Back to Journals » Infection and Drug Resistance » Volume 14
Polymyxin E–Resistant Gram-Negative Bacteria in Tunisia and Neighboring Countries: Are There Commonalities?
Authors Dziri O, Dziri R, El Salabi AA, Alawami Jnr AA, Ksouri R, Chouchani C
Received 3 July 2021
Accepted for publication 13 August 2021
Published 16 November 2021 Volume 2021:14 Pages 4821—4832
DOI https://doi.org/10.2147/IDR.S327718
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Héctor Mora-Montes
Olfa Dziri,1– 3 Raoudha Dziri,1 Allaaeddin A El Salabi,4,5 Alhussain A Alawami,4 Riadh Ksouri,6 Chedly Chouchani1– 3
1Laboratory of Microorganisms and Active Biomolecules, Faculty of Sciences of Tunis, University of Tunis El Manar, Tunis, Tunisia; 2Laboratory of Research in Sciences and Technology of Environment, Higher Institute of Sciences and Technologies of Environment of Borj Cédria, University of Carthage, Hammam-Lif, Tunisia; 3Joint Service Unit for Research Genomic Platform, Higher Institute of Environmental Sciences and Technologies of Environment of Borj Cédria, Center of Biotechnology of Borj Cédria, Hammam-Lif, Tunisia; 4Infection Control and Patient Safety Office, New Marwa Hospital, Benghazi, Libya; 5Department of Environmental Health, Faculty of Public Health, University of Benghazi, Benghazi, Libya; 6Laboratory of Aromatic and Medicinal Plants, Center of Biotechnology of Borj Cédria, Hammam-Lif, Tunisia
Correspondence: Chedly Chouchani Email [email protected]
Abstract: The current global dissemination of polymyxin E resistance constitutes a real public health threat because of the restricted therapeutic options. This review provides a comprehensive assessment of the epidemiology of polymyxin E–resistant bacteria, with special reference to colistin-resistant Gram-negative bacteria in Tunisia and neighboring countries, based on available published data to January 2020. We aimed to determine their prevalence by species and origin, shedding light on the different genes involved and illustrating their genetic support, genetic environment, and geographic distribution. We found that colistin resistance varies considerably among countries. A majority of the research has focused on Algeria (13 of 32), followed by Tunisia (nine of 32), Egypt (nine of 32), and Libya (one of 32). All these reports showed that colistin-resistant Gram-negative bacteria were dramatically disseminated in these countries, as well as in African wildlife. Moreover, high prevalence of these isolates was recorded from various sources (humans, animals, food products, and natural environments). Colistin resistance was mainly reported among Enterobacteriaceae, particularly Klebsiella pneumoniae and Escherichia coli. It was associated with chromosomal mutations and plasmid-mediated genes (mcr). Four mcr variants (mcr 1, mcr 2, mcr 3, and mcr8), mobilized by several plasmid types (IncHI2, IncP, IncFIB, and IncI2), were detected in these countries and were responsible for their rapid spread. Countrywide dissemination of high-risk clones was also observed, including E. coli ST10 and K. pneumoniae ST101 and ST11. Intensified efforts to raise awareness of antibiotic use and legalization thereon are required in order to monitor and minimize the spread of multidrug-resistant bacteria.
Keywords: Gram-negative bacteria, polymyxin E resistance, chromosomal mutations, mcr genes, plasmids
Introduction
Colistin is an ancient antibiotic known also as polymyxin E, it was discovered in 1949 and introduced to human medicine in the 1950s. It has bactericidal activity against most Gram-negative bacteria by targeting lipopolysaccharides in the outer membrane.1 In the early 1980s, it was abandoned and replaced by other antibiotics because of its neurotoxicity and nephrotoxicity.1 However, the global spread of multidrug-resistant bacterial infections in the 21st century led to the revival of colistin in clinical practice.1,2 Indeed, it is considered one of the last-resort antibiotics used for the treatment of severe infections, caused in particular by carbapenem-resistant Gram-negative bacteria.3,4 Several studies have revised and evaluated the colistin dosing regimen, as well as the best way to use and minimize its toxicity, sos as to ensure its safety and efficacy.5–8 Colistin consumption has been continuously increasing since 2000 worldwide, including low-income countries.9
The use of colistin has not been restricted only to human medicine. It has also been used in veterinary medicine for decades, even for the prevention of microbial infections.10 Unfortunately, in several regions, such as China, India, and Africa, it has been used without strict regulation in animal feed as a growth promoter, eg, for livestock, poultry production, and aquaculture.10–13 Excessive use of colistin has led to the emergence and spread of colistin-resistant Gram-negative bacteria, constituting an emerging phenomenon and a real threat to public health.14 High rates of colistin resistant Gram-negative species, especially Enterobacteriaceae, Pseudomonas aeruginosa, and Acinetobacter baumannii, have been recorded throughout the globe. They were isolated from different origins notably animals, food products of animal origin, humans, and natural environments.14,15 The dissemination of these isolates constitutes a real concern and a potential risk to human health. In fact, they can be transmitted to healthy humans through the food chain.15 Facing this problem, several countries have banned the use of or tightened polices on colistin use in animal husbandry to reduce and monitor this crisis.12 Some species of Proteus, Providencia, Morganella, Neisseria, and Serratia are naturally resistant to colistin.16 These species should not be ignored, because they could cause severe infections with limited treatment options.
