Back to Journals » OncoTargets and Therapy » Volume 12
Notch Intracellular Domain (NICD) Expression and Clinical Manifestations of Second Primary Tumor at Esophagus in Patients with Head and Neck Squamous Cell Carcinoma
Authors Anekpuritanang T , Pongsapich W , Watcharadilokkul T , Ngaotepprutaram P , Pithuksurachai P, Bunbanjerdsuk S
Received 18 August 2019
Accepted for publication 2 December 2019
Published 17 December 2019 Volume 2019:12 Pages 11175—11181
DOI https://doi.org/10.2147/OTT.S227745
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Leo Jen-Liang Su
Tauangtham Anekpuritanang,1 Warut Pongsapich,2 Tanasarun Watcharadilokkul,2 Premyot Ngaotepprutaram,2 Paveena Pithuksurachai,2 Sacarin Bunbanjerdsuk3
1Department of Pathology, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok, Thailand; 2Department of Otorhinolaryngology, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok, Thailand; 3Section for Translational Medicine, Faculty of Medicine Ramathibodi Hospital, Mahidol University, Bangkok, Thailand
Correspondence: Warut Pongsapich
Department of Otorhinolaryngology Faculty of Medicine Siriraj Hospital, Mahidol University, 2 Wanglang Road, Bangkoknoi, Bangkok 10700, Thailand
Tel +66 2-419-8047
Fax +66 2-419-8044
Email [email protected]
Introduction: Second primary tumor (SPT) is a major factor that affects the survival of head and neck squamous cell carcinoma (HNSCC) patients, and the esophagus is a common site. Detection of SPT is essential for optimal HNSCC treatment planning and follow-up. Mutation of the NOTCH1 gene is common in head and neck cancer. However, details relating to Notch signaling and clinical outcomes among different primary tumors are still inconclusive. This study aimed to identify the role of the Notch signaling pathway in HNSCC, and to compare NOTCH1 expression in HNSCC compared between those with and without SPT at esophagus while focusing on the Notch intracellular domain (NICD).
Methods: Twenty-three cases of esophageal SPT and 47 non-SPT controls that were treated at Siriraj Hospital during 2006–2017 were included. Patient information and clinical outcomes were analyzed. NICD expression demonstrated by immunohistochemistry technique in formalin-fixed paraffin-embedded specimens was studied.
Results: Mean age of SPT and non-SPT was 55.13 and 62.09 years, respectively, and 94.3% of patients were male. Regarding SPT detection, 82.6% were synchronous and 17.4% were metachronous. There was significantly more active smoking among SPT than among non-SPT (87.0% vs 51.1%, p=0.01). Active alcohol use was also significantly greater among SPT than among non-SPT (87.0% vs 61.7%; p=0.04). Hypopharynx was the most common primary tumor site among SPT. Three-year and 5-year survival among SPT patients was 38.0% and 25.3%, respectively. NICD expression was absent in 52.2% of SPT, and in 53.3% of non-SPT. NICD expression intensity was mostly weak or moderate.
Conclusion: Active smoking and alcohol use were found to be significantly associated with SPT development. A high percentage of NICD inactivation was noted in HNSCC with no significant difference between groups. The Notch signaling pathway is involved in HNSCC tumorigenesis, but may not be a suitable molecular marker for SPT development.
Keywords: NOTCH, second primary tumor, esophagus, head and neck cancer, synchronous cancer
Introduction
Second primary tumor (SPT) is prevalent in patients with head and neck squamous cell carcinoma (HNSCC). From a 10-year post-treatment study of index cancer, Leon et al reported that the second primary tumor originated at other sites within the field of the head and neck (35%–73%), followed by lung (15%–32%), and esophagus (9%).1 Nonetheless, esophagus is one of the most common sites of SPT in HNSCC, especially when hypopharynx is the primary site.2 Despite the improved survival rate among certain groups of patients following recent advances in HNSCC treatment, SPT remains a major factor that influences long-term survival in this patient population.3 To improve treatment outcomes, the detection of second primary cancer is essential. Optimized pretreatment screening would facilitate the simultaneous detection of both the primary tumor and the second primary tumor.
Oncogenesis in HNSCC is multifactorial, with both environmental and genomic factors playing a contributory role. The most common molecular disorder that causes HNSCC is TP53;4 however, the mechanisms of carcinogenesis remain unclear. Lee et al reported NOTCH1 to be a commonly mutated gene in patients with head and neck cancer with a prevalence of 12%–15%.4,5 However, the role of Notch1 signaling in cancer development is tissue-specific. The Notch signaling pathway has dual biological roles, with either pro-carcinogenic or anti-carcinogenic effect.6 Abnormal expression of Notch receptors has been differently reported in various types of epithelial malignant lesions.
The Notch signaling pathway plays an important role in cell-cell communication, and in modulating epithelial-mesenchymal transition, which controls multiple cell differentiation processes during embryonic and adult life.7 Activation of the Notch1 receptor subsequently translocates the Notch intracellular domain (NICD) into the nucleus to regulate the expression of target genes. Dysregulation of the Notch signaling pathway was reported to be associated with adverse clinical characteristics, such as lymph node metastasis, vascular invasion, and chemo-resistance.8 Non-physiologic expression of NICD, which is an active form of Notch1 in the primary tumor, may indicate SPT development in HNSCC patients.
Enhanced understanding of Notch signaling in HNSCC may improve treatment planning and patient outcomes. Accordingly, the aims of this study were to identify the role of the Notch signaling pathway in HNSCC, and to compare NOTCH1 expression in HNSCC compared between those with and without SPT at the esophagus while focusing on the Notch intracellular domain (NICD).
Patients and Methods
Patients
A total of 1427 HNSCC cases were treated at the Department of Otorhinolaryngology, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok, Thailand during the 2006–2017 study period. Twenty-three SPTs at the esophagus and forty-seven non-SPTs were matched by the index primary cancer. The primary sites included oral cavity, oropharynx, hypopharynx, and larynx (nasopharynx excluded). Using the criteria proposed by Warren and Gates,9 second primary tumor was defined as two tumors diagnosed as malignant by histologic examination that are separated by normal non-neoplastic mucosa. In the present series, the lower limit was set at 2 cm. In this study, second primary tumor also included the development of a tumor after 5 years of treatment.
Demographic and clinical data, including age, gender, TNM staging, primary site of lesion, histopathologic differentiation, and total follow-up time were recorded. Lifestyle risk factors, such as cigarette smoking and alcohol consumption, were further classified according to Hosokawa et al.10 Cigarette smoking status was defined as “never smoked”, “active smoker” (defined as current cigarette smoker or stop smoking for less than 10 years), or “stopped smoking” (defined as stopped smoking for ≥10 years). Alcohol consumption was defined as “never drank alcohol”, “occasional drinker”, or “daily drinker”. A person who consumes alcohol daily was categorized as a “daily drinker”, whereas a person who consumes alcohol less often than daily was categorized as an “occasional drinker”.
Patients aged 18 years and over at the time of diagnosis were included. Those with HNSCC diagnosed without SPT received follow-up for a period of at least 3 years after the primary tumor diagnosis. The pathologic specimens of HNSCC were verified by an experienced head and neck pathologist. Exclusion criteria were subjects with non-squamous cell carcinoma, post-radiation treatment, and/or low-quality formalin-fixed paraffin-embedded (FFPE) tissues.
This retrospective study was conducted in accordance with the ethical principles of the Declaration of Helsinki. All personally identifiable information was removed and confidentially recorded as anonymous data. Since the level of research did not exceed the minimum risk to subjects, the requirement to obtain written informed consent was waived. The protocol for this study was approved by the Siriraj Institutional Review Board (SIRB) of the Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok, Thailand (protocol number 110/2559 [EC1]).
Immunohistochemistry
FFPE samples from 23 HNSCCs with SPT and 47 HNSCCs without SPT were obtained from the Department of Pathology, Faculty of Medicine Siriraj Hospital, Mahidol University. Sections (5 µm) of FFPE specimens were mounted on poly-L-lysine coated slides and processed for conventional histologic assessment by immunohistochemistry (IHC). Activated Notch signaling pathway was determined by intranuclear staining of NICD. For antigen retrieval, tissue sections were deparaffinized in target retrieval solution (10x, pH 6.0) at 95°C for 40 mins. Sections were incubated with 3% hydrogen peroxide (H2O2) for 10 mins, 2% bovine serum albumin (BSA) for 20 mins, and at room temperature with primary polyclonal anti-human antibody of Notch1 (ab 8925; Abcam, Cambridge, United Kingdom) for 1 hr. The appropriate secondary antibodies were then applied and counterstained with hematoxylin. A single-blinded analysis of stained tissue slides was performed via microscopic evaluation by a single pathologist (TA).
Scoring of Immunohistochemical Staining
IHC results for Notch1 proteins were in accordance with the scoring criterion proposed by Tripathi et al.11 Briefly, protein expression was semi-quantified using a 0–3 scoring system to indicate the intensity of cell staining (0 for no staining, 1 for weak staining, 2 for moderate staining, and 3 for intense staining) (Figure 1). A scoring system of 1–4 was used to rate the percentage (range: 0–100%) of positively stained cells of any intensity (0–10% = 0; 11–30% = 1; 31–50% = 2; 51–70% = 3, and 71–100% = 4). Overexpression was defined for tumors that show both an intensity score of ≥2 and a positive staining score of ≥3.
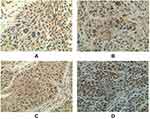 |
Figure 1 Intensity grading of NICD staining: Absence (A), Weak (B), Moderate (C), and Intense (D). |
Statistical Analysis
Factors affecting survival rate, including age, gender, tumor stage, smoking status, alcohol consumption status, and site of primary tumor, were analyzed by unpaired t-test, chi-square test, or Fisher’s exact test. Categorical data are presented as frequency and percentage, and continuous data are presented as mean ± standard deviation. A p-value of ≤0.05 was considered statistically significant. Survival time of HNSCC with SPT, which was defined as the interval from the date of initial presentation at our center to the date of the last follow-up or death, was calculated using Kaplan–Meier method. All data were analyzed using PASW Statistics for Windows, version 18.0 (SPSS, Inc., Chicago, IL, USA).
Results
The demographic and clinical characteristics of the SPT and non-SPT groups are shown in Table 1. The mean age was significantly lower in the SPT group than in the non-SPT group (55.13 vs 62.09 years, respectively; p=0.02). Regarding SPT detection, 82.6% were synchronous and 17.4% were metachronous. The average duration until detection of the secondary tumor was 37 months. The 3- and 5-year survival rates were 38.0% and 25.3%, respectively, with a median survival time of 2 years (Figure 2).
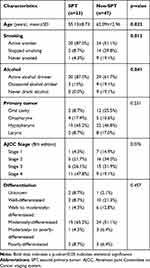 |
Table 1 Demographic and Clinical Characteristics of Study Patients |
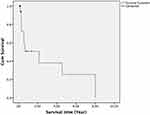 |
Figure 2 The 3- and 5-year survival rates of SPT patients. |
Regarding lifestyle risk factors, the percentage of patients that were active smokers in the SPT group and the non-SPT group was 87.0% and 51.1%, respectively (p=0.01). The proportion of those who had stopped smoking in the SPT group and the non-STP group was 8.7% and 29.8%, respectively (p=0.013). Regarding active alcohol use, 87.0% of patients in the SPT group and 61.7% of patients in the non-SPT group were daily alcohol drinkers (p=0.041).
NICD expression in HNSCC was noted in 11 cases with SPT (47.8%), and in 21 cases without SPT (44.7%). The intensity and percentage of expression between groups is shown in Table 2.
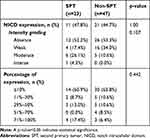 |
Table 2 Notch Intracellular Domain (NICD) Expression |
Discussion
The development of secondary tumor in HNSCC, particularly esophageal carcinoma, is common. Regarding the location of SPT in head and neck carcinoma, it is more likely that a patient with oropharyngeal or hypopharyngeal carcinoma will develop esophageal cancer than oral cavity or laryngeal carcinomas. The risk factors for developing SPT include age, male gender, cigarette smoking, alcohol use, and betel nut consumption.10,12,13 These carcinogens cause multiple sites of mutation, resulting in “mutant clone”. In addition, via intercellular signaling, the aberrant “microenvironment” selectively promotes the mutant lineages that lead to tumorigenesis. In this study, patients with active cigarette smoking and alcohol consumption were at significantly higher risk for having synchronous and/or metachronous SPT.
Analysis of TCGA data relative to smoking and HPV status has been performed.14 Different genomic portraits were demonstrated between these two exposome components. High-risk HPV infection is now considered a leading cause of virus-associated HNSCC carcinogenesis. From our previous study, only 6 percent of HPV-related cancer was detected in the oropharynx. No detectable high-risk HPV was demonstrated at other subsites.15,16 Due to a very low prevalence of HPV-related HNSCC in our population, no significant association with SPT was established.
Despite modern treatment modalities, the survival rate of patients with SPT remains unsatisfactory. HNