Back to Journals » Infection and Drug Resistance » Volume 12
Rapid simultaneous detection of blaoxa-23, Ade-B, int-1, and ISCR-1 in multidrug-resistant Acinetobacter baumannii using single-tube multiplex PCR and high resolution melting assay
Authors Sun H, Xiao G, Zhang J, Pan Z, Chen Y, Xiong F
Received 1 March 2019
Accepted for publication 13 May 2019
Published 7 June 2019 Volume 2019:12 Pages 1573—1581
DOI https://doi.org/10.2147/IDR.S207225
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Hengbiao Sun,1,* Gang Xiao,1,* Jing Zhang,1 Zuhan Pan,1 Youming Chen,1 Fu Xiong,2
1Department of Clinical Laboratory, The Third Affiliated Hospital of Southern Medical University, Guangzhou 510630, People’s Republic of China; 2Department of Medical Genetics, School of Basic Medical Sciences, Southern Medical University, Guangzhou 510515, People’s Republic of China
*These authors contributed equally to this work
Objective: The aim of this study was to develop a multiplex PCR system for the rapid and simultaneous detection of blaoxa-23, Ade-B, int-1, and ISCR-1 genes in multidrug-resistant Acinetobacter baumannii (MDRAB) using high resolution melting (HRM) assay.
Methods: Four pairs of primers were designed, and PCR amplification products were sequenced and compared with NCBI GeneBank sequences to ensure primer specificity. Multiplex PCR was performed using a dedicated HRM reagent, and melting curves and temperatures were able to distinguish the four genes. This method was subsequently used to detect these genes in 79 MDRAB isolates from the Third Affiliated Hospital of Southern Medical University in southern China.
Results: Using the HRM assay, 73 out of 79 isolates were found to carry both blaoxa-23 and Ade-B, one isolate carried int-1, two isolates carried both int-1 and ISCR-1, and three isolates carried Ade-B, int-1, and ISCR-1. No isolates carried all four genes.
Conclusion: Compared with traditional resistance gene detection methods–PCR and agarose gel electrophoresis-based resistance gene detection methods–the multiplex PCR and HRM assay method was simple, rapid, highly efficient, and cost-effective. Our results showed that blaoxa-23 and Ade-B were the main resistance genotypes in MDRAB.
Keywords: blaoxa-23, Ade-B, int-1, ISCR-1, multiplex PCR, high resolution melting
Introduction
Acinetobacter baumannii is an aerobic, non-fermenting, Gram-negative opportunistic pathogen that causes pneumonia, secondary meningitis, and a variety of other infections following incisions or wounds to the skin and soft tissues.1–3 The ability of A. baumannii to survive under a wide range of environmental conditions for a long time ensures this species is a prevalent nosocomial pathogen worldwide, especially in the intensive care, neurosurgery, and burns departments of hospitals.4–6 Owing to the ability of A. baumannii to develop and/or acquire a variety of resistance mechanisms, currently susceptible bacteria may soon evolve into multidrug-resistant A. baumannii (MDRAB) that are resistant to a wide variety of antibacterial agents including β-lactams, cephems, carbapenems, aminoglycosides, tetracyclines, fluoroquinolones, and folate pathway inhibitors.7 MDRAB infections can ultimately lead to increased morbidity and mortality.
