Back to Journals » Infection and Drug Resistance » Volume 9
Optimizing the detection of methicillin-resistant Staphylococcus aureus with elevated vancomycin minimum inhibitory concentrations within the susceptible range
Authors Phillips C , Wells N, Martinello M, Smith S, Woodman R, Gordon D
Received 7 March 2016
Accepted for publication 31 March 2016
Published 31 May 2016 Volume 2016:9 Pages 87—92
DOI https://doi.org/10.2147/IDR.S107961
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Cameron J Phillips,1–3 Nicholas A Wells,4 Marianne Martinello,4 Simon Smith,4 Richard J Woodman,5 David L Gordon2,4
1SA Pharmacy, Flinders Medical Centre, Bedford Park, SA, Australia; 2Department of Microbiology and Infectious Diseases, School of Medicine, Flinders University, Adelaide, SA, Australia; 3School of Pharmacy and Medical Sciences, University of South Australia, Adelaide, SA, Australia; 4SA Pathology, Microbiology and Infectious Diseases, Flinders Medical Centre, Bedford Park, SA, Australia; 5Flinders Centre for Epidemiology and Biostatistics, School of Medicine, Flinders University, Adelaide, SA, Australia
Background: Determination of vancomycin minimum inhibitory concentration (MIC) can influence the agent used to treat methicillin-resistant Staphylococcus aureus (MRSA) infection. We studied diagnostic accuracy using E-test and VITEK® 2 against a gold standard broth microdilution (BMD) methodology, the correlation between methods, and associations between vancomycin MIC and MRSA phenotype from clinical isolates.
Methods: MRSA isolates were obtained from April 2012 to December 2013. Vancomycin MIC values were determined prospectively on all isolates by gradient diffusion E-test and automated VITEK® 2 . The Clinical and Laboratory Standards Institute reference BMD method was performed retrospectively on thawed frozen isolates. Diagnostic accuracy for detecting less susceptible strains was calculated at each MIC cutoff point for E-Test and VITEK® 2 using BMD ≥1 µg/mL as a standard. The correlation between methods was assessed using Spearman’s rho (ρ). The association between MRSA phenotype and MIC for the three methods was assessed using Fisher’s exact test.
Results: Of 148 MRSA isolates, all except one (E-test =3 µg/mL) were susceptible to vancomycin (MIC of ≤2 µg/mL) irrespective of methodology. MICs were ≥1.0 µg/mL for 9.5% of BMD, 50.0% for VITEK® 2 , and 27.7% for E-test. Spearman’s ρ showed weak correlations between methods: 0.29 E-test vs VITEK® 2 (P=0.003), 0.27 E-test vs BMD (P=0.001), and 0.31 VITEK® 2 vs BMD (P=0.002). The optimal cutoff points for detecting BMD-defined less susceptible strains were ≥1.0 µg/mL for E-test and VITEK® 2 . E-test sensitivity at this cutoff point was 0.85 and specificity 0.29, while VITEK® 2 sensitivity and specificity were 0.62 and 0.51, respectively. Multiresistant MRSA strains tended to have higher MIC values compared to nonmultiresistant MRSA or epidemic MRSA 15 phenotypes by E-test (Fisher’s exact P<0.001) and VITEK® 2 (Fisher’s exact P<0.001).
Conclusion: Overall diagnostic accuracy and correlations between MIC methods used in routine diagnostic laboratories and the gold standard BMD showed limited overall agreement. This study helps optimize guidance on the effective use of vancomycin.
Keywords: MIC, MRSA, sensitivity, specificity, susceptibility, vancomycin
A letter to the Editor has been recieved and published for this article.
