Back to Journals » International Journal of Nanomedicine » Volume 15
Publication Trends in Exosomes Nanoparticles for Cancer Detection
Authors Ale Ebrahim S , Ashtari A, Zamani Pedram M, Ale Ebrahim N , Sanati-Nezhad A
Received 24 January 2020
Accepted for publication 12 May 2020
Published 26 June 2020 Volume 2020:15 Pages 4453—4470
DOI https://doi.org/10.2147/IJN.S247210
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Anderson Oliveira Lobo
Saba Ale Ebrahim,1,* Amirhossein Ashtari,2,* Maysam Zamani Pedram,3– 5,* Nader Ale Ebrahim,6,* Amir Sanati-Nezhad4,5,*
1School of Electrical Engineering, Iran University of Science and Technology, Tehran, Iran; 2Department of Electrical Engineering, Politecnico di Milano, Milan, Italy; 3Faculty of Electrical Engineering, K. N. Toosi University of Technology, Tehran, Iran; 4Department of Mechanical and Manufacturing Engineering, University of Calgary, Calgary, Alberta T2N 1N4, Canada; 5Center for Bioengineering Research and Education, Biomedical Engineering Program, University of Calgary, Calgary, Alberta T2N 1N4, Canada; 6Research and Technology Department, Alzahra University, Vanak, Tehran, Iran
*These authors contributed equally to this work
Correspondence: Maysam Zamani Pedram; Amir Sanati-Nezhad
Email [email protected]; [email protected]
Background: Exosomes are small vesicles produced by almost all cells in the body and found in all biofluids. Cancer cell-derived exosomes are known to have distinct, measurable signatures, applicable for early cancer diagnosis. Despite the present bibliometric studies on “Cancer detection” and “Nanoparticles”, no single study exists to deal with “Exosome” bibliometric study.
Methods: This bibliometric work investigated the publication trends of “Exosomes” nanoparticles and its application in cancer detection, for the literature from 2008 to July 2019. The data were collected from the Web of Science Core Collection. There were variant visual maps generated to show annual publication, most- relevant authors, sources, countries, topics and keywords. The network analysis of these studies was investigated to evaluate the research trends in the field of exosomes. In addition, the data were qualitatively analyzed according to 22 top-cited articles, illustrating the frequently used subjects and methods in exosomes research area.
Results: The results showed that the documents in this field have improved the citation rate. The top-relevant papers are mostly published in Scientific Reports journal which has lost its popularity after 2017, while today, Analytical Chemistry is leading in publishing the most articles related to exosomes. The documents containing keywords of plasma, cells, cancer, biomarkers, and vesicles as keywords plus, are more likely to be published in PLoS One journal. The clustering of the keywords network showed that the keyword theme of “extracellular vesicles” has the highest centrality rate. In global research, USA is the most corresponding country, followed by China, Korea and Australia. Based on the qualitative analysis, the published documents with at least 50 citations have used exosome release, cargo, detection, purification and secretion, as their targets and applied cell culture or isolation as their methods.
Conclusion: The bibliometric study on exosomes nanoparticles for cancer detection provides a clear vision of the future research direction and identifies the potential opportunities and challenges. This may lead new researchers to select the proper subfields in exosome-related research fields.
Keywords: exosomes, cancer detection, nanoparticles, microvesicles, bibliometrics, research productivity
Introduction
Conventional techniques for cancer detection (eg, computed tomography, advanced imaging and tissue biopsy) are associated with many challenges, such as high cost, invasive surgical procedures, low sensitivity and high false-positive rate.1,2 To address these challenges, exosome-based liquid biopsy has been introduced in the last decade for early cancer detection, given its minimally invasive and highly sensitive and specific performance.2 The unique characteristics of cancer cell-derived exosomes make them a potential biomarker for early cancer diagnosis, detection of highly metastatic cancer cells, and assessment of cancer heterogeneity. Exosomes are also markers for tracking cancer patient’s response to treatments and detection of resistance mechanisms against treatment, contributing to precise and personalized cancer treatment (Table 1).3 Exosomes shed from cancer cells have shown to be highly predictive for cancer detection since they contain the biomolecules that are reflective of oncogenic signaling in cancer cells of origin.4 Exosomal proteins can reliably distinguish cancer type and stage for some types of cancers.5,6
 |
Table 1 Exosomal Biomarkers for Diagnosis of Different Cancers |
Bibliometric is defined as the application of mathematical and statistical methods to evaluate scholarly outputs.7 The bibliometric research to date has focused on natural products against cancer,8 nanotechnology applied in oncology,9 neurotoxicity of nanoparticles,10 and drug delivery and magnetic nanoparticles.11 Although extensive research has been carried out on the bibliometric study of “Cancer detection” and “Nanoparticles”, no single study exists to deal with “Exosome” bibliometric study. This study aims to explore the research status on “Exosome” research field in “Cancer detection” and “Nanoparticles”, from the beginning to the current year (January 2020) by bibliometric approaches.
Bibliometric approach has been used to measure scientific progress in many disciplines for the systematic analysis of publications.12 For research evaluation, some other indicators were needed. Citation analysis along with peer review ensured better judgment in countless cases.13 Nowadays, several tools have apparently made it easier to produce a bibliometric report.14 This ranges from databases such as Web of Science Core Collection (WoS), SCOPUS, and Google Scholar that have added incorporated reference-handling capabilities.14–16 Google Scholar provides free access to scholarly documents of all types, which makes it questioned due to sporadic coverage.17 WoS and Scopus are the most extensive databases in different scientific fields, used for searching the literature.18 Therefore, in this research SCOPUS and WoS databases are compared against the coverage of “Exosome” research and the best one is selected for the study.
