some autoimmune diseases, both males and females are equally susceptible, or the diseases have a tendency to occur more often in males. Nevertheless, overall, a majority of autoimmune diseases occur in women. Despite extensive studies in past decades, the etiology of autoimmune diseases and the mechanism of female prevalence of autoimmune diseases remain elusive. Multiple factors including genetic predisposition, epigenetic dysregulation, and hormonal and environmental influence are thought to contribute to the pathogenesis of autoimmune disease. These factors likely interact with each other to confer susceptibility to autoimmune disease. Sex differences in immune responses and prevalence of autoimmune diseases are attributed to direct differences in male and female genome and sex hormone production, as well as sex-specific epigenetic regulation and gene expression.8–11 One of the important epigenetic mechanisms is microRNA (miRNA) regulation of gene expression at the posttranscriptional level, which is a focus of this review in relation to sex differences in autoimmune diseases.
miRNAs are a group of endogenously expressed, small, non-protein-coding RNAs (about 22 nucleotides [nts] in size) that regulate gene expression posttranscriptionally. Since the discovery of the first miRNA, lin-4, in nematode Caenorhabditis elegans in 1993, the importance of miRNAs in the regulation of various biological functions including stem cell development and differentiation, organogenesis, early embryo development, and immune responses has been highly appreciated.12–15 Dysregulated miRNA expression profiles have been observed in various human diseases, including female-predominant autoimmune diseases, such as systemic lupus erythematosus (SLE), rheumatoid arthritis (RA), Sjögren’s syndrome, and multiple sclerosis, implying their contribution to the etiology of these diseases.13,16–21 It is plausible that miRNAs are differentially expressed between males and females in the context of autoimmune diseases, leading to sex differences in disease susceptibility and severity. In support of this view, our laboratory recently demonstrated the sex differences in the expression of lupus-associated miRNAs in immune cells from NZB/WF1 mice, which manifest lupus predominantly in females.22 In this review, we summarize the sexual dimorphism of miRNA expression that is observed in diverse gonadal and somatic tissues and further discuss the potential involvement of sexually dimorphic miRNA regulation in the development of sex-biased autoimmune disease, an aspect not addressed thus far.
miRNA biogenesis and function
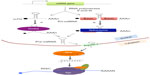
So far, 24,521 hairpin precursor miRNAs entries, which produce 30,424 mature miRNAs in 206 species, have been registered in the miRNA database (miRBase [http://www.mirbase.org], version 20, released June 2013). The majority of miRNA genes (about 80%) are located in the intergenic and intronic regions of protein-coding and non-protein-coding genes. The remaining 20% of the miRNA genes are located in the exonic regions of noncoding transcripts, of which some are considered a “mix” (either exonic or intronic miRNA, depending on the alternative splicing pattern).23,24 As illustrated in Figure 1, miRNA genes are transcribed by RNA polymerase II or III to produce a long primary transcript (pri-miRNA), which may contain single (monocistronic) or clusters (polycistronic) of hairpin loop structures. The long pri-miRNA transcript is initially processed in the nucleus by a microprocessor complex comprising nuclear RNase III enzyme Drosha, generating a 60–70-nts-long stem-loop intermediate structure called precursor miRNA (pre-miRNA). The pre-miRNA is transported by the nucleocytoplasmic shuttle protein exportin 5 to the cytoplasm, where it is further cleaved by the RNase III enzyme, Dicer, to yield a short, imperfectly matched miRNA/miRNA* duplex. To regulate target gene expression, the miRNA/miRNA* duplex is loaded onto the Argonaute (Ago) protein to form the final effector complex, RNA-induced silencing complex (RISC). In RISC, only a single strand of miRNA duplex will remain as functional miRNA to interact with the target mRNA to regulate target gene expression; the other strand (miRNA*) will be degraded.25–27
Although not common, there are some special miRNA genes called mirtrons that are encoded within the intronic region of genes, but their biogenesis and maturation bypass Drosha processing in the nucleus.28–30 After being transcribed, the mirtron is spliced out from primary transcript by spliceosome and then further goes through lariat debranching to generate pre-miRNA (Figure 1). Similar to classical miRNA, mirtron-derived pre-miRNA is also further cleaved by Dicer in the cytoplasm to generate functionally mature miRNAs.
The mature, single-stranded miRNA usually binds to the 3′ untranslated region (UTR) sequence of the target mRNA in the RISC, repressing target gene expression through either translational inhibition or mRNA destabilization. Although the perfect complementarity between miRNA and target mRNA is not necessary for miRNA-mediated gene regulation, the seed region of 2–8 nts located at the 5′ end of miRNA is critical for target gene recognition.31 One miRNA can target hundreds of genes in mammalian cells, but the inhibitory effect of a miRNA on each target gene is typically mild or moderate.32,33 Although miRNA is largely identified as negative regulator of gene expression, recent studies revealed that in certain circumstances, miRNA could also stimulate gene expression via either directly binding to the target gene or indirectly, suppressing the suppressor of gene expression.34,35 In addition to interacting with the 3′ UTR, miRNA may also bind to the 5′ UTR of target mRNA to promote or repress target gene expression.35,36
miRNAs are ubiquitously expressed and evolutionally conserved in different species. Computational genome analysis revealed that miRNAs compose about 3% of human genome, and they are computationally predicted to regulate 30%–90% of human genes.31,37–39 The crucial role of miRNAs in the regulation of almost all biological pathways in both physiological and pathological conditions has been extensively documented in numerous reports.13,14,40,41 Importantly, miRNAs have now emerged as new diagnostic markers and therapeutic targets for various diseases.42–44
miRNA regulation of immunity and autoimmunity
The expression and function of miRNAs is tightly regulated during immune cell development (lineages commitment and differentiation) and function (innate and adaptive immune responses). The proper expression and function of miRNAs are prerequisite for immune system homeostasis. Interruption or malfunctioning of miRNA regulatory machinery could cause the breakdown of immune homeostasis and self-tolerance and, consequently, lead to the development of autoimmunity. Disruption of miRNA biogenesis by depleting Dicer or Drosha in early lymphocyte progenitors impaired T- and B-cell development and function.45–48 In mice with Dicer or Drosha deficiency, specifically in the lymphocyte progenitor cells, there were reduced T-cell numbers in the central and peripheral lymphoid organs, a complete developmental block of B cell at the pro- to pre-B transition stage, impaired antibody diversity, aberrant T helper cell differentiation, and cytokine production.45–48 Several research groups reported that depletion of Dicer or Drosha specifically in T regulatory cells (Tregs) in mice led to a complete loss of immunoregulatory functions of Tregs, resulting in the development of inflammatory diseases and fatal autoimmunity.47,49–51 These studies with Dicer- or Drosha-deficient mice revealed the important contribution of general miRNA biogenesis to immune system development and function. Moreover, functional contribution of individual miRNA to immunity and autoimmunity has also been well-documented in numerous research reports and has been extensively reviewed elsewhere.13,14,17,40,52–63 Therefore, this review will not discuss the functional significance of individual miRNA in immunity and autoimmunity. Rather, we focus on the sexual dimorphism of miRNA expression and its potential contribution to sex differences in immune responses and autoimmune diseases.