Generally, bacteria acquire resistance to colistin via mutations of chromosomal genes, eg, mgrB, and some operons and regulators, such as pmrABC, leading to lipopolysaccharide modifications.17 The plasmid-mediated colistin gene mcr1 was reported for the first time in 2015 in China.18 Since then, it has disseminated dramatically around the world and become the most commonly reported mechanism implicated in colistin resistance.14 The emergence of this gene is considered an alarming problem, because it contributes to the rapid dissemination of colistin-resistant isolates by horizontal gene transfer. So far, ten MCR variants have been identified, taking into consideration a heterogenous geographic distribution.12,14
This review summarizes the epidemiology of colistin-resistant bacteria isolated from different origins in Tunisia and neighboring countries and determines their prevalence based on available published data to January 2020. We also review the genes involved in colistin resistance, illustrating their genetic support, genetic environment as well as geographic distribution.
Prevalence and Origins of Colistin-Resistant Isolates
All studies that had investigated the dissemination of colistin-resistant Gram-negative bacteria in Tunisia and neighboring countries — Algeria, Libya, Egypt, Morocco, and Mauritania — to January 2020 are summarized in Table 1. Algeria had the most published papers (n=13), followed by Tunisia (n=9), Egypt (n=9), and Libya (n=1). However, there were no published data from Morocco or Mauritania (Figure 1). The lack of such data does not mean an absence of colistin-resistant isolates in these two countries, but could be explained by the absence of surveillance programs and diagnostic microbiology laboratories. Furthermore, a Moroccan study showed that colistin was one of the most used antibiotics in broiler production and overdosed in most administrations.19
 |  |  |
Table 1 Characteristics of polymyxin E–resistant Gram-negative bacteria detected in Tunisia and neighboring countries to January 2020 |
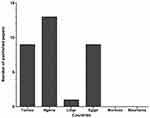 |
Figure 1 Papers reporting colistin resistance in each country to January 2020. |
As shown in Figure 2A, the highest prevalence of colistin-resistant Gram-negative bacteria was recorded in Tunisia (58.09%), followed by Egypt (24.76%), Algeria (14.29%), and Libya (2.86%). The distribution of colistin-resistant isolates per origin showed a higher prevalence in animals than humans, suggesting the major role of animals in the dissemination of colistin resistance (Figure 2B). In sum, 59.52% of colistin-resistant bacteria were isolated from animals (chickens, bovids, camels, Barbary macaques, and wild birds), and 20.48% and 2.86% were isolated from clinical specimens and healthy humans, respectively (Figure 2B, and Table 1). The detection of colistin-resistant isolates in healthy humans has recently been reported in Egypt, which is of concern.15 These findings could be explained by the transmission of colistin resistance via the food chain, which deepens the antibiotic-resistance crisis and its challenges.15 Colistin-resistant isolates were detected in foodstuffs (10%), such as raw bovine milk, Karish cheese, raw chicken meat, and beef sausage. Colistin-resistant bacteria were also observed in agricultural soil and manure (3.81%), as well as water sources (3.33%, Figure 2B and Table 1).
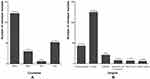 |
Figure 2 (A) Distribution of colistin resistant isolates per country and (B) per origin. |
In the main, four colistin-resistant species were identified in these four countries. Colistin resistance was frequently observed among Enterobacteriaceae, with Escherichia coli (n=154) dominant, followed by Klebsiella pneumoniae (n=45). However, few colistin-resistant isolates belonging to the nonfermentative Gram-negative bacteria Acinetobacter baumannii (n=4) and Pseudomonas aeruginosa (n=7) were recorded (Tables 1 and 2). We also noticed that colistin-resistant K. pneumoniae and A. baumannii were isolated mainly from clinical specimens. Obviously, when the number of samples increased, the number of resistant isolates did also.