Background
Vancomycin remains the antibiotic of choice for treating serious infection with methicillin-resistant Staphylococcus aureus (MRSA) and other serious Gram-positive infections despite its continuous use for over half a century.1,2 However, some have called into question “how long vancomycin may remain an effective therapy”.3,4 In recent years, there have been a number of new agents licensed by the US Food and Drug Administration to treat resistant infection with Gram-positive bacteria, including MRSA; however, it is essential to reserve these agents for when vancomycin is no longer effective.5,6
Prudent management of the way in which vancomycin is used in therapy is by prompt identification of the organism and testing of antibiotic susceptibility, which, along with optimizing dosing and serum concentration monitoring, may help ensure that vancomycin is not abandoned prematurely.7 There are a number of methods used to determine minimum inhibitory concentration (MIC); however, broth microdilution (BMD) remains the gold standard.8 The MIC along with vancomycin exposure measured as area under the concentration curve is the key pharmacokinetic–pharmacodynamic index used to optimize bacterial killing and clinical outcomes with vancomycin therapy.9,10 An area under the concentration curve/MIC index target of 400 mg/L × hour is recommended for contemporary vancomycin dosing.11 In the mid-2000s, the US Clinical Laboratory Standards Institute (CLSI) redefined the vancomycin MIC susceptibility breakpoint for S. aureus to ≤2 µg/mL.12 However, since that time, there have been a number of individual studies that have demonstrated associations between isolates with vancomycin MIC in the susceptible range and patient outcomes.13 Varying methods for determining MIC have been used in these studies, which makes extrapolation of results to routine clinical management challenging. Important consideration must be given to the method used to determine MIC, and decisions for treatment should be based upon the optimal cutoff points for the various methods. A meta-analysis of 14 papers with 2,439 patients with susceptible MRSA infection clearly defined high vancomycin MIC as ≥1 µg/mL by BMD and ≥1.5 µg/mL by E-test. This meta-analysis, which included patients with bloodstream and nonbloodstream infection, found a treatment failure risk ratio of 1.40 (95% confidence interval =1.15–1.71) and overall mortality risk ratio of 1.42 (confidence interval =1.08–1.87) for those with high vancomycin MIC.14
Although BMD remains the gold standard for measuring vancomycin MIC, this method is time consuming, labor intensive, and requires a high level of skill for consistent results. Alternative methodologies to determine vancomycin MIC such as the automated VITEK® 2 (BioMérieux Inc, Durham NC, USA) and gradient diffusion E-test are frequently used in diagnostic laboratories; however, these methods produce varying results in comparison to each other and to BMD.15
Inappropriate interpretation and overestimation of the MIC may cause unnecessary use of other agents when vancomycin would still be effective. Assessing the diagnostic accuracy of commonly used MIC methods compared against BMD would assist in meaningful interpretation of MIC values from each method and application of this information to treatment. Unnecessary abandonment of vancomycin for newer antibiotics in patients with MRSA infection with higher yet susceptible MICs (≥1 and ≤2 µg/mL) will potentially promote the emergence of resistance to these agents. Furthermore, the strength of the association among the vancomycin method, MIC, and MRSA phenotype is unclear.
The aim of this study was to measure the diagnostic accuracy of E-test and VITEK® 2 vancomycin MIC determination for clinical MRSA isolates compared against a BMD standard. A secondary aim was to explore the strength of the association between MIC and MRSA antibiotic resistance phenotype.
Materials and methods
Study design, data collection, and ethical approval
MRSA clinical isolates were obtained from hospitalized patients aged ≥18 years during the process of usual care between April 2012 and December 2013. The study was conducted at Flinders Medical Centre, a 550-bed teaching hospital in Adelaide, Australia. The pathology database ULTRA Laboratory Information System, Release 2.5C (Cirdan, Lisburn, Northern Ireland) was used to identify patient isolates during the study period. The study was approved by the Southern Adelaide Clinical Human Research Ethics Committee (approval number 123.12). A waiver of consent was granted with the approval as the participants were not exposed to any risk of harm. The waiver of consent was consistent with the Australian Government National Statement of Ethical Conduct in Human Research 2007.