Sexually dimorphic gene expression and sex differences in immune responses
There are marked differences in immune capabilities between males and females.4,64 Compared with their male counterparts, females typically mount stronger innate and adaptive immune responses to various pathogens and toxins. The better immune capability presumably contributes to the greater survival rate in females compared with males. However, the enhanced immune capability in females is a double-edged sword. Females also tend to respond vigorously to self antigens and, consequently, become more susceptible to a majority of autoimmune inflammatory diseases. The sex differences in immune responses are attributed to multiple factors including a direct genetic difference such as the X chromosome and X chromosome-linked genes, sex hormones, and sex-specific regulation of immune-related genes.2,4,10,65
The salient difference between the male and female genome is that the male possesses one X chromosome and one Y chromosome, whereas the female has two copies of the X chromosome. There are about 1,100 genes in the human X chromosome, but only about 100 genes in the human Y chromosome. Among them, only 54 genes are homologous between the Y and X chromosomes.10,66 While the initial sex determination is dependent on the Y chromosome-specific Sry gene, subsequently, X-chromosome and X-chromosome linked genes, together with sex hormones are critical contributors to the differences in physiology, behavior, gene expression/regulation, and immune functionality between males and females.67–69 Because the female possesses two copies of the X chromosome, during the early stage of female embryonic development, one X chromosome is randomly inactivated (XCI) to balance the dosage of X chromosome-linked genes between the male and female. However, about 15% of X chromosome-linked genes might escape XCI and therefore contribute to the sexual dimorphism of X chromosome-linked genes.70 In addition to XCI escape, other mechanisms such as skewed XCI (nonrandom silencing of the X chromosome) and imprinting expression of X chromosome genes may also contribute to the sex differences in the expression of X chromosome-linked genes.10
Many immune-related genes are located on the X chromosome, including toll-like receptor (TLR) family members TLR7 and TLR8, Foxp3 gene (key transcription factor regulating Tregs development and function), CD40 (cluster of differentiation 40) ligand (CD40L, a B-cell costimulatory molecule), key components of TLR and nuclear factor–kappa B signaling pathways such as Bruton’s tyrosine kinase, interleukin 1 (IL-1) receptor–associated kinase 1, IκB kinase γ, cytokine receptors such as IL-9 receptor (IL-9R), IL-2 receptor γ chain (IL-2RG), and IL-13 receptor α1 chain (IL13RA1), among others.10,69 The differential expression or mutation of these immune-related, X chromosome-linked genes could contribute to the sex differences in immune response and disease incidence. For example, varied X chromosome-linked TLR7 gene copy number, increased TLR7 expression, and TLR7 functional polymorphisms are associated with SLE susceptibility in both human and murine lupus.71,72 Demethylation and overexpression of X chromosome-linked CD40L in CD4+ T cells from female lupus patients is believed to contribute significantly to the striking female predominance of lupus.73 In contrast, the pathogenesis of immune dysregulation, polyendocrinopathy, enteropathy, and X-linked syndrome, a rare inflammatory disease that occurs exclusively in males, is attributed to the mutation of the X chromosome-linked Foxp3 gene.69
Although the sex chromosome-linked genes contribute significantly to sex dimorphism of gene expression, they can only count for a small number of sexually dimorphic genes. In fact, many sexually dimorphic genes are located on autosomes, which possess 95% of the annotated human genome. The differential expression of autosomal genes between males and females is subjected to X chromosome-mediated epigenetic controls and sex hormone regulation.68,74 The contribution of sexually dimorphic expression of immune-related protein-coding genes to sex differences in immune responses and in susceptibility to autoimmune disease is now appreciated. With recognition of the immune regulatory role of miRNAs, we propose that miRNAs are differentially expressed in immune cells between males and females, which disparately regulate immune-linked genes in males and females and subsequently contribute to the sex differences in immune capabilities and in susceptibility to autoimmune diseases.
Sexually dimorphic miRNA expression in gonadal and nongonadal (somatic) tissues
The miRNA expression has been extensively profiled in testis and ovary in various species.75–81 In a recent report, 770 known and 5 novel miRNAs were identified in normal human testis.82 Further computational bioinformatic analysis indicated that select miRNAs expressed in testis regulate many important biological pathways that are implicated in spermatogenesis, including meiosis and p53-related pathways.82 A study using ovine fetuses identified a number of differentially expressed miRNAs in testis and ovary during gestation. These were further predicted to target genes that have been previously implicated in mammalian gonadal development such as ESR1, CYP19A1, and SOX9.80 Differential expression of miRNAs in porcine testis and ovary was also reported.83 After a further survey of the expression of X chromosome-linked miRNAs in different normal porcine tissues, the authors predicted that X chromosome-linked miRNAs were expressed in a testis-preferential or testis-specific pattern.83 By using a small RNA cloning method, 141 miRNAs were identified in testis and 122 miRNAs were identified in mouse ovary.78,79 In a separate study, 49 miRNAs were exclusively detected in adult testis and 48 miRNAs were exclusively detected in ovary.77 There were also 24 miRNAs that were differentially expressed in testis and ovary with at least fivefold changes. The authors further suggested that most male-biased expressions of miRNAs were located on the X chromosome.77 Overall, these studies documented that miRNAs are differentially expressed in testis and ovary in diverse species, suggesting the important role of miRNA in driving gonadal tissue development and function.
Sex differences of miRNA expression have been observed not only in gonadal tissues but also in diverse somatic tissues such as liver,84 lung,85 and brain.74,86,87 Among the somatic tissues, the brain is the most extensively studied organ to address the contribution of sexually dimorphic miRNA expression to sex differences in brain development and function. In a recent report, by using the RNA-Seq method, the spatial–temporal expression of miRNA in a human developing brain spanning from infancy to adolescence was studied.87 The authors demonstrated that 40 miRNAs displayed significant sex-biased expression pattern in the prefrontal cortex, particularly during adolescence. Moreover, these sex-biased miRNAs in the human prefrontal cortex targeted genes that are related to Wnt signaling and transforming growth factor-beta pathways, which are implicated in neurological disorders with marked sex bias in prevalence.87 A different study has shown that 149 miRNAs were differentially expressed between male and female mouse neonatal brain.74 Of these 149 sex-biased miRNAs, 47 miRNAs were regulated by a sex chromosome-mediated mechanism, and 72 miRNAs were regulated by sex hormones.74,88 Overall, the mentioned studies documented that miRNAs are differentially expressed between males and females in both gonadal and nongonadal tissues, suggesting the important role of miRNA in the sex differences of diverse physiological system development and function.
Sex-specific regulation of miRNA expression in response to stimuli
In addition to the inherent sex differences in miRNA expression, males and females also display sex-specific miRNA expression in response to environmental stimuli and/or pathological changes. For example, a study has shown that ionizing radiation induced sex-specific miRNA expression change in the mouse brain.89 Specifically, miR-29 family members (miR-29a and miR-29c) were exclusively downregulated in female, but not male, mouse frontal cortex brain region in response to ionizing radiation. Of relevance, miR-29 was confirmed to target DNA methyltransferases (DNMT)3a and DNMT3, which provided a mechanistic understanding of ionizing radiation induction of epigenetic changes in brain.89 Further, sex-specific regulation of the X-linked inhibitor of apoptosis (XIAP) gene in response to stroke was known to contribute significantly to sex differences in cerebral ischemia.90 A recent study revealed that miR-23a, which was differentially regulated in male and female ischemic brains, targeted XIAP expression directly in the brain of mice.90 Specifically, miR-23a was decreased in the male ischemic brain but significantly increased in female ischemic brains when compared with their sham controls. The increased expression of miR-23a was associated with reduced expression of XIAP protein in female ischemic brains. This study provided direct evidence for the contribution of sex-specific regulation of miRNA to the sex differences in response to stroke.90 In addition, males and females have different pharmacologic and toxicologic responses to drugs, which may be partly caused by the sex differences in the expression of cytochrome P450 superfamily members, including Cyp2b9, a 16α-hydroxylase enzyme specifically expressed in the female mouse liver.91 The cytochrome P450 superfamily is a large group of enzymes that catalyze the metabolism of drugs and carcinogens. A current study revealed that multiple miRNAs were associated with the sexual dimorphism of Cyp2b9 expression. There was a negative association between the expression levels of these miRNAs and Cyp2b9 in mouse liver. The female-specific expression of the Cy2b9 gene may be contributed towards by the reduced expression of these miRNAs in female mouse liver when compared with the male.91 Interestingly, the sex-specific miRNA expression in response to stress is also observed in insects. The red flour beetle Tribolium castaneum exhibited both sex- and stressor-specific expression of immune genes and miRNAs. Strikingly, more miRNAs were induced in female T. castaneum in response to stress, thereby supporting the theory of female immune advantage in response to bacterial antigen challenge.92 Because miRNA plays an important role in regulation of gene expression, the sex-specific regulation of miRNA in response to stimuli may subsequently lead to sex-specific control of gene expression and cellular functions.
Sexually dimorphic miRNA expression in immune cells in the context of autoimmune disease
It is now well acknowledged that miRNAs have a critical role in immunity and autoimmunity.13,14,40,93,94 Nevertheless, there is paucity of data with regard to sex differences in miRNA expression in cells of the immune system. The potential contribution of sexually dimorphic miRNA expression to sex differences in immune response and autoimmune disease is under-investigated thus far. SLE is a prototypic autoimmune disease with marked female bias. Numerous dysregulated miRNAs have been identified in both human and murine lupus, thereby implicating their role in lupus pathogenesis.13,17,19 To determine the contribution of sexually dimorphic miRNA expression to the sex bias of autoimmune disease, we recently investigated the sex differences in the expression of lupus-associated miRNAs, including the miR-182-96-183 cluster, miR-155, miR-31, miR-148a, miR-127, and miR-379, in splenocytes of male and female NZB/WF1 mice.16,22 We found that the sex differences in the expression of aforementioned lupus-associated miRNAs were markedly evident after the onset of lupus, especially at 30 weeks of age when female NZB/WF1 mice manifested moderate to severe lupus. Before the onset of lupus, only miR-127 and miR-379 displayed slight but significant increase in female NZB/WF1 mice when compared with age-matched male counterparts.22 To our knowledge, this is the first report to show sexual differential expression of miRNA in splenocytes in the context of autoimmune disease. The significance of these lupus-associated miRNAs in lupus pathogenesis has been suggested in other recent reports. Increased miR-148a level in lupus T cells was linked to lupus pathogenesis because it targeted DNMT1, resulting in DNA hypomethylation and induction of autoimmune-associated genes in lupus T cells.95 miR-31 was shown to target Foxp3, suggesting its role in the regulation of Tregs development or function.96 Unlike in murine lupus, where miR-31 was upregulated, there was reduced miR-31 expression level in human lupus peripheral blood T cells. Further, miR-31 was shown to regulate IL-2 production in human lupus T cells by targeting ras homolog family member A (RhoA).97 miR-182 is a dominant miRNA in the regulation of T helper cell activation and proliferation. Inhibition of miR-182 in T cells suppressed ovalbumin (OVA)-induced arthritis in mice, which suggests a potential link between miR-182 dysregulation and autoimmune inflammation.61 miR-155 is one of the most extensively studied miRNAs that plays an essential role in the regulation of both innate and adaptive immune response.57,63,98,99 Dysregulated miR-155 expression or function has also been implicated in the pathogenesis of various autoimmune-related disorders.58,100–102 There is limited knowledge with regard to the immune regulatory function of miR-127 and miR-379. A recent study revealed that miR-127 inhibited lung inflammation by targeting immunoglobulin G (IgG) FcγR1 (CD64).103 Together, our recent finding of a differential expression of lupus-associated miRNA between male and female lupus mice suggests a potential contribution of sexually dimorphic miRNA expression to female predominance of lupus. However, there is a pressing need to mechanistically determine the contribution of these sexually dimorphic miRNAs to the sex differences in autoimmune disease development and progression.
Mechanism of sexual dimorphism of miRNA expression
X chromosome-linked miRNAs (X-linked miRNAs)
X chromosome and X chromosome-linked genes contribute significantly to the sex differences in immunity and autoimmunity.10,68,69 The potential contribution of X chromosome-linked miRNAs to sex differences in immune response and autoimmunity is beginning to be appreciated, although direct experimental data are still lacking.10,104,105 It is noteworthy that the X chromosome is highly enriched in miRNAs. One hundred and thirteen miRNAs, representing about 7% of total human miRNAs, are located on the X chromosome, whereas only 2 miRNAs have been identified so far on the Y chromosome.104,105 Although the function of most X chromosome-linked miRNAs remains unclear, some X chromosome-linked miRNAs have been shown to be involved in immune regulation. The most well-studied X chromosome-linked miRNA is miR-223. miR-223 is highly expressed in bone marrow myeloid cell lineages and plays a key role in the regulation of granulocytes differentiation and function.106–108 Reduced expression of miR-223 was commonly observed in various types of cancer and was correlated with the progression of these cancers.109,110 Altered expression of miR-223 is also associated with different autoimmune pathological conditions. miR-223 was increased in human RA or type 2 diabetes patients111,112 but decreased in human lupus patients.113
In addition to miR-223, several other X chromosome-linked miRNAs such as miR-221/222 cluster, miR-98, miR-106a, miR-424, and miR-18b, have also been implicated in immune regulation. miR-221/222 was increased in synovial fibroblasts from tumor necrosis factor transgenic mice and from human RA patients.114 miR-221 was shown to regulate proliferation and survival of immune cells.115,116 miR-221 expression level was inversely correlated with the expression of cell cycle inhibitor protein, cyclin-dependent kinase inhibitor 1B (p27kip1), during dendritic cell development and maturation. Silencing miR-221 resulted in p27kip1 protein accumulation and dendritic cell apoptosis.115 In a different study, miR-221 was shown to be highly upregulated in activated CD4+ T lymphocytes.116 Inhibition of miR-221 in CD4+ T lymphocytes promoted cell proliferation by suppressing proliferation suppressors such as insulin receptor substrate 2, phosphatidylinositol 3-kinase regulatory subunit alpha, I-kappa-B kinase epsilon, and FBJ murine osteosarcoma viral oncogene homolog (FOS).116 miR-98 was reduced in lipopolysaccharide (LPS)-activated macrophages, and negatively regulated LPS-induced IL-10 production in macrophages, by directly targeting IL-10 3′ UTR.117 Enhancing miR-98 function with miR-98 mimic inhibited TLR4-triggered IL-10 production and promoted cyclooxygenase 2 production, but did not affect tumor necrosis factor α and IL-6. Further, the authors showed that LPS-mediated downregulation of miR-98 contributed to endotoxin tolerance in macrophages.117 In addition, four other X chromosome-linked miRNAs, miR-106a, miR-424, miR-503, and miR-542, have been shown to be involved in monocyte differentiation.104
Importantly, a recent report revealed that 18 X chromosome-linked miRNAs including miR-188, miR-421, miR-503, Let-7f2, and miR-98 were overexpressed in CD4+ T cells from female lupus patients when compared with male lupus patients.105,118 These data add support to the view that X chromosome-linked miRNAs contribute to the sex differences in the susceptibility and severity of lupus. Interestingly, miRNA microarray analysis of murine lupus splenocytes has shown that two X chromosome-linked miRNAs, miR-222 and miR-98, were dysregulated in lpr lupus mice when compared with control mice.16 miR-222 was upregulated in lpr lupus mice, whereas miR-98 was downregulated.16 In addition, we also found that miR-223 was significantly increased in lpr lupus mice but not in NZB/WF1 mice when compared with control mice (Rujuan Dai, unpublished data 2010). However, whether miR-222, miR-98, and miR-223 are differentially expressed between male and female lupus mice is not known so far.
Sex hormone regulation of miRNAs in the immune system
In addition to the direct genetic contribution by the X and Y chromosomes, sex hormones markedly influence the sex differences in gene expression and immune function. The sexual dimorphic expression of immune-related genes after puberty in mice strongly suggested a role of sex hormones in sex-specific gene regulation.106 The profound effect of sex hormones on immune system development and function has been extensively documented.2,64,65,119–126 There are an increasing number of reports indicating that sex hormones regulate miRNA expression in various normal human tissues, including the lymphoid organ, and also in hormone responsive cancer cells, via the activation of the hormone nuclear receptors,74,127–130 which suggests the involvement of miRNA in sex hormone-mediated regulation of biological functions. It has been shown that sex hormones regulate miRNA expression at either the transcriptional or the posttranscriptional level. We next discuss recent data with regard to the regulatory effect of estrogen and androgen on miRNA expression.
Estrogen regulation of miRNA expression
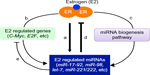
After estrogenic activation, estrogen receptor (ER)α and ERβ regulate the expression of estrogen-responsive genes through genomic and nongenomic mechanisms.121,131 Estrogen regulation of miRNA expression via activation of ER is important for fine-tuning of estrogen-mediated cellular responses.127,132–134 It has been shown that estrogen treatment induced 21 miRNAs and repressed expression of 7 miRNAs in MCF-7 breast cancer cells. These estrogen-regulated miRNAs could potentially affect the expression of hundreds of estrogen-regulated and non-estrogen-regulated protein-coding genes at the posttranscriptional level.132 Importantly, estrogen-induced miR-21, miR-98, and let-7 miRNA family members target C-Myc and E2F, two estrogen-regulated transcription factors that play a critical role in the regulation of ERα activity and estrogen responses.132 Estrogen was also reported to upregulate pri-miR-17-92 in MCF-7 cells.133 The induction of pri-miR-17-92 upon estrogenic stimulation was dependent on the direct transcriptional regulation by estrogen-induced c-Myc.133 Further, estrogen-induced miR-17-92 family members such as miR-18a, miR-19b, and miR-20b targeted and reduced ERα protein expression.133 Together, these reports demonstrated that estrogen-activated ERs, estrogen-regulated miRNAs, and estrogen-regulated mRNAs (protein-coding genes) form a negative feedback circuit to fine-tune estrogen action and estrogen-mediated cellular responses (Figure 2).
Akin to estrogen regulation of the protein-coding gene, estrogen also regulates miRNA expression through genomic and nongenomic mechanisms. Estrogen-bound ERα binds directly to the estrogen receptor binding site (ERE) on the promoter region of estrogen-responsive miRNAs, such as miR-21, miR-23a, miR-221/222, to stimulate the expression of these miRNAs.132,135 Nevertheless, not all estrogen-regulated miRNAs contain ERE sites on their promoter. As we mentioned earlier, the induction of miR-17-92 by estrogen was dependent on the estrogen-induced coactivator c-Myc, as miR-17-92 does not have ERE on its promoter.133 In addition, estrogen may also regulate miRNA expression/function at the posttranscriptional level by affecting pri-miRNA processing (Figure 2). Estrogen-bound ERα can interact with Drosha-containing microprocessor complexes and attenuate the processing of pri-miRNAs into pre-miRNAs, leading to the inhibition of a set of miRNAs in the mouse uterus.136 In addition, estrogen was shown to regulate the expression of Exportin-5, Dicer, and Ago 2, which are key components of miRNA biogenesis and functional pathways.132,137,138 Moreover, it has been shown that estrogen reduced expression levels of a set of mature miRNAs without affecting the expression of these miRNAs at precursor level, providing direct experimental evidence of estrogen regulation of miRNA at the posttranscriptional level.136
Our laboratory was the first to demonstrate that estrogen regulated miRNA expression in splenocytes from normal C57BL/6 mice, which contributed to estrogen-mediated promotion of inflammation.130 It is noteworthy that several estrogen-regulated miRNAs, such as miR-146a, miR-125b, and miR-148a, have been implicated in autoimmune disease pathogenesis. This suggests that estrogen might contribute to the sex bias of autoimmune disease and affect disease course by regulating miRNA expression and function. In a recent follow-up study, we addressed this question directly with a female-biased NZB/WF1 lupus model. We found that estrogen treatment of orchidectomized male NZB/WF1 mice promoted the expression of lupus-associated miRNAs, including miR-182-96-183 cluster, miR-379, and miR-148a, but did not affect miR-155 in splenocytes.22 To further substantiate the role of estrogen and other female hormones in the expression of lupus-associated miRNAs, it would be interesting to determine the expression of these miRNAs during pregnancy when estrogen and other female hormones are markedly increased. Together, estrogen regulation of miRNA provides a new paradigm for understanding estrogen-induced immune modulation and potential contribution to the sex differences in autoimmune disease.
Androgen regulation of miRNA expression
Androgen usually functions through the activation of androgen receptors (ARs) to regulate the expression of androgen-responsive genes. Androgen and the AR signaling pathway is critical for male reproductive organ function, such as spermatogenesis. The AR signaling pathway is also involved in the development of prostate cancer (CaP). A recent study demonstrated that testosterone regulated the expression of a group of miRNAs in mouse Sertoli cells, of which some are preferentially expressed in testis.139 One of the androgen-responsive miRNAs, miR-471, targeted Forkhead/winged-helix transcription factor (FoxD1) and the desmosomal cadherin (Dsc1) gene, which are highly expressed in Sertoli cells and germ cells and are important for androgen-mediated regulation of spermatogenesis.139 Numerous miRNAs have been implicated in the development and progression of CaP.127,140 Deep sequencing revealed that 83 miRNAs were differentially expressed in an androgen-dependent (LNCaP) and androgen-independent (LNCaP-AI) prostate cancer cell line, suggesting the involvement of miRNAs in the transition of prostate cancer cells from androgen-dependent state to androgen-independent state.141 Further bioinformatic analysis suggested that most of these differentially expressed miRNAs target genes involved in mitogen-activated protein kinase (MAPK) pathways, which are implicated in the promotion of tumor growth and in the development of hormone-refractory disease.141 In a different report, miR-125b was shown to differentially express in LNCaP and LNCap-AI cell line, as well as in benign and malignant prostate tissues.142 The authors further reported that androgen upregulated miR-125b expression via direct binding of AR to its promoter. Administration of synthetic miR-125b mimics stimulated androgen-independent growth of LNCaP cells by reducing the expression of the proapoptotic protein, Bcl-2 antagonist killer 1.142 Moreover, androgen was shown to promote the expression of miR-21, miR-19a, miR-27a, and miR-133 in CaP cancer cells through the direct binding of activated AR to the promoter of these miRNAs.143–145
In addition to regulating male reproductive organ function, androgen and AR signaling also play an important role in regulation of somatic tissues development and function. It was shown that testosterone induced the expression of 6 miRNAs in mouse liver, of which most were induced via nongenomic mechanism.146 Androgens are usually considered as immune suppressors, thereby exerting protective effect on different autoimmune diseases such as SLE, multiple sclerosis, and RA.147–149 Nevertheless, further investigation is warranted to investigate AR regulation of miRNA expression in immune cells, which may provide a plausible explanation for AR-mediated suppressive effect on SLE.