Methodology
A bibliometric study was conducted to evaluate the publication trends and find the insight of publications on “Exosome”. A title search of “Exosome” was carried out on both well-known databases, SCOPUS and WoS, to select the preferred database for data collection. The number of documents on SCOPUS was 1665 which was less than the number of documents with the same search on WoS (1869 records). WoS is the most appropriate influential, extensive, and trustworthy database for literature retrieval and analysis.18,19 Therefore, the data were collected from WoS database base of the title search of “Exosome”. A title search of “Exosome” in WoS returned 1869 documents on 30 July 2019. The entire documents of 1869 received total citations of 64,520 and average citations per paper of 34.52. The trends in the number of publications in the “Exosome” field grow dramatically, as shown in Figure 1.
 |
Figure 1 Publication trends in Exosome research field from 1989 to 2019 (red dotted line: the prediction trends, blue line: the original trends). |
The search was refined by the topic search of “Cancer Detection*” and “Nanoparticle*” which therefore reached 166 documents indexed in the Science Citation Index Expanded (SCIEXPANDED), Social Sciences Citation Index (SSCI), Arts & Humanities Citation Index (A&HCI), and Emerging Sources Citation Index (ESCI) of WoS. These 166 relevant documents have received the average citations per item of 29.33, which is slightly less than “Exosome” citations. This is due to the nature of new emerging research areas, as the first related document was published in 2008. The final 166 sets of documents were analyzed by Bibliometrix-package (http://www.bibliometrix.org/), which is an R-Tool for science mapping.20
Table 2 shows a summary of the primary information on collected Bibliometric data. There were 135 articles, 23 review papers, and 8 editorials, meeting abstracts, and corrections. To map the present subtopics of the exosome-based research field, especially nanoparticles and cancer detection, 22 top-cited articles have been selected (from the total of 135 articles) for qualitative analysis. The quantitative and qualitative analysis of the data comes in the following sections.
 |
Table 2 Summary of the Main Information of Collected Bibliometric Data |
Quantitative Analysis
Analysis of Publication Years
Figure 2 shows the yearly scientific production of published papers about exosomes nanoparticles and cancer detection, within the period of 2008 to 31st July 2019. Within this period, there are 166 scientific productions including 135 articles, 23 review papers, two proceedings papers and six others. The number of articles published has totally increased from the minimum one document in 2008 to 46 documents in 2018. There are two turning points during this decade. In 2014 and again in 2018, the number of publications were duplicated compared to the prior year. The distribution of exosome nanoparticles and exosome cancer detection publications are split into two stages. In the first stage, from 2008 to 2013, there are only a few numbers of scientific productions. However, there is a publication growth explosion from 2014 to 2019. Although the number of published documents decreased from 28 in 2016 to 23 in 2017, more researchers focused in this area in 2018.
 |
Figure 2 Annual scientific production of exosomes nanoparticles for cancer detection research field within 2008 to 2018 period. |
The average article citations per year from 2008 to 31st July 2019 are shown in Figure 3. According to the trend, the highest number of average citations per year, were collected by six articles published in 2012. The publications of 2017 had 15 citations on average, while the number of published papers decreased.
 |
Figure 3 Average article citations per year of exosomes nanoparticles for cancer detection research field within 2008 to 2018 period. |
There are 166 citing documents such as articles, reviews, and conference proceedings in this bibliographic collection. Figure 4 shows the most local cited documents in the period of 2008 to 2019. Local citations measure the number of records citations received from papers involved in the analyzed set. As the figure shows, the top two highly local citations belong to the articles published in 2016. These articles gathered 15 and 11 local citations, respectively, though the leading average article citations occurred in 2012 (Figure 3).
 |
Figure 4 Top 20 most local cited documents published on exosomes nanoparticles for cancer detection research field. |
Analysis of Authors
There is a total number of 1102 research works publishing their scientific accomplishments in the field of exosome nanoparticles and cancer detection from 2008 to 2019. This fact means, there are 6.64 authors and 7.55 co-authors per document in our analysis collection. Figure 5 shows the most relevant authors’ production over time from 2013 to 2019. The red line signifies the author’s timeline. Zhang HG published six articles in the field of exosome nanoparticles or cancer detection from 2013 to 2018. The author has the most protracted timeline among other authors. The bubble size is related to the number of documents published during the years. The bubble color intensity is also related to the total citations per year. Deng ZB, Lee J and Miller D are the authors with the highest publications in the field of exosomes, with five publications during 2013 to 2018. There are also several authors with two highly cited papers in 2013. According to the graph, more researchers are working on exosomes and getting interested in this area.
Analysis of Sources
There are 108 sources such as journals, books, conference proceeding series and others in this bibliographic collection. Figure 6 shows the most relevant sources in exosomes publications applied in the field of nanoparticles and cancer detection. Each of the sources has published one or more documents in this analyzed set. The open-access journal of Scientific Reports is the top critical journal in the field of exosomes, as it published about 11 documents from 2008 to 2019. The PLoS One journal is the second top journal with nine publication counts. The Analytical Chemistry journal has approximately five and the other journals have less than five published papers in this analysis. The three top journals in the figure are significant for researchers in the field of exosomes to consider first for submitting their articles.