Genomics analysis revealed that MDRAB contain several large resistance islands that contain 24 genes associated with antibiotic resistance and 16 genes associated with resistance to heavy metal salts or quaternary ammonium compounds commonly used in disinfectants. These resistance genes were reportedly acquired from Pseudomonas aeruginosa, Salmonellaspp., Escherichia coli, and other Gram-negative bacteria.8 Multiple mechanisms are involved in resistance in A. baumannii, including the production of inactivating enzymes, deletion, or mutation of outer membrane proteins, expression of multidrug efflux systems, changing or protecting the target sites of antibacterial agents, and transferring resistance genes through mobile genetic elements.9 The blaoxa-23 gene is the most common carbapenem resistance gene in MDRAB clinical isolates from most regions of the world. This gene encodes a carbapenem-hydrolyzing class D oxacillinase, and production of this enzyme is not inhibited by clavulanic acid.10–12 Multidrug efflux systems such as Ade-ABC have a tripartite structure comprising an inner membrane component (AdeA), an antibacterial agent transport component (AdeB), and an outer membrane component (AdeC). Resistance to β-lactams, erythromycin, aminoglycosides, tetracyclines, and chloramphenicol is dependent on overexpression of Ade-ABC.13–15 The class 1 integron int-1 is a three-part bacterial mobile genetic element found in A. baumannii. The 5ʹ conserved region of int-1 encodes a site-specific integrase, the central variable region encodes several gene cassettes, and there is also a 3ʹ conserved region.16 The class 1 integron can capture resistance genes and integrate the associated gene cassettes into the A. baumannii genome to generate MDRAB.17 Insertion sequence common region 1(ISCR-1) is a special mobile genetic element that is similar to IS91-like and shares a sustaining transpose rolling-circle mechanism.18 Like the class 1 integron, ISCR-1 has the ability to capture resistance genes from other bacteria.19 The ability to detect OXA-23, Ade-ABC, class 1 integron, and ISCR-1 resistance genes is crucial for identifying and treating MDRAB.
Recent improvements in the molecular methods used to detect resistance genes have decreased turnaround time and increased the specificity and sensitivity of testing procedures.20,21 There are existing molecular assays that can detect blaoxa-23, Ade-B, int-1, and ISCR-1, but these are time-consuming and unable to detect all four resistance genes in a single assay.19,22 Although detection of these genes may not revolutionize clinical treatment, their identification will assist epidemiological investigations and help to control hospital-acquired infections. Our objective was to develop a single-tube multiplex PCR assay that can detect these four genes simultaneously using a dedicated high resolution melting (HRM) assay reagent. The established method was used to characterize the four resistance genes in MDRAB isolates collected in 2014 from the Third Affiliated Hospital of Southern Medical University in southern China.
Materials and methods
Definition of MDRAB
MDRAB is defined as non-susceptible to at least one agent in ≥3 antimicrobial categories of aminoglycosides, antipseudomonal carbapenems, antipseudomonal fluoroquinolones, antipseudomonal penicillins + β-lactamase inhibitors, extended-spectrum cephalosporins, folate pathway inhibitors, penicillins + β-lactamase inhibitors, and tetracyclines.23
Bacterial strains and antibiotic susceptibility testing
After exclusion of strains isolated repeatedly from one patient, a total of 87 MDRAB strains were isolated from various clinical specimens at the Third Affiliated Hospital of Southern Medical University, a comprehensive teaching hospital in southern China. All isolates were identified and tested for drug sensitivity using Microscan Walkaway 40 Plus (Beckman Coulter, West Sacramento, USA), and the results were interpreted according to the Clinical and Laboratory Standards Institute 2017 guidelines.
DNA isolation
Total DNA was extracted from well-established colonies using the Rapid Ezup Column Genomic DNA Extraction Kit (Sangon Biotech, Shanghai, China) according to the recommendations of the manufacturer. DNA was eluted with 100 μL CE buffer and stored at –80°C.
Primer design
With reference to sequences from the NCBI database (
 | Table 1 Primers used in this study |
Preliminary screening of blaoxa-23, Ade-B, int-1, and ISCR-1
Four sets of preliminary primers were used to characterize the four genes in the 8 of 87 MDRAB isolates. Each 20 μL PCR contained 10 μL of 2× PCR master mix (Generay Biotech, Shanghai, China), 0.6 μL of 10 μM forward and reverse primers, 1.4 μL of template DNA, and 7.4 μL of double-distilled water. Amplification was performed using the following conditions: initial denaturation at 95°C for 3 mins, followed by 40 cycles of denaturation (95°C, 30 s), annealing (56°C for blaoxa-23, 55°C for Ade-B, 59°C for int-1, 56°C for blaoxa-23, 45 s), extension (72°C, 1 min), and a final extension at 72°C for 5 mins. To validate the PCR products they were separated by agarose gel electrophoresis, stained with GoldView TM(nontoxic) and photographed on a UV transilluminator.
