Back to Journals » OncoTargets and Therapy » Volume 13
Progression of Thyroid Carcinoma Is Promoted by the m6A Methyltransferase METTL3 Through Regulating m6A Methylation on TCF1
Authors Wang K, Jiang L, Zhang Y, Chen C
Received 15 October 2019
Accepted for publication 4 February 2020
Published 21 February 2020 Volume 2020:13 Pages 1605—1612
DOI https://doi.org/10.2147/OTT.S234751
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Leo Jen-Liang Su
Kejin Wang, Lin Jiang, Yan Zhang, Chao Chen
Department of Head and Neck Surgery, Cancer Hospital of University of Chinese Academy of Sciences, Key Laboratory of Head & Neck Cancer Translational Research of Zhejiang Province, Hangzhou, People’s Republic of China
Correspondence: Chao Chen
Department of Head and Neck Surgery, Cancer Hospital of University of Chinese Academy of Sciences, Key Laboratory of Head & Neck Cancer Translational Research of Zhejiang Province, No. 1, Banshan East Road, Hangzhou 310022, People’s Republic of China
Tel +86 571 8812 2222
Email [email protected]
Objective: This study aims to uncover the progression of thyroid carcinoma influenced by the m6A methyltransferase METTL3 through regulating m6A methylation on TCF1 mRNA and the activated Wnt pathway.
Methods: Thyroid carcinoma tissues and paracancerous ones were collected for detecting levels of METTL3 and TCF1. Potential correlation between levels of METTL3 and TCF1 was analyzed by Pearson analysis. Survival of thyroid carcinoma patients influenced by METTL3 level was assessed by Kaplan–Meier method. Regulatory effect of METTL3 on migratory ability in TPC-1 cells was examined by wound healing assay. The interaction between METTL3 with TCF1 and IGF2BP2 was verified by RNA-Binding Protein Immunoprecipitation (RIP) assay. Meanwhile, the activity of the Wnt pathway was reflected by TOP/FOP-Flash. At last, rescue experiments were conducted to clarify the involvement of TCF1 in phenotype changes of thyroid carcinoma cells that were regulated by METTL3.
Results: METTL3 and TCF1 were upregulated in thyroid carcinoma. Similarly, METTL3 was highly expressed in thyroid carcinoma cells as well. Kaplan–Meier method uncovered poor prognosis in thyroid carcinoma patients expressing a high level of METTL3. Silence of METTL3 inhibited migratory ability and Wnt activity in TPC-1 cells. RIP assay confirmed the interaction between TCF1 and METTL3 or IGF2BP2. Moreover, METTL3 positively regulated the enrichment abundance of TCF1 in anti-IGF2BP2. Rescue experiments demonstrated that TCF1 was responsible for METTL3-regulated thyroid carcinoma progression via the m6A methylation.
Conclusion: Upregulated m6A methyltransferase METTL3 promotes the progression of thyroid carcinoma through m6A methylation on TCF1.
Keywords: thyroid carcinoma, m6A, methyltransferase METTL3, TCF1, migration
Introduction
The prevalence of thyroid carcinoma accounts for 1% in malignancies. In particular, the subtype of papillary carcinoma with low malignancy and well prognosis is the most common one.1 The incidence of thyroid carcinoma varies a lot in different regions, races and genders. Females are more affected by thyroid carcinoma than males, with the female-male incidence ratio of (2–4): 1.2,3 Thyroid carcinoma occurs in any age, which typically affects young adults.4 It is necessary to clarify the molecular mechanism underlying the occurrence and progression of thyroid carcinoma.
m6A methylation is the most abundant mRNA modification in eukaryotes, which is involved in various biological processes, including tissue development, stem cell differentiation, DNA damage repair, etc.5,6 In addition, m6A methylation is closely linked to cancer.7 Cancer drugs targeting m6A methylation are promising in the clinical application.8,9 Abnormally expressed enzymes involved in m6A methylation may result in pathological conditions.10,11 Under the assistance of methyltransferases (ie METTL3, METTL14 and WTAP), methylation modification presents on the N6-Methyladenosine.7,12
RNAs are the core components of the genome and the linkage between genetic information (DNAs) and proteins.13,14 METTL3 is widely expressed and highly conserved in the majority of eukaryotes.15 During the tumor progression, m6A methylation exerts a dual role, which could either promote or alleviate the malignant progression.16 It is reported that m6A methylation is involved in many types of cancer, and it displays certain diagnostic and therapeutic potentials.17,18
Transcription factors are protein molecules that can specifically bind to upstream of the 5ʹ end of the gene, thereby ensuring target gene expression with a specific intensity.19 TCF1 is expressed in multiple isoforms. All isoforms of TCF1 share the common HDAC and DNA binding domains.20,21 This study aims to uncover the progression of thyroid carcinoma promoted by the m6A methyltransferase METTL3 through regulating m6A methylation on TCF1 mRNA.
Methods
Subjects
Thyroid carcinoma tissues and paracancerous ones were surgically resected from thyroid carcinoma patients undergoing surgery in Cancer Hospital of the University of Chinese Academy of Sciences. None of the enrolled patients had the medical history of preoperative anti-tumor therapy. Tissue samples were immediately frozen in liquid nitrogen and stored at −80°C. Patients and their families in this study have been fully informed. This study was approved by the Ethics Committee of Cancer Hospital of the University of Chinese Academy of Sciences, and conducted in accordance with the Declaration of Helsinki. All samples received written informed consent from the patients.
Cell Culture
Normal thyroid cell line (Nthy-ori3-1) and thyroid carcinoma cell lines (TPC-1 and BC-PAP) were purchased from ATCC (Manassas, VA, USA). Cells were cultured in low-glucose Dulbecco’s modified eagle medium (DMEM) (Thermo Fisher Scientific, Waltham, MA, USA) containing 10% fetal bovine serum (FBS) (Gibco, Rockville, MD, USA) and 1% penicillin-streptomycin.
Plasmid Construction and Cell Transfection
TCF1 cDNA was cloned into pcDNA3.0, that was, pcDNA3.0-TCF1. Cells were transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) at 60% confluence. Fresh medium was replaced 6 hrs later. Transfected cells for 48 h were collected for functional experiments.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
TRIzol (Invitrogen, Carlsbad, CA, USA) was applied for isolating cellular RNA, which was quantified using a spectrometer. RNA was reversely transcribed into cDNA using the PrimeScript RT reagent Kit (Takara, Tokyo, Japan). SYBR Premix Ex Taq TM (Takara, Tokyo, Japan) was utilized for qRT-PCR for 35–40 cycles.
Western Blot
Cellular protein was isolated and electrophoresed. Protein samples were loaded on polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA, USA). Subsequently, non-specific antigens were blocked in 5% skim milk for 2 hrs. Membranes were reacted with primary and secondary antibodies for the indicated time. Band exposure and analyses were finally conducted.
CCK-8 Assay
1×103 cells per well were inoculated in a 96-well plate. After cell culture for 6, 24, 48, 72 and 96 h, 10 μL of CCK-8 reagent was added in each well, respectively. Following 1-h incubation at 37°C, optimal density at 450 nm was measured. Each group had five replicates.
RNA-Binding Protein Immunoprecipitation (RIP) Assay
RIP assay was performed following the procedures of Millipore Magna RIP Kit. Cells were incubated with the input, corresponding antibodies or anti–IgG at 4°C overnight. A protein-RNA complex was obtained after capturing intracellular specific proteins by the antibody. Subsequently, proteins were digested by proteinase K to extract RNAs. During the experiment, the magnetic beads were repeatedly washed with RIP washing buffer to remove non-specific adsorption as much as possible. The immunoprecipitant RNAs were finally quantified by qRT-PCR.
Wound Healing Assay
All instruments were required for 30-min ultraviolet radiation, including a ruler, a marker pen and a 200 μL pipette. Five lines with 0.5–1 cm interval were depicted on the back of a 6-well plate using a marker pen. Subsequently, 5×105 cells/well were inoculated in a 6-well plate. An artificial wound was created in the confluent cell monolayer using a 200 μL pipette tip. Wound closure was captured at 0 and 24 h, respectively.
TOP/FOP-Flash Luciferase Reporter Assay
TOP/FOP-Flash luciferase reporter assay was conducted as previously reported.18 Cells were inoculated in a 24-well plate and transfected with TOP/FOP plasmids (Simo Biomedical Technology, Shanghai, China). Cells were then lysed for determining relative luciferase activity.
Statistical Analyses
SPSS 16.0 (SPSS IBM, Armonk, NY USA) was used for all statistical analyses. Data were expressed as mean ± SD. A comparison between multiple groups was done using a one-way ANOVA test, followed by Post Hoc Test (Least Significant Difference). Survival analysis was conducted by Kaplan–Meier method, followed by Log rank test for comparing differences between curves. Pearson analysis was applied for assessing the relationship between expression levels of METTL3 and TCF1 in thyroid carcinoma tissues. P<0.05 indicated the significant difference.
Results
METTL3 and TCF1 Were Upregulated in Thyroid Carcinoma
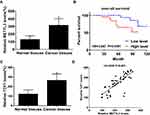
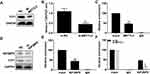
In thyroid carcinoma tissues (n=48), METTL3 and TCF1 were markedly upregulated than those of paracancerous ones (n=24) (Figure 1A and C). Kaplan–Meier curves revealed poor survival in thyroid carcinoma patients expressing a high level of METTL3 (Figure 1B). In addition, expression levels of METTL3 and TCF1 were positively correlated in thyroid carcinoma tissues (Figure 1D). After analyzing their clinical data, it was found that METTL3 level was correlated to tumor size, TNM staging and lymphatic metastasis rate in thyroid carcinoma patients, while unrelated to age and sex (Table 1). It is suggested that METTL3 and TCF1 were involved in the progression of thyroid carcinoma.
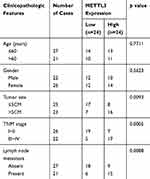 |
Table 1 Association of METTL3 Expression with Clinicopathologic Characteristics of Thyroid Carcinoma |
Knockdown of METTL3 Suppressed Migratory Ability in Thyroid Carcinoma
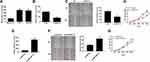
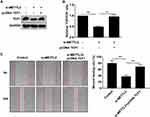
Consistently, METTL3 was highly expressed in thyroid carcinoma cells than that of normal thyroid cells (Figure 2A). Transfection efficacy of si-METTL3 was firstly verified in TPC-1 cells (Figure 2B). As wound healing assay results reflected, migratory ability was remarkably inhibited in TPC-1 cells transfected with si-METTL3 than those of controls (Figure 2C). Subsequently, we found that transfection of si-METTL3 markedly decreased proliferative ability in TPC-1 cells (Figure 2D). On the contrary, transfection of pcDNA-METTL3 effectively upregulated METTL3 level in thyroid carcinoma cells (Figure 2E). Overexpression of METTL3, as expected, enhanced migratory ability (Figure 2F) and proliferative rate (Figure 2G) in TPC-1 cells. It is considered that METTL3 aggravated the malignant progression of thyroid carcinoma through stimulating tumor cell metastasis.
METTL3 Induced m6A Methylation on TCF1 mRNA and Thus Upregulated Protein Level of TCF1
Interestingly, transfection of si-METTL3 downregulated protein level of TCF1 in TPC-1 cells (Figure 3A). Moreover, knockdown of METTL3 decreased TOP/FOP ratio, suggesting the blocked activity of Wnt (Figure 3B). We, therefore, speculated their potential interaction. As RIP assay illustrated, TCF1 was mainly enriched in anti-METTL3 than that of anti–IgG (Figure 3C). Protein level of IGF2BP2 was downregulated in TPC-1 cells transfected with si–IGF2BP2 (Figure 3D). In TPC-1 cells, protein level of TCF1 was downregulated after transfection of si–IGF2BP2 (Figure 3D). RIP assay further verified the interaction between TCF1 and IGF2BP2 (Figure 3E). More importantly, knockdown of METTL3 greatly decreased the enrichment abundance of TCF1 in anti–IGF2BP2 (Figure 3F). The above data demonstrated that METTL3 induced m6A methylation on TCF1 mRNA and thus upregulated protein level of TCF1.
METTL3 Activated the Wnt Pathway Through TCF1
Since the interaction between METTL3 and TCF1 has been confirmed, we speculated whether TCF1 was involved in the progression of thyroid carcinoma regulated by METTL3. We first tested the transfection efficacy of pcDNA TCF1 by Western blot (Figure 4A). TOP/FOP-Flash results pointed out that overexpression of TCF1 partially reversed the inhibitory effect of downregulated METTL3 on the activity of Wnt (Figure 4B). Similarly, the inhibited migratory ability in thyroid carcinoma cells transfected with si-METTL3 was abolished by overexpression of TCF1 (Figure 4C). Collectively, METTL3 activated the Wnt pathway through m6A methylation on TCF1 mRNA, thus aggravating the progression of thyroid carcinoma.
Discussion
Thyroid carcinoma is a highly prevalent endocrine tumor, with the incidence of 7.7/100,000. In recent years, its incidence has been risen globally.22 Thyroid carcinoma has multiple subtypes presenting different clinical and pathological features.23 Target drugs on specific genes related to thyroid carcinoma, including BRAF and RET, contribute to improve the clinical outcome of affected people.24 After active treatment of surgical resection combined radioactive iodine and thyroid hormone suppression therapy, thyroid carcinoma patients usually achieve a satisfied prognosis.25
Previous studies have shown that RNA methylation accounts for over 60% of all RNA modifications, including m6A, m1A and m5C, etc. Among them, m6A is the most popular one.10,26 In eukaryotes, the 5ʹ end cap and the 3ʹ end poly(A) modification exert an important role in transcriptional regulation, while the internal modification of mRNA contributes to maintain mRNA stability.27 Currently, three kinds of enzymes have been identified in m6A methylation, including demethylases, methylases and methylation recognition enzymes.28 Among them, m6A methyltransferase complex is composed of METTL3, METTL14 and WTAP.29 m6A methylation is discovered to affect tumor progression.30 In this paper, METTL3 was found to be upregulated in thyroid carcinoma tissues. METTL3 level predicted a poor prognosis in thyroid carcinoma patients as an oncogene. In vitro experiments further confirmed its promotive effect on migratory ability in thyroid carcinoma cells.
The initiation process of eukaryotic transcription is very complicated and often requires the assistance of various protein factors. A transcription initiation complex formed by transcription factor and RNA polymerase II participates in the process of transcription initiation.31 TCF1 is encoded by the Tcf7 gene, which is the downstream effector of the classical Wnt pathway.32,33 Our analyses showed that TCF1 was upregulated in thyroid carcinoma tissues. Importantly, METTL3 induced m6A methylation on TCF1 mRNA, thus aggravating the progression of thyroid carcinoma by activating the Wnt pathway.
Conclusions
Upregulated m6A methyltransferase METTL3 promotes the progression of thyroid carcinoma through m6A methylation on TCF1 mRNA.
Funding
International Cooperation Project of the Zhejiang Basic Research Program (LGJ18H160002) funded this study.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Cabanillas ME, McFadden DG, Durante C. Thyroid cancer. Lancet. 2016;388(10061):2783–2795. doi:10.1016/S0140-6736(16)30172-6
2. Roman BR, Morris LG, Davies L. The thyroid cancer epidemic, 2017 perspective. Curr Opin Endocrinol Diabetes Obes. 2017;24(5):332–336. doi:10.1097/MED.0000000000000359
3. Carling T, Udelsman R. Thyroid cancer. Annu Rev Med. 2014;65:125–137. doi:10.1146/annurev-med-061512-105739
4. Araque D, Bleyer A, Brito JP. Thyroid cancer in adolescents and young adults. Future Oncol. 2017;13(14):1253–1261. doi:10.2217/fon-2017-0024
5. Yu J, Chen M, Huang H, et al. Dynamic m6A modification regulates local translation of mRNA in axons. Nucleic Acids Res. 2018;46(3):1412–1423. doi:10.1093/nar/gkx1182
6. Liu Z, Zhang J. Most m6A RNA modifications in protein-coding regions are evolutionarily unconserved and likely nonfunctional. Mol Biol Evol. 2017. doi:10.1093/molbev/msx320
7. Liu ZX, Li LM, Sun HL, Liu SM. Link between m6A modification and cancers. Front Bioeng Biotechnol. 2018;6:89. doi:10.3389/fbioe.2018.00089
8. Zhang SY, Zhang SW, Liu L, Meng J, Huang Y. m6A-driver: identifying context-specific mRNA m6A methylation-driven gene interaction networks. PLoS Comput Biol. 2016;12(12):e1005287. doi:10.1371/journal.pcbi.1005287
9. Chen M, Wei L, Law CT, et al. RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology. 2018;67(6):2254–2270. doi:10.1002/hep.29683
10. Sun T, Wu R, Ming L. The role of m6A RNA methylation in cancer. Biomed Pharmacother. 2019;112:108613. doi:10.1016/j.biopha.2019.108613
11. Hong K. Emerging function of N6-methyladenosine in cancer. Oncol Lett. 2018;16(5):5519–5524. doi:10.3892/ol.2018.9395
12. Dai D, Wang H, Zhu L, Jin H, Wang X. N6-methyladenosine links RNA metabolism to cancer progression. Cell Death Dis. 2018;9(2):124. doi:10.1038/s41419-017-0129-x
13. Re A, Joshi T, Kulberkyte E, Morris Q, Workman CT. RNA-protein interactions: an overview. Methods Mol Biol. 2014;1097:491–521. doi:10.1007/978-1-62703-709-9_23
14. Cammas A, Millevoi S. RNA G-quadruplexes: emerging mechanisms in disease. Nucleic Acids Res. 2017;45(4):1584–1595. doi:10.1093/nar/gkw1280
15. Lin S, Choe J, Du P, Triboulet R, Gregory RI. The m(6)A methyltransferase METTL3 promotes translation in human cancer cells. Mol Cell. 2016;62(3):335–345. doi:10.1016/j.molcel.2016.03.021
16. Li X, Tang J, Huang W, et al. The M6A methyltransferase METTL3: acting as a tumor suppressor in renal cell carcinoma. Oncotarget. 2017;8(56):96103–96116. doi:10.18632/oncotarget.21726
17. Cai X, Wang X, Cao C, et al. HBXIP-elevated methyltransferase METTL3 promotes the progression of breast cancer via inhibiting tumor suppressor let-7g. Cancer Lett. 2018;415:11–19. doi:10.1016/j.canlet.2017.11.018
18. Du M, Zhang Y, Mao Y, et al. MiR-33a suppresses proliferation of NSCLC cells via targeting METTL3 mRNA. Biochem Biophys Res Commun. 2017;482(4):582–589. doi:10.1016/j.bbrc.2016.11.077
19. Papavassiliou KA, Papavassiliou AG. Transcription factor drug targets. J Cell Biochem. 2016;117(12):2693–2696. doi:10.1002/jcb.25605
20. Gullicksrud JA, Li F, Xing S, et al. Differential requirements for Tcf1 long isoforms in CD8(+) and CD4(+) T cell responses to acute viral infection. J Immunol. 2017;199(3):911–919. doi:10.4049/jimmunol.1700595
21. Xing S, Li F, Zeng Z, et al. Tcf1 and Lef1 transcription factors establish CD8(+) T cell identity through intrinsic HDAC activity. Nat Immunol. 2016;17(6):695–703. doi:10.1038/ni.3456
22. Yip L, Sosa JA. Molecular-directed treatment of differentiated thyroid cancer: advances in diagnosis and treatment. JAMA Surg. 2016;151(7):663–670. doi:10.1001/jamasurg.2016.0825
23. Chmielik E, Rusinek D, Oczko-Wojciechowska M, et al. Heterogeneity of thyroid cancer. Pathobiology. 2018;85(1–2):117–129. doi:10.1159/000486422
24. Wang J, Chi S, Huang Z, et al. Comprehensive characterization of differentially expressed genes in thyroid cancer. Future Oncol. 2017;13(24):2159–2169. doi:10.2217/fon-2017-0168
25. Callender GG, Carling T, Christison-Lagay E, Udelsman R. Surgery for thyroid cancer. Endocrinol Metab Clin North Am. 2014;43(2):443–458. doi:10.1016/j.ecl.2014.02.011
26. Meyer KD, Jaffrey SR. Rethinking m(6)A readers, writers, and erasers. Annu Rev Cell Dev Biol. 2017;33:319–342. doi:10.1146/annurev-cellbio-100616-060758
27. Bartosovic M, Molares HC, Gregorova P, Hrossova D, Kudla G, Vanacova S. N6-methyladenosine demethylase FTO targets pre-mRNAs and regulates alternative splicing and 3ʹ-end processing. Nucleic Acids Res. 2017;45(19):11356–11370. doi:10.1093/nar/gkx778
28. Du Y, Hou G, Zhang H, et al. SUMOylation of the m6A-RNA methyltransferase METTL3 modulates its function. Nucleic Acids Res. 2018;46(10):5195–5208. doi:10.1093/nar/gky156
29. Yue Y, Liu J, He C. RNA N6-methyladenosine methylation in post-transcriptional gene expression regulation. Genes Dev. 2015;29(13):1343–1355. doi:10.1101/gad.262766.115
30. Engel M, Chen A. The emerging role of mRNA methylation in normal and pathological behavior. Genes Brain Behav. 2018;17(3):e12428. doi:10.1111/gbb.12428
31. Simicevic J, Deplancke B. Transcription factor proteomics – tools, applications, and challenges. Proteomics. 2017;17:3–4. doi:10.1002/pmic.201600317
32. Jeevan-Raj B, Gehrig J, Charmoy M, et al. The transcription factor Tcf1 contributes to normal NK cell development and function by limiting the expression of granzymes. Cell Rep. 2017;20(3):613–626. doi:10.1016/j.celrep.2017.06.071
33. Hrckulak D, Kolar M, Strnad H, Korinek V. TCF/LEF transcription factors: an update from the internet resources. Cancers (Basel). 2016;8:7. doi:10.3390/cancers8070070
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.