Back to Journals » Research and Reports in Urology » Volume 7
One-year monitoring of an oligonucleotide fluorescence in situ hybridization probe panel laboratory-developed test for bladder cancer detection
Authors Tinawi-Aljundi R, King L, Knuth S, Gildea M, Ng C, Khal J, Dion J, Young C, Schervish E, Frontera JR, Hafron J, Kernen K, Di Loreto R, Aurich-Costa J
Received 11 December 2014
Accepted for publication 12 February 2015
Published 9 April 2015 Volume 2015:7 Pages 49—55
DOI https://doi.org/10.2147/RRU.S79085
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Jan Colli
Rima Tinawi-Aljundi,1 Lauren King,2 Shannon T Knuth,2 Michael Gildea,2 Carrie Ng,2 Josh Kahl,2 Jacqueline Dion,2 Chris Young,2 Edward W Schervish,1 J Rene Frontera,1 Jason Hafron,1 Kenneth M Kernen,1 Robert Di Loreto,1 Joan Aurich-Costa2
1Michigan Institute of Urology, St Claire Shores, MI, USA; 2Cellay, Inc., Cambridge, MA, USA
Background: Previously, we had developed and manufactured an oligonucleotide fluorescence in situ hybridization (OligoFISH) probe panel based on the most clinically sensitive chromosomes found in a reference set of bladder carcinoma cases. The panel was clinically validated for use as a diagnostic and monitoring assay for bladder cancer, reaching 100% correlation with the results of the UroVysion test. After 1 year of using this probe panel, we present here the comparison of cytology, cystoscopy, and pathology findings to the OligoFISH probe panel results to calculate its clinical performance.
Materials and methods: In order to calculate clinical performance, we compared the OligoFISH results to the cytology and cystoscopy/pathology findings for 147 initial diagnoses and 399 recurrence monitorings. Finally, we compared clinical performance to published values for the UroVysion test, including both low- and high-grade tumors.
Results: Chromosomes 3, 6, 7, and 20 were highly involved in bladder carcinoma aneuploidy. At the initial diagnosis, we obtained 90.5% (95% confidence interval [CI]: 84.5%–94.7%) accuracy, 96.8% sensitivity (95% CI: 91.0%–99.3%), 79.2% specificity (95% CI: 65.9%–87.8%), 89.2% positive predictive value (PPV; 95% CI: 81.5%–94.5%), and 93.3% negative predictive value (NPV; 95% CI: 81.7%–97.3%). When monitoring for recurrence, we obtained 85.2% accuracy (95% CI: 81.3%–88.5%), 82.0% sensitivity (95% CI: 76.0%–87.1%), 88.4% specificity (95% CI: 83.2%–92.5%), 87.7% PPV (95% CI: 82.1%–92.0%), and 83.0% NPV (95% CI: 77.3%–87.8%). When looking at low- and high-grade tumors, the test showed 100% sensitivity for high-grade tumors (95% CI: 92.5%–100%) and 87.5% sensitivity (95% CI: 68.8%–95.5%) for low-grade tumors. All the clinical parameters for the OligoFISH panel were higher than the UroVysion test's published performance. We found significantly higher clinical sensitivity and NPV at initial diagnosis and significantly higher specificity and PPV for recurrence.
Conclusion: The OligoFISH probe panel is a fast, easy, and reproducible test for bladder cancer diagnosis and monitoring, with excellent clinical performance and utility.
Keywords: UroVysion, FISH, urologic oncology, bladder neoplasm
Introduction
Whole chromosome gains are the most common type of aberration in cancer cells.1 Aneuploidy as a marker of cancer was first proposed in 1914.2 The UroVysion Bladder Cancer Kit (Abbott Laboratories, Abbott Park, IL, USA) assay was developed based on these principles.3 The UroVysion Bladder Cancer Kit is a fluorescence in situ hybridization (FISH) assay that reveals aneuploidy for chromosomes 3, 7, 17 as well as deletion of 9p21 from urothelial cells in voided urine.
Based on case–control and cohort studies, the UroVysion test is capable of detecting bladder cancer. UroVysion has high sensitivity (81%) and specificity (96%) for high-grade tumors but lower sensitivity (36%–57%) for low-grade tumors.4 Many authors have modified the scoring criteria of the US Food and Drug Administration–approved test in order to increase sensitivity; however, it remains low for low-grade tumors and usually results in a decrease in specificity.5–7 UroVysion’s low sensitivity for the detection of low-grade bladder tumors was the impetus to evaluate other combinations of chromosomal probes in the hopes of developing a more sensitive and specific test for both high- and low-grade bladder tumors. Also, since OligoFISH® probes hybridize in 10 minutes, this probe panel improves turnaround time by not having to hybridize overnight.8,9
Many have investigated chromosomal abnormalities for bladder carcinomas, focusing on a limited number of chromosomes of interest.10–12 The Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer1 relates chromosomal aberrations to tumor characteristics. Using this database, we determined the individual involvement of all 24 chromosomes in the database’s bladder carcinoma cases and manufactured a panel of Same Day OligoFISH probes based on the most clinically-sensitive chromosomes, resulting in a test that is highly sensitive and specific for both high- and low-grade tumors.13 The probe panel was analytically and clinically validated in a previous report.13 Briefly, for the clinical validation, the results with the OligoFISH test were compared to UroVysion results. Initially, a pilot study including five known UroVysion-negative and five UroVysion-positive cases were analyzed with the OligoFISH panel scoring 500 cells. The maximum number of positive cells in the negative cases was used to calculate the higher interval at 95% confidence, which will constitute our normal cutoff value. By determining how many cells needed to be scored in the positive cases for the cases to be called positive, we determined the number of cells that need to be scored in order to provide a valid result. Once these scoring criteria were determined, the test was validated in a series of 20 UroVysion-positive and 20 UroVysion-negative cases in order to determine that concordance between the two tests reached 100%, allowing us to claim the same clinical performance between them. However, the true clinical performance of the test should be calculated by comparing it to the gold standard, which in this case is cystoscopy and pathology.
Herein, we report the clinical performance of this test after being used in our clinic for 1 year by comparing the OligoFISH test results to cytology, cystoscopy, and pathology findings. We also show the comparison of its clinical performance to the published performance of the UroVysion test.
Materials and methods
Sample collection and OligoFISH procedure
A urine specimen (30–60 mL) was collected and preservative was added to the urine (2:1 urine:preservative) to prevent bacterial growth and preserve cells and DNA (Hologic, Inc., Bedford, MA, USA). Preserved samples were stored at 4°C until use. Urothelial slides were prepared using the ThinPrep machine (Hologic, Inc.) following the manufacturer’s instructions. The slides were partially digested in IsoThermal Protease (Cellay, Inc., Cambridge, MA, USA) at room temperature for 15 minutes followed by 5 minutes incubation in Detergent Solution (Cellay, Inc.). The cells were lightly fixed for 5 minutes in 1% formaldehyde and rinsed in phosphate-buffered saline (PBS). Cellular DNA was denatured in IsoThermal Denaturing Solution (Cellay, Inc.) at room temperature for 10 minutes. Slides were dehydrated in 85% and 100% alcohol for 1 minute each and air dried. Slides were hybridized with 3 μL of OligoFISH probe mix, coverslips applied, and incubated for 10 minutes at 37°C. Slides were washed in 2× saline sodium citrate (SSC) under agitation to float off the coverslips, and then incubated in IsoThermal Wash Solution (2× SSC and Isothermal Denaturing Reagent) for 5 minutes at room temperature to remove unbound or nonspecifically hybridized probes. The slides were rinsed in 2× SSC and mounted with Antifade with 4′,6-diamidino-2-phenylindole dihydrochloride (DAPI) and analyzed with an epifluorescence microscope.
Scoring criteria
The scoring criteria were established during the clinical validation.13 The normal cutoff was determined to be three abnormal cells maximum and the scoring criteria are:
- score at least 100 cells, skipping polymorphonuclear leukocytes;
- a positive cell must show at least one chromosomal gain (Figure 1);
- if ≥4 positive cells are found, the sample is positive;
- if <4 positive cells are found in at least 100 cells, the sample is negative;
- if ≥4 positive cells are scored in at least 25 cells, analysis can be stopped, the sample is positive;
- if three positive cells are found, analysis is extended until a fourth positive cell is found or there are no more cells to score;
- if the sample is negative and <100 cells can be scored, the analysis is invalid.
