Back to Journals » Pharmacogenomics and Personalized Medicine » Volume 10
Prevalence of clinically actionable genotypes and medication exposure of older adults in the community
Authors Daneshi N, Holliday E, Hancock S , Schneider JJ, Scott RJ, Attia J, Milward EA
Received 2 October 2016
Accepted for publication 6 December 2016
Published 27 January 2017 Volume 2017:10 Pages 17—27
DOI https://doi.org/10.2147/PGPM.S123719
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Martin H Bluth
Nilofar Daneshi,1,2 Elizabeth Holliday,3,4 Stephen Hancock,3,4 Jennifer J Schneider,1,2 Rodney J Scott,1,2,5 John Attia,3,4 Elizabeth A Milward1,2
1Faculty of Health and Medicine, School of Biomedical Sciences and Pharmacy, 2Faculty of Health and Medicine, Centre for Bioinformatics, Biomarker Discovery and Information-Based Medicine, The University of Newcastle, Callaghan, 3Clinical Research Design, IT and Statistical Support Unit, Hunter Medical Research Institute, 4Centre for Clinical Epidemiology and Biostatistics, School of Medicine and Public Health, Faculty of Health, University of Newcastle, 5Hunter Area Pathology Service, John Hunter Hospital, Newcastle, NSW, Australia
Abstract: This study analyzed clinically actionable pharmacogenotypes for clopidogrel, warfarin, statins, thiopurines, and tacrolimus using microarray data for 2121 participants (55–85 years) from the Australian Hunter Community Study (HCS). At least 74% of participants (95% confidence interval [CI]: 72%–76%) had strong level evidence for at least one medium- or high-risk actionable genotype that would trigger a change in standard therapy under current international recommendations. About 14% of these participants (95% CI: 12%–16%) were taking medication potentially affected by the genotype in question. Furthermore, ~2.6% of all participants with medication data (95% CI: 1.4%–3.8%) had a high-risk clinically actionable genotype for a medication to which they were exposed. This represents a considerable number of people at the population level. Although relationships between genotype and health outcomes remain contentious, pharmacogenotyping of multiple variants simultaneously may have considerable potential to improve medication safety and efficacy for older people in the community.
Keywords: actionable genotype, community, older adults, pharmacogenomics, pre-emptive genotyping, single-nucleotide polymorphism
Introduction
Clinical pharmacogenomics involves predicting drug response and delivering tailored therapy based on each individual’s unique genome.1–4 On the basis of their genotype profile – single-nucleotide polymorphisms (SNPs) or other genetic variants – individuals may benefit from a change in standard therapy that may involve an increase or decrease in dose or an alternative treatment. Genotypes for which there is considered to be robust evidence justifying changes in standard therapy are sometimes termed “clinically actionable”.5–7
Systematic reviews of published studies by internationally recognized bodies such as the Pharmacogenomics Knowledgebase (PharmGKB),6 the Clinical Pharmacogenetics Implementation Consortium (CPIC),8 and the US Food and Drug Administration (FDA)9 have provided compelling evidence for a number of drug–gene combinations for which nonstandard medication regimens may be recommended based on genotype. Ideally, to avoid delays in clinical decision-making, pharmacogenomic test results would be available in the earliest stages of the clinical process to aid the prescribing decision. This would allow the health care professional to simultaneously assess the patient’s genotype along with other clinical factors to better predict patients’ response to drugs.2,10,11 This is one facet of what is now being termed “precision medicine” and could ultimately lead to better efficacy, cost–benefit ratio, and fewer adverse effects.12–15 Some medical centers are already using array-based testing or other technologies to implement “pre-emptive” clinical pharmacogenomics, with results available to the clinician prior to prescribing, as opposed to indication-based testing based on medication exposure.2,5,16–18
The Pharmacogenomic Resource for Enhanced Decisions in Care and Treatment (PREDICT) program at Vanderbilt University Medical Center is implementing pharmacogenomics into clinical practice by focusing on common clinical gene variants that may lead to important drug–gene interactions (DGIs). In 2014, a pre-emptive, panel-based pharmacogenomic study arising from the PREDICT program was published by Van Driest et al.5
This study investigated gene variants that were deemed clinically actionable based on institutionally approved clinical decision support advisors for five common DGIs in a clinical group of 10,044 cardiovascular disease (CVD) or cancer patients of mixed race.5 It was found that 91% had at least one actionable genotype and at least 5% had one high-risk genotype. Of the patients who had one or more actionable genotypes, 42% were exposed to a risk-associated medication or medication class.
Cardiac and cancer patients such as those investigated in the PREDICT study are likely to be prescribed a relatively high number of medications. This increases the chance of exposure to a medication for which genotype-driven clinical action would be recommended. Ideally, as noted above, pharmacogenomic test results should be available pre-emptively before prescription. Yet, pre-emptive genotyping may encounter opposition on the ground that is inappropriate to perform pharmacogenomic testing before patients present in the clinic, since many results will never become actionable for people not prescribed relevant medication during their lifetime.
To provide more insight into the extent to which earlier pre-emptive genotyping may be justified, before patients attend a hospital, we have performed a comparable study in an elderly community cohort from the Hunter Valley region in Australia. This group was selected both because genome-wide microarray data were already available for many of the participants and because there are higher rates of polypharmacy among elderly people,19–21 who are also likely to have higher rates of CVD and cancer medication usage than the general population, while not as high as those attending hospital clinics.
This study provides important new information on the proportions of people in the general population likely to have combinations of pharmacogenomic variants that may eventually affect their health care. The findings substantiate results from clinical samples from different populations,5,22 showing that most people are likely to have clinically actionable pharmacogenotypes, irrespective of the population under consideration.
This paper also provides insights into the proportions of individuals on select cardiovascular drugs that prescribers may expect to have clinically actionable SNPs, including detailed assessment of warfarin genotype actionability based on FDA recommendations that take into account potential interactions between the CYP2C9 and VKORC1 genotypes. We show that the proportions with clinically actionable genotypes are substantial even when just considering a small number of drugs, providing evidence that pre-emptive testing has the potential to be of substantial value in improving the lifetime health care of people in the general population.
Methods
The Hunter Community Study (HCS) is an ongoing initiative investigating factors affecting the health of people of European ancestry from Newcastle, New South Wales, Australia, and the surrounding Hunter region. Details of the HCS have been reported previously.23,24 Briefly, ethics approval for the study was obtained from the Hunter New England Local Health District and University of Newcastle Human Research Ethics Committees. Initially, 9748 potential participants were randomly selected from the electoral roll, of whom 7575 subjects responded. Of these, 258 were not eligible to participate (148 did not speak English, 18 had moved to an aged care facility, and the remaining 92 were deceased). From the remaining cohort, 3253 gave informed written consent to participate in the study. Information was available on baseline social demographics, age and sex distribution, and medical history, including self-reported medication data.24 We considered the five drugs used in the PREDICT study – clopidogrel, warfarin, statins, thiopurine, and tacrolimus.5 For the purpose of this study, all statins were included when determining statin exposure rates for participants.
This present study examined 2121 HCS participants for whom genotyping data had previously been obtained using Affymetrix Axiom Kaiser microarrays.25 These comprised 1060 females and 1061 males (Table 1). The Axiom Kaiser arrays were not specifically tailored for pharmacogenomics, and only one gene variant (rs4149056) assessed in the PREDICT study was directly measured by this array. The other gene variants (rs4244285, rs4149056, rs1799853, rs1057910, rs9923231, rs1800460, rs1142345, and rs776746) assessed in the PREDICT study were imputed from two reference panels, HapMap Phase II and the 1000 Genomes European reference panel. All other SNPs looked at in the PREDICT study for the five DGIs were checked for potential proxies with a relaxed r2 value that might be highly correlated and present in our data set; none could be found.
  | Table 1 Group demographics and medication exposure Note: *A total of 2121 participants had genotype data, and of these, medication history was available for 1963 participants. |
In total, three SNPs (rs1346268, rs2231142, and rs4149056) were directly genotyped using the Affymetrix Axiom Kaiser array. Imputation through the 1000 Genomes and HapMap Phase II European reference panels was used for 10 other SNPs (rs776746, rs1057910, rs1142345, rs1719247, rs1799853, rs1800460, rs2032582, rs4244285, rs4693075, and rs9923231). All SNPs had high imputation accuracy, with the estimated genotype correlation to the true genotype (imputation r2) being >0.95. Medication data were available for 1963 participants.24
Clinically actionable genotypes for the five drugs of interest were determined according to PharmGKB and CPIC guidelines, based on previously published studies in the literature.5,6,8,26–33 We first interrogated actionable DGIs relevant to populations of European descent and deemed to have a strong level of evidence under CPIC guidelines. We then went on to consider other actionable DGIs for which CPIC guidelines cite a moderate level of evidence.
For most of the actionable genotypes investigated, the SNPs involved are the minor allelic variant; however, for three SNPs relevant to statins, the actionable allele is the major allele, present in over half of the population samples examined in the studies considered by PharmGKB/CPIC and in our study. These SNPs are rs2231142 on ABCG2, listed in PharmGKB as the major allele for both the “white” and “Asian” races, together with rs1346268 on GATM (55.5%) and rs1719247 upstream of GATM (55.4%), both listed in PharmGKB as major alleles for the white race (major alleles for other ethnicities are not yet well described in the literature). These three SNPs were present in our study at frequencies of 77.1%, 55.5%, and 55.4%, respectively. (Note that because SNP rs1719247 lies outside gene boundaries, it was assigned to the closest gene [GATM] using the single-nucleotide polymorphism database [dbSNP]). Assuming standard statin doses are optimal for the “average” patient, that is, the majority of the population, these variants must necessarily have relatively small effects compared with other factors and this was taken into account in the analysis, as further described below.
Note also that the nature of the actionable genotypes can differ for different SNPs. For instance, either homozygosity or heterozygosity for the rs4149056 SNP variant of the solute carrier organic anion transporter family, member 1B1 (SLCO1B1) gene would prompt recommendations for change in standard simvastatin therapy, as both genotypes for this SNP are deemed actionable. In contrast, for some other SNPs such as rs2032582 on the ATP-binding cassette, subfamily B (MDR/TAP), member 1 gene (ABCB1), only variant homozygosity may be actionable, with standard doses for simvastatin being suitable for both heterozygotes and wild-type homozygotes.
Results
The age and gender demographics for the study participants are presented in Table 1. Because this study has used existing genome-wide microarray data, we were unable to replicate the PREDICT design in full. Nonetheless, using both imputed as well as directly genotyped variants, we have been able to assess the prevalence of clinically actionable genotypes for six of the gene variants reported in the PREDICT study and compare this to medicine exposure for the five drugs considered in the study.5 In addition, we have expanded the analysis to include other relevant cardiovascular drug–gene combinations for which there is a moderate level of evidence according to PharmGKB and CPIC guidelines.6,8
We first assessed variants for which there was strong evidence for clinically actionable medium- or high-risk effects, as further described below. Prevalence is presented in two forms, either relative to the group with actionable genotypes or relative to the group exposed to a given medicine. The former may be considered more relevant from the perspective of the value of pre-emptive genotyping, as it gives insights into what proportions are likely to benefit from advance knowledge of genotype as a result of subsequently being prescribed relevant drugs. The latter may be considered more relevant from the clinical perspective, as it gives practitioners knowledge of the proportion of patients prescribed a particular medicine that would be predicted to require a change of dose or change of drug.
Clopidogrel
Actionable genotypes (strongly evidenced)
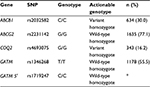
A considerable proportion of participants (~27%; 576/2121; 95% confidence interval [CI]: 25%–29%) had strongly evidenced (PharmGKB Level 1A) medium- or high-risk actionable genotypes for resistance to clopidogrel, associated with SNP rs4244285 on the cytochrome P450, family 2, subfamily C, polypeptide 19 (CYP2C19) gene (Table 2). In total, ~2% (46/2121; 95% CI: 1%–3%) of participants were homozygous for the actionable SNPs for clopidogrel and were considered to be at high-risk of clopidogrel resistance.5,28,29,34
Medication exposure
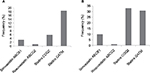
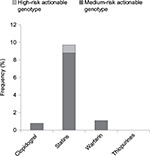
The medication history of the participants was analyzed to determine the number of participants with actionable genotypes who were exposed to each medication or medication class of interest. Of the participants from the cohort with an actionable genotype for clopidogrel, ~3% (15/576; 95% CI: 2%–4%) were exposed to clopidogrel, representing ~0.8% (15/1963; 95% CI: 0.4%–1.2%) of all participants for whom medication data were available. Conversely, ~3% (54/1963; 95% CI: 2%–4%) of all participants with available medication data were taking clopidogrel and ~28% of these (15/54; 95% CI: 16%–40%) had actionable genotypes with moderate risk of resistance to clopidogrel, although none of the participants exposed to clopidogrel were among the ~2% of participants with high-risk genotypes. Figure 1 shows the actionable and high-risk genotypes of the drug–gene pairs of interest.
Statins
Actionable genotypes (strongly evidenced)
For the statins, we first considered pharmacogenomic effects for which there is strong evidence (PharmGKB Level 1A or 2A) before going on to consider less strongly evidenced effects below. In total, ~30% of participants (627/2121; 95% CI: 28%–32%) had strongly evidenced medium- or high-risk actionable genotypes of SNP rs4149056 in the SLCO1B1 gene that may contribute to myopathy and rhabdomyolysis in response to the statins simvastatin, atorvastatin, rosuvastatin, pravastatin, or fluvastatin (Table 2). In total, ~3% of participants (61/2121; 95% CI: 2%–4%) were homozygous for the actionable SNP and were considered to have high-risk genotypes.5,32
Medication exposure
Of the participants from the cohort with strongly evidenced actionable genotypes for statins, ~30% (190/627; 95% CI: 26%–34%) were exposed to one of the above statins, representing ~10% (190/1963; 95% CI: 9%–11%) of all participants with available medication data. Conversely, ~33% (641/1963; 95% CI: 31%–35%) of all participants with medication data were taking statins and ~30% (190/641; 95% CI: 26%–34%) of these had actionable genotypes, including ~3% (17/641; 95% CI: 2%–4%) with the high-risk homozygous genotype (Figure 1).
Actionable genotypes (moderate evidence)
The rs4149056 variant of SLCO1B1 discussed above is a minor allelic variant present in less than half of our cohort. However, as described in Methods, three of the five genotypes for which there is moderate evidence for actionability (PharmGKB Level 2A or 2B) are major alleles, present in over half of our cohort (Table 3). These effects are likely to be substantially affected by other as yet unknown modifiers. This issue is considered in more detail in the context of warfarin below.
In addition, two of the major allelic variants with moderate evidence levels for statin actionability, the GATM rs1346268 variant and the upstream GATM 5′ rs1719247 variant, were observed in an almost identical subset of participants (Table 3). These, therefore, appear to constitute a haplotype. This is consistent with a previous report that SNPs rs1346268 and rs1719247 are in linkage disequilibrium (r2>0.7).35 For our analysis, these SNPs were therefore considered as a single genotype, which was found to be present in ~56% of participants (1178/2121; 95% CI: 54%–58%).
