Back to Journals » Infection and Drug Resistance » Volume 10
Peptide consensus sequence determination for the enhancement of the antimicrobial activity and selectivity of antimicrobial peptides
Authors Almaaytah A, Ajingi Y, Abualhaijaa A, Tarazi S, Alshar’i N , Al-Balas Q
Received 3 August 2016
Accepted for publication 11 November 2016
Published 29 December 2016 Volume 2017:10 Pages 1—17
DOI https://doi.org/10.2147/IDR.S118877
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Ammar Almaaytah,1 Ya’u Ajingi,2 Ahmad Abualhaijaa,2 Shadi Tarazi,2 Nizar Alshar’i,3 Qosay Al-Balas3
1Department of Pharmaceutical Technology, Faculty of Pharmacy, Jordan University of Science and Technology, Irbid, Jordan; 2Department of Applied Biological Sciences, Faculty of Science and Arts, Jordan University of Science and Technology, Irbid, Jordan; 3Department of Medicinal Chemistry, Faculty of Pharmacy, Jordan University of Science and Technology, Irbid, Jordan
Abstract: The rise of multidrug-resistant bacteria is causing a serious threat to the world’s human population. Recent reports have identified bacterial strains displaying pan drug resistance against antibiotics and generating fears among medical health specialists that humanity is on the dawn of entering a post-antibiotics era. Global research is currently focused on expanding the lifetime of current antibiotics and the development of new antimicrobial agents to tackle the problem of antimicrobial resistance. In the present study, we designed a novel consensus peptide named “Pepcon” through peptide consensus sequence determination among members of a highly homologous group of scorpion antimicrobial peptides. Members of this group were found to possess moderate antimicrobial activity with significant toxicity against mammalian cells. The aim of our design method was to generate a novel peptide with an enhanced antimicrobial potency and selectivity against microbial rather than mammalian cells. The results of our study revealed that the consensus peptide displayed potent antibacterial activities against a broad range of Gram-positive and Gram-negative bacteria. Our membrane permeation studies displayed that the peptide efficiently induced membrane damage and consequently led to cell death through the process of cell lysis. The microbial DNA binding assay of the peptide was found to be very weak suggesting that the peptide is not targeting the microbial DNA. Pepcon induced minimal cytotoxicity at the antimicrobial concentrations as the hemolytic activity was found to be zero at the minimal inhibitory concentrations (MICs). The results of our study demonstrate that the consensus peptide design strategy is efficient in generating peptides.
Keywords: peptide, scorpion, venom, peptide design, antimicrobial peptides, consensus sequence
Introduction
Antimicrobial peptides (AMPs) are produced by multicellular organisms that supply them with a formidable defense mechanism against invading pathogens.1 AMPs display a net positive charge ranging between +2 and +9 with hydrophobic residues amounting to 30% in proportion of the total amino acids residues found in the mature peptide.2 These positively charged amino acid residues are separated by hydrophobic regions and patches which enable them to fold into amphiphilic and amphipathic conformations during contact with target membranes.3 AMPs are potentially active against a broad range of microorganisms including Gram-positive and Gram-negative bacteria, yeast, fungi, and enveloped viruses.2 AMPs show an efficient broad spectrum of activity against multidrug-resistant strains and are believed to exhibit a very low probability of generating bacterial resistance against them.2 The broad spectrum activity and diverse mechanism of action associated with AMPs have qualified them to be considered as a potential source of novel therapeutic antibiotics. One of the major drawbacks of AMPs responsible for hindering the development of this class of bioactive molecules is related to AMPs low antibacterial target selectivity and high toxicity against mammalian cells.4 Current research is focused on overcoming the issue of AMPs-induced mammalian cell toxicity.
Scorpion venom is composed of a complex cocktail of biologically active peptides.5 The scorpion venom peptides are classified into disulfide-bridged peptides, which usually target membrane bound ion channels and the smaller group of non-disulfide-bridged peptides (NDBPs). The NDBPs are a novel group that have gained interest only recently. NDBPs are 13–50 amino acids long, structurally diverse and display a minimal degree of sequence similarities.6 The scorpion NDBPs have been reported to display bradykinin potentiating, hemolytic, antimicrobial, and immune modulating activities.5
Recently, the scorpion NDBPs have been classified based in principle on their pharmacological activity, peptide length, and sequence into five different subfamilies, with group number four being the largest comprising of peptide members having chain lengths of 13–19 amino acids with all members displaying a degree of antimicrobial activity. This group of peptides is thought to be synthesized and secreted as a result of pathogenic invasion or environmental pressure imposed on the different scorpion species.7
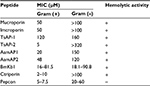
Peptides belonging to group four of scorpion NDBPs (Table 1) were shown to possess weak antimicrobial activities and significant hemolytic and cytotoxic activity against mammalian cells and human erythrocytes, an issue that limits their application for further antimicrobial and clinical development.