 |
Figure 6 Top 20 most relevant sources by the number of documents published on exosomes nanoparticles for cancer detection research. |
Figure 7 shows the number of sources occurrences in the time span of 2008 to 2019. The graph shows the best sources of dynamics in the field of exosomes nanoparticles and cancer detection. Yearly publications on the top journals has increased as the time passes. Moreover, the Scientific Reports journal growth slowed down after 2017. Today, the journal of Analytical Chemistry is the leader in publishing the most relevant articles. On the other hand, Lab On a Chip journal is publishing exosome-related papers in a downward trend, since 2016. Although ASC Nano had no occurrence in 2014, its presentation has exceeded the PLoS One as the second most relevant source in exosomes in 2019.
 |
Figure 7 Annual occurrences of top five most relevant sources in exosomes nanoparticles for cancer detection research field within 2008 to 2018 period (method: loess smoothing). |
Analysis of Countries
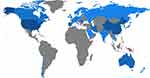
There are 166 papers in the field of exosome cancer detection and nanotechnology, distributed among 43 countries. In Figure 8, top 20 countries were ranked by their count of scientific productions. The red lines demonstrate the publications rate by corresponding author’s country, wherein at least one foreign co-author exits. The blue lines represent the number of publications by authors from the same country. These are called Multiple Countries publication (MCP) and Single Country Publication (SCP), respectively. The USA with 63, China with 40, and South Korea with 14 publications are considered as the top three most relevant countries. The USA has by far the most international collaboration. Two North American countries, 11 European countries, six Asian countries and one Oceania country, are ranked as countries with highest publications in the area of exosomes.
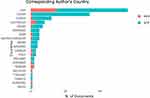 |
Figure 8 Top 20 corresponding author’s country (red line: Multiple Countries Publication (MCP), blue line, Single Country Publication (SCP)). |
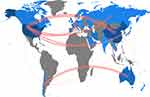
Figure 9 gives a complete picture of the number of authors affiliated with the country of publication. Figure 10 shows the number of joint documents published by the top countries in the field of exosomes nanoparticles and cancer detection. The blue color intensity, in both figures, is proportional to the number of affiliated authors with each country. Every blue color indicates a number of related authors ranging from the darkest blue with 223 authors in the USA, to the lightest with the minimum of one author in Sweden. The USA and China are the two central research powers in exosome nanoparticles and cancer detection research fields. South Korea, Australia and Spain are the second most highly productive countries. The red line thickness in Figure 10 is proportional to the number of joint publications within the countries. The thickest of every red link between countries indicates a number of collaborated papers ranging from the highest thickness of six joint documents between the USA and China, to the lowest thickness of one joint document between Australia and Japan. As Figure 10 illustrates, China and USA are very collaborative in scientific productions. Interestingly, Australia and Chile have too many mutual articles in exosomes area. Overall, USA seems to be the hub country of any published document, as there are lots of scientific links between the USA and other countries.
Analysis of Topics
To analyze the main topics of exosome publications on nanoparticles and cancer detection, keywords networks are used. Keywords networks represent the co-occurrences among bibliographic dataset. It is likely to highlight the variant themes by clustering the keywords networks. Each keyword belongs to only one theme. A thematic map is a specific plot able to represent each theme. Figure 11 shows the topics of exosome research field focused on cancer detection and nanoparticles in a thematic map. Each bubble shows one keyword network cluster. The cluster name is the word with the highest existence rate. Therefore, invitro, stem-cells, apoptosis, dendritic cells, membrane-vesicles, ovarian-cancer, mechanism, microvesicles, biomarkers and extracellular vesicles are the most relevant theme indicators.
 |
Figure 11 Thematic map of keywords network clusters in exosomes nanoparticles for cancer detection research field (bubble size: the clusters word occurrences). |
The bubble size is relative to the cluster word occurrences, and its position depends on the cluster centrality and density. Centrality and density show the theme importance and theme improvement in the exosomes research area, respectively. Therefore, highly developed and isolated themes are on the top left, motor themes are on the top right, emerging or declining themes are on the bottom left, and primary and transversal themes are on the bottom right of the thematic topic map. Keywords such as in-vitro, stem-cells, apoptosis, membrane vesicles and dendritic cells, are the five cluster representatives with a few numbers of occurrences. They are called as highly developed and isolated themes for their low level of importance and high level of improvement.
On the other side, keywords such as ovarian cancer, mechanism, microvesicles, biomarkers and extracellular vesicles, are the five cluster representatives called basic and transversal themes. Extracellular vesicles’ keywords theme is the most important theme with the highest centrality, among others. Microvesicles, extracellular vesicles and biomarkers are the top mostly occurred keywords theme and more likely to be used in future exosomes research.
Analysis of Keywords
Three-field plots are created to give a general view of the articles’ keywords referring to the application of nanoparticles and cancer detection in exosomes. Figures 12 and 13 show the three-field plots, which mostly focus on the top keywords. Figure 12 is formed by selecting the three main metadata fields, keywords as the middle field, authors as the left field and sources as the right field. It shows the relationship among top keywords, top authors and top journals. As it is shown in Figure 12, Salomon C, Boriachek K, Nguyen NT, Shiddiky MJA, Weissleder R, Kim JA, Rhee WJ, Lee JH, Lee J and Liu L, have used almost every top keyword in their publications.
The top relevant keywords such as exosomes, extracellular vesicles, microRNA, prostate cancer, cancer, exosome, diagnosis, liquid biopsy, breast cancer, microfluidics and pancreatic cancer are the most frequently used keywords. While, microvesicles, exosomes, biomarkers, extracellular vesicles, nanoparticles, lung cancer, biomarkers, nanoparticle tracking analysis, inflammation, breast cancer, microfluidics, pancreatic cancer and microRNAs are mostly the top journal articles’ keywords. Sources such as Nano Letters, Small, Journal of Controlled Release, Journal of Translational Medicine, Advanced Materials, Molecular Cancer, ACS Nano, Cancers, Biosensors & Bioelectronics, Nanomedicine-Nanotechnology Biology and Medicine, miRNA, Oncotarget, Micromachines and Frontiers in Pharmacology are the top journals publishing articles about exosomes.