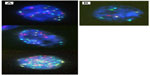  | Figure 1 Examples of urothelial cells (A) with chromosome gains and (B) a normal cell. |
Analytical performance of the four-chromosome OligoFISH probe panel
The analytical performance of the OligoFISH panel was validated based on schema previously described for hybridizing probes.14–16 Briefly, the probe panel was validated against normal blood cells in metaphase from five chromosomally normal individuals to determine analytical specificity and sensitivity.
One-year surveillance of the OligoFISH test
We identified diagnostic events as positive or negative by cytology, cystoscopy, and pathology. For cytology, reports of ‘positive’ or ‘suspicious’ were scored positive, and others were scored negative.
Cystoscopy/pathology scoring was more complicated. A ‘positive’ score results when a) tumors reported by cystoscopy are confirmed by pathology or b) the pathology report indicates the presence of tumors in the absence of a cystoscopy report. A ‘negative’ score results when a) cystoscopy reports no lesions, and thus no biopsy/pathology is performed, or b) cystoscopy reports tumors that are not confirmed by pathology.
By comparing these diagnostic results to the OligoFISH results, we determined the true positives (TP), true negatives (TN), false positives (FP), and false negatives (FN), and calculated the clinical performance of the OligoFISH assay. Because the data does not follow a Gaussian distribution, the 95% confidence interval (CI) was calculated using the Clopper–Pearson method.17
Finally, in the initial diagnostic patient group, clinical sensitivity was determined for both low-and high-grade tumors when the grade was reported in the pathology report. Pathology descriptions of high-grade tumors include: papillary urothelial carcinoma, papillary transition cell carcinoma, and transitional cell carcinoma in situ; carcinoma in situ; urothelial carcinoma in situ; and infiltrating urothelial carcinoma and poorly differentiated adenocarcinoma. Pathology descriptions of low-grade tumors include: papillary transitional cell carcinoma, low-grade papillary urothelial carcinoma, noninvasive, and papillary urothelial neoplasm of low malignant potential.18
Clinical performance calculations and statistical methods
By comparing the results of OligoFISH and cytology, cystoscopy, and pathology, all the initial diagnosis and monitoring events were classified as TP, FP, TN, and FN. In order to calculate clinical performance, the following formulas were used:
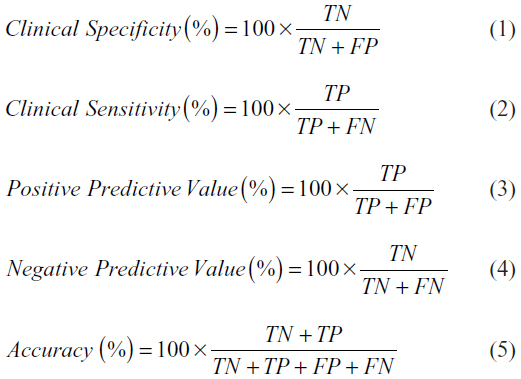

In order to calculate the 95% CIs for these frequencies or percentages, we used the Clopper–Pearson method, since they are binomial variables; as they are all far from 50%, the normal approximation would not be appropriate.17 We calculated the intervals using Excel (Microsoft Corporation, Redmond, WA, USA) using the following formulas based on the β distribution:

where α=0.05 for 95% confidence; x = number of successes (ex TP for sensitivity, or TN for specificity), and n = total number of cases used in the calculation.
Results
Analytical performance of the four-chromosome OligoFISH probe panel
Analytical specificity
Cytogenetic slides from peripheral blood from five chromosomally normal males were hybridized with the probe panel and 20 metaphases were captured, analyzed in reverse DAPI banding (similar to G banding; Figure 2), and each probe’s position determined. Analytical specificity was calculated as the percentage of metaphases with probes hybridizing to the correct loci. Analytical specificity was 100%.
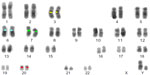  | Figure 2 Example of one of the metaphases fully karyotyped using reverse DAPI staining. |
Analytical sensitivity
On the above slides, 200 interphase nuclei were scored. Analytical sensitivity was calculated as the percentage of nuclei with the correct number of signals, and for each probe, was greater than 98%, as recommended by the American College of Genetic Medicine (Table 1).19
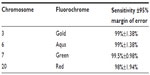  | Table 1 Analytical sensitivity of OligoFISH probes |
One-year surveillance of the OligoFISH Test
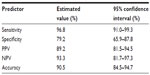
OligoFISH testing, along with cytology, cystoscopy, and upper tract imaging, was ordered for the initial diagnosis of bladder cancer and to monitor recurrence in patients previously diagnosed with bladder cancer. To determine its clinical performance, we retrospectively reviewed the medical records of patients who had the OligoFISH test performed. Data transfer was authorized by the Western Institutional Review Board and no informed consent was required. Results of the diagnostic analysis of 147 patient urine samples using the OligoFISH panel are presented in Table 2; the test’s accuracy results are in Table 3. For the initial diagnosis, the OligoFISH probe panel reached 90.5% accuracy (95% CI: 84.5%–94.7%) with a clinical sensitivity of 96.8% (95% CI: 91.0%–99.3%) and specificity of 79.2% (95% CI: 65.9%–87.8%; Table 4).
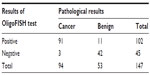  | Table 2 OligoFISH results in patients with suspected bladder cancer |
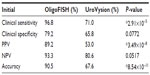  | Table 4 Clinical performance comparison between OligoFISH bladder cancer and UroVysion |
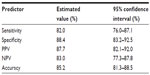
When analyzing the diagnostic events in patients being monitored for recurrence of bladder cancer (399 events), we reached an accuracy of 85.2% (95% CI: 81.3%–88.5%) with a clinical sensitivity of 82.0% (95% CI: 76.0%–87.1%) and a specificity of 88.4% (95% CI: 83.2%–92.5%) (Tables 5 and 6).
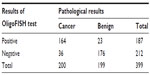  | Table 5 OligoFISH diagnostic events in monitoring bladder cancer recurrence |
Comparison of clinical performance of OligoFISH to UroVysion
Many different authors have reported different clinical performances for the UroVysion test, with sensitivities ranging from 68.3% to 89.2%.20 We compared the frequencies of TP, FP, TN, and FN for the OligoFISH panel and the UroVysion kit approval letter (Table 4).4 When comparing all the frequencies together (accuracy), the OligoFISH test demonstrated significant differences from UroVysion (P=8.54×10–11). When comparing only cancer patients (clinical sensitivity) and patients with a positive predictive value (PPV), we also observed significant differences between the two tests (P=2.90×10–5 and P=3.49×10–8, respectively). When comparing benign patients (clinical specificity) and patients with a negative predictive value (NPV), the differences between the two tests were not statistically significant (P=0.0772 and 0.0517, respectively), despite finding higher values. To analyze the clinical performance of OligoFISH to monitor recurrence, we compared cytology and cystoscopy/pathology for 399 diagnostic events to OligoFISH results as described above (Tables 5 and 6). When compared to UroVysion, the differences in accuracy for both tests are significant (P=1.39×10–8). Although all clinical parameter values were higher for OligoFISH than UroVysion, we found statistically significant differences only for clinical specificity (P=1.31×10–6) and PPV (P=4.02×10–10). Clinical sensitivity and NPV P-values were P=0.06 and P=0.62, respectively (Table 7).
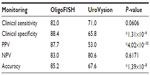  | Table 7 Clinical performance comparison between OligoFISH bladder cancer and UroVysion4 for monitoring the recurrence of bladder cancer |
Detection of low-grade tumors
Based on clinical studies, in low-grade bladder tumors, UroVysion has a low PPV.4 To determine the clinical performance of the assay by grade, we identified 24 and 46 patients with low- and high-grade tumors, respectively (Table 8). Twenty-one out of 24 low-grade tumors had a positive OligoFISH result, with a clinical sensitivity of 87.5% (95% CI: 68.8%–95.5%). All 46 high-grade tumors had a positive OligoFISH test (clinical sensitivity =100% [95% CI: 92.5%–100%]).
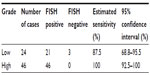  | Table 8 Clinical sensitivity of the OligoFISH probe panel by grade of the lesion in the initial diagnosis |
Discussion
Aneuploidy has been recognized as a hallmark of cancer for more than 100 years,2 and recent studies are focusing on how these changes arise and how they profile cancer genomes.21,22 The Mitelman Database is a very valuable tool to analyze the cytogenetic changes in cancer. Mitelman’s team has accumulated a large database encompassing all the cases published for many years in all human cancers.1 By using this database to choose the chromosomes mostly involved, we were able to devise a powerful probe panel for bladder cancer detection. Development of the probe panel was fully based on the karyotypes of published cases of bladder carcinoma collected by Dr Mitelman’s team. Using this database, we were able not only to calculate aneusomy rates for each one of the chromosomes, but also to calculate their involvement independent from each other. These analyses brought us to select chromosomes 3, 6, 7, and 20 as the four chromosomes most independently involved in most of the aneuploidies found in the bladder cancer cases contained in the database.
The multiprobe UroVysion FISH test was developed over 10 years ago to overcome the shortcomings of urine cytology. During this time, UroVysion has been shown to help in certain clinical situations but has also been shown to have significant weaknesses. The test detects and quantifies chromosomes 3, 7, and 17 and the locus 9p21 in fixed and stained cells obtained from urine using four colors of fluorescently labeled DNA probes using a fluorescence microscope.4 In our hands, we found chromosomes 3, 6, 7, and 20 to be the most informative. When we compared the clinical performance parameters for the OligoFISH bladder cancer probe panel to published results for the same parameters for the UroVysion Bladder Cancer Kit, the OligoFISH panel had higher clinical performance, especially with the detection of more low-grade tumors.4 There are several reasons that could be hindering UroVysion’s performance. First, only three chromosomes are being scored for aneuploidy, instead of four in our probe panel. Although, in the Mitelman database most of the chromosomes were involved in aneuploidy in bladder cancer and more in general in human carcinomas, our four chromosomes seem to be the most involved in bladder cancer. Second, by selecting cells to be scored based on their abnormal DAPI pattern, UroVysion could be overlooking cells with little or no morphological changes that would be normally scored as low-grade tumors, thus hindering sensitivity for these tumors. Finally, despite the strong rationale on monitoring the absence of gene p16 as a necessary step for aneuploid cells to continue dividing, the homozygous deletion of this gene is not a very common observation. Furthermore, gene p16 has been shown to be silenced not only by deletion but by methylation, which renders the use of an FISH probe clinically irrelevant to monitor this gene’s function.23,24
Most of the OligoFISH test is conducted at room temperature and we found the test easy to perform. The test can be performed in a single day, and we found it had a higher clinical performance and a reduced test turnaround time compared to UroVysion. Moreover, the OligoFISH test demonstrated to be a very reliable tool to both diagnose and monitor bladder cancer in voided urine.
Conclusion
Previously, we validated the OligoFISH 3, 6, 7, and 20 chromosome probe panel for screening patient voided urine samples for bladder cancer.13 After 1 year of using this probe panel clinically, we evaluated its clinical performance on initial diagnoses and monitoring recurrence events. We found a statistically significant higher clinical sensitivity and NPV at initial diagnosis and statistically significant higher specificity and PPV when monitoring for recurrence compared to the UroVysion test. The UroVysion test has poor sensitivity for low-grade lesions (54.5%) but high sensitivity for high-grade tumors (88.9%).4 The OligoFISH test had high sensitivity for low-grade lesions (87.5%) and detected all high-grade tumors (100%).
Acknowledgment
We would like to thank Amy Juodawlkis for her editorial help.
Disclosure
All the authors from Cellay are current or past employees. Joan Aurich-Costa is a stockholder of Cellay.
References
Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer [database on the Internet]. Mitelman F, Johansson B, Mertens F, editors. Available from: http://cgap.nci.nih.gov/Chromosomes/ Mitelman; 2011. Accessed February 17, 2015. | |
Boveri T. Concerning the origin of malignant tumours by Theodor Boveri. Translated and annotated by Henry Harris. J Cell Sci. 2008; 121 Suppl 1:1–84. | |
Halling KC, Kipp BR. Bladder cancer detection using FISH (UroVysion assay). Adv Anat Pathol. 2008;15(5):279–286. | |
US Food and Drug Administration, Office of In Vitro Diagnostic Device Evaluation and Safety Center for Devices and Radiological Health. 510(k) Summary: Safety and Effectiveness Information for the UroVysionTM Bladder Cancer Recurrence Kit approval letter, December 19, 2003. Available from http://www.accessdata.fda.gov/cdrh_docs/pdf3/k033982.pdf. Accessed March 4, 2015.. | |
Halling KC, King W, Sokolova IA, et al. A comparison of BTA stat, hemoglobin dipstick, telomerase and Vysis UroVysion assays for the detection of urothelial carcinoma in urine. J Urol. 2002;167(5):2001–2006. | |
Moatamed NA, Apple SK, Bennett CJ, et al. Exclusion of the uniform tetraploid cells significantly improves specificity of the urine FISH assay. Diagn Cytopathol. 2013;41(3):218–225. | |
Ferra S, Denley R, Herr H, Dalbagni G, Jhanwar S, Lin O. Reflex UroVysion testing in suspicious urine cytology cases. Cancer. 2009; 117(1):7–14. | |
Aurich-Costa J, Zamechek L, Keenan P, Bradley S. Oligo fluorescence in situ hybridization (Oligo-FISH), a new strategy for enumerating chromosomes in interphase nuclei. Fertil Steril. 2007;88:S86. | |
Aurich-Costa J, Ng C, Selvaggio S, Colls P, Bradley S. Oligonucleotide (ODN) fluorescence in situ hybridization (Oligo-FISH) and conventional FISH allow enumeration of 24 chromosomes in 6 successive hybridizations performed in a single day. Fertil Steril. 2009;92(3):S50. | |
Zhang FF, Arber DA, Wilson TG, Kawachi MH, Slovak ML. Toward the validation of aneusomy detection by fluorescence in situ hybridization in bladder cancer: comparative analysis with cytology, cytogenetics, and clinical features predicts recurrence and defines clinical testing limitations. Clin Cancer Res. 1997;3(12 Pt 1):2317–2328. | |
Yokogi H, Wada Y, Moriyama-Gonda N, Igawa M, Ishibe T. Genomic heterogeneity in bladder cancer as detected by fluorescence in situ hybridization. Br J Urol. 1996;78(5):699–703. | |
Sokolova IA, Halling KC, Jenkins RB, et al. The development of a multitarget, multicolor fluorescence in situ hybridization assay for the detection of urothelial carcinoma in urine. J Mol Diagn. 2000;2(3):116–123. | |
Tinawi-Aljundi R, Frontera R, Diloreto R. Clinical validation of a novel, same day oligoFISH chromosome panel for bladder carcinoma detection in urine. J Mol Diagn. 2012(14):718. | |
Wiktor AE, Van Dyke DL, Stupca PJ, et al. Preclinical validation of fluorescence in situ hybridization assays for clinical practice. Genet Med. 2006;8(1):16–23. | |
Reddy KS, Tripodi J. Contemplations on preclinical validation of fluorescence in situ hybridization probe assay for paraffin-embedded tissues in hematologic disorders. Cancer Genet Cytogenet. 2008; 183(1):1–5. | |
Dewald G, Stallard R, Al Saadi A, et al. A multicenter investigation with interphase fluorescence in situ hybridization using X- and Y-chromosome probes. Am J Med Genet. 1998;76(4):318–326. | |
Newcombe RG. Two-sided confidence intervals for the single proportion: comparison of seven methods. Stat Med. 1998;17(8):857–872. | |
Eble JN, Sauter G, Epstein JI, Sesterhenn IA. World Health Organization Classification of Tumors, Pathology and Genetics, Tumors of the Urinary System and Male Genital Organs. Lyon: IARC Press; 2004. | |
American College of Genetic Medicine. Standards and Guidelines for Clinical Genetics Laboratories. Bethesda, MD: American College of Genetic Medicine; 2009. Available from: http://www.acmg.net/StaticContent/SGs/Section_E_2011.pdf. Accessed February 17, 2015. | |
Ho CC, Tan WP, Pathmanathan R, Tan WK, Tan HM. Fluorescence-in-situ-hybridization in the surveillance of urothelial cancers: can use of cystoscopy or ureteroscopy be deferred? Asian Pac J Cancer Prev. 2013;14(7):4057–4059. | |
Davoli T, Xu AW, Mengwasser KE, et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell. 2013;155(4):948–962. | |
Tusell L, Pampalona J, Soler D, Frias C, Genescà A. Different outcomes of telomere-dependent anaphase bridges. Biochem Soc Trans. 2010;38(6):1698–1703. | |
Chan MW, Chan LW, Tang NL, et al. Hypermethylation of multiple genes in tumor tissues and voided urine in urinary bladder cancer patients. Clin Cancer Res. 2002;8(2):464–470. | |
Markl ID, Cheng J, Liang G, Shibata D, Laird PW, Jones PA. Global and gene-specific epigenetic patterns in human bladder cancer genomes are relatively stable in vivo and in vitro over time. Cancer Res. 2001;61(15):5875–5884. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.