Susceptibility testing
All MRSA isolates were tested to determine the vancomycin MIC. Isolate susceptibility to vancomycin was defined by the CLSI breakpoint of MIC ≤2 µg/mL. Automated VITEK® 2 System Version 05.04 (BioMérieux Inc.) sensitivity testing was performed on fresh isolates during routine processing. Gradient diffusion E-test (BioMérieux Inc.) was used according to the manufacturer’s instructions. Reading of E-test was conducted independently by a senior medical scientist, with the result confirmed by a medical microbiologist; any disagreement was adjudicated by a third reader. BMD was performed using thawed frozen isolates. Frozen isolates were stored (−20°C for 6–12 months) to enable batched processing. Vancomycin hydrochloride (Sigma-Aldrich, St. Louis, MO, USA) was sourced to prepare the stock solution. Susceptibility was tested at vancomycin concentrations 0.25–8.0 µg/mL in twofold dilutions according to the CLSI methodology.16
Validation of MIC results obtained using BMD method was performed concurrently using S. aureus ATCC 29213 and Escherichia coli ATCC 25922 as controls for every set of tests. MICs were determined after a period of 24 hours of incubation at 35°C in oxygen. Reading of BMD was performed independently by a senior medical scientist, a medical microbiologist, and a specialist pharmacist. Where a difference in reading the MIC occurred, a consensus of two readers was required.
MRSA resistance phenotype
Phenotyping was determined from antibiogram testing using VITEK® 2 AST-612 (BioMérieux Inc.). Three distinct phenotypes were recognized. Nonmultiresistant MRSA isolates were defined as those resistant to <3 non-<-lactam antibiotic classes, while multiresistant MRSA (mMRSA) isolates were defined as those resistant to ≥3 non-<-lactam antibiotic classes.17 Epidemic MRSA 15 was separately defined by resistance to ciprofloxacin ± erythromycin antibiotic.18
Statistical analysis
Data were stored in Microsoft Excel and were reported using descriptive statistics. Spearman’s rho (r) correlation coefficients were used to assess the strength of the association between the methods used to determine MIC. Specificity, sensitivity, and area under the receiver operating characteristic curve measured as C-statistic were used to calculate diagnostic accuracy for VITEK® 2 and E-test MIC methods for detecting strains with MIC ≥1 µg/mL by using BMD as the reference MIC value. The C-statistic is a measure of discrimination and reports the global test accuracy, ie, for all cutoff points combined. The reference MIC methodology (BMD) and MIC value were selected as they were shown to be independent predictors of poor clinical outcomes.14,19 Fisher’s exact test was used to assess whether MRSA phenotype was associated with MIC concentrations for each of the three MIC methods. Stata version 14.0 (StataCorp LP, College Station, TX, USA) was used for all statistical analyses.
Results
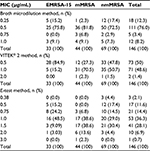
A total of 148 isolates were obtained from 111 patients during the study period. The clinical isolates were sourced from multiple anatomical sites, with skin and soft tissue and respiratory sites featuring prominently and 10% of isolates being from blood or central nervous system (Table 1). All MRSA isolates, with the exception of one isolate with E-test of 3 µg/mL (1 µg/mL by VITEK® 2 and BMD), were susceptible to vancomycin (≤2 µg/mL) by all the three methods. The distribution of MIC values by methodology is shown in Table 2. The percentage of isolates with MIC ≤0.5 µg/mL was 90.5%, 50%, and 28% by BMD, VITEK® 2, and E-test, respectively. MIC values ≥1 µg/mL were observed in 9.5% by BMD, 50% by VITEK® 2, and 72% by E-test.
  | Table 1 Anatomical region clinical isolate obtained Abbreviation: CSF, cerebrospinal fluid. |
Correlation of MIC methodologies
Spearman’s rho (r) correlation coefficients between the three methods were significant but weak; 0.29 for E-test vs VITEK® 2 (P=0.003), 0.27 for E-test vs BMD (P=0.001), and 0.31 for BMD vs VITEK® 2 (P=0.002).