Figure 13 is created by selecting another three metadata fields, keywords plus as the middle field, authors as the left field and sources as the right field. It shows the relationship among top keywords plus, top authors and top journals. Keywords plus are words or phrases that frequently appear in the article references titles and generated automatically by computer. In Figure 13, compared to Figure 12, there are more authors mentioned with more keywords. Analyzing top keywords plus represents hot research topics in exosomes field of research and helps the reader to find research frontiers.
As it is shown in Figure 13, articles related to in-vivo, are only published in the Scientific Reports journal. This journal also has a high number of publications in plasma, drug delivery, cells, extracellular vesicles, cancer and nanoparticles. Keywords plus such as plasma, cells, cancer, biomarkers and vesicles are more likely to be published in PLoS One journal. There is a spectacular reception of extracellular vesicles, as every top author has used it in their articles with a vast number of available publishers. 80% of top authors in exosomes have been working on cancer, where their publications are mainly published in Scientific Reports, PLoS One and Analytical Chemistry journals. Authors as Weissleder R has published articles on microvesicles in the journals of ACS Nano and Lab on a Chip. Therefore, Figure 13 is very suitable in finding the subjects and topics of the articles published in the top journals, in order to provide guidance to submit a paper to a specific journal.
Figure 14 shows the relationship among keywords of the publication set by designing a conceptual structure map. The two dimensions of the map represents the average position of the articles included in each keyword, and the midpoint of the map represents the center of the exosomes nanoparticles for cancer detection research field.21 In a conceptual structure map, every document’s words are linked via one network. This co-word network structure helps the readers to understand research fields’ covered topics and find the research fronts (top new highlighted matters). Data reduction techniques (factorial analysis) can detect subfields, separately from the network analysis. Correspondence analysis (CA), as a dimensionality reduction technique, is chosen in the creation of the conceptual structure map. Factorial approaches aim to reduce data dimensionality and represent it in a low-dimensionality space.
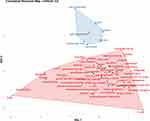 |
Figure 14 Conceptual structure map of keywords in exosomes nanoparticles for cancer detection publications (Dim.1 and Dim.2: the average position of the articles included in each keyword). |
As Figure 14 illustrates, each color represents a cluster of words which is known by classified clustering. Therefore, keywords are divided into two clusters. The blue cluster consists of seven keywords and shows cell-proliferation, gene-expression, messenger RNAs, mechanism, delivery, inhibition, and growth due to its color. Approximately all keywords in this cluster are far from each other. On the other hand, the red cluster contains 55 keywords and is more prominent than the blue cluster. Keyword sets such as stem-cells and serum, cancer and cells, ovarian-cancer and extracellular vesicles, identification and microvesicles, chip and messenger-RNA, plasma and quantification, proteomic analysis and antigen, proteins and expression, and pancreatic-cancer and transferrin receptor are close to each other.
Figure 15 shows another type of conceptual structure graph of keywords called dendrogram. This figure contains the same information as in Figure 14, with a different view. Similarly, the conceptual structure dendrogram shows two clusters of keywords. The height measures the distance among words or clusters of words. Every dendrogram describes a partition, while split in a right place. Distant words are keywords with different topics, which are not usually included in the same articles.
 |
Figure 15 Conceptual structure dendrogram of keywords in exosomes nanoparticles for cancer detection publications (height: the distance among clusters of words). |
Sometimes the reader needs to quickly perceive the most prominent terms in his/her research area. Word cloud is a visual illustration of keywords metadata which instantly remarks the top words. Top keywords plus, top author’s keywords, top title words and top abstract words are represented in Figures 16–19, respectively. Keywords plus are extracted by article reference titles. These words are capable of reaching the article’s in-depth content. Author’s keywords are a list of words that suites the article from authors’ point of view. There is total of 659 keywords plus and 330 author’s keywords produced for this analysis. Although keywords plus and author’s keywords have the same effect in exploring knowledge from the bibliometric analysis, author’s keywords are more inclusive in providing subjects. The top title and abstract words are driven from abstracts or titles cleaned of any punctuation or trivial terms, like paper, study, work, data etc. Figure 16 visualizes the keywords plus in the field of exosomes from 2008 to 2019. The font size or color of these single words shows their importance. The keywords plus occurrences ranging from 58 to four are mentioned in Figure 16. Microvesicles, cells, cancer and biomarkers are the top ranked terms indicated in the picture and the best keywords plus in our research area.
 |
Figure 16 Word cloud of top keywords plus in exosomes nanoparticles for cancer detection publications (font size: word occurrences). |
 |
Figure 17 Word cloud of top author’s keywords in exosomes nanoparticles for cancer detection publications (font size: word occurrences). |
 |
Figure 18 Word cloud of top title words in exosomes nanoparticles for cancer detection publications (font size: word occurrences). |
 |
Figure 19 Word cloud of top abstract words in exosomes nanoparticles for cancer detection publications (font size: word occurrences). |
Figure 17, shows the author’s keywords, ranging from the total number of 41 to the minimum of two occurrences, with exosomes, exosome, biomarker, microRNA and extracellular vesicles on top of them. Figure 18 shows the words that appeared in titles, where exosome, cancer, cells, detection, exosome-like and nanoparticles are the most relevant keywords. The title word occurrence is ranged from as high as 111 to as low as four times. Figure 19 shows the words appeared in abstracts, most frequently exosomes, exosome, cells, cancer, cell and patients. The top abstract words are frequently (from the word exosomes with 549 to pancreatic with 40) appeared in Exosome-related publications. It is interesting to know that the top two or three results in figures, come straight from the terms used for collecting the data and are very bold and massive. In conclusion of these word clouds, surprisingly, word clouds of author’s, title or abstract keywords are not following the pattern in keywords plus word cloud. Therefore, authors should attend to use more relevant words, as illustrated in the word cloud of keywords plus (such as microvesicles), in the title, abstract or as keywords of the publications.