Several methods have been applied to enhance the selectivity of AMPs and improve their therapeutic index including AMP congeners, AMP mimetics, peptide libraries, and hybrid AMPs.8,9 A semi-rational design employing consensus peptide design has proven to be a popular and successful strategy in designing peptides with enhanced stability with regard to their physicochemical properties.10 The consensus design is based on the use of evolutionary information derived from homologous peptide sequences.11 Consensus design has been effectively used to generate a consensus protein with improved potency, stability, and immunogenicity profile, one successful example is the alignment of 25 interferon-α subtypes, which generated interferon alfacon-1 that has been developed successfully as a therapeutic drug and received US Food and Drug Administration approval in 1997.12
In this study, we have designed a novel consensus AMP named “Pepcon” by employing the homologous peptides of scorpion NDBPs belonging to group number four as a platform for amino acid consensus sequence determination and analysis for the prediction and design of a universal potent AMP with increased activity and selectivity toward microbial cells and an increased selectivity index that would minimize the problem of cell mammalian toxicity. We investigated the antimicrobial activity of the rationally designed peptide against standard representative strains of Gram-positive and Gram-negative bacteria in addition to bacterial strains displaying multidrug resistance. Additionally, the killing kinetics and the molecular mechanism of action responsible for the antimicrobial activity were evaluated using time-kill and beta-galactosidase assays. The toxicity of the peptide and its target selectivity were evaluated using hemolytic assays.
Materials and methods
Bioinformatic analysis and peptide design
Homologous members of group number four of scorpion NDBPs were shown to possess moderate antibacterial activities against Gram-positive and Gram-negative bacteria and induced a significant degree of hemolysis against mammalian cells. These peptides were employed as a platform for consensus sequence design and prediction, which is based on calculating the order of the most frequent amino acid found at each position in the homologous peptides belonging to group four of scorpion NDBPs. Multiple sequence alignment and consensus sequence determination for the peptides were performed using ClustalW2 software (European Molecular Biology Laboratory, European Bioinformatics Institute, Cambridgeshire, UK). The generated consensus sequence was named Pepcon and was subjected to peptide synthesis in order to proceed with the in vitro antimicrobial assays.
Peptide synthesis and purification
Synthetic replicates of the short consensus sequence peptide named Pepcon were synthesized by GL Biochem Ltd. (Shanghai, China). Purification and identification of the synthetic peptide were performed using high-performance liquid chromatography (HPLC) with (>95%) purity and electrospray ionization mass spectrometry.
Microorganisms
The antimicrobial activity of the peptide Pepcon was tested and determined against four Gram-positive bacterial strains, including two standard strains of Staphylococcus epidermis (ATCC 12228), S. aureus (ATCC 29213), in addition to two methicillin-resistant strains of S. aureus (ATCC 43300, 33591). Additionally, the antimicrobial activity of the peptide was evaluated against five strains of Gram-negative bacteria including Escherichia coli (ATCC 25922), Salmonella enterica (ATCC 10708), Pseudomonas aeruginosa (ATCC 27853), Acinetobacter baumannii (ATCC 19606), and Klebsiella pneumoniae (ATCC 13883).
Minimum inhibitory concentrations (MICs) and minimum bactericidal concentrations (MBCs)
The MIC and the MBC values of the peptide were tested and determined according to the method previously described by Wiegand et al.13 Ten microliters aliquot of thawed bacteria immersed in warm water was cultured using surface spreading into a fresh pre-sterilized nutrient agar plates and then incubated at 37°C for 18–24 hours. The next day, three to five pure colonies from the overnight culture were transferred into a sterile tube containing 10 mL of Mueller Hinton Broth (MHB) and vortexed accordingly. Turbidity was adjusted to 0.5 McFarland standards (108 CFU/mL) by measuring absorbance at 625 nm using a spectrophotometer.
The bacterial suspension was utilized within 30 min after turbidity adjustment to avoid duplication of the bacterial cells. The bacterial suspension was then mixed thoroughly and diluted by 100-fold, 50 µL of the diluted suspension was added to each well in a sterile 96 well plate including the positive and negative controls.
Nine different concentrations of the peptide were prepared each from a single stock and 50 µL was taken from each well and added to all wells containing the bacterial suspension, the plate was then incubated at 37°C for 18–24 hours. The following day, the plates were examined for any growth and inhibition was determined by measuring absorbance at 570 nm using a microplate reader (BioTek Instruments, Inc., Winooski VT, USA).
The MBC was then determined by withdrawing 10 µL aliquots from both clear and positive control wells, after which they were transferred into a pre-sterilized labeled agar plates and incubated overnight to determine the MBC. The following day, plates were examined for bacterial growth and the wells displaying no growth were considered to be the values of the corresponding MBC.
Time-kill studies against exponentially and stationary phase growing bacteria
Time-kill studies for exponentially growing bacterial strains were done according to the methods previously described by Liu et al.14 Bacterial strains were cultured in MHB and incubated overnight at 37°C, the following day, turbidity was adjusted to McFarland standards. Adjusted bacterial suspension was incubated at 37°C with shaking at 220 rpm to allow bacterial cells to reach the exponential phase. The culture was then diluted in fresh MHB to adjust the concentration to approximately 107 CFU/mL, after which 150 µL of the bacterial suspension was added to each well of the 96 well plate.
Nine peptide solutions were prepared in MHB ranging from 1MIC, 2MIC, 3MIC, and 4MIC. Each peptide solution was prepared individually from a single stock solution and all concentrations were prepared with twice the desired concentration as a twofold dilution will occur in the 96 well plate. Afterward, 150 µL of each peptide concentration was added to nine wells of the 96 well plate.
Rates of killing were determined by measuring the reduction in viable bacterial count at 0–60 min (10 min apart), 120 min apart, and finally at 480 min with a fixed concentration of the peptide. At each time point, 10 µL aliquots were withdrawn from each well and serially diluted into pre-sterilized nutrient agar plates, and afterward, the plates were incubated overnight at 37°C.
For the exponentially growing bacteria, the bacterial strains were cultured overnight in MHB. Bacterial suspensions were diluted into phosphate-buffered saline (PBS) to adjust cell density to approximately 1×106 CFU/mL and then 150 µL of bacterial suspension was added to each well of the 96 well plate.
Four peptide solutions were prepared into PBS, ranging from 1MIC, 2MIC, 3MIC, and 4MIC.
Rates of killing were determined by measuring the reduction in viable bacterial count as mentioned previously.
β-Galactosidase assay
The β-galactosidase assay was performed according to the method described by Luca et al.15 E. coli cells were grown at 37°C in lactose broth (to induce E. coli lac operon to express β-galactosidase enzyme gene). The culture was removed from the incubation chamber when the bacterial growth reached approximately 0.8 optical density (OD) at 590 nm. Turbidity was adjusted to 0.5 McFarland standards by measuring the absorbance at 625 nm using a spectrophotometer. E. coli suspension was diluted into PBS to adjust the concentration to approximately 1×106 CFU/100 µL. Then, 100 µL of this suspension was added to each well of the 96 well plate, also 30 µL of ortho-nitrophenyl (o-nitrophenol)-β-galactoside (ONPG) was added to each well.
Four peptide solutions were prepared into PBS ranging from 1MIC, 2MIC, 3MIC, and 4MIC. The hydrolysis of ONPG to o-nitrophenol by β-galactosidase enzyme was monitored at 405 nm using enzyme-linked immunosorbent assay (ELISA) over different time points.
Extraction of genomic DNA
The isolation of genomic DNA from E. coli was performed according to the Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA) protocol. Extracted DNA was considered to be pure if an optical ratio of DNA was in the range of OD 260/OD 280 ≥ 1.8. The concentration of purified E. coli DNA, the absorbance at 260 nm, and the absorbance at 280 nm were calculated using the Thermo Scientific™ NanoDrop™ 1000 (Thermo Fisher Scientific Inc., Waltham, MA, USA) and a DNA rehydration solution (Tris-ethylenediaminetetraacetic acid) was used to calibrate the NanoDrop 1000 system.
DNA gel retardation
Sixty µm of the peptide was dissolved into PBS, seven DNA/peptide samples were prepared, and the peptide/DNA ratios were calculated for each sample. Samples were incubated at room temperature for 10 min, after adding 2 µL of loading dye, the migration of DNA was assessed by agarose gel electrophoresis using 0.8% agarose gel prepared by adding 1.5 g of agarose into 150 mL of 1× tris borate ethylenediaminetetraacetic acid buffer (Sigma Aldrich, St. Louis, MO, USA). Gel retardation was visualized under UV illumination using a Gel Doc_IT 310 imaging system.
Hemolysis assay
Two milliliters of human blood (Sigma Aldrich) was placed into a 50 mL centrifuge tube followed by centrifugation at 3000 × g for 5 min, after discarding the supernatant, the cell pellet was re-suspended in 25 mL of PBS and centrifuged as above. Finally, the cell pellet was re-suspended in a pre-sterile tube containing 50 mL PBS to reach a final concentration of 4% red blood cells (RBCs). Eight peptide solutions with different concentrations were prepared into PBS, and then 2 mL of each peptide concentration was added to 2 mL of erythrocyte suspension.
Positive controls were prepared by adding 5 µL of Triton X-100 (Santa Cruz Biotechnology, Dallas, TX, USA) to 2 mL of RBC suspension (positive control), and negative controls were prepared by adding 2 mL of RBC suspension only (negative control), the suspension was incubated with the peptides for 60 min at 37°C. After the incubation step, the tubes were gently vortexed, and 1 mL of each sample was removed and placed into a pre-sterilized Eppendorf tubes and then centrifuged for 5 min at 3000 × g and 200 µL of each supernatant was placed into a well of a 96 well plate. Absorbance was measured at λ=550 nm with the aid of an ELISA reader.
The percentage hemolysis was calculated according to the following equation:
%Hemolysis = (A _ AO) / (AX _ AO) × 100
where A = absorbance of test solution, AO = absorbance of negative control, and AX = absorbance of positive control.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Results
Peptide design
The peptide employed in this study was designed based on consensus sequence determination method which is based on the computed order of the most frequent amino acid that is present at every position after multiple sequence alignment has been carried out between the parent peptides that were all proven to exert potential antimicrobial activities at different concentrations but suffered from low therapeutic indices as discussed previously in the “Introduction” section. Employing multiple sequence alignment according to the method described previously, a novel peptide analogue named Pepcon composed of 18 amino acid residues was derived from the selected homologous peptides of group number four of scorpion NDBPs by employing consensus sequence determination and as shown in Table 2.
Synthesis and purification of Pepcon peptide
The Pepcon peptide was synthesized according to the solid-phase method and standard Fmoc chemistry by GL Biochem, Shanghai, China. The purity of peptides was assessed by HPLC and its identity was confirmed by performing positive electrospray ionization mass spectrometric analysis. Pepcon was synthesized with high purity of >95%. The synthetic analogue of Pepcon displays a major HPLC and mass peak in the +2 charge state of 941.68 Da (Figures S1 and S2).
Bacterial susceptibility assay
The peptide proved to exhibit strong antimicrobial activity against all the tested bacterial pathogens. It was able to inhibit the growth of the entire Gram-positive bacterial strains employed in the study including both methicillin-resistant S. aureus (ATCC 43300), S. aureus (ATCC 33591) in addition to the control strain of S. aureus (ATCC 29213) at a MIC value of 5 µM, Pepcon also displayed moderate activity against S. epidermidis (ATCC 12228) with a MIC of 7.5 µM. Table 3 displays the MIC values of Pepcon against the tested bacterial strains. Additionally, Pepcon displayed an inhibitory activity against all the Gram-negative bacterial strains studied, with A. baumannii (ATCC 19606) having the lowest MIC of 20 µM, while S. enterica (ATCC 10708) and K. pneumoniae (ATCC 13883) were inhibited at a concentration of 60 µM respectively. The synthetic peptide inhibited the growth of E. coli (ATCC 25922) at a MIC of 50 µM while P. aeruginosa was inhibited at a MIC of 40 µM.
  | Table 3 Minimum inhibitory concentrations (MICs) of the consensus peptide Pepcon against the tested bacterial strains |
The bactericidal activity of Pepcon peptide revealed that the MBC is the same as the MIC, and this observation clearly displays that Pepcon exerts a bactericidal behavior rather than bacteriostatic against the pathogenic strains tested (data not shown).
Activity of Pepcon against logarithmic growing bacteria
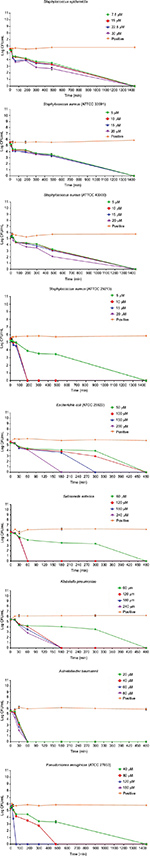
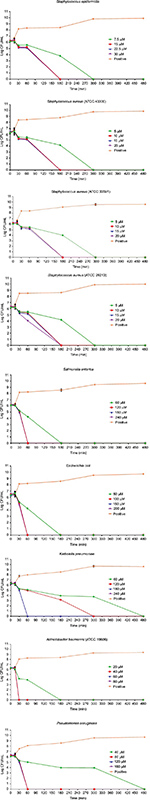
The results of the killing kinetics of Pepcon against logarithmic growing bacterial pathogens are shown in Figure 1. The peptide was able to completely eliminate (99.9% killing) almost all of bacterial strains tested within 60–180 min of exposure after incubation at 37°C with the corresponding concentrations of the peptide. At twofold MIC, all the Gram-positive strains employed in this study were eradicated within 180 min of exposure. Furthermore, at twofold MIC, E. coli, S. enterica, and P. aeruginosa were eliminated within 60 min of incubation, while at twofold MIC, A. baumannii was totally eradicated within 30 min of exposure. Nevertheless, threefold MIC was required by K. pneumoniae before it became completely eliminated after 60 min of exposure at 37°C. The killing kinetics of Pepcon is in line with that of other cationic AMPs reported previously and signifies the rapid bactericidal action of Pepcon against the bacterial pathogens.16
Activity of Pepcon against stationary growing bacteria
Time-kill assay for the stationary phase slow growing bacterial strains is represented in Figure 2. Pepcon inhibited the growth of S. aureus (ATCC 29213) within 200 min of exposure at twofold MIC concentration, while the other Gram-positive bacterial strains tested took up to 1400 min before they have been eradicated. Pepcon exhibited a bactericidal activity (99.9%) of killing E. coli at fourfold MIC within 180 min and K. pneumoniae at twofold MIC at the same time, respectively. The peptide totally eradicated the growth of S. enterica at twofold concentration of the MIC within 60 min, likewise P. aeruginosa was killed at threefold MIC within the same period. The most sensitive of all the bacterial strains to the peptide was the Gram-negative A. baumannii strain, which was completely eliminated within 60 min of incubation with the peptide at onefold MIC concentration. The potential ability of Pepcon to entirely eradicate the bacterial pathogens tested at different times of exposure over a varying range of concentrations indicates that it exhibits a broad antimicrobial spectrum with potent cytolytic and bactericidal activity.
β-Galactosidase assay
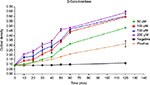
The conversion of ONGP to galactose and o-nitrophenol due to the action of β-galactosidase enzyme as a result of membrane destruction and an incremental increase in cytoplasmic release of intracellular components and permeability, was monitored at 405 nm spectrophotometrically (Figure 3). Pepcon, at varying degrees of concentrations, caused consistent increase in E. coli cytoplasmic membrane permeability as shown by the fast rise in the values of OD over time, which is equivalent to o-nitrophenol formation. The absorbance increased with increased time in the dose-dependent manner and reached its peak value after 60 min of incubation for the three concentrations employed in this study.
DNA binding assay
The purity and concentration of the genomic DNA extracted from E. coli were determined using the Nano drop 1000 system. Seven DNA samples were been prepared by adding different volumes of the peptide from a single stock to 4 µL of 95.9 ng/µL DNA. At the end of the incubation time, the samples were run on 1.5% agarose gel and the migration was monitored using a Gel Doc_ IT 310 imaging system. Binding of the peptide to DNA was analyzed by the ability of the peptide to inhibit the migration of the DNA along the agarose in comparison to the control DNA sample without the peptide. The results revealed that there was no retardation in the migration of DNA by the peptide which is indicative that Pepcon lacks the ability to bind DNA and excludes the possibility that peptide is causing microbial death through binding to intracellular targets (Figure 4).
Hemolytic assay
To determine the toxicity of the peptide against mammalian cells, the hemolytic activity was assessed against 4% human erythrocytes (RBCs) at concentrations ranging from 1 µM to 100 µM of the peptide. Absorbance was measured at λ=550 nm using an ELISA reader after 1 and 2 hours of incubation, respectively.
Pepcon displayed 0% hemolytic activity when incubated for both 1 and 2 hour periods at concentrations of 1 µM to 40 µM. At a concentration of 60 µM, the percentage hemolysis reported was 4.9% and 6.5%. At 80 µM, percentage hemolysis was found to be of 22.3% and 22%, respectively. At 100 µM, the percentage hemolysis reported was 37.7% and 39.2% (Table 4). These results indicated that the peptide was inducing mild hemolysis at the concentrations required for the inhibition of bacterial growth.
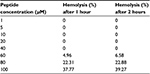  | Table 4 Hemolytic activity of Pepcon on human erythrocytes after 60 and 120 minutes of incubation |
Discussion
Hadrurin from the venom of the scorpion Hadrurus aztecus and Scorpine from the venom of the scorpion Pandinus imperator were the first AMPs discovered from scorpion venoms.16 Since then, a rapid increase in the isolation and recognition of scorpion venom multipurpose AMPs from different species has been reported in literature.5 Prior to the discovery of the first AMPs within scorpion venoms, little knowledge was known concerning the natural innate immune defense of scorpion venom glands toward protection against bacterial infection. It has been hypothesized that scorpions spray venom-loaded peptides on their body surfaces as means of cleaning and probably to protect themselves against microbial invasion.17 Scorpion venom peptides are classified into two main major groups, the disulfide-bridged peptide, which usually target membrane bound ion channels, and the NDBPs, which exhibit multifunctional biological activities.18,19 The structural and functional activities and sequence analysis of most scorpion NDBPs indicate that the scorpion AMPs are members of the widely distributed family of cationic peptides found in nature which has been recognized and isolated from plants, and insects, to more complex immune systems and highly advanced mammals.20 However, scorpion AMPs that were identified from the different scorpion species seem to display only moderate antimicrobial activity in addition to being highly cytotoxic to normal mammalian cells and do not seem to display any selectivity toward microbial cells. The environmental pressure faced by scorpions could explain the mild antimicrobial selectivity of scorpion AMPs, as scorpion venoms serve to protect the host organism from predators and aim to capture prey, scorpions venom AMPs did not diverge through gene duplication to elicit a selective antimicrobial behavior against bacteria as this did not meet the needs demanded by the environmental pressure applied on these organisms.
In this project, we implemented a design strategy to enhance the antimicrobial activity and the selectivity of scorpion AMPs by employing amino acid consensus sequence identification as a platform for the design of peptides with an enhanced therapeutic index when compared with the parent peptides. The strategy employed depended on aligning the sequences of the parent peptides and the calculation of the similar sequence motifs most commonly found on each amino acid position. The resultant peptide named Pepcon displayed potent antibacterial activities against a broad range of Gram-positive and Gram-negative bacterial strains that were employed in this study. Pepcon was able to inhibit the growth of S. epidermidis at a concentration of 7.5 µM and the rest of the Gram-positive bacteria, including standard (ATCC 29213) and methicillin-resistant S. aureus (ATCC: 33591, 43300), were all inhibited at MIC of 5 µM. Additionally, Pepcon managed to completely inhibit the growth of S. enterica and K. pneumoniae at a concentration of 60 µM each, while the growth of E. coli and P. aeruginosa was inhibited at a concentration of 50 µM and 40 µM, respectively. A. baumannii was the most susceptible strain to the peptide among all the Gram-negative strains employed in this study, the growth of which was inhibited at a concentration of 20 µM, this indicates that the synthetic peptide exhibits an enhanced activity in comparison to the parent peptides, the majority of which have higher MIC values greater than that of Pepcon (5–300 µM). Pepcon displayed a higher potency against Gram-positive bacterial strains compared with the Gram-negative bacteria in a similar manner with the parent peptides. The difference in antimicrobial activity, which was directed against Gram-positive bacteria, can best be explained due to the fact that Gram-positive bacteria only possess a thick peptidoglycan single lipid bilayer composed of teichuronic or teichoic acids, while Gram-negative bacteria contain an additional outer membrane that is made up of lipopolysaccharides rich in highly acidic lipids which can efficiently decrease the net negative charge on the bacterial membrane and neutralize the cationic charge found on the scorpion AMPs which is responsible for the initial step required for antimicrobial inhibition, and that is the binding of the cationic peptide electrostatically on the surface membrane of target cells.20 The concentration of the novel peptide needed to inhibit the growth of the Gram-negative organisms is significantly lower than that of the parent peptides (Table 2). This clearly shows a marked increase in the activity of the designed synthetic peptide over the parent peptides it was derived from. The presence of the positively charged amphipathic feature, coupled with moderate hydrophobicity of Pepcon, fit the structure of the selective cationic peptides which allows them to fold into the precise conformation required to display selective toxicity against planktonic unicellular organisms’ membranes rather than mammalian cells. 21
The data obtained for Pepcon from the time-kill studies of the logarithmic growing bacteria showed that the peptide managed to kill all the Gram-positive bacteria within 180 min of incubation at twofold the MIC of each strain. Likewise, when tested against the Gram-negative bacteria, the peptide completely eradicated the growth of all strains within 60 min of exposure at twofold the MIC. The results of time-kill assay for the stationary phase growing bacteria are shown graphically in Figure 2. At twofold the MIC, Pepcon eliminated the growth of S. aureus within 200 min of incubation, but 1400 min were required to totally kill the other Gram-positive strains used in the study, this is probably attributed to the fact that most Staphylococcus species employed in this study are resistant to multiple conventional antibiotics. At fourfold the MIC, Pepcon managed to eradicate the growth of E. coli within 180 min and K. pneumoniae was totally killed at twofold its MIC within the same time span. The ability of the peptide to potentially eliminate the growth of all the bacterial strains rapidly and in a short time period proved that Pepcon exhibits a bactericidal behavior due to the rapid killing kinetics and is consistent with other results from previously mentioned cationic AMPs.