The C-statistic was weak for both E-test (0.5428) and VITEK® 2 (0.5815) (Table 3). Sensitivity and specificity for detection of an MIC ≥1 µg/mL by BMD were calculated for each possible cutoff point for the E-test and VITEK® 2 methods (Table 3). The optimum cutoff point for the E-test was ≥1.0 µg/mL, with a sensitivity of 0.85 and specificity of 0.29, while the optimum cutoff point for VITEK® 2 was also ≥1.0 µg/mL, with corresponding values of 0.62 and 0.51, respectively.
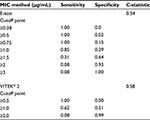  | Table 3 Sensitivity and specificity of E-test and VITEK® 2 for detection of a minimum inhibitory concentration (MIC) ≥1 µg/mL by broth microdilution (BMD) |
Breakdown by phenotype
There was no significant association between MRSA phenotype and the BMD MICs (P=0.15), although it appeared that there were relatively fewer mMRSA phenotypes than expected at BMD =0.25 µg/mL (2.3%), and relatively more mMRSA than expected at BMD =0.75 µg/mL (6.8%) based on observed percentages for both epidemic MRSA 15 and nonmultiresistant MRSA (Table 4). There was a significant association between MRSA phenotype and the VITEK® 2 MIC categories (P<0.001), with a lower than expected percent of mMRSA at VITEK® 2 =0.5 µg/mL (27.3%) and a higher than expected percent of mMRSA at VITEK® 2 =1.0 µg/mL (70.5%) based on the observed percentages for the other two MRSA phenotypes. There was also a significant association between phenotype and the E-test MICs (P<0.001), with a lower than expected percent of mMRSA at E-test =0.5 µg/mL MICs and 0.75 µg/mL MICs (0%–7%) and a higher than expected percent of mMRSA at E-test =1.5 µg/mL and E-test =2 µg/mL (39% and 14%) based on the observed percentages for the other two MRSA phenotypes.
Discussion
In this study of MRSA isolates that were susceptible to vancomycin, we found only a weak level of agreement between the accepted gold standard BMD and two commonly used methods to determine vancomycin MIC (VITEK® 2 and E-test). This weak agreement is consistent with the findings of other authors.20 Although some authors have reported that E-test does not produce higher MIC than other methods,21 we found higher E-test MIC values than either BMD or VITEK® 2, which concurs with the results of other studies.22,23
Patient outcomes are worse for MRSA infection with susceptible yet higher vancomycin MICs. Van Hal et al24 in a systematic review of 22 papers on the significance of vancomycin MIC reported that MIC ≥1.5 µg/mL was associated with worse clinical outcomes than those with <1.5 µg/mL; however, this MIC range was not ascribed to any one MIC method. In this study, we used valid statistical approaches to compare susceptible MIC values that are obtained in routine care from several commonly used methods.
As BMD is acknowledged as the gold standard method for MIC testing, and a BMD MIC ≥1 µg/mL has been associated with poor clinical outcomes,14,19 we compared the diagnostic accuracy of VITEK® 2 and E-test using BMD ≥1 µg/mL as the defined cutoff point for indicating reduced susceptibility. The value of sensitivity and specificity was assessed at the various E-test and VITEK® 2 MIC categories. For the MRSA strains used in this study, the optimum E-test and VITEK® 2 cutoff points for detection of reduced susceptibility were achieved at ≥1 µg/mL (E-test: sensitivity 0.85, specificity 0.29; VITEK® 2: sensitivity 0.62, specificity 0.51). These cutoff points need confirmation in a larger and more diverse dataset but provide novel and practical guidance toward assessing MIC results obtained from differing methodologies. These findings should prove useful to both diagnosticians and clinicians in evaluating test results for commonly employed MIC methodologies.