Conclusion and perspectives
The disparities between males and females in immune capabilities, in susceptibility to diverse diseases, and in responses to drugs are apparent. Nevertheless, the sex factor is usually neglected in biomedical research and in drug clinic trials, which has led to the withdrawal of 8 of 10 prescription drugs in the US market.150 Thankfully, the importance of defining gene regulation and functional mechanism in the context of different sexes and pathological conditions is increasingly appreciated by biomedical researchers and pharmaceutical drug development scientists.4,69,151 Dissecting the sex differences in autoimmune disease shall provide optimal disease management for male and female patients and will aid us in developing efficient, sex-based therapy.
Despite their short history due to recent discovery, miRNAs have provided us with a new paradigm for understanding genome regulation and function in diverse biological systems. miRNAs have emerged as key regulators of immune system development and immune responses and have been linked to the development of various immune disorders, including autoimmune diseases. Given that most autoimmune diseases display marked sex differences, it is the time to incorporate sex factors in studies pertaining to miRNA regulation and functions. Further investigations are warranted to determine the functional contribution of sexual dimorphism of miRNA expression to the sex differences in autoimmune disease. It is also equally important to delineate the underlying basis of sex differential miRNA expression.
Epigenetic mechanisms such as DNA methylation also play an important role in the regulation of both non protein-coding miRNA and protein-coding mRNA expression. Sex-specific epigenetic control of miRNAs could also contribute to the sexual dimorphism of miRNA expression. It should be noted that sex chromosomes, sex hormones, sex-specific environmental exposures, and other sex-biased factors do not work as individual components. Rather, they interact with each other to cause sex-biased action in gene networks and, consequently, lead to sex differences in physiological traits and health outcomes.151 We believe that incorporating sex factors in miRNA and autoimmune research is critical for better understanding of the sex bias of autoimmune diseases. This is a prerequisite for developing a sex-based, efficient, miRNA interfering therapeutic approach to treat autoimmune diseases with marked sex bias.
Acknowledgments
This work was supported by the American Autoimmune Related Diseases Association, National Institutes of Health T35 grant 9T350DO1187-06, the Lupus Foundation of America Novel Pilot Project (208-11-110B-033-918-1), the Alliance for Lupus Research (award 219636), and Virginia-Maryland Regional College of Veterinary Medicine Intramural Research Competition Grant 17385.
Disclosure
The authors report no conflicts of interest in this work.
References
Ansar Ahmed S, Hissong BD, Verthelyi D, Donner K, Becker K, Karpuzoglu-Sahin E. Gender and risk of autoimmune diseases: possible role of estrogenic compounds. Environ Health Perspect. 1999; 107 Suppl 5:681–686. | |
Ansar Ahmed S, Penhale WJ, Talal N. Sex hormones, immune responses, and autoimmune diseases. Mechanisms of sex hormone action. Am J Pathol. 1985;121(3):531–551. | |
Whitacre CC. Sex differences in autoimmune disease. Nat Immunol. 2001;2(9):777–780. | |
Klein SL. Immune cells have sex and so should journal articles. Endocrinology. 2012;153(6):2544–2550. | |
Shapira Y, Agmon-Levin N, Shoenfeld Y. Defining and analyzing geoepidemiology and human autoimmunity. J Autoimmun. 2010;34(3): J168-J17. | |
National Institutes of Health, Autoimmune Disease Coordinating Committee. Progress in Autoimmune Disease Research. Bethesda, MD: National Institutes of Health; 2005. Available from: http://www.niaid.nih.gov/topics/autoimmune/documents/adccfinal.pdf. Accessed March 2005. | |
Walsh SJ, Rau LM. Autoimmune diseases: a leading cause of death among young and middle-aged women in the United States. Am J Public Health. 2000;90(9):1463–1466. | |
Strickland FM, Hewagama A, Lu Q, et al. Environmental exposure, estrogen and two X chromosomes are required for disease development in an epigenetic model of lupus. J Autoimmun. 2012;38(2–3):J135–J143. | |
Invernizzi P, Pasini S, Selmi C, Gershwin ME, Podda M. Female predominance and X chromosome defects in autoimmune diseases. J Autoimmun. 2009;33(1):12–16. | |
Libert C, Dejager L, Pinheiro I. The X chromosome in immune functions: when a chromosome makes the difference. Nat Rev Immunol. 2010;10(8):594–604. | |
Lamason R, Zhao P, Rawat R, et al. Sexual dimorphism in immune response genes as a function of puberty. BMC Immunol. 2006;7:2. | |
Bernstein E, Kim SY, Carmell MA, et al. Dicer is essential for mouse development. Nat Genet. 2003;35(3):215–217. | |
Dai R, Ansar Ahmed S. MicroRNA, a new paradigm for understanding immunoregulation, inflammation, and autoimmune diseases. Transl Res. 2011;157(4):163–179. | |
Baltimore D, Boldin MP, O’Connell RM, Rao DS, Taganov KD. MicroRNAs: new regulators of immune cell development and function. Nat Immunol. 2008;9(8):839–845. | |
Zhang B, Pan X, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302(1):1–12. | |
Dai R, Zhang Y, Khan D, et al. Identification of a common lupus disease-associated microRNA expression pattern in three different murine models of lupus. PLoS One. 2010;5(12):e14302. | |
Shen N, Liang D, Tang Y, de Vries N, Tak PP. MicroRNAs – novel regulators of systemic lupus erythematosus pathogenesis. Nat Rev Rheumatol. 2012;8(12):701–709. | |
Alevizos I, Illei GG. MicroRNAs in Sjögren’s syndrome as a prototypic autoimmune disease. Autoimmun Rev. 2010;9(9):618–621. | |
Ceribelli A, Yao B, Dominguez-Gutierrez PR, Nahid MA, Satoh M, Chan EK. MicroRNAs in systemic rheumatic diseases. Arthritis Res Ther. 2011;13(4):229. | |
Du C, Liu C, Kang J, et al. MicroRNA miR-326 regulates TH-17 differentiation and is associated with the pathogenesis of multiple sclerosis. Nat Immunol. 2009;10(12):1252–1259. | |
Ode F Jr, Moore CS, Kennedy TE, Antel JP, Bar-Or A, Dhaunchak AS. MicroRNA dysregulation in multiple sclerosis. Front Genet. 2013;3:311. | |
Dai R, McReynolds S, Leroith T, Heid B, Liang Z, Ansar Ahmed S. Sex differences in the expression of lupus-associated miRNAs in splenocytes from lupus-prone NZB/WF1 mice. Biol Sex Differ. 2013;4(1):19. | |
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 2004;14(10A):1902–1910. | |
Kim VN, Nam JW. Genomics of microRNA. Trends Genet. 2006;22(3): 165–173. | |
Gregory RI, Yan KP, Amuthan G, et al. The Microprocessor complex mediates the genesis of microRNAs. Nature. 2004;432(7014): 235–240. | |
Lee Y, Ahn C, Han J, et al. The nuclear RNase III Drosha initiates microRNA processing. Nature. 2003;425(6956):415–419. | |
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–297. | |
Ruby JG, Jan CH, Bartel DP. Intronic microRNA precursors that bypass Drosha processing. Nature. 2007;448(7149):83–86. | |
Okamura K, Hagen JW, Duan H, Tyler DM, Lai EC. The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell. 2007;130(1):89–100. | |
Berezikov E, Chung WJ, Willis J, Cuppen E, Lai EC. Mammalian mirtron genes. Mol Cell. 2007;28(2):328–336. | |
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120(1):15–20. | |
Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP. The impact of microRNAs on protein output. Nature. 2008;455(7209):64–71. | |
Selbach M, Schwanhäusser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N. Widespread changes in protein synthesis induced by microRNAs. Nature. 2008;455(7209):58–63. | |
Vasudevan S, Tong Y, Steitz JA. Switching from repression to activation: microRNAs can up-regulate translation. Science. 2007; 318(5858):1931–1934. | |
Da Sacco L, Masotti A. Recent Insights and Novel Bioinformatics Tools to Understand the Role of MicroRNAs Binding to 5′ Untranslated Region. Int J Mol Sci. 2012;14(1):480–495. | |
Lee I, Ajay SS, Yook JI, et al. New class of microRNA targets containing simultaneous 5′-UTR and 3′-UTR interaction sites. Genome Res. 2009;19(7):1175–1183. | |
Engels BM, Hutvagner G. Principles and effects of microRNA-mediated post-transcriptional gene regulation. Oncogene. 2006;25(46): 6163–6169. | |
Griffiths-Jones S. The microRNA Registry. Nucleic Acids Res. 2004; 32(Database issue):D109–D111. | |
Miranda KC, Huynh T, Tay Y, et al. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell. 2006;126(6):1203–1217. | |
O’Connell RM, Rao DS, Chaudhuri AA, Baltimore D. Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol. 2010;10(2):111–122. | |
Krützfeldt J, Stoffel M. MicroRNAs: a new class of regulatory genes affecting metabolism. Cell Metab. 2006;4(1):9–12. | |
Zhang B, Farwell MA. microRNAs: a new emerging class of players for disease diagnostics and gene therapy. J Cell Mol Med. 2008;12(1): 3–21. | |
Kosaka N, Iguchi H, Ochiya T. Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010;101(10):2087–2092. | |
van Rooij E, Purcell AL, Levin AA. Developing microRNA therapeutics. Circ Res. 2012;110(3):496–507. | |
Cobb BS, Nesterova TB, Thompson E, et al. T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. J Exp Med. 2005;201(9):1367–1373. | |
Koralov SB, Muljo SA, Galler GR, et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell. 2008;132(5):860–874. | |
Chong MM, Rasmussen JP, Rudensky AY, Littman DR. The RNAseIII enzyme Drosha is critical in T cells for preventing lethal inflammatory disease. J Exp Med. 2008;205(9):2005–2017. | |
Muljo SA, Ansel KM, Kanellopoulou C, Livingston DM, Rao A, Rajewsky K. Aberrant T cell differentiation in the absence of Dicer. J Exp Med. 2005;202(2):261–269. | |
Zhou X, Jeker LT, Fife BT, et al. Selective miRNA disruption in T reg cells leads to uncontrolled autoimmunity. J Exp Med. 2008;205(9): 1983–1991. | |
Liston A, Lu LF, O’Carroll D, Tarakhovsky A, Rudensky AY. Dicer-dependent microRNA pathway safeguards regulatory T cell function. J Exp Med. 2008;205(9):1993–2004. | |
Cobb BS, Hertweck A, Smith J, et al. A role for Dicer in immune regulation. J Exp Med. 2006;203(11):2519–2527. | |
Taganov KD, Boldin MP, Baltimore D. MicroRNAs and immunity: tiny players in a big field. Immunity. 2007;26(2):133–137. | |
Lodish HF, Zhou B, Liu G, Chen CZ. Micromanagement of the immune system by microRNAs. Nat Rev Immunol. 2008;8(2):120–130. | |
Gantier MP, Sadler AJ, Williams BR. Fine-tuning of the innate immune response by microRNAs. Immunol Cell Biol. 2007;85(6):458–462. | |
Sonkoly E, Ståhle M, Pivarcsi A. MicroRNAs and immunity: novel players in the regulation of normal immune function and inflammation. Semin Cancer Biol. 2008;18(2):131–140. | |
Lu LF, Boldin MP, Chaudhry A, et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell. 2010;142(6): 914–829. | |
Rodriguez A, Vigorito E, Clare S, et al. Requirement of bic/microRNA-155 for normal immune function. Science. 2007;316(5824):608–611. | |
O’Connell RM, Kahn D, Gibson WS, et al. MicroRNA-155 promotes autoimmune inflammation by enhancing inflammatory T cell development. Immunity. 2010;33(4):607–619. | |
Li QJ, Chau J, Ebert PJ, et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell. 2007;129(1):147–161. | |
Xiao C, Srinivasan L, Calado DP, et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat Immunol. 2008;9(4):405–414. | |
Stittrich AB, Haftmann C, Sgouroudis E, et al. The microRNA miR-182 is induced by IL-2 and promotes clonal expansion of activated helper T lymphocytes. Nat Immunol. 2010;11(11):1057–1062. | |
Taganov KD, Boldin MP, Chang KJ, Baltimore D. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci U S A. 2006;103(33):12481–12486. | |
Thai TH, Calado DP, Casola S, et al. Regulation of the germinal center response by microRNA-155. Science. 2007;316(5824):604–608. | |
Marriott I, Huet-Hudson YM. Sexual dimorphism in innate immune responses to infectious organisms. Immunol Res. 2006;34(3):177–192. | |
Ansar Ahmed S, Karpuzoglu E, Khan D. Effects of sex steroids on innate and adaptive immunity. In: Klein SL, Roberts CW, editors. Sex Hormones and Immunity to Infection. Berlin: Springer-Verlag; 2010:19–52. | |
Ross MT, Grafham DV, Coffey AJ, et al. The DNA sequence of the human X chromosome. Nature. 2005;434(7031):325–337. | |
Wilhelm D, Koopman P. The makings of maleness: towards an integrated view of male sexual development. Nat Rev Genet. 2006;7(8): 620–631. | |
Wijchers PJ, Festenstein RJ. Epigenetic regulation of autosomal gene expression by sex chromosomes. Trends Genet. 2011;27(4):132–140. | |
Fish EN. The X-files in immunity: sex-based differences predispose immune responses. Nat Rev Immunol. 2008;8(9):737–744. | |
Carrel L, Willard HF. X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature. 2005;434(7031): 400–404. | |
García-Ortiz H, Velázquez-Cruz R, Espinosa-Rosales F, Jiménez-Morales S, Baca V, Orozco L. Association of TLR7 copy number variation with susceptibility to childhood-onset systemic lupus erythematosus in Mexican population. Ann Rheum Dis. 2010;69(10):1861–1865. | |
Shen N, Fu Q, Deng Y, et al. Sex-specific association of X-linked Toll-like receptor 7 (TLR7) with male systemic lupus erythematosus. Proc Natl Acad Sci U S A. 2010;107(36):15838–15843. | |
Lu Q, Wu A, Tesmer L, Ray D, Yousif N, Richardson B. Demethylation of CD40LG on the inactive X in T cells from women with lupus. J Immunol. 2007;179(9):6352–6358. | |
Morgan CP, Bale TL. Sex differences in microRNA regulation of gene expression: no smoke, just miRs. Biol Sex Differ. 2012;3(1):22. | |
Hossain MM, Ghanem N, Hoelker M, et al. Identification and characterization of miRNAs expressed in the bovine ovary. BMC Genomics. 2009;10:443. | |
Juanchich A, Le Cam A, Montfort J, Guiguen Y, Bobe J. Identification of differentially expressed miRNAs and their potential targets during fish ovarian development. Biol Reprod. 2013;88(5):128. | |
Mishima T, Takizawa T, Luo SS, et al. MicroRNA (miRNA) cloning analysis reveals sex differences in miRNA expression profiles between adult mouse testis and ovary. Reproduction. 2008;136(6):811–822. | |
Ro S, Park C, Sanders KM, McCarrey JR, Yan W. Cloning and expression profiling of testis-expressed microRNAs. Dev Biol. 2007;311(2): 592–602. | |
Ro S, Song R, Park C, Zheng H, Sanders KM, Yan W. Cloning and expression profiling of small RNAs expressed in the mouse ovary. RNA. 2007;13(12):2366–2380. | |
Torley KJ, da Silveira JC, Smith P, et al. Expression of miRNAs in ovine fetal gonads: potential role in gonadal differentiation. Reprod Biol Endocrinol. 2011;9:2. | |
Bannister SC, Tizard ML, Doran TJ, Sinclair AH, Smith CA. Sexually dimorphic microRNA expression during chicken embryonic gonadal development. Biol Reprod. 2009;81(1):165–176. | |
Yang Q, Hua J, Wang L, et al. MicroRNA and piRNA profiles in normal human testis detected by next generation sequencing. PLoS One. 2013;8(6):e66809. | |
Li M, Liu Y, Wang T, et al. Repertoire of porcine microRNAs in adult ovary and testis by deep sequencing. Int J Biol Sci. 2011;7(7):1045–1055. | |
Cheung L, Gustavsson C, Norstedt G, Tollet-Egnell P. Sex-different and growth hormone-regulated expression of microRNA in rat liver. BMC Mol Biol. 2009;10:13. | |
Mujahid S, Logvinenko T, Volpe MV, Nielsen HC. miRNA regulated pathways in late stage murine lung development. BMC Dev Biol. 2013;13:13. | |
Pak TR, Rao YS, Prins SA, Mott NN. An emerging role for microRNAs in sexually dimorphic neurobiological systems. Pflugers Arch. 2013; 465(5):655–667. | |
Ziats MN, Rennert OM. Identification of differentially expressed microRNAs across the developing human brain. Mol Psychiatry. Epub August 6, 2013. | |
Morgan CP, Bale TL. Early prenatal stress epigenetically programs dysmasculinization in second-generation offspring via the paternal lineage. J Neurosci. 2011;31(33):11748–11755. | |
Koturbash I, Zemp F, Kolb B, Kovalchuk O. Sex-specific radiation-induced microRNAome responses in the hippocampus, cerebellum and frontal cortex in a mouse model. Mutat Res. 2011;722(2):114–118. | |
Siegel C, Li J, Liu F, Benashski SE, McCullough LD. miR-23a regulation of X-linked inhibitor of apoptosis (XIAP) contributes to sex differences in the response to cerebral ischemia. Proc Natl Acad Sci U S A. 2011;108(28):11662–11667. | |
Xie X, Miao L, Yao J, et al. Role of multiple microRNAs in the sexually dimorphic expression of Cyp2b9 in mouse liver. Drug Metab Dispos. 2013;41(10):1732–1737. | |
Freitak D, Knorr E, Vogel H, Vilcinskas A. Gender- and stressor-specific microRNA expression in Tribolium castaneum. Biol Lett. 2012;8(5):860–863. | |
Ceribelli A, Satoh M, Chan EK. MicroRNAs and autoimmunity. Curr Opin Immunol. 2012;24(6):686–691. | |
Pauley KM, Cha S, Chan EK. MicroRNA in autoimmunity and autoimmune diseases. J Autoimmun. 2009;32(3–4):189–194. | |
Pan W, Zhu S, Yuan M, et al. MicroRNA-21 and microRNA-148a contribute to DNA hypomethylation in lupus CD4+ T cells by directly and indirectly targeting DNA methyltransferase 1. J Immunol. 2010;184(12):6773–6781. | |
Rouas R, Fayyad-Kazan H, El Zein N, et al. Human natural Treg microRNA signature: role of microRNA-31 and microRNA-21 in FOXP3 expression. Eur J Immunol. 2009;39(6):1608–1618. | |
Fan W, Liang D, Tang Y, et al. Identification of microRNA-31 as a novel regulator contributing to impaired interleukin-2 production in T cells from patients with systemic lupus erythematosus. Arthritis Rheum. 2012;64(11):3715–3725. | |
Vigorito E, Perks KL, Abreu-Goodger C, et al. microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity. 2007;27(6):847–859. | |
O’Connell RM, Taganov KD, Boldin MP, Cheng G, Baltimore D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc Natl Acad Sci U S A. 2007;104(5):1604–1609. | |
Kurowska-Stolarska M, Alivernini S, Ballantine LE, et al. MicroRNA-155 as a proinflammatory regulator in clinical and experimental arthritis. Proc Natl Acad Sci U S A. 2011;108(27):11193–11198. | |
Leng RX, Pan HF, Qin WZ, Chen GM, Ye DQ. Role of microRNA-155 in autoimmunity. Cytokine Growth Factor Rev. 2011;22(3):141–147. | |
Divekar AA, Dubey S, Gangalum PR, Singh RR. Dicer insufficiency and microRNA-155 overexpression in lupus regulatory T cells: an apparent paradox in the setting of an inflammatory milieu. J Immunol. 2011;186(2):924–930. | |
Xie T, Liang J, Liu N, et al. MicroRNA-127 inhibits lung inflammation by targeting IgG Fcγ receptor I. J Immunol. 2012;188(5):2437–2444. | |
Pinheiro I, Dejager L, Libert C. X-chromosome-located microRNAs in immunity: might they explain male/female differences? The X chromosome-genomic context may affect X-located miRNAs and downstream signaling, thereby contributing to the enhanced immune response of females. Bioessays. 2011;33(11):791–802. | |
Hewagama A. Role of X-Chromosome encoded miRNAs in autoimmunity: suppressing the suppressor and female predisposition. Rheumatol Curr Res. 2013;3(1):118. | |
Fazi F, Rosa A, Fatica A, et al. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell. 2005;123(5):819–831. | |
Johnnidis JB, Harris MH, Wheeler RT, et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature. 2008;451(7182):1125–1129. | |
Pulikkan JA, Dengler V, Peramangalam PS, et al. Cell-cycle regulator E2F1 and microRNA-223 comprise an autoregulatory negative feedback loop in acute myeloid leukemia. Blood. 2010;115(9): 1768–1778. | |
Stamatopoulos B, Meuleman N, Haibe-Kains B, et al. microRNA-29c and microRNA-223 down-regulation has in vivo significance in chronic lymphocytic leukemia and improves disease risk stratification. Blood. 2009;113(21):5237–5245. | |
Wong QW, Lung RW, Law PT, et al. MicroRNA-223 is commonly repressed in hepatocellular carcinoma and potentiates expression of Stathmin1. Gastroenterology. 2008;135(1):257–269. | |
Fulci V, Scappucci G, Sebastiani GD, et al. miR-223 is overexpressed in T-lymphocytes of patients affected by rheumatoid arthritis. Hum Immunol. 2010;71(2):206–211. | |
Lu H, Buchan RJ, Cook SA. MicroRNA-223 regulates Glut4 expression and cardiomyocyte glucose metabolism. Cardiovasc Res. 2010;86(3):410–420. | |
Wang H, Peng W, Ouyang X, Li W, Dai Y. Circulating microRNAs as candidate biomarkers in patients with systemic lupus erythematosus. Transl Res. 2012;160(3):198–206. | |