 |
Table 2 Distribution of colistin-resistance genes per species and country |
Generally, these species constituted the main cause of the most worrisome infections occurring in health-care settings and the community.20 In Tunisia, colistin-resistant K. pneumoniae and A. baumannii isolates were recovered from clinical specimens of patients hospitalized in coastal cities.21–23 Colistin-resistant E. coli isolates were detected among chickens and bovids on northern farms, in addition to camels on a southern farm.24–26 As described in Table 1, colistin-resistant E. coli was also detected in food of animal origin.26,27
On the other hand, Algerian studies reported the isolation of these three species from clinical specimens of patients hospitalized in different governorates of the country.28–30 However, only E. coli isolates were detected in other sources, including animals,31,32 agricultural soil and manure,33 in addition to seawater.34
The detection of colistin-resistant E. coli in Barbary macaques in the north of Algeria constituted the first report of such isolates in the wildlife of Africa.32 A second African study describing the isolation of colistin-resistant isolates in wild animals has recently been published in Egypt. It showed the detection of three colistin-resistant species — E. coli, K. pneumoniae, and P. aeruginosa — among wild birds.15 Colistin-resistant P. aeruginosa isolates of other origins were not detected in Egypt nor in the other countries included in this review (Table 1). The presence of colistin-resistant Gram-negative bacteria in the wildlife samples is a real concern. In fact, wildlife plays a major role in the global dissemination of multidrug-resistant bacteria, including colistin-resistant isolates. As such, wild birds are considered reservoirs and vectors for the transmission of antibiotic-resistant genes.35 Moreover, the health situation in Egypt has become more critical since the discovery of colistin-resistant E. coli and K. pneumoniae in healthy humans and water.15 Egyptian studies have also described the detection of colistin-resistant E. coli, K. pneumoniae, and A. baumannii in clinical specimens collected from hospitalized patients. Colistin-resistant E. coli isolates have been recovered from chickens,36,37 bovids,38 and food of animal origin.39,40 To date, only one study has reported the detection of colistin-resistant isolates in Libya. These isolates were recovered from clinical specimens of hospitalized patients in Tripoli and were identified as K. pneumoniae.41
All these reports confirmed the dissemination of colistin-resistant Gram-negative bacteria in Tunisia and neighboring countries, with high prevalence of various origin. The question is whether there are commonalities or even links among the detected colistin-resistant isolates at genetic and clonal levels. This is elucidated upon in the two next sections.
Colistin-Resistance Genes and Genetic Environments
As shown in Tables 1 and 2, several colistin-resistance genes were detected, showing a heterogeneous distribution among countries and species. mcr was observed more often than chromosomal mutations. Indeed, we noted the detection of four mcr variants — mcr1, mcr2, mcr3, and mcr8 — with of mcr1 dominant (75.48%). Obviously, this observation is limited to the published reports included in our study and does not reflect the real scenery of colistin resistance, because of the lack of reporting in several governorates in these countries. More epidemiological studies are required to clarify the health situation and get closer to reality.
These four mcr genes are distantly related, as shown by the phylogenetic tree of the nine variants of mcr genes in Figure 3. This phylogenetic tree was performed using the Clustal Omega program (https://www.ebi.ac.uk/services) after collecting the nucleotide sequences from the GenBank database (https://www.ncbi.nlm.nih.gov), and distances were calculated using the neighbor-joining algorithm (Figure 3).
 |
Figure 3 Phylogenetic tree of the nine variants of mcr genes generated using the Clustal Omega program (https://www.ebi.ac.uk/services). Distances were calculated using the neighbor-joining algorithm. |
mcr1 was predominant among E. coli isolates, showing high prevalence in Tunisia (63.125%), followed by Egypt (23.75%), and Algeria (13.125%). The emergence of mcr1 among K. pneumoniae and P. aeruginosa isolates was observed only in Egypt (Table 2). Likewise, the detection of mcr2 and the coexpression of mcr1 and mcr2 were described only in Egypt among E. coli, K. pneumoniae, and P. aeruginosa.15 The variants mcr3 and mcr8 were described only in Algeria among E. coli and K. pneumoniae, respectively, as demonstrated in Table 2. As such, mcr genes constitute the main mechanism conferring resistance to colistin among E. coli isolates. However, these genes were rarely detected among K. pneumoniae (n=13) and P. aeruginosa (n=7).
mcr is characterized by a variety of plasmid types or even associated with transposons, ensuring its rapid spread worldwide.42 The present study showed that mcr1 was carried mainly by the IncHI2 replicon in Tunisia (Table 1). To our knowledge, the IncP plasmid type harboring mcr1 has been described in only one report in this country. This was detected in E. coli isolated from fecal samples of healthy chickens.43 However, in Algeria, mcr1 was detected in various plasmid groups, noting IncFIB, IncI2, and IncHI2A.34,44,45 Interestingly, the two plasmid types harboring mcr1, IncHI2 and IncI2, were also identified among E. coli isolated from food of animal origin in Egypt.39,40 A recent study showed that plasmids belonging to three incompatibility groups — IncI2, IncX4, and IncHI2 — are involved in the global dissemination of mcr genes.46 The IncI2 plasmid is commonly reported in Asia and America, and the IncHI2 plasmid frequently reported in Europe and Africa.46 This gene is also associated with a transposable element, namely the insertion sequence ISApl.40,43
Colistin-resistant A. baumannii was rarely isolated in these countries: there was a clear absence of mcr genes among these isolates (Table 2). In contrast, dissemination of colistin-resistant A. baumannii harboring plasmid-encoded mcr genes has recently been described in Spain, China, and Iraq.14,47 We found that colistin resistance among the detected A. baumannii clinical isolates was usually due to mutated chromosomal genes, including pmrB and the pmrCAB operon (Table 1). The mutated pmrB gene leads to colistin resistance by alteration of lipid A, which is a component of lipopolysaccharide. This mechanism has been detected in Tunisia and Algeria.22,29 Also, mutations in the pmrCAB operon have been identified in Egypt.48
Colistin resistance through mutations of chromosomal genes was also observed in K. pneumoniae isolates, with mutated pmrA and pmrB detected in clinical specimens in Algeria. However, mutated mgrB constituted the main mechanism conferring resistance to colistin among K. pneumoniae in Tunisia and Libya. This mechanism has rarely been described in Algeria and Egypt (Tables 1 and 2). This species constitutes the main reservoir of colistin-resistance genes, including both acquired and mutated ones.
Sequence Types
Several colistin-resistant bacterial clones were disseminated among the countries included in the present study, as shown in Table 1. Thirteen distinct sequence types of colistin-resistant E. coli were detected in Tunisia, with four clones predominant: ST10, ST162, ST57, and ST69. Colistin-resistant isolates assigned to ST10 originated from chickens and bovids, whereas ST162 isolates were detected among chickens and camels. ST57 and ST69 were isolated from chickens. The other disseminated clones originated from chickens also: ST2179, ST349, ST1011, ST1642, ST117, ST398, ST2220, ST5416, and ST5686.
Seven sequence types of colistin-resistant E. coli were identified in Algeria, with ST405 the most abundant. This clone was isolated from different origins, including clinical specimens, Barbary macaques, and agricultural soil and manure. Regarding the latter, three other colistin-resistant E. coli clones were detected — ST10, ST345, and ST155. In addition, two E. coli clones, ST23 and ST115, were identified in seawater. However, only one colistin-resistant E. coli clone (ST48) was isolated from chickens in this country (Table 1).
There were four major colistin-resistant E. coli clones in Egypt: ST10, ST1011, ST69, and ST101. ST10 was also recorded in Tunisia and Algeria, while ST1011 and ST69 were detected in Tunisia from different origins (Table 1). A recent report recorded the dominance of E. coli ST10 in animal and water. This clone is considered the most common ST of those harboring mcr1 worldwide.46
Many colistin-resistant K. pneumoniae belonging to different sequence types were detected in Tunisia (ST11, ST15, ST101, ST147, and ST392), Algeria (ST101, ST2620, ST3242, and ST336), Egypt (ST11 and ST1399), and Libya (ST101), and were mainly isolated from clinical specimens (Table 1). It is worth noting that colistin-resistant K. pneumoniae ST101 was disseminated in three neighboring countries — Tunisia, Algeria, and Libya. Infections caused by such clones have been described in Italy and Serbia.49 We also noticed the spread of colistin resistant ST11 isolates in both Tunisia and Egypt. These two sequence types belonged to the same clonal complex (CC11) and were considered high-risk clones in terms of morbidity and mortality rates.49 The isolates assigned to CC11 usually show a multidrug-resistance profile and also harbor numerous antibiotic-resistance genes, especially those coding for ESBLs and carbapenemases, thus restricting therapeutic options.50 Countrywide dissemination of the multidrug-resistant K. pneumoniae ST147 and ST15, particularly carbapenemase producers, has been reported in Mediterranean countries, including Tunisia.4,51
Regarding the uncommon colistin-resistant A. baumannii, we noted the detection of two circulated clones — ST641 in Tunisia and the international clone ST2 in Algeria — according to Pasteur’s scheme.22,29 However, it should be noted that other sequence types might have emerged, despite the lack of sequence-type identification in the Egyptian report.48
Impact of High-Throughput Sequencing in the Detection of Colistin-Resistance Genes
The seriousness of the antibiotic-resistance crisis has prompted researchers to develop sophisticated methods to facilitate the determination of antibiotic-resistance mechanisms in a rapid and more precise manner. Herein, we are talking about whole-genome sequencing using high-throughput technologies, which have achieved significant progress. This method could be considered a scientific revolution in molecular biology, including the antibiotic-resistance field. It is becoming a potent tool of choice to identify resistance genes, especially those implicated in colistin resistance.52 This method seems to be a key to the revelation of new colistin-resistance mechanisms by detection of unknown mutations of chromosomal genes and new variants of mcr genes,52 and also leads to precise and in-depth research using bioinformatic analysis to determinate the genetic environment of the detected genes and their genetic support.52 Several reports using whole-genome sequencing to investigate the colistin-resistance mechanism, have been performed in Tunisia,21,22 Algeria,45 and Egypt.39,40 It is very important to note the use of this method in low-income countries, such as Tunisia, Algeria, and Egypt. This step is an indicator of the scientific progress in these countries and proves the significance of efforts of researchers to improve the quality of the health-care system in their countries, despite the difficult financial conditions.