In our study, we observed significant variations in vancomycin MIC by phenotype using E-test and VITEK® 2, but not with BMD. Specifically, mMRSA strains had higher MIC values than expected. Since mMRSA strains are more likely to be hospital associated, these strains are likely to spread in an environment of higher vancomycin usage than the other “community-acquired” strains. It is unclear why these differences were not detected by BMD, but the clustering of BMD MICs at 0.5 µg/mL may have limited the ability to detect strain differences.
Guidance on treatment of MRSA infection is based on clinical response to vancomycin rather than MIC.25 However, if vancomycin MIC is a determinant of antibiotic selection, our findings provide useful guidance to better understanding of MIC results obtained through reference and routine laboratory methods.
The main limitations of our study were that the clinical isolates were obtained from a single geographical region (hospital catchment) and that there were also a relatively small number of isolates which were all in the susceptible range.
Conclusion
The level of agreement between MIC determination by BMD, E-test, and VITEK® 2 was relatively weak. The estimated sensitivity and specificity of the methods provide guidance on the best MIC cutoff points to use to interpret the results of each method. This permits more objective evaluation of test results obtained from routine methods and selection of vancomycin when appropriate.
Acknowledgment
The authors thank the staff of the Microbiology laboratory, Department of Microbiology and Infectious Diseases, Flinders Medical Centre for their assistance.
Author contributions
CJP, NAW, and DLG designed the study. CJP, NAW, SS, and MM collected data and performed laboratory work. RJW performed statistical analysis. All authors were involved in the drafting and approval of the final manuscript.
Disclosure
The salary of CJP was part funded by a National Health and Medical Research Council of Australia, Translating Research into Practice Fellowship (1035960) during this study. The authors report no conflicts of interest in this work.
References
Moellering RC. Vancomycin: a 50-year reassessment. Clin Infect Dis. 2006;42(Supp1):S3–S4. | ||
Rubinstein E, Keynan Y. Vancomycin revisited – 60 years later. Front Public Health. 2014;2:217. | ||
Deresinski S. Vancomycin and Staphylococcus aureus – an antibiotic enters obsolescence. Clin Infect Dis. 2007;44(12):1543–1548. | ||
Deresinski S. Vancomycin: does it still have a role as an antistaphylococcal agent? Expert Rev Anti Infect Ther. 2007;5(3):393–401. | ||
Yu T, Stockmann C, Balch AH, Spigarelli MG, Sherwin CM. Evolution of interventional vancomycin trials in light of new antibiotic development in the USA, 1999–2012. Int J Antimicrob Agents. 2014;43(3):215–222. | ||
Perez F, Salata RA, Bonomo RA. Current and novel antibiotics against resistant Gram-positive bacteria. Infect Drug Resist. 2008;1:27–44. | ||
Kollef MH. Limitations of vancomycin in the management of resistant staphylococcal infections. Clin Infect Dis. 2007;45(Suppl 3):S191–S195. | ||
Dhand A, Sakoulas G. Reduced vancomycin susceptibility among clinical Staphylococcus aureus isolates (‘the MIC Creep’): implications for therapy. F1000 Med Rep. 2012;4:4. | ||
Moise-Broder PA, Forrest A, Birmingham MC, Schentag JJ. Pharmacodynamics of vancomycin and other antimicrobials in patients with Staphylococcus aureus lower respiratory tract infections. Clin Pharmacokinet. 2004;43(13):925–942. | ||
Lodise TP, Drusano GL, Zasowski E, et al. Vancomycin exposure in patients with methicillin-resistant Staphylococcus aureus bloodstream infections: how much is enough? Clin Infect Dis. 2014;59(5):666–675. | ||