Pandis I, Ospelt C, Karagianni N, et al. Identification of microRNA-221/222 and microRNA-323-3p association with rheumatoid arthritis via predictions using the human tumour necrosis factor transgenic mouse model. Ann Rheum Dis. 2012;71(10):1716–1723. | |
Lu C, Huang X, Zhang X, et al. miR-221 and miR-155 regulate human dendritic cell development, apoptosis, and IL-12 production through targeting of p27kip1, KPC1, and SOCS-1. Blood. 2011;117(16): 4293–4303. | |
Grigoryev YA, Kurian SM, Hart T, et al. MicroRNA regulation of molecular networks mapped by global microRNA, mRNA, and protein expression in activated T lymphocytes. J Immunol. 2011;187(5): 2233–2243. | |
Liu Y, Chen Q, Song Y, et al. MicroRNA-98 negatively regulates IL-10 production and endotoxin tolerance in macrophages after LPS stimulation. FEBS Lett. 2011;585(12):1963–1968. | |
Hewagama A, Gorelik G, Patel D, et al. Overexpression of X-linked genes in T cells from women with lupus. J Autoimmun. 2013;41:60–71. | |
Verthelyi D. Sex hormones as immunomodulators in health and disease. Int Immunopharmacol. 2001;1(6):983–993. | |
Bouman A, Heineman MJ, Faas MM. Sex hormones and the immune response in humans. Hum Reprod Update. 2005;11(4):411–423. | |
Khan D, Cowan C, Ansar Ahmed S. Estrogen and signaling in the cells of immune system. Adv Neuroimmune Biol. 2012;3(1):73–93. | |
Straub RH. The complex role of estrogens in inflammation. Endocr Rev. 2007;28(5):521–574. | |
Dai R, Phillips RA, Karpuzoglu E, Khan D, Ansar Ahmed S. Estrogen regulates transcription factors STAT-1 and NF-kappaB to promote inducible nitric oxide synthase and inflammatory responses. J Immunol. 2009;183(11):6998–7005. | |
Calippe B, Douin-Echinard V, Laffargue M, et al. Chronic estradiol administration in vivo promotes the proinflammatory response of macrophages to TLR4 activation: involvement of the phosphatidylinositol 3-kinase pathway. J Immunol. 2008;180(12): 7980–7988. | |
Karpuzoglu E, Fenaux JB, Phillips RA, Lengi AJ, Elvinger F, Ansar Ahmed S. Estrogen up-regulates inducible nitric oxide synthase, nitric oxide, and cyclooxygenase-2 in splenocytes activated with T cell stimulants: role of interferon-gamma. Endocrinology. 2006;147(2): 662–671. | |
Ghisletti S, Meda C, Maggi A, Vegeto E. 17beta-estradiol inhibits inflammatory gene expression by controlling NF-kappaB intracellular localization. Mol Cell Biol. 2005;25(8):2957–2968. | |
Yang Z, Wang L. Regulation of microRNA expression and function by nuclear receptor signaling. Cell Biosci. 2011;1(1):31. | |
Paris O, Ferraro L, Grober OM, et al. Direct regulation of microRNA biogenesis and expression by estrogen receptor beta in hormone-responsive breast cancer. Oncogene. 2012;31(38):4196–4206. | |
Klinge CM. Estrogen Regulation of MicroRNA Expression. Curr Genomics. 2009;10(3):169–183. | |
Dai R, Phillips RA, Zhang Y, Khan D, Crasta O, Ansar Ahmed S. Suppression of LPS-induced Interferon-gamma and nitric oxide in splenic lymphocytes by select estrogen-regulated microRNAs: a novel mechanism of immune modulation. Blood. 2008;112(12):4591–4597. | |
Björnström L, Sjöberg M. Mechanisms of estrogen receptor signaling: convergence of genomic and nongenomic actions on target genes. Mol Endocrinol. 2005;19(4):833–842. | |
Bhat-Nakshatri P, Wang G, Collins NR, et al. Estradiol-regulated microRNAs control estradiol response in breast cancer cells. Nucleic Acids Res. 2009;37(14):4850–4861. | |
Castellano L, Giamas G, Jacob J, et al. The estrogen receptor-alpha-induced microRNA signature regulates itself and its transcriptional response. Proc Natl Acad Sci U S A. 2009;106(37):15732–15737. | |
Klinge CM. miRNAs and estrogen action. Trends Endocrinol Metab. 2012;23(5):223–233. | |
Di Leva G, Gasparini P, Piovan C, et al. MicroRNA cluster 221–222 and estrogen receptor alpha interactions in breast cancer. J Natl Cancer Inst. 2010;102(10):706–721. | |
Yamagata K, Fujiyama S, Ito S, et al. Maturation of microRNA is hormonally regulated by a nuclear receptor. Mol Cell. 2009;36(2): 340–347. | |
Nothnick WB, Healy C, Hong X. Steroidal regulation of uterine miRNAs is associated with modulation of the miRNA biogenesis components Exportin-5 and Dicer1. Endocrine. 2010;37(2):265–273. | |
Cheng C, Fu X, Alves P, Gerstein M. mRNA expression profiles show differential regulatory effects of microRNAs between estrogen receptor-positive and estrogen receptor-negative breast cancer. Genome Biol. 2009;10(9):R90. | |
Panneerdoss S, Chang YF, Buddavarapu KC, et al. Androgen-responsive microRNAs in mouse Sertoli cells. PLoS One. 2012;7(7): e41146. | |
Coppola V, De Maria R, Bonci D. MicroRNAs and prostate cancer. Endocr Relat Cancer. 2010;17(1):F1–F17. | |
Xu G, Wu J, Zhou L, et al. Characterization of the small RNA transcriptomes of androgen dependent and independent prostate cancer cell line by deep sequencing. PLoS One. 2010;5(11):e15519. | |
Shi XB, Xue L, Yang J, et al. An androgen-regulated miRNA suppresses Bak1 expression and induces androgen-independent growth of prostate cancer cells. Proc Natl Acad Sci U S A. 2007;104(50): 19983–19988. | |
Ribas J, Ni X, Haffner M, et al. miR-21: an androgen receptor-regulated microRNA that promotes hormone-dependent and hormone-independent prostate cancer growth. Cancer Res. 2009;69(18): 7165–7169. | |
Mo W, Zhang J, Li X, et al. Identification of novel AR-targeted microRNAs mediating androgen signalling through critical pathways to regulate cell viability in prostate cancer. PLoS One. 2013;8(2): e56592. | |
Fletcher CE, Dart DA, Sita-Lumsden A, Cheng H, Rennie PS, Bevan CL. Androgen-regulated processing of the oncomir miR-27a, which targets Prohibitin in prostate cancer. Hum Mol Genet. 2012;21(14):3112–3127. | |
Delić D, Grosser C, Dkhil M, Al-Quraishy S, Wunderlich F. Testosterone-induced upregulation of miRNAs in the female mouse liver. Steroids. 2010;75(12):998–1004. | |
Palaszynski KM, Loo KK, Ashouri JF, Liu HB, Voskuhl RR. Androgens are protective in experimental autoimmune encephalomyelitis: implications for multiple sclerosis. J Neuroimmunol. 2004;146(1–2):144–152. | |
Roubinian JR, Talal N, Greenspan JS, Goodman JR, Siiteri PK. Delayed androgen treatment prolongs survival in murine lupus. J Clin Invest. 1979;63(5):902–911. | |
Röntzsch A, Thoss K, Petrow PK, Henzgen S, Bräuer R. Amelioration of murine antigen-induced arthritis by dehydroepiandrosterone (DHEA). Inflamm Res. 2004;53(5):189–198. | |
Simon V. Wanted: women in clinical trials. Science. 2005;308(5728): 1517. | |
Arnold AP, Lusis AJ. Understanding the sexome: measuring and reporting sex differences in gene systems. Endocrinology. 2012;153(6):2551–2555. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.