Figure 20 shows the number of top keywords in exosome-based cancer detection or nanotechnology in the time span of 2008 to 31st July 2019. Extracellular vesicles as a research topic is overgrowing since 2013, reaching 15 occurrences in 2019. Keywords such as cells, cancer, biomarkers, expression and nanoparticles, are getting more visible in recent publications. Nanoparticle tracking analysis and microvesicles have lost their popularity since 2016 and 2017. Most commonly used keywords are increasing yearly, and research on these topics is also enlarging. Due to the dramatic rise in frequency from 2008 to 2019, a vast amount of research work is predicted on these topics in the exosomes research field.
 |
Figure 20 Annual occurrences of top keywords in exosomes nanoparticles for cancer detection research field within 2008 to 2018 period (method: loess smoothing). |
Qualitative Analysis
There are 166 papers in the field of exosomes which are related to cancer detection and nanoparticles, published from 2008 to July 2019. It is considered that 16.8% of the total scientific productions have equal to or more than 50 citations. This number points the high-quality papers within our bibliographic dataset. Therefore, there are 50 articles with at least 50 citations published in 2008–2019. In this section, the data is qualitatively analyzed according to 22 top-cited articles. Supplementary Table shows the most/least studied subjects and methods in exosomes research field.
Subjects
The frequently used subjects in top-cited exosome articles include exosomes, cancer/experimental cells, targeting and protective mechanism. The most relevant article topics followed by their subtopics are mentioned in Supplementary Table. Every top article has discussed or somehow used exosomal protein in its case study. About 68.1% of the authors in this analysis has focused on exosomal membrane and exosomal markers as their articles’ subjects. Although microRNA is a typical author’s keyword and previously was seen in the top keywords’ graph, it has been occurred in half of the top-cited articles.
Research subjects such as tumor-derived exosomes, serum-derived exosomes, liposomes (synthetic lipid vesicles), exosome-like vesicles (ELVs), exosome-like nanoparticles and extracellular vesicles (EVs) have also attracted almost half the authors of most cited documents. There are only limited papers that use all exosomal protein, microRNA, tumor-derived exosomes, exosomal membrane, serum-derived exosomes, microvesicles, liposomes, and exosome-like vesicles (ELVs) subject keywords together.22,23 Microvesicles, as the most critical term in the total articles’ keywords plus, seems to have a moderately rare usage here. Multivesicular endosomes and exosome-derived DNA are quite rare headings in this collection set, as just a few titles are focusing on them.22,24,25
Human cancer cells were regularly observed by authors. Breast cancer cells, prostate cancer cells, and lung cancer cells are the extremely examined cells in these highly cited papers. Supplementary Table illustrates that almost every top cited article experimenting on cancer cells uses tumor-derived exosomes. Scientific documents with the minimum number of 50 citations target exosome release, exosome cargo, exosome detection, exosome purification, and exosome secretion. 77.2% of the articles aimed to purify exosomes, 68.1% of them considered released exosomes, and only 45.4% of them detect the exosomes. Interestingly, exosome cargo comes with exosome release in every article’s purpose. There are a few targeting titles in our top collection in exosome secretion which has been applied by five articles.22,23,26-28 Serum exosome protein, exosome-based drug delivery, and nanoparticle-mediated protein delivery concluded to be the best protective mechanisms in highly cited papers. Nearly every article with a protective exosome-based drug delivery works with exosome-like nanoparticles, which makes them very related.25,29-31
Methods
The analysis of exosomes based methods are divided into two groups, characterization and quantification of exosomes and statistical analysis. As Supplementary Table shows, only a few numbers of articles misses cell culture or exosome isolation methods. The reason refers to frequently usage of cell cultivation in articles with a running experiment. Most cited papers characterized the isolated exosomes by nanoparticle tracking analysis (NTA), Western blot analysis (WB), staining, electron microscopy, quantitative analysis, and fluorescence detection methods. These are exosomes characterization and qualification methods utilized by more than half of the analyzed set. Among these, the exosomes preparation method of electron microscopy has attracted 19 (out of 22) top-cited articles’ authors. Meanwhile, confocal microscopy, used for micrometre ranged cells, has rarely been found in the top five papers.25,29,30,32,33
Other representation methods such as plasma collection, RNA extraction, flow cytometry, dynamic light scattering, and real-time PCR have been applied to 45.4% of the total papers with higher than 50 citations. It is notable that, articles monitoring the PCR reaction in real-time have often used RNA extraction preparation techniques for exosomes.27,33-37 As it is shown in Supplementary Table, statistical analyses were mostly carried out using ANOVA statistical analysis and Student’s T-test. Microarray statistical analysis, Tukey HSD statistical analysis, and Mann–Whitney test were captured in 45.4% of the top articles’ statistical analysis. Two related authors were interested in using parametric analysis as their statistical analysis.23,38 Nearly every top-cited article in the field of exosomes nanoparticles and cancer detection, has displayed the error bars as the standard error of the mean (SEM).
In this final section, 22 exosome based articles with the highest WoS times cited count were reviewed. Exosomal protein has been accounted for the most popular material in exosomes research. Most of the analyzed references have nominated breast and prostate cancer cells as their experimental cells. There is a huge percent of research studies that applied exosome isolation. Cell culture and electron microscopy are the other focused characterization methods in top-cited articles. Finally, the related authors ran the student’s T-rest as the statistical analysis and plotted the standard error of the mean as a result in every top article.
Conclusion
Bibliometric analysis uses mathematical and statistical methods to evaluate scholarly outputs. Although extensive research has been carried out on the bibliometric study of “Cancer detection” and “Nanoparticles”, there is no study dealing with “Exosome” bibliometric analysis. This work focuses on the bibliometric analysis of exosomes research literature from the year 2008 to 2019 with WoS data source. The results show that exosomes and applications of exosome-based detection are progressing in recent years. Annual publication in exosomes nanoparticles and cancer detection research field continue to increase due to turning points in 2014 and 2018. Researchers from USA, China, Korea, Australia and Spain contributed the most to the publications. The USA and China were ranked first and second in quantity of paper outputs. Scientific Reports, PLoS One, and Analytical Chemistry were identified as the most relevant journals in exosomes research field. The basic and transversal themes in our bibliographic set are representative keywords such as ovarian-cancer, mechanisms, microvesicles, biomarkers and extracellular vesicles. Keywords like cancer and biomarkers are improving their visibility and microvesicles are losing their occurrence in recent years. Furthermore, the qualitative analysis nominated breast and prostate cancer cells as the top experimental cells and exosome isolation as the most frequent research method. It is expected that the outcomes from this work provide a better vision on the future research direction and identify the potential opportunities for translation of the exosome research into clinical applications.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Patz EF
2. Liu C, Yang Y, Wu Y. Recent advances in exosomal protein detection via liquid biopsy biosensors for cancer screening, diagnosis, and prognosis. AAPS J. 2018;20(2):41. doi:10.1208/s12248-018-0201-1
3. Panagiotara A, Markou A, Lianidou ES, Patrinos GP, Katsila T. Exosomes: a cancer theranostics road map. Public Health Genomics. 2017;20(2):116–125. doi:10.1159/000478253
4. Melo SA, Luecke LB, Kahlert C, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523(7559):177–182. doi:10.1038/nature14581
5. He M, Crow J, Roth M, Zeng Y, Godwin AK. Integrated immunoisolation and protein analysis of circulating exosomes using microfluidic technology. Lab Chip. 2014;14(19):3773–3780. doi:10.1039/C4LC00662C
6. Szajnik M, Derbis M, Lach M, et al. Exosomes in plasma of patients with ovarian carcinoma: potential biomarkers of tumor progression and response to therapy. Gynecol Obstet (Sunnyvale). 2013;003.
7. Elaish MM, Shuib L, Ghani NA, Mujtaba G, Ale Ebrahim N. A bibliometric analysis of m-learning from topic inception to 2015. Int J Mob Learn Organ. 2019;13(1):91–112. doi:10.1504/IJMLO.2019.096470
8. Du J, Tang XL. Natural products against cancer: a comprehensive bibliometric study of the research projects, publications, patents and drugs. J Cancer Res Ther. 2014;10(5):27. doi:10.4103/0973-1482.139750
9. Dong X, Qiu X-C, Liu Q, Jia J. Bibliometric analysis of nanotechnology applied in oncology from 2002 to 2011. Tumor Biol. 2013;34(6):3273–3278. doi:10.1007/s13277-013-1032-4
10. Su B, Guan Q, Yu S. The neurotoxicity of nanoparticles: a bibliometric analysis. Toxicol Ind Health. 2018;34(12):922–929. doi:10.1177/0748233718804973
11. Ale Ebrahim S, Ashtari A, Pedram MZ, Ale Ebrahim N. Publication trends in drug delivery and magnetic nanoparticles. Nanoscale Res Lett. 2019;14(1):164. doi:10.1186/s11671-019-2994-y
12. Kalantari A, Kamsin A, Kamaruddin HS, et al. A bibliometric approach to tracking big data research trends. J Big Data. 2017;4(30):1–18. doi:10.1186/s40537-017-0088-1
13. Das A-K. Introduction to Research Evaluation Metrics and Related Indicators. In: Sen BK, Mishra S, editors. Open Access for Researchers, Module 4: Research Evaluation Metrics. UNESCO, Paris, 7, place de Fontenoy, 75352 Paris 07 SP, France: United Nations Educational, Scientific and Cultural Organization; 2015.1–18.
14. Ellegaard O, Wallin J. The bibliometric analysis of scholarly production: how great is the impact? Scientometrics. 2015;105:1–23.
15. Martín-Martín A, Orduna-Malea E, Thelwall M, Delgado López-Cózar E. Google scholar, Web of Science, and Scopus: a systematic comparison of citations in 252 subject categories. J Informetr. 2018;12(4):1160–1177. doi:10.1016/j.joi.2018.09.002
16. Bergman EML. Finding citations to social work literature: the relative benefits of using Web of Science, Scopus, or Google Scholar. J Acad Librariansh. 2012;38(6):370–379. doi:10.1016/j.acalib.2012.08.002
17. Mongeon P, Paul-Hus A. The journal coverage of web of science and scopus: a comparative analysis. Scientometrics. 2015;106:1–16.
18. Aghaei Chadegani A, Salehi H, Yunus MM, et al. A comparison between two main academic literature collections: web of science and scopus databases. Asian Soc Sci. 2013;9(5):18–26.