DNA sequencing and actual melting temperature (tm) determination
To ensure the specificity of the primers used for the multiplex PCR with HRM assay, the four sets of the primers were used to detect the four genes that were amplified using the preliminary primers. Reactions were similar to the preliminary screening conditions but contained 10 μL of 2× HRM FAST PCR master mix (Kapa Biosystems, Cape Town, South Africa), 0.6 μL of 10 μM forward and reverse primers, 1.4 μL of template DNA, 2 μL of 25 mM Mgcl2, and 5.4 μL of double-distilled water. Amplifications were performed as described above but with an annealing temperature of 56°C for all four sets of primers. PCR products were separated by agarose gel electrophoresis and sequenced, and data were analyzed using the Basic Local Alignment Search Tool (BLAST,
In addition to determining the actual melting temperature (Tm) using HRM analysis with the LightScanner System (Idaho Technology Inc., Cape Town, South Africa), melting curves were performed by measuring the decrease in fluorescent signal with increasing temperature by ramping at 0.02°C/s with 25 acquisitions/°C. Tm is defined as the temperature at which 50% of the amplified product dissociated into single-stranded DNA, and the actual Tm of the four genes was used as a reference to determine which gene was present in amplification reactions from the isolates.
Development of the multiplex PCR and HRM assay
Template DNA used in multiplex PCR was a mixture of isolates, and these were tested for the presence of blaoxa-23, Ade-B, int-1, and ISCR-1. Multiplex PCR was performed in single-tube 20 μL reactions containing 10 μL of 2× HRM FAST PCR master mix, 0.4 μL of 10 μM forward and reverse primers (total of 3.2 μL), 1.4 μL of compound template DNA, 2 μL of 25 mM Mgcl2, and 3.4 μL of double-distilled water. To ensure the sensitivity and specificity of multiplex PCR, annealing was tested at 54°C, 56°C, 58°C, and 60°C. All other reaction conditions were as described above. Melting curves from all multiplex PCR amplification products were generated using a LightScanner System, and products were separated by agarose gel electrophoresis following HRM analysis.
Screening of clinical isolates
Seventy-nine of the remaining 87 MDRAB strains were screened for the presence of the four genes using the multiplex PCR and HRM assay described above. We also tested for the presence of the four genes using traditional PCR and agarose gel electrophoresis-based methods (one PCR reaction detecting one gene) to verify the accuracy of the multiplex PCR and HRM assay.
Results
Preliminary screening for blaoxa-23, Ade-B, int-1, and ISCR-1
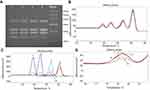
Preliminary screening of eight MDRAB isolates for the four genes was performed to provide positive controls for developing multiplex PCR of unknown samples. Agarose gel electrophoresis demonstrated the presence of Ade-B, int-1, and ISCR-1 in strain 1, and blaoxa-23 and Ade-B in all of the other seven strains (Figure 1).
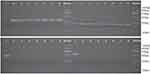 | Figure 1 Agarose gel electrophoresis of the PCR amplification products of the four genes from eight MDRAB isolates using preliminary primers. |
DNA sequencing and differentiation of the four genes using HRM curve analysis
Strains 1 and 2 were used as positive controls for establishing the multiplex PCR with HRM assay. Four sets of the primers were used to detect the four genes in these strains, and agarose gel electrophoresis resulted in bands of 139, 390, 234, and 187 bp as expected (Figure 2). The BLAST results confirmed that the sequences of the PCR amplification products were the same as those previously published.
 | Figure 2 Agarose gel electrophoresis of PCR amplification products of the four genes using multiplex PCR primers. |
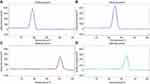
The PCR amplification products were differentiated by HRM curve analysis. The presence of blaoxa-23 was revealed by a single dominant peak with a melting temperature of 79.5°C in the melting curve analysis (Figure 3A), while Ade-B int-1 and ISCR-1 corresponded to single dominant peaks at 82.4°C, 90.3°C, and 87.1°C, respectively (Figure 3B–D). All melting temperatures were lower than predicted, and differences in the actual melting temperatures allowed the four genes to be easily distinguished by HRM curve analysis (Figure 4).
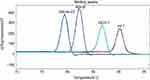 | Figure 4 Melting curves of the four genes form HRM analysis plotted on the same graph. Abbreviation: HRM, high resolution melting. |
Development of the multiplex PCR and HRM assay
Multiplex PCR amplification products with different annealing temperatures were separated by agarose gel electrophoresis and photographed, and melting curve analysis was performed after electrophoresis. Multiplex PCR amplification generated four products with sizes corresponding to the four genes (Figure 5A). Additionally, HRM analysis showed that the melting curves of multiplex PCR amplification products produced four peaks with the same melting temperature as the four genes (Figure 5B–C). Agarose gel electrophoresis showed that the blaoxa-23 amplification product decreased with increasing annealing temperature (Figure 5A), and the melting curve of blaoxa-23 similarly decreased with increasing annealing temperature (Figure 5D). Since the blaoxa-23 peak was the lowest of all four peaks, we selected 54°C as the most appropriate annealing temperature to improve the sensitivity of blaoxa-23 detection in the multiplex PCR with HRM assay.
Screening clinical isolates for the presence of the four genes
Seventy-nine MDRAB clinical isolates were screened for the presence of the four genes using the multiplex PCR with HRM assay. From the melting curves, the HRM assay identified blaoxa-23 and Ade-B in 73/79 isolates, while int-1 was present one isolate, both int-1 and ISCR-1 were present in two isolates, and Ade-B, int-1 and ISCR-1 were present in three isolates (Figure 6). No isolates were found to be carried all four genes (Figure 6). The presence of the four genes detected using the multiplex PCR with HRM assay was consistent with data from traditional methods (Figure7, Table 2).
 | Table 2 Presence of the four genes in clinical isolates detected by multiplex PCR with HRM assay and traditional method |
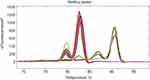 | Figure 6 Melting curves of the four genes detected in 79 clinical isolates using the single-tube multiplex PCR with HRM assay. Abbreviation: HRM, high resolution melting. |
Discussion
A. baumannii causes nosocomial infections in hospitals and health care departments and is difficult to treat in multidrug-resistant organisms that often carry OXA-23, Ade-ABC, Class 1 integron, and ISCR-1.9,24 Accurate and rapid detection of blaoxa-23, Ade-B, int-1, and ISCR-1 in clinical microbiology laboratories is important for prevention and treatment of MDRAB.25 Conventional assays for screening these genes are expensive and time-consuming. In contrast, HRM analysis is a rapid, cost-effective, and high-throughput method. We, therefore, sought to develop a rapid assay for detecting these genes without the need to perform agarose gel electrophoresis in this study.
We designed two sets of primers for each gene and PCR amplification products were sequenced to ensure the specificity of the multiplex PCR with HRM assay. The melting temperatures of the amplified products were different enough to allow us to distinguish between them.
Compared with traditional phenotypic assays, our multiplex PCR with HRM assay has a number of advantages; (1) It is significantly faster and more high-throughput. All four genes are detected simultaneously in the multiplex PCR with HRM assay in a single tube, minimizing contamination and improving efficiency since an agarose gel electrophoresis step is not required. The multiplex PCR with HRM assay could test 96 clinical isolates in 2 hrs 30 mins. (2) The novel assay is cost-effective. The approximate cost of this assay from DNA template extraction to multiplex PCR with HRM analysis is $2 per sample for the detection of all four genes. (3) The novel assay is non-destructive and environmentally friendly since no harmful ethidium bromide is used because an agarose gel electrophoresis step is not performed.
The four genes are known to play an important role in the evolution if resistance in A. baumannii, and in MDRAB in particular. In the 79 clinical isolates tested, blaoxa-23 and Ade-B were the most common, in accordance with published literature.26,27 Our results also demonstrate the co-existence of ISCR-1 and int-1 in bacteria. This may be related to the fact that ISCR1 can insert DNA into the 3ʹ conserved regions of class 1 integrons to generate complex class 1 integrons carrying the sul1, resX, and trIb resistance genes.24,28
In conclusion, we have developed a single-tube multiplex PCR with HRM assay which can reliably detect the presence of blaoxa-23, Ade-B, int-1, and ISCR-1 separately or in combination. This assay is rapid, cost-effective, non-destructive, easy to set-up, and has high-throughput in nature.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Towner KJ. Acinetobacter: an old friend, but a new enemy. J Hosp Infect. 2009;73(4):355–363. doi:10.1016/j.jhin.2009.03.032
2. Poulakou G, Renieris G, Sabrakos L, et al. Daptomycin as adjunctive treatment for experimental infection by Acinetobacter baumannii with resistance to col1istin. Int J Antimicrob AG. 2019;53(2):190–194. doi:10.1016/j.ijantimicag.2018.10.024
3. Schuertz KF, Tuon FF, Palmeiro JK, et al. Bacteremia and meningitis caused by OXA-23-producing Acinetobacter baumannii – molecular characterization and susceptibility testing for alternative antibiotics. Braz J Microbiol. 2018;49:199–204. doi:10.1016/j.bjm.2018.04.002
4. Chuang Y, Cheng A, Sun H, et al. Microbiological and clinical characteristics of Acinetobacter baumannii bacteremia: implications of sequence type for prognosis. J Infect. 2019;78(2):106–112. doi:10.1016/j.jinf.2018.10.001
5. Rocha IV, Xavier DE, Almeida KRHD, Oliveira SRD, Leal NC. Multidrug-resistant Acinetobacter baumannii clones persist on hospital inanimate surfaces. Braz J Infect Dis. 2018;22(5):438–441. doi:10.1016/j.bjid.2018.08.004
6. Jain M, Sharma A, Sen MK, Rani V, Gaind R, Suri JC. Phenotypic and molecular characterization of Acinetobacter baumannii isolates causing lower respiratory infections among ICU patients. Microb Pathog. 2019;128:75–81. doi:10.1016/j.micpath.2018.12.023
7. Lee M, Chen T, Lee Y, et al. Dissemination of multidrug-resistant Acinetobacter baumannii carrying BlaOxA-23 from hospitals in central Taiwan. J Microbiol Immunol Infect. 2013;46(6):419–424. doi:10.1016/j.jmii.2012.08.006
8. Fournier PE, Vallenet D, Barbe V, et al. Comparative genomics of multidrug resistance in Acinetobacter baumannii. PLoS Genet. 2006;2(1):e7. doi:10.1371/journal.pgen.0020007
9. Jamal S, Al Atrouni A, Rafei R, Dabboussi F, Hamze M, Osman M. Molecular mechanisms of antimicrobial resistance in acinetobacter baumannii, with a special focus on its epidemiology in Lebanon. J Glob Antimicrob Re. 2018;15:154–163. doi:10.1016/j.jgar.2018.05.022
10. Charfi-Kessis K, Mansour W, Ben Haj Khalifa A, et al. Multidrug-resistant Acinetobacter baumannii strains carrying the blaOxA-23 and the blaGES-11 genes in a neonatology center in Tunisia. Microb Pathog. 2014;74:20–24. doi:10.1016/j.micpath.2014.07.003
11. Da Silva KE, Maciel WG, Croda J, et al. A high mortality rate associated with multidrug-resistant Acinetobacter baumannii ST79 and ST25 carrying OXA-23 in a Brazilian intensive care unit. PLoS One. 2018;13(12):e209367. doi:10.1371/journal.pone.0209367
12. Petrović T, Uzunović S, Barišić I, et al. Arrival of carbapenem-hydrolyzing-oxacillinases in Acinetobacter baumannii in Bosnia and Herzegovina. Infect Genet Evol. 2018;58:192–198. doi:10.1016/j.meegid.2017.12.021
13. Costa EM, Silva S, Vicente S, Veiga M, Tavaria F, Pintado MM. Chitosan as an effective inhibitor of multidrug resistant Acinetobacter baumannii. Carbohydr Polym. 2017;178:347–351. doi:10.1016/j.carbpol.2017.09.055
14. Lee S, Oh MH, Yun SH, et al. Genomic characterization of extensively drug-resistant Acinetobacter baumannii strain, KAB03 belonging to ST451 from Korea. Infect Genet Evol. 2018;65:150–158. doi:10.1016/j.meegid.2018.07.030
15. Almasaudi SB. Acinetobacter spp. as nosocomial pathogens: epidemiology and resistance features. Saudi J Biol Sci. 2018;25(3):586–596. doi:10.1016/j.sjbs.2016.02.009
16. Poey ME, Laviña M. Horizontal transfer of class 1 integrons from uropathogenic Escherichia coli to E. coli K12. Microb Pathog. 2018;117:16–22. doi:10.1016/j.micpath.2018.02.006
17. Liu C, Tang CY, Chang K, Kuo H, Liou M. A comparative study of class 1 integrons in acinetobacter baumannii. Gene. 2014;544(1):75–82. doi:10.1016/j.gene.2014.04.047
18. Rui Y, Lu W, Li S, Cheng C, Sun J, Yang Q. Integrons and insertion sequence common region 1 (ISCR1) of carbapenem-non-susceptible gram-negative bacilli in fecal specimens from 5000 patients in southern China. Int J Antimicrob AG. 2018;52(5):571–576. doi:10.1016/j.ijantimicag.2018.06.015
19. Zheng F, Sun J, Cheng C, Rui Y. Molecular characteristics of carbapenem-resistant gram-negative bacteria in Southern China. Microb Drug Resist. 2015;21(2):178–185. doi:10.1089/mdr.2014.0085
20. Yong TB, Hashim R, Noor AM, Hamzah SH, Ahmad N. Identification of brucella spp. isolated from human brucellosis in Malaysia using high-resolution melt (HRM) analysis. Diagn Micr Infect Dis. 2015;81(4):227–233. doi:10.1016/j.diagmicrobio.2014.12.012
21. Pasanen T, Koskela S, Mero S, et al. Rapid molecular characterization of Acinetobacter baumannii clones with rep-PCR and evaluation of carbapenemase genes by new multiplex PCR in hospital district of helsinki and uusimaa. PLoS One. 2014;9(1):e85854. doi:10.1371/journal.pone.0085854
22. Lee Y, Yum JH, Kim C, et al. Role of OXA-23 and AdeABC efflux pump for acquiring carbapenem resistance in an Acinetobacter baumannii strain carrying the blaOXA-66 gene. Ann Clin Lab Sci. 2010;40(1):43–48.
23. Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. doi:10.1111/j.1469-0691.2011.03570.x
24. Guan X, He L, Hu B, et al. Laboratory diagnosis, clinical management and infection control of the infections caused by extensively drug-resistant gram-negative bacilli: a Chinese consensus statement. Clin Microbiol Infect. 2016;22:S15–25. doi:10.1016/j.cmi.2015.11.004
25. Roth AL, Hanson ND. Rapid detection and statistical differentiation of KPC gene variants in Gram-negative pathogens by use of high-resolution melting and ScreenClust analyses. J Clin Microbiol. 2013;51(1):61–65. doi:10.1128/JCM.02193-12
26. Lin M, Lin Y, Tu C, Lan C. Distribution of different efflux pump genes in clinical isolates of multidrug-resistant Acinetobacter baumannii and their correlation with antimicrobial resistance. J Microbiol Immunol Infect. 2017;50(2):224–231. doi:10.1016/j.jmii.2015.04.004
27. Pournaras S, Dafopoulou K, Del Franco M, et al. Predominance of international clone 2 OXA-23-producing-Acinetobacter baumannii clinical isolates in Greece, 2015: results of a nationwide study. Int J Antimicrob AG. 2017;49(6):749–753. doi:10.1016/j.ijantimicag.2017.01.028
28. Ramírez MS, Vilacoba E, Stietz MS, et al. Spreading of AbaR-type genomic islands in multidrug resistance acinetobacter baumannii strains belonging to different clonal complexes. Curr Microbiol. 2013;67(1):9–14. doi:10.1007/s00284-013-0326-5
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.