Total prevalence of actionable genotypes for variants with moderate evidence of effects, as well as the prevalence of actionable genotypes in participants exposed to statins, is shown in Table 3 and Figure 2. Where an actionable genotype is specific to a particular statin, prevalence has been determined for exposure to this statin only. For example, a major allelic variant associated with SNP rs2231142 of ABCG2 has moderate evidence levels specifically for rosuvastatin actionability and, although present in ~77% of participants (1635/2121; 95% CI: 75%–79%), only ~1% of these (17/1635; 95% CI: 0.5%–1.5%) were exposed to rosuvastatin. In total, there were 554 participants with one or more moderately evidenced actionable genotypes who were exposed to statins potentially affected by those genotypes. Of these, over half (329 participants) had two or more such genotypes.
Warfarin
Actionable genotypes (strong evidence)
PREDICT approach
In the PREDICT analysis for warfarin, variant homozygosity and heterozygosity for each of the cytochrome P450, family 2, subfamily C, polypeptide 9 (CYP2C9) SNPs rs1799853 and rs1057910 and for the vitamin K epoxide reductase complex, subunit 1 (VKORC1) SNP rs9923231 were all considered actionable, with either medium- or high-risks variants (Table 2). Using this approach, ~74% (1568/2121; 95% CI: 72%–76%) of our cohort were considered to have one or more actionable gene variants for warfarin (Table 4), similar to the figure of 69% reported for the PREDICT sample.5 Of the SNPs above, only the CYP2C9 SNP rs1057910 was regarded as high risk in the PREDICT paper5,36–38 consistent with other studies.34–36 In total, ~0.8% (18/2121; 95% CI: 0.4%–1.2%) of our full cohort had this genotype.
The above approach is at odds with the concept that the standard dose should be correct for the “average” patient and appropriate in at least 50% of the population, implying that <50% of the population would usually be expected to have actionable genotypes for any drug. This approach also does not take into account potential interactions between the CYP2C9 and VKORC1 genotypes.
FDA approach
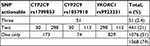
To address these issues, genotype actionability for warfarin was also determined using the FDA table, which considers both CYP2C9 and VKORC1 genotypes simultaneously.39 The rationale underlying this is that, for any patient, the actionability of a given genotype for either one of these genes potentially depends on their genotype for the other gene. Using this approach, ~35% (740/2121; 95% CI: 33%–37%) of the cohort had actionable genotypes for warfarin (Table 5). This is under 50%, as would be expected, although still representing a relatively large proportion of the group.
This subgroup comprises all participants with 1) the VKORC1 genotype with strongest effect (TT)39 irrespective of CYP2C9 genotype, 2) either of the two VKORC1 CT genotypes with strong or intermediate effect (TT or CT, respectively)39 in combination with the CYP2C9*2 actionable genotype, or 3) the CYP2C9*3 actionable genotype, irrespective of VKORC1 or CYP2C9*2 genotype. Note that *2 corresponds to rs1799853 and *3 corresponds to rs1057910.
Note that although the FDA label notes that deficiency of protein C or its cofactor, protein S, can affect tissue necrosis, there are no strong-level genotype-based recommendations for the associated genes in the warfarin drug label and these have not been considered in the present analysis.
Medication exposure
PREDICT approach
Of the participants from the cohort with an actionable genotype for warfarin using the PREDICT approach, ~3% (41/1568; 95% CI: 2% –4%) were exposed to warfarin, representing ~2% (41/1963; 95% CI: 1%–3%) of all participants with medication data. Only one of the participants exposed to warfarin was from the ~0.8% (18/2121; 95% CI: 0.4%–1.2%) of participants with high-risk genotypes. Conversely, ~3% (57/1963; 95% CI: 2%–4%) of the participants with medication data were taking warfarin and ~72% of these (41/57; 95% CI: 60%–84%) had actionable genotypes.
FDA approach
Genotype actionability for the participants taking warfarin was also calculated using the FDA table, as described above.39 Of the participants from the cohort with an actionable genotype for warfarin based on the FDA approach, ~3% (23/740; 95% CI: 2%–4%) were exposed to warfarin, representing ~1.2% (23/1963; 95% CI: 0.7%–1.7%) of participants with medication data. As above, only one of the participants exposed to warfarin was from the ~0.8% (18/2121; 95% CI: 0.4%–1.2%) of participants with high-risk genotypes. Conversely, ~3% (57/1963; 95% CI: 2%–4%) of all participants with medication data were taking warfarin, and 40% (23/57; 95% CI: 27%–53%) of these had actionable genotypes. This is shown in Figure 1.
Thiopurines
Actionable genotypes (strong evidence)
Approximately 9% of participants (193/2121; 95% CI: 8%–10%) had strongly evidenced (PharmGKB Level 1A) medium- or high-risk actionable genotypes for thiopurines associated with the SNPs rs1800460 and rs1142345 of the thiopurine S-methyltransferase (TPMT) gene (Table 2). In accordance with the PREDICT analysis, individuals were considered high risk if they were homozygous for either or both of these two actionable SNPs (6/2121) or if they were heterozygous for both of these two actionable SNPs5,30,31 (compound heterozygous; 185/2121), that is, ~9% of participants (191/2121; 95% CI: 8%–10%). Two participants were classified as medium risk based on heterozygosity for a single actionable SNP.
Medication exposure
Of the participants from the cohort with an actionable genotype for thiopurines, none were exposed to thiopurines (Figure 1). Conversely, two participants from the total cohort were taking thiopurines and neither of these had actionable genotypes.
Tacrolimus
Actionable genotypes (strong evidence)
Tacrolimus is not considered by PharmGKB to have a high-risk genotype based on current evidence but 11% of participants (239/2121; 95% CI: 10%–12%) had strongly evidenced (PharmGKB Level 1A) medium-risk actionable genotypes for tacrolimus associated with SNP rs776746 on the cytochrome P450, family 3, subfamily A, polypeptide 5 (CYP3A5) gene (Table 2). However, no participants from the total community cohort were taking tacrolimus.
Combined risks for all investigated DGIs
Of the 2121 participants who were genotyped or imputed for actionable SNPs with strong clinical evidence (with warfarin assessed using the FDA table, as above), in total 306 (~14%; 95% CI: 12%–16%) had at least one high-risk gene variant, with or without additional medium-risk variants, and a further 1273 (~60%; 95% CI: 58%–62%) had at least one medium-risk gene variant that would be considered clinically actionable for the drugs of interest, without any high-risk variants. That is, ~74% of participants (95% CI: 72%–76%) had actionable genotypes with strong evidence of effects on one or more of the drugs under consideration and approximately one in five of these participants had at least one high-risk genotype. (Using the PREDICT approach, ~90% (95% CI: 89%–91%) of the participants had actionable genotypes for one or more of the drugs under consideration and approximately one in six of these had at least one high-risk genotype, that is, again 306 or ~14%.)
Of the participants who had strong level evidence for at least one medium- or high-risk actionable genotypes, ~14% (218/1579; 95% CI: 12%–16%) were exposed to the relevant drug, representing ~11% (218/1963; 95% CI: 10%–12%) of all participants with medication data. For these analyses, warfarin was assessed using the FDA table; the PREDICT approach yielded comparable proportions (data not shown).
Conversely, ~35% (687/1963; 95% CI: 33%–37%) of the participants with medication data were using one or more of the drugs considered here (clopidogrel, statins, warfarin, and thiopurines) and, of these, ~32% (218/687; 95% CI: 29%–35%) had at least one medium- or high-risk actionable genotype with strong level evidence. Approximately 2.6% (18/687; 95% CI: 1.4%–3.8%) of all participants with medication data had a high-risk clinically actionable genotype for a medication to which they were exposed.
Ethical statement
Ethics approval for the study was obtained from the Hunter New England Local Health District and University of Newcastle Human Research Ethics Committees.
Discussion
Our analysis has revealed that the majority of people in this elderly community cohort (74% or more) have at least one clinically actionable pharmacogenotype that, based on CPIC guidelines, would trigger a change in standard therapy. In total, about a third of this cohort of community-dwelling elderly Australians were taking clopidogrel, statins or warfarin, with statin usage accounting for most of this group, consistent with widespread prescription of statins for the prevention of cardiovascular events.40,41 Immunosuppressant drugs such as thiopurines and tacrolimus, which are mainly indicated for leukemia or organ transplant recipients, were not used by most of the community-dwelling participants, who were presumably relatively healthy compared with patient populations such as that analyzed in the PREDICT study.5
From the microarray data available on our cohort, we were able to analyze eight SNPs with strong levels of evidence, just over half of the 15 variants measured by the PREDICT study.5 In total, the variants we were unable to assess comprised only 1.8% of the total variant prevalence in the PREDICT study5 since the variants that our study covered corresponded to the more prevalent variants analyzed (presumably, in part, because these are more likely to be imputable).
The total prevalence of participants with actionable genotypes in our study was virtually identical to that in the PREDICT paper5 (90% vs. 91%), when warfarin was analyzed in the same way and, although lower (74%), was still large when warfarin actionability was instead calculated according to the FDA guidelines.39
Suarez-Kurtz22 from the Brazilian Pharmacogenomics network found similar prevalences of ~93%–95%22 after inclusion of additional gene variants relevant to the Brazilian population and exclusion of irrelevant variants. The high prevalences across different groups in these studies and our own, including non-Hispanic European-Americans, African-Americans, South Americans of Hispanic descent and Australians of Anglo-Celtic descent, provide strong evidence that most people are likely to have clinically actionable pharmacogenotypes, irrespective of the population under consideration.
In the PREDICT study,5 45% of patients having actionable genotypes with strong level evidence were exposed to risk-associated medications (42% of the entire cohort). As would be expected, medication exposure was considerably lower in our community cohort. Even so, the opportunities for DGIs were surprisingly common with over 10% of the cohort having strongly evidenced actionable genotypes also being exposed to risk-associated medications. This suggests that pharmacogenomic testing of older members of the community has the potential to help improve medication safety and efficacy for considerable numbers of people.5,42
Verbeurgt et al43 investigated a cohort of 1143 patients ranging in age from 18 to 89 years and found that ~15% of their patient population had an actionable genotype and were taking the medication of interest. It was not possible to compare directly with our study due to possible methodological differences. However, their findings appear similar to our study.
Although prevalences of specific variants will differ between different populations, as noted elsewhere,5,22,43,44 making it important to try to carry out the fullest possible characterization of relevant alleles in the population under study, the findings from our study and similar studies5,22,43 are generally in agreement. Factors other than differences in population genetics may also be involved, the most obvious being environmental factors. Such factors will become clearer as information becomes available for larger sample sizes across different population groups.
One important feature of our study is the supplementary analysis of warfarin pharmacogenomics based on the FDA guidelines.39 This takes into account potential interactions between the CYP2C9 and VKORC1 genotypes and is consistent with the concept that the standard dose should be correct for the “average” patient and appropriate in at least 50% of the population, as opposed to the majority of the population requiring a change in standard therapy. Using this approach, 35% of participants had actionable genotypes for warfarin, which still represents a considerable proportion of warfarin patients.
Another recent study led by Shahabi et al,45 published in 2016, has used additional gene variants in calculating warfarin actionability. These authors systematically evaluated publicly available pharmacogenomic data and proposed that nine variants across three genes (CYP2C9, VKORC1, and CYP4F2) significantly affect warfarin dose response, which they incorporated into a dosing algorithm. While the results of this analysis appear promising, the additional variants that were used in the new algorithm are not yet supported by sufficient evidence to meet PHARMGKB criteria. Therefore, these were not included in our own analysis, which we note may therefore have provided a more conservative (lower) estimate of people potentially at risk. A more extensive warfarin analysis could be explored in the future. While the utility of genomic information in improving clinical management remains to be confirmed, the balance of current evidence around warfarin dosing suggests that the use of clinico-genomic algorithms potentially benefits patients for whom genetic information is available.46–48
Our study was limited with respect to the number of community-dwelling participants and the number of participants with missing medication data, which was obtained by self-report. In addition, the population was limited to European ethnicity and the findings may not be generally applicable to other ethnicities. This study was also limited in that some relevant SNPs could not be genotyped or imputed, limiting the number of DGIs which could be examined, and other SNPs were not genotyped but were imputed using reference panels; however, all such SNPs had high imputation accuracy. Having more extensive customized arrays specific for pharmacogenomic testing for the population background or, ultimately, using RNAseq techniques could further enhance this study.
Subsequent studies should include larger numbers of participants and extend into younger age ranges and wider populations of mixed ethnic backgrounds.5 This may be facilitated in the future by customized arrays that are specific and precise for pharmacogenomic testing and the participant population under investigation. More research is required into the extent to which multifactorial DGI pharmacogenomics correlates both with pharmacokinetic measurements of active drug levels in different individuals and with clinical outcomes.
In conclusion, this study has shown that most older people in the community are likely to have at least one actionable genotype. Since many are also exposed to one or more medications that may be affected by these genotypes, prescribers need to give greater consideration to the possibility of clinically relevant DGIs in this group. Our findings affirm that pre-emptive genotyping is likely to have strong potential to improve medication safety, efficacy, and health outcomes.1,49 Further investigations correlating genotypes and medication exposures to adverse reactions and other outcomes in older people appear justified.
Acknowledgments
The authors would like to thank the men and women participating in the HCS as well as all the staff, investigators, and collaborators who have supported or been involved in the project to date. The authors also acknowledge the contributions of Mark McEvoy, Wayne Smith, Catherine D’Este, Janine Duke, Roseanne Peel, Peter Schofield, Julie Byles, David Henry, Ben Ewald, and Derek Smith. A special thanks to Alison Koschel and Debbie Quain who were instrumental in setting up the pilot study and initial phase of the project.
The University of Newcastle provided $300,000 from its Strategic Initiatives Fund and $600,000 from the Gladys M Brawn Senior Research Fellowship scheme; Vincent Fairfax Family Foundation, a private philanthropic trust, provided $195,000; The Hunter Medical Research Institute provided media support during the initial recruitment of participants; and Dr Anne Crotty, Prof. Rodney Scott and Associate Professor Levi provided financial support towards freezing costs for the long-term storage of participant blood samples.
Disclosure
The authors report no conflicts of interest in this work.
References
Relling MV, Evans WE. Pharmacogenomics in the clinic. Nature. 2015;526(7573):343–350. | ||
Dunnenberger HM, Crews KR, Hoffman JM, et al. Preemptive clinical pharmacogenetics implementation: current programs in five US medical centers. Annu Rev Pharmacol Toxicol. 2015;55:89–106. | ||
Haga SB, LaPointe NM. The potential impact of pharmacogenetic testing on medication adherence. Pharmacogenomics J. 2013;13(6):481–483. | ||
Daneshi N, Graham M, Biswas M, Milward E. Gene scene-health’s new frontier. Aust Pharm. 2015;34(5):27–30. | ||
Van Driest SL, Shi Y, Bowton EA, et al. Clinically actionable genotypes among 10,000 patients with preemptive pharmacogenomic testing. Clin Pharmacol Ther. 2014;95(4):423–431. | ||
The Pharmacogenomics Knowlegebase (PharmGKB) [homepage on the Internet]; 2016. Available from: https://www.pharmgkb.org/. Accessed February 5, 2016. | ||
Chang JS, Pham DA, Dang MT, Lu Y, VanOsdol S, Shin J. Evaluation of popular drug information resources on clinically useful and actionable pharmacogenomic information. J Med Libr Assoc. 2016;104(1):58–61. | ||
The Pharmacogenomics Knowlegebase (PharmGKB). Clinical Pharmacogenetics Implementation Consortium (CPIC) [webpage on the Internet]; 2016. Available from: https://www.pharmgkb.org/page/cpic. Accessed February 14, 2016. | ||
Food and Drug Adminstration (FDA) [homepage on the Internet]; 2016. Available from: http://www.fda.gov/. Accessed February 14, 2016. | ||
Haga SB, Moaddeb J. Comparison of delivery strategies for pharmacogenetic testing services. Pharmacogenet Genomics. 2014;24(3):139–145. | ||
Kitzmiller JP, Groen DK, Phelps MA, Sadee W. Pharmacogenomic testing: relevance in medical practice: why drugs work in some patients but not in others. Cleve Clin J Med. 2011;78(4):243–257. | ||
Adams SA, Petersen C. Precision medicine: opportunities, possibilities, and challenges for patients and providers. J Am Med Inform Assoc. 2016;23(4):787–790. | ||
Twilt M. Precision medicine: the new era in medicine. EBioMedicine. 2016;4:24–25. | ||
Collins FS, Varmus H. A new initiative on precision medicine. N Engl J Med. 2015;372(9):793–795. | ||
Lander ES. Cutting the gordian helix – regulating genomic testing in the era of precision medicine. N Engl J Med. 2015;372(13):1185–1186. | ||
Ji Y, Skierka JM, Blommel JH, et al. Preemptive pharmacogenomic testing for precision medicine: a comprehensive analysis of five actionable pharmacogenomic genes using next-generation DNA sequencing and a customized CYP2D6 genotyping cascade. J Mol Diagn. 2016;18(3):438–445. | ||
Milward EA, Daneshi N, Johnstone DM. Emerging real-time technologies in molecular medicine and the evolution of integrated ‘pharmacomics’ approaches to personalized medicine and drug discovery. Pharmacol Ther. 2012;136(3):295–304. | ||
Shahandeh A, Johnstone DM, Atkins JR, et al. Advantages of array-based technologies for pre-emptive pharmacogenomics testing. Microarrays. 2016;5(2):12. | ||
Malinovska A, Bingisser R, Nickel CH. [Adverse drug events of older patients presenting in the emergency department]. Ther Umsch. 2015;72(11–12):673–677. German. | ||
Chen TF. Pharmacist-led home medicines review and residential medication management review: the Australian model. Drugs Aging. 2016;33(3):199–204. | ||
Fulton MM, Allen ER. Polypharmacy in the elderly: a literature review. J Am Acad Nurse Pract. 2005;17(4):123–132. | ||
Suarez-Kurtz G. Clinically actionable genotypes among Brazilians. Clin Pharmacol Ther. 2014;96(1):25–26. | ||
Holliday EG, Maguire JM, Evans TJ, et al; Australian Stroke Genetics Collaborative, International Stroke Genetics Consortium, Wellcome Trust Case Control Consortium 2. Common variants at 6p21.1 are associated with large artery atherosclerotic stroke. Nat Genet. 2012;44(10):1147–1151. | ||
McEvoy M, Smith W, D’Este C, et al. Cohort profile: The Hunter Community study. Int J Epidemiol. 2010;39(6):1452–1463. | ||
Holliday EG, Attia J, Hancock S, et al. Genome-wide association study identifies two novel genomic regions in irritable bowel syndrome. Am J Gastroenterol. 2014;109(5):770–772. | ||
Birdwell KA, Decker B, Barbarino JM, et al. Clinical pharmacogenetics implementation consortium (CPIC) guidelines for CYP3A5 genotype and tacrolimus dosing. Clin Pharmacol Ther. 2015;98(1):19–24. | ||
Johnson JA, Gong L, Whirl-Carrillo M, et al; Clinical Pharmacogenetics Implementation Consortium. Clinical pharmacogenetics implementation consortium guidelines for CYP2C9 and VKORC1 genotypes and warfarin dosing. Clin Pharmacol Ther. 2011;90(4):625–629. | ||