The capability of the novel peptide to permeate the bacterial membranes was assessed by the ability of the peptide to damage bacterial membranes and causes the release of the intracellular components of E. coli cells and thus causing the conversion of ONPG to galactose and o-nitrophenol by the β-galactosidase enzyme. The fast rate increase observed in the values of OD with time is equivalent to the rate at which o-nitrophenol is formed and indicates that the Pepcon is responsible for significant membrane damage coupled with a significant release of the bacterial intracellular components, such as the β-galactosidase enzyme. Membrane permeation is not the only destructive factor that governs the activity of AMPs against microbial cells, several reports indicated that AMPs could induce multiple inhibitory effects by affecting the functions of other cell structures, such as nucleic acids, the cell wall, and cellular proteins.1–3 For instance, the binding action of Buforin IIb to either DNA or RNA results into an excessive release of calcein from liposomes which is coupled with cell lysis and death.22 To determine if Pepcon was responsible for the antimicrobial activity due to other mechanisms other than membrane permeability, we assessed the ability of the peptide to bind bacterial DNA. According to our results, Pepcon displayed weak DNA binding ability against genomic bacterial DNA and caused no DNA binding retardation at all (Figure 4). This data suggest that DNA is not one of the targets responsible for the antibacterial activity of Pepcon and could also indicate the peptide exhibits weak DNA binding abilities and a low potential to cause genotoxicity to mammalian cells.
The toxicity of Pepcon toward mammalian cells was examined by determining the hemolytic activity of the peptide against human erythrocytes. The peptide induced 0% hemolysis at a concentration range of 1 µM–40 µM, while at a concentration of 60 µM, the percentage hemolysis reported was 4.9% and 6.5%, at 80 µM, it was found to be of 22.3% and 22%. At 100 µM, the percentage hemolysis was 37.7% and 39.2% (Table 4). The low hemolytic activity of Pepcon observed at these concentrations which correspond to the peptide’s MIC values is an indication of its selective toxicity against prokaryotic membrane over that of eukaryotes, in contrast to the parent peptides that were all shown to exhibit potent hemolytic activity, Pepcon was found to induce mild hemolysis against human RBCs even at the highest concentration of 100 µM. The results from the hemolytic assay proved that the synthetic peptide is highly active and exerts a very weak, if any, hemolytic activity even at high concentrations when compared to the parent peptides. In general, the consensus design method employed in this study proved to be successful in generating peptides with enhanced therapeutic index over the parent peptides. This design strategy could be implemented in future studies for the generation of several novel peptide analogues with enhanced performance over the AMPs found and identified in nature.
Acknowledgment
This study was funded by the Deanship of Research at the Jordan University of Science and Technology (grant number 85/2014).
Disclosure
All of the authors involved in this manuscript declare that they have no conflicts of interest in this work. The authors have no affiliations with or involvement in any organization or entity with any financial interest (such as honoraria; educational grants; participation in speakers’ bureaus; membership, employment, consultancies, stock ownership, or other equity interest; and expert testimony or patent-licensing arrangements) in the subject matter or materials discussed in this manuscript.
References
Fjell CD, Hiss JA, Hancock RE, Schneider G. Designing antimicrobial peptides: form follows function. Nat Rev Drug Discov. 2011;11(1):37–51. | ||
Zasloff M. Antimicrobial peptides of multicellular organisms. Nature. 2002;415(6870):389–395. | ||
Hancock RE, Sahl HG. Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nat Biotechnol. 2006;24(12):1551–1557. | ||
Almaaytah A, Zhou M, Wang L, Chen T, Walker B, Shaw C. Antimicrobial/cytolytic peptides from the venom of the North African scorpion, Androctonus amoreuxi: biochemical and functional characterization of natural peptides and a single site-substituted analog. Peptides. 2012;35(2):291–299. | ||
Almaaytah A, Albalas Q. Scorpion venom peptides with no disulfide bridges: a review. Peptides. 2014;51:35–45. | ||
Zeng XC, Corzo G, Hahin R. Scorpion venom peptides without disulfide bridges. IUBMB Life. 2005;57(1):13–21. | ||
Gao B, Tian C, Zhu S. Inducible antibacterial response of scorpion venom gland. Peptides. 2007;28(12):2299–2305. | ||
Peters BM, Shirtliff ME, Jabra-Rizk MA. Antimicrobial peptides: primeval molecules or future drugs? PLoS Pathog. 2010;6(10):e1001067. | ||
Tharad M, Samuchiwal SK, Bhalla K, Ghosh A, Kumar K, Kumar S, Ranganathan A. A three-hybrid system to probe in vivo protein-protein interactions: application to the essential proteins of the RD1 complex of M. tuberculosis. PloS One. 2011;6(11):e27503. | ||
Porebski BT, Buckle AM. Consensus protein design. Protein Eng Des Sel. 2016:29(7):245–251. | ||
Steipe B. Evolutionary approaches to protein engineering. In: Combinatorial Chemistry in Biology. Springer: Berlin Heidelberg; 1999:55–86. | ||
Jacobs SA, Diem MD, Luo J, et al. Design of novel FN3 domains with high stability by a consensus sequence approach. Protein Eng Des Sel. 2012;25(3):107–117. | ||
Wiegand I, Hilpert K, Hancock RE. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc. 2008;3(2):163–175. | ||
Liu Y, Knapp KM, Yang L, Molin S, Franzyk H, Folkesson A. High in vitro antimicrobial activity of β-peptoid–peptide hybrid oligomers against planktonic and biofilm cultures of Staphylococcus epidermidis. Int J Antimicrob Agents. 2013;41(1):20–27. | ||
Luca V, Stringaro A, Colone M, Pini A, Mangoni ML. Esculentin (1-21), an amphibian skin membrane-active peptide with potent activity on both planktonic and biofilm cells of the bacterial pathogen Pseudomonas aeruginosa. Cell Mol Life Sci. 2013;70(15):2773–2786. | ||
Conde R, Zamudio FZ, Rodríguez MH, Possani LD. Scorpine, an anti-malaria and anti-bacterial agent purified from scorpion venom. FEBS Lett. 2000;471(2–3):165–168. | ||
Torres-Larios A, Gurrola GB, Zamudio FZ, Possani LD. Hadrurin, a new antimicrobial peptide from the venom of the scorpion Hadrurus aztecus. Eur J Biochem. 2000;267(16):5023–5031. | ||
Zhijian C, Feng L, Yingliang W, Xin M, Wenxin L. Genetic mechanisms of scorpion venom peptide diversification. Toxicon. 2006;47(3):348–355. | ||
Papo N, Shai Y. Host defense peptides as new weapons in cancer treatment. Cell Mol Life Sci. 2005;62(7–8):784–790. | ||
Kacprzyk L, Rydengård V, Mörgelin M, et al. Antimicrobial activity of histidine-rich peptides is dependent on acidic conditions. Biochim Biophys Acta. 2007;1768(11):2667–2680. | ||
Glukhov E, Stark M, Burrows LL, Deber CM. Basis for selectivity of cationic antimicrobial peptides for bacterial versus mammalian membranes. J Biol Chem. 2005;280(40):33960–33967. | ||
Hao G, Shi YH, Tang YL, Le GW. The membrane action mechanism of analogs of the antimicrobial peptide Buforin 2. Peptides. 2009;30(8):1421–1427. |
Supplementary materials
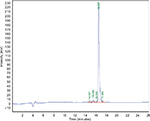  | Figure S1 Analytical RP-HPLC chromatogram of the synthetic consensus peptide Pepcon. Abbreviation: RP-HPLC, reversed-phase high-performance liquid chromatography. |
  | Figure S2 Positive electrospray ionization mass spectrometric analysis of the synthetic peptide Pepcon showing a major peak in the +2 charge state of 941.68 Da. |
 © 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.