Rybak MJ, Lomaestro BM, Rotschafer JC, et al. Vancomycin therapeutic guidelines: a summary of consensus recommendations from the infectious diseases Society of America, the American Society of Health-System Pharmacists, and the Society of Infectious Diseases Pharmacists. Clin Infect Dis. 2009;49(3):325–327. | ||
Tenover FC, Moellering RC Jr. The rationale for revising the Clinical and Laboratory Standards Institute vancomycin minimal inhibitory concentration interpretive criteria for Staphylococcus aureus. Clin Infect Dis. 2007;44(9):1208–1215. | ||
Holland TL, Fowler VG Jr. Vancomycin minimum inhibitory concentration and outcome in patients with Staphylococcus aureus bacteremia: pearl or pellet? J Infect Dis. 2011;204(3):329–331. | ||
Jacob JT, DiazGranados CA. High vancomycin minimum inhibitory concentration and clinical outcomes in adults with methicillin-resistant Staphylococcus aureus infections: a meta-analysis. Int J Infect Dis. 2013;17(2):e93–e100. | ||
Hsu DI, Hidayat LK, Quist R, et al. Comparison of method-specific vancomycin minimum inhibitory concentration values and their predictability for treatment outcome of methicillin-resistant Staphylococcus aureus (MRSA) infections. Int J Antimicrob Agents. 2008;32(5):378–385. | ||
Clinical and Laboratory Standards Institute. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically: Approved Standard: Ninth Edition M07-A9. Wayne, PA: CLSI;2012. | ||
Tong SY, Bishop EJ, Lilliebridge RA, et al. Community-associated strains of methicillin-resistant Staphylococcus aureus and methicillin-susceptible S. aureus in indigenous Northern Australia: epidemiology and outcomes. J Infect Dis. 2009;199(10):1461–1470. | ||
Nimmo GR, Coombs GW, Pearson JC, et al. Methicillin-resistant Staphylococcus aureus in the Australian community: an evolving epidemic. Med J Aust. 2006;184(8):384–388. | ||
Wi YM, Kim JM, Joo EJ, et al. High vancomycin minimum inhibitory concentration is a predictor of mortality in methicillin-resistant Staphylococcus aureus bacteraemia. Int J Antimicrob Agents. 2012;40(2):108–113. | ||
van Hal SJ, Barbagiannakos T, Jones M, et al. Methicillin-resistant Staphylococcus aureus vancomycin susceptibility testing: methodology correlations, temporal trends and clonal patterns. J Antimicrob Chemother. 2011;66(10):2284–2287. | ||
Khatib R, Riederer K, Shemes S, Musta AC, Szpunar S. Correlation of methicillin-resistant Staphylococcus aureus vancomycin minimal inhibitory concentration results by Etest and broth microdilution methods with population analysis profile: lack of Etest overestimation of the MIC. Eur J Clin Microbiol Infect Dis. 2013;32(6):803–806. | ||
Keel RA, Sutherland CA, Aslanzadeh J, Nicolau DP, Kuti JL. Correlation between vancomycin and daptomycin MIC values for methicillin-susceptible and methicillin-resistant Staphylococcus aureus by 3 testing methodologies. Diag Microbiol Infect Dis. 2010;68(3):326–329. | ||
Mason EO, Lamberth LB, Hammerman WA, Hulten KG, Versalovic J, Kaplan SL. Vancomycin MICs for Staphylococcus aureus vary by detection method and have subtly increased in a pediatric population since 2005. J Clin Microbiol. 2009;47(6):1628–1630. | ||
Van Hal SJ, Lodise TP, Paterson DL. The clinical significance of vancomycin minimum inhibitory concentration in Staphylococcus aureus infections: a systematic review and meta-analysis. Clin Infect Dis. 2012;54(6):755–771. | ||
Liu C, Bayer A, Cosgrove SE, et al. Clinical practice guidelines by the infectious diseases society of America for the treatment of methicillin-resistant Staphylococcus aureus infections in adults and children: executive summary. Clin Infect Dis. 2011;52(3):285–292. |
 © 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.