Conclusion
Colistin is considered a beacon in the darkness of the antibiotic-resistance challenge.12 It is used as the last-resort antibiotic for the treatment of multidrug-resistant Gram-negative bacterial infections. However, extensive use has led to the emergence and dissemination of colistin resistant Gram-negative bacteria, which constitute a real threat to public health. The flexibility of the bacterial genome and its ability to adapt and resist colistin has prompted researchers to find other solutions.
As described in this review, colistin resistant Gram-negative bacteria were disseminated in Tunisia and neighboring countries, showing high prevalence with different origins (humans, animals, food products, and natural environments). The detection of colistin-resistant isolates in the African wildlife is a real concern. Both plasmid-mediated genes and chromosomal mutations were involved in colistin resistance. We found that colistin resistance in K. pneumoniae was caused mainly by chromosomal mutations. However, it was often caused by mcr in E. coli. Colistin resistance varies considerably among countries, and there is countrywide dissemination of several clones. These findings emphasize the seriousness of antibiotic resistance in these countries, requiring urgent interventions to monitor and control the situation before it becomes worse, in order to avoid pandrug resistance and a therapeutic impasse. Preserving human health and natural wealth is everyone’s responsibility. When crises intensify, every small detail can carry a great meaning and achieve a great goal.
Acknowledgment
The authors want to thank all researchers for their dedicated work in the surveillance of antibiotic resistance.
Funding
This research received funding from the Center of Biotechnology of Borj Cédria for publication fees payment.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Falagas M, Kasiakou S, Saravolatz L. Colistin: the revival of polymyxins for the management of multidrug-resistant gram-negative bacterial infections. Clin Infect Dis. 2005;40(9):1333–1341. doi:10.1086/429323
2. Nasnas R, Saliba G, Hallak P. The revival of colistin: an old antibiotic for the 21st century. Pathol Biol. 2009;57(3):229–235. doi:10.1016/j.patbio.2007.09.013
3. Deris ZZ. The multidrug-resistant Gram-negative superbugs threat require intelligent use of the last weapon. Malays J Med Sci. 2015;22(5):1–6.
4. Dziri O, Dziri R, Ali El Salabi A, et al. Carbapenemase producing Gram-Negative bacteria in tunisia: history of thirteen years of challenge. Infect Drug Resist. 2020;13:4177–4191. doi:10.2147/IDR.S259562
5. Jung S, Chung E, Jun M, et al. Differences in colistin administration and bacterial and treatment outcomes in critically ill patients. Sci Rep. 2019;9(1):8781. doi:10.1038/s41598-019-44965-y
6. Dalfino L, Puntillo F, Mosca A, et al. High-dose, extended-interval colistin administration in critically ill patients: is this the right dosing strategy? A preliminary study. Clin Infect Dis. 2012;54(12):1720–1726. doi:10.1093/cid/cis286
7. Luque S, Grau S, Valle M, et al. Differences in pharmacokinetics and pharmacodynamics of colistimethate sodium (CMS) and colistin between three different CMS dosage regimens in a critically ill patient infected by a multidrug-resistant Acinetobacter baumannii. Int J Antimicrob Agents. 2013;42(2):178–181. doi:10.1016/j.ijantimicag.2013.04.018
8. Nation R, Li J. Colistin in the 21st century. Curr Opin Infect Dis. 2009;22(6):535–543. doi:10.1097/QCO.0b013e328332e672
9. Klein E, Van Boeckel T, Martinez E, et al. Global increase and geographic convergence in antibiotic consumption between 2000 and 2015. Proc Natl Acad Sci U S A. 2018;115(15):E3463–E3470. doi:10.1073/pnas.1717295115
10. Kumar H, Chen B, Kuca K, et al. Understanding of colistin usage in food animals and available detection techniques: a review. Animals. 2020;10(10):1892. doi:10.3390/ani10101892
11. Kempf I, Jouy E, Chauvin C. Colistin use and colistin resistance in bacteria from animals. Int J Antimicrob Agents. 2016;48(6):598–606. doi:10.1016/j.ijantimicag.2016.09.016
12. Shen Y, Zhang R, Schwarz S, et al. Farm animals and aquaculture: significant reservoirs of mobile colistin resistance genes. Environ Microbiol. 2020;22(7):2469–2484. doi:10.1111/1462-2920.14961
13. Van T, Yidana Z, Smooker M, et al. Antibiotic use in food animals worldwide, with a focus on Africa: pluses and minuses. J Glob Antimicrob Resist. 2020;20:170–177. doi:10.1016/j.jgar.2019.07.031
14. Luo Q, Wang Y, Xiao Y. Prevalence and transmission of mobilized colistin resistance (mcr) gene in bacteria common to animals and humans. Biosafety Health. 2020;2(2):71–78. doi:10.1016/j.bsheal.2020.05.001
15. Ahmed Z, Elshafiee E, Khalefa H, et al. Evidence of colistin resistance genes (mcr-1 and mcr-2) in wild birds and its public health implication in Egypt. Antimicrob Resist Infect Control. 2019;8:197. doi:10.1186/s13756-019-0657-5
16. Zhang H, Hou M, Xu Y, et al. Action and mechanism of the colistin resistance enzyme MCR-4. Commun Biol. 2019;2:36. doi:10.1038/s42003-018-0278-1
17. Aghapour Z, Gholizadeh P, Ganbarov K, et al. Molecular mechanisms related to colistin resistance in Enterobacteriaceae. Infect Drug Resist. 2019;12:965–975. doi:10.2147/IDR.S199844
18. Liu Y, Wang Y, Walsh T, et al. Emergence of plasmid-mediated colistin resistance mechanism mcr-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16(2):161–168. doi:10.1016/S1473-3099(15)00424-7
19. Rahmatallah N, El Rhaffouli H, Lahlou Amine I, et al. Consumption of antibacterial molecules in broiler production in Morocco. Vet Med Sci. 2018;4(2):80–90. doi:10.1002/vms3.89
20. Morris S, Cerceo E. Trends, epidemiology, and management of multi-drug resistant gram-negative bacterial infections in the hospitalized setting. Antibiotics. 2020;9(4):196. doi:10.3390/antibiotics9040196
21. Jaidane N, Bonnin R, Mansour W, et al. Genomic insights into colistin-resistant Klebsiella pneumoniae from a Tunisian teaching hospital. Antimicrob Agents Chemother. 2018;62(2):e01601–17. doi:10.1128/AAC.01601-17
22. Jaidane N, Naas T, Mansour W, et al. Genomic analysis of in vivo acquired resistance to colistin and rifampicin in Acinetobacter baumannii. Int J Antimicrob Agents. 2018;51(2):266–269. doi:10.1016/j.ijantimicag.2017.10.016
23. Mansour W, Haenni M, Saras E, et al. Outbreak of colistin-resistant carbapenemase-producing Klebsiella pneumoniae in Tunisia. J Glob Antimicrob Resist. 2017;10:88–94. doi:10.1016/j.jgar.2017.03.017
24. Saidani M, Messadi L, Chaouechi A, et al. High genetic diversity of enterobacteriaceae clones and plasmids disseminating resistance to extended-spectrum cephalosporins and colistin in healthy chicken in Tunisia. Microb Drug Resist. 2019;25(10):1507–1513. doi:10.1089/mdr.2019.0138
25. Saidani M, Messadi L, Mefteh J, et al. Various Inc-type plasmids and lineages of Escherichia coli and Klebsiella pneumoniae spreading blaCTX-M-15, blaCTX-M-1 and mcr-1 genes in camels in Tunisia. J Glob Antimicrob Resist. 2019;19:280–283. doi:10.1016/j.jgar.2019.05.007
26. Hassen B, Saloua B, Abbassi M, et al. mcr-1 encoding colistin resistance in CTX-M-1/CTX-M-15-producing Escherichia coli isolates of bovine and caprine origins in Tunisia. First report of CTX-M-15-ST394/D E. coli from goats. Comp Immunol Microbiol Infect Dis. 2019;67:101366. doi:10.1016/j.cimid.2019.101366
27. Hassen B, Abbassi M, Ruiz-Ripa L, et al. High prevalence of mcr-1 encoding colistin resistance and first identification of blaCTX-M-55 in ESBL/CMY-2-producing Escherichia coli isolated from chicken faeces and retail meat in Tunisia. Int J Food Microbiol. 2020;318:108478. doi:10.1016/j.ijfoodmicro.2019.108478
28. Berrazeg M, Hadjadj L, Ayad A, et al. First detected human case in Algeria of mcr-1 plasmid-mediated colistin resistance in a 2011 Escherichia coli isolate. Antimicrob Agents Chemother. 2016;60(11):6996–6997. doi:10.1128/AAC.01117-16
29. Bakour S, Olaitan A, Ammari H, et al. Emergence of colistin-and carbapenem-resistant Acinetobacter baumannii ST2 clinical isolate in Algeria: first case report. Microb Drug Resist. 2015;21(3):279–285. doi:10.1089/mdr.2014.0214
30. Yousfi H, Hadjadj L, Dandachi I, et al. Colistin-and carbapenem-resistant Klebsiella pneumoniae Clinical_Isolates: algeria. Microb Drug Resist. 2019;25(2):258–263. doi:10.1089/mdr.2018.0147
31. Chabou S, Leangapichart T, Okdah L, et al. Real-time quantitative PCR assay with Taqman® probe for rapid detection of MCR-1 plasmid-mediated colistin resistance. New Microbes New Infect. 2016;13:71–74. doi:10.1016/j.nmni.2016.06.017
32. Bachiri T, Lalaoui R, Bakour S, et al. First report of the plasmid-mediated colistin resistance gene mcr-1 in Escherichia coli ST405 isolated from wildlife in Bejaia, Algeria. Microb Drug Resist. 2018;24(7):890–895. doi:10.1089/mdr.2017.0026
33. Touati M, Hadjadj L, Berrazeg M, et al. Emergence of Escherichia coli harboring mcr-1 and mcr-3 gene in North West Algerian farmlands. J Glob Antimicrob Resist. 2019;21:132–137.
34. Drali R, Berrazeg M, Zidouni L, et al. Emergence of mcr-1 plasmid-mediated colistin-resistant Escherichia coli isolates from seawater. Sci Total Environ. 2018;642:90–94. doi:10.1016/j.scitotenv.2018.05.387
35. Wang J, Ma Z, Zeng Z, et al. The role of wildlife (wild birds) in the global transmission of antimicrobial resistance genes. Zool Res. 2017;38(2):55–80. doi:10.24272/j.issn.2095-8137.2017.024
36. Barbieri N, Nielsen D, Wannemuehler Y, et al. mcr-1 identified in avian pathogenic Escherichia coli (APEC). PLoS One. 2017;12(3):e0172997. doi:10.1371/journal.pone.0172997
37. Moawad A, Hotzel H, Neubauer H, et al. Antimicrobial resistance in Enterobacteriaceae from healthy broilers in Egypt: emergence of colistin-resistant and extended-spectrum β-lactamase-producing Escherichia coli. Gut Pathog. 2018;10:39. doi:10.1186/s13099-018-0266-5
38. Khalifa H, Ahmed A, Oreiby A, et al. Characterisation of the plasmid-mediated colistin resistance gene mcr-1 in Escherichia coli isolated from animals in Egypt. Int J Antimicrob Agents. 2016;47(5):413–414. doi:10.1016/j.ijantimicag.2016.02.011
39. Sadek M, Poirel L, Nordmann P, et al. Draft Genome Sequence of a mcr-1/IncI2-carrying multidrug-resistant Escherichia coli B1: ST101 isolated from meat and meat products in Egypt. J Glob Antimicrob Resist. 2019;20:41–42. doi:10.1016/j.jgar.2019.11.015
40. Hammad A, Hoffmann M, Gonzalez-Escalona N, et al. Genomic features of colistin resistant Escherichia coli ST69 strain harboring mcr-1 on IncHI2 plasmid from raw milk cheese in Egypt. Infect Genet Evol. 2019;73:126–131. doi:10.1016/j.meegid.2019.04.021
41. Kieffer N, Ahmed M, Elramalli A, et al. Colistin-resistant carbapenemase-producing isolates among Klebsiella spp. and Acinetobacter baumannii in Tripoli, Libya. J Glob Antimicrob Resist. 2018;13:37–39. doi:10.1016/j.jgar.2018.02.020
42. Wang R, van Dorp L, Shaw L, et al. The global distribution and spread of the mobilized colistin resistance gene mcr-1. Nat Commun. 2018;9(1):1179. doi:10.1038/s41467-018-03205-z
43. Maamar E, Alonso C, Hamzaoui Z, et al. Emergence of plasmid-mediated colistin-resistance in CMY-2-producing Escherichia coli of lineage ST2197 in a Tunisian poultry farm. Int J Food Microbiol. 2018;269:60–63. doi:10.1016/j.ijfoodmicro.2018.01.017
44. Yanat B, Machuca J, Yahia R, et al. First report of the plasmid-mediated colistin resistance gene mcr-1 in a clinical Escherichia coli isolate in Algeria. Int J Antimicrob Agents. 2016;48(6):760–761. doi:10.1016/j.ijantimicag.2016.09.003
45. Berrazeg M, Deriet A, De Keersmaecker S, et al. Whole-genome sequencing of multidrug-resistant Escherichia coli strains harboring the mcr-1 gene, isolated from seawater of the algiers coast in Algeria. Microbiol Resour Announc. 2019;8(34):e00638–19. doi:10.1128/MRA.00638-19
46. Elbediwi M, Li Y, Paudyal N, et al. Global burden of colistin-resistant bacteria: mobilized colistin resistance genes study (1980–2018). Microorganisms. 2019;7(10):461. doi:10.3390/microorganisms7100461
47. Al-Kadmy I, Ibrahim S, Al-Saryi N, et al. Prevalence of genes involved in colistin resistance in Acinetobacter baumannii: first report from Iraq. Microb Drug Resist. 2020;26(6):616–622. doi:10.1089/mdr.2019.0243
48. Abdulzahra A, Khalil M, Elkhatib W. First report of colistin resistance among carbapenem-resistant Acinetobacter baumannii isolates recovered from hospitalized patients in Egypt. New Microbes New Infect. 2018;26:53–58. doi:10.1016/j.nmni.2018.08.007
49. Can F, Menekse S, Ispir P, et al. Impact of the ST101 clone on fatality among patients with colistin-resistant Klebsiella pneumoniae infection. J Antimicrob Chemother. 2018;73(5):1235–1241. doi:10.1093/jac/dkx532
50. Mshana S, Fritzenwanker M, Falgenhauer L, et al. Molecular epidemiology and characterization of an outbreak causing Klebsiella pneumoniae clone carrying chromosomally located blaCTX-M-15 at a German University-Hospital. BMC Microbiol. 2015;15:122. doi:10.1186/s12866-015-0460-2
51. Mairi A, Pantel A, Sotto A, et al. OXA-48-like carbapenemases producing Enterobacteriaceae in different niches. Eur J Clin Microbiol Infect Dis. 2018;37(4):587–604. doi:10.1007/s10096-017-3112-7
52. World Health Organization. Global Antimicrobial Resistance Surveillance System (GLASS): the detection and reporting of colistin resistance. Licence: CC BY-NC-SA 3.0 IGO. World Health Organization; 2018. Available from: https://apps.who.int/iris/handle/10665/277175.
53. Grami R, Mansour W, Mehri W, et al. Impact of food animal trade on the spread of mcr-1-mediated colistin resistance, Tunisia, July 2015. Euro Surveill. 2016;21(8):30144. doi:10.2807/1560-7917.ES.2016.21.8.30144
54. Chabou S, Leulmi H, Rolain JM. Emergence of mcr-1-mediated colistin resistance in Escherichia coli isolates from poultry in Algeria. J Glob Antimicrob Resist. 2019;16:115–116. doi:10.1016/j.jgar.2018.12.012
55. Nabti LZ, Sahli F, Hadjadj L, et al. Autochthonous case of mobile colistin resistance gene mcr-1 from a uropathogenic Escherichia coli isolate in Sétif Hospital, Algeria. J Glob Antimicrob Resist. 2019;19:356–357. doi:10.1016/j.jgar.2019.10.006
56. Belbel Z, Lalaoui R, Bakour S, et al. First report of colistin resistance in an OXA-48- and a CTX-M-15 producing Klebsiella pneumoniae clinical isolate in Algeria due to PmrB protein modification and mgrB inactivation. J Glob Antimicrob Resist. 2018;14:158–160. doi:10.1016/j.jgar.2018.06.014
57. Nabti LZ, Sahli F, Ngaiganam EP, et al. Development of real-time PCR assay allowed describing the first clinical Klebsiella pneumoniae isolate harboring plasmid-mediated colistin resistance mcr-8 gene in Algeria. J Glob Antimicrob Resist. 2019;20:266–271.
58. Elnahriry SS, Khalifa H, Soliman A, et al. Emergence of plasmid-mediated colistin resistance gene mcr-1 in a clinical Escherichia coli isolate from Egypt. Antimicrob Agents Chemother. 2016;60(5):3249–3250. doi:10.1128/AAC.00269-16
59. Zafer M, El-Mahallawy H, Abdulhak A, et al. Emergence of colistin resistance in multidrug-resistant Klebsiella pneumoniae and Escherichia coli strains isolated from cancer patients. Ann Clin Microbiol Antimicrob. 2019;18(1):40. doi:10.1186/s12941-019-0339-4
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