19. Ghanbari Baghestan A, Khaniki H, Kalantari A, et al. A crisis in “Open Access”: should communication scholarly outputs take 77 years to become open access? SAGE Open. 2019;9(3):1–8. doi:10.1177/2158244019871044
20. Aria M, Cuccurullo C. bibliometrix: an R-tool for comprehensive science mapping analysis. J Informetr. 2017;11(4):959–975. doi:10.1016/j.joi.2017.08.007
21. Cuccurullo C, Aria M, Sarto F. Foundations and trends in performance management. A twenty-five years bibliometric analysis in business and public administration domains. Scientometrics. 2016;108(2):595–611. doi:10.1007/s11192-016-1948-8
22. Lehmann SB, Paine M, Brooks A, et al. Senescence-associated exosome release from human prostate cancer cells. Cancer Res. 2008;68:7864–7871. doi:10.1158/0008-5472.CAN-07-6538
23. Valencia K, Luis-Ravelo D, Bovy N, et al. miRNA cargo within exosome-like vesicle transfer influences metastatic bone colonization. Mol Oncol. 2014;8. doi:10.1016/j.molonc.2014.01.012
24. Allenson K, Castillo J, San Lucas F, et al. High prevalence of mutant KRAS in circulating exosome-derived DNA from early-stage pancreatic cancer patients. Ann Oncol. 2017;28(4):741–747. doi:10.1093/annonc/mdx004
25. Yim N, Ryu S-W, Choi K, et al. Exosome engineering for efficient intracellular delivery of soluble proteins using optically reversible protein–protein interaction module. Nat Commun. 2016;7:12277. doi:10.1038/ncomms12277
26. Soo CY, Song Y, Zheng Y, et al. Nanoparticle tracking analysis monitors microvesicle and exosome secretion from immune cells. Immunology. 2012;136:192–197. doi:10.1111/j.1365-2567.2012.03569.x
27. Hornick NI, Huan J, Doron B, et al. Serum Exosome MicroRNA as a minimally-invasive early biomarker of AML. Sci Rep. 2015;5:11295. doi:10.1038/srep11295
28. Yamashita T, Kamada H, Kanasaki S, et al. Epidermal growth factor receptor localized to exosome membranes as a possible biomarker for lung cancer diagnosis. Pharmazie. 2013;68:969–973.
29. Kim MS, Haney M, Zhao Y, et al. Development of exosome-encapsulated paclitaxel to overcome MDR in cancer cells. Nanomedicine. 2016;12:655–664.
30. Ju S, Mu J, Dokland T, et al. Grape exosome-like nanoparticles induce intestinal stem cells and protect mice from DSS-induced colitis. Mol Ther. 2013;21. doi:10.1038/mt.2013.64
31. Hood J, Scott M, Wickline S. Maximizing exosome colloidal stability following electroporation. Anal Biochem. 2013;448:41–49.
32. Saari H, Lazaro-Ibañez E, Viitala T, Vuorimaa E, Siljander P, Yliperttula M. Microvesicle- and exosome-mediated drug delivery enhances the cytotoxicity of Paclitaxel in autologous prostate cancer cells. J Control Release. 2015;220. doi:10.1016/j.jconrel.2015.09.031
33. Mu J, Zhuang X, Wang Q, et al. Interspecies communication between plant and mouse gut host cells through edible plant derived exosome-like nanoparticles. Mol Nutr Food Res. 2014;58. doi:10.1002/mnfr.201300729
34. King HW, Michael MZ, Gleadle JM. Hypoxic enhancement of exosome release by breast cancer cells. BMC Cancer. 2012;12(1):421. doi:10.1186/1471-2407-12-421
35. Helwa I, Cai J, Drewry MD, et al. A comparative study of serum exosome isolation using differential ultracentrifugation and three commercial reagents. PLoS One. 2017;12(1):e0170628. doi:10.1371/journal.pone.0170628
36. Hannafon BN, Trigoso YD, Calloway C, et al. Plasma exosome microRNAs are indicative of breast cancer. Breast Cancer Res. 2016;18:90. doi:10.1186/s13058-016-0753-x
37. Rodríguez M, Silva J, López‐Alfonso A, et al. Different exosome cargo from plasma/bronchoalveolar lavage in non-small-cell lung cancer. Genes Chromosomes Cancer. 2014;53:713–724.
38. Madhavan B, Yue S, Galli U, et al. Combined evaluation of a panel of protein and miRNA serum‐exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int J Cancer. 2015;136(11):2616–2627. doi:10.1002/ijc.29324
39. Shender VO, Pavlyukov MS, Ziganshin RH, et al. Proteome–metabolome profiling of ovarian cancer ascites reveals novel components involved in intercellular communication. Mol Cell Proteomics. 2014;13(12):3558–3571. doi:10.1074/mcp.M114.041194
40. Im H, Shao H, Park YI, et al. Label-free detection and molecular profiling of exosomes with a nano-plasmonic sensor. Nat Biotechnol. 2014;32(5):490. doi:10.1038/nbt.2886
41. Sun Y, Huo C, Qiao Z, et al. Comparative proteomic analysis of exosomes and microvesicles in human saliva for lung cancer. J Proteome Res. 2018;17(3):1101–1107. doi:10.1021/acs.jproteome.7b00770
42. Li Y, Zhang Y, Qiu F, Qiu Z. Proteomic identification of exosomal LRG1: a potential urinary biomarker for detecting NSCLC. Electrophoresis. 2011;32(15):1976–1983. doi:10.1002/elps.201000598
43. Huang A, Dong J, Li S, et al. Exosomal transfer of vasorin expressed in hepatocellular carcinoma cells promotes migration of human umbilical vein endothelial cells. Int J Biol Sci. 2015;11(8):961. doi:10.7150/ijbs.11943
44. Nilsson J, Skog J, Nordstrand A, et al. Prostate cancer-derived urine exosomes: a novel approach to biomarkers for prostate cancer. Br J Cancer. 2009;100(10):1603–1607. doi:10.1038/sj.bjc.6605058
45. Tian Y, Ma L, Gong M, et al. Protein profiling and sizing of extracellular vesicles from colorectal cancer patients via flow cytometry. ACS Nano. 2018;12(1):671–680. doi:10.1021/acsnano.7b07782
46. Zhang H, Deng T, Liu R, et al. Exosome-delivered EGFR regulates liver microenvironment to promote gastric cancer liver metastasis. Nat Commun. 2017;8(1):1–11. doi:10.1038/s41467-016-0009-6
47. Madhankumar A, Mrowczynski OD, Patel SR, et al. Interleukin-13 conjugated quantum dots for identification of glioma initiating cells and their extracellular vesicles. Acta Biomater. 2017;58:205–213. doi:10.1016/j.actbio.2017.06.002
48. Skog J, Würdinger T, Van Rijn S, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 2008;10(12):1470–1476. doi:10.1038/ncb1800
49. Peinado H, Alečković M, Lavotshkin S, et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med. 2012;18(6):883. doi:10.1038/nm.2753
50. Zhou J, Gong G, Tan H, et al. Urinary microRNA-30a-5p is a potential biomarker for ovarian serous adenocarcinoma. Oncol Rep. 2015;33(6):2915–2923. doi:10.3892/or.2015.3937
51. Taylor D, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol. 2008;110:13–21. doi:10.1016/j.ygyno.2008.04.033
52. Cappellesso R, Tinazzi A, Giurici T, et al. Programmed cell death 4 and micro RNA 21 inverse expression is maintained in cells and exosomes from ovarian serous carcinoma effusions. Cancer Cytopathol. 2014;122(9):685–693. doi:10.1002/cncy.21442
53. Fang T, Lv H, Lv G, et al. Tumor-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat Commun. 2018;9(1):1–13. doi:10.1038/s41467-017-02583-0
54. Thakur BK, Zhang H, Becker A, et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 2014;24(6):766–769. doi:10.1038/cr.2014.44
55. Rodríguez M, Bajo-Santos C, Hessvik NP, et al. Identification of non-invasive miRNAs biomarkers for prostate cancer by deep sequencing analysis of urinary exosomes. Mol Cancer. 2017;16(1):156. doi:10.1186/s12943-017-0726-4
56. Bryant R, Pawlowski T, Catto J, et al. Changes in circulating microRNA levels associated with prostate cancer. Br J Cancer. 2012;106(4):768–774. doi:10.1038/bjc.2011.595
57. Huang X, Yuan T, Liang M, et al. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant prostate cancer. Eur Urol. 2015;67(1):33–41. doi:10.1016/j.eururo.2014.07.035
58. McKiernan J, Donovan MJ, O’Neill V, et al. A novel urine exosome gene expression assay to predict high-grade prostate cancer at initial biopsy. JAMA Oncol. 2016;2(7):882–889. doi:10.1001/jamaoncol.2016.0097
59. Li Y, Zheng Q, Bao C, et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 2015;25(8):981–984. doi:10.1038/cr.2015.82
60. Teng Y, Ren Y, Hu X, et al. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat Commun. 2017;8(1):1–16. doi:10.1038/ncomms14448
61. Joshi GK, Deitz-McElyea S, Liyanage T, et al. Label-free nanoplasmonic-based short noncoding RNA sensing at attomolar concentrations allows for quantitative and highly specific assay of microRNA-10b in biological fluids and circulating exosomes. ACS Nano. 2015;9(11):11075–11089. doi:10.1021/acsnano.5b04527
62. Lai X, Wang M, McElyea SD, Sherman S, House M, Korc M. A microRNA signature in circulating exosomes is superior to exosomal glypican-1 levels for diagnosing pancreatic cancer. Cancer Lett. 2017;393:86–93. doi:10.1016/j.canlet.2017.02.019
63. Zhao R, Zhang Y, Zhang X, et al. Exosomal long noncoding RNA HOTTIP as potential novel diagnostic and prognostic biomarker test for gastric cancer. Mol Cancer. 2018;17(1):68. doi:10.1186/s12943-018-0817-x
64. Qu L, Ding J, Chen C, et al. Exosome-transmitted lncARSR promotes sunitinib resistance in renal cancer by acting as a competing endogenous RNA. Cancer Cell. 2016;29(5):653–668. doi:10.1016/j.ccell.2016.03.004
65. Lou G, Song X, Yang F, et al. Exosomes derived from miR-122-modified adipose tissue-derived MSCs increase chemosensitivity of hepatocellular carcinoma. J Hematol Oncol. 2015;8(1):122. doi:10.1186/s13045-015-0220-7
66. Conigliaro A, Costa V, Dico AL, et al. CD90+ liver cancer cells modulate endothelial cell phenotype through the release of exosomes containing H19 lncRNA. Mol Cancer. 2015;14(1):155. doi:10.1186/s12943-015-0426-x
67. Sohn W, Kim J, Kang SH, et al. Serum exosomal microRNAs as novel biomarkers for hepatocellular carcinoma. Exp Mol Med. 2015;47(9):e184. doi:10.1038/emm.2015.68
68. Sugimachi K, Matsumura T, Hirata H, et al. Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. Br J Cancer. 2015;112(3):532–538. doi:10.1038/bjc.2014.621
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.