Scott SA, Sangkuhl K, Stein CM, et al; Clinical Pharmacogenetics Implementation Consortium. Clinical pharmacogenetics implementation consortium guidelines for CYP2C19 genotype and clopidogrel therapy: 2013 update. Clin Pharmacol Ther. 2013;94(3):317–323. | ||
Scott SA, Sangkuhl K, Gardner EE, et al; Clinical Pharmacogenetics Implementation Consortium. Clinical pharmacogenetics implementation consortium guidelines for cytochrome P450-2C19 (CYP2C19) genotype and clopidogrel therapy. Clin Pharmacol Ther. 2011;90(2):328–332. | ||
Relling MV, Gardner EE, Sandborn WJ, et al; Clinical Pharmacogenetics Implementation Consortium. Clinical pharmacogenetics implementation consortium guidelines for thiopurine methyltransferase genotype and thiopurine dosing. Clin Pharmacol Ther. 2011;89(3):387–391. | ||
Relling MV, Gardner EE, Sandborn WJ, et al; Clinical Pharmacogenetics Implementation Consortium. Clinical pharmacogenetics implementation consortium guidelines for thiopurine methyltransferase genotype and thiopurine dosing: 2013 update. Clin Pharmacol Ther. 2013;93(4):324–325. | ||
Wilke RA, Ramsey LB, Johnson SG, et al; Clinical Pharmacogenomics Implementation Consortium (CPIC). The clinical pharmacogenomics implementation consortium: CPIC guideline for SLCO1B1 and simvastatin-induced myopathy. Clin Pharmacol Ther. 2012;92(1):112–117. | ||
Owen RP, Gong L, Sagreiya H, Klein TE, Altman RB. VKORC1 pharmacogenomics summary. Pharmacogenet Genomics. 2010;20(10):642–644. | ||
Mega JL, Hochholzer W, Frelinger AL, et al. Dosing clopidogrel based on CYP2C19 genotype and the effect on platelet reactivity in patients with stable cardiovascular disease. JAMA. 2011;306(20):2221–2228. | ||
Mangravite LM, Engelhardt BE, Medina MW, et al. A statin-dependent QTL for GATM expression is associated with statin-induced myopathy. Nature. 2013;502(7471):377–380. | ||
Ogg MS, Brennan P, Meade T, Humphries SE. CYP2C9*3 allelic variant and bleeding complications. Lancet. 1999;354(9184):1124. | ||
Ablin J, Cabili S, Lagziel A, Peretz H. Warfarin therapy in a patient homozygous for the CYP2C9 3 allele. Isr Med Assoc J. 2002;4(2):139–141. | ||
Steward DJ, Haining RL, Henne KR, et al. Genetic association between sensitivity to warfarin and expression of CYP2C9*3. Pharmacogenetics. 1997;7(5):361–367. | ||
Food and Drug Ddminstration (FDA) [webpage on the Internet]. Coumadin Tablets (Warfarin Sodium Tablets, USP) Crystalline Coumadin for Injection (Warfarin Sodium for Injection, USP); 2016. Available from: http://www.accessdata.fda.gov/drugsatfda_docs/label/2010/009218s108lbl.pdf. Accessed March 25, 2016. | ||
National Prescribing Servive (NPS) Medicinewise [webpage on the Internet]. Benefits and risks of statin use in older people; 2016. Available from: http://www.nps.org.au/topics/ages-life-stages/for-individuals/older-people-and-medicines/for-health-professionals/evidence-and-guidelines/using-statins. Accessed June 26, 2016. | ||
National Prescribing Servive (NPS) Medicinewise [webpage on the Internet]. Statins revisited: appropriate patient selection is key; 2016. Available from: http://www.nps.org.au/publications/health-professional/health-news-evidence/2013/statins-absolute-CVD-risk. Accessed August 10, 2016. | ||
Bell GC, Crews KR, Wilkinson MR, et al. Development and use of active clinical decision support for preemptive pharmacogenomics. J Am Med Inform Assoc. 2014;21(e1):e93–e99. | ||
Verbeurgt P, Mamiya T, Oesterheld J. How common are drug and gene interactions? Prevalence in a sample of 1143 patients with CYP2C9, CYP2C19 and CYP2D6 genotyping. Pharmacogenomics. 2014;15(5):655–665. | ||
Van Driest SL, Roden DM. Response to “clinically actionable genotypes among Brazilians”. Clin Pharmacol Ther. 2014;96(1):26. | ||
Shahabi P, Scheinfeldt LB, Lynch DE, et al. An expanded pharmacogenomics warfarin dosing table with utility in generalised dosing guidance. Thromb Haemost. 2016;116(2):337–348. | ||
Daneshi N, Milward EA, Graham M. An update on warfarin pharmacogenomics: part 1 of 1. Aust Pharm. 2015;34(5):52–56. | ||
Daneshi N, Milward EA, Graham M. An update on warfarin pharmacogenomics: part 2. Aust Pharm. 2015;34(6):46–49. | ||
Li X, Yang J, Wang X, Xu Q, Zhang Y, Yin T. Clinical benefits of pharmacogenetic algorithm-based warfarin dosing: meta-analysis of randomized controlled trials. Thromb Res. 2015;135(4):621–629. | ||
Schildcrout JS, Denny JC, Bowton E, et al. Optimizing drug outcomes through pharmacogenetics: a case for preemptive genotyping. Clin Pharmacol Ther. 2012;92(2):235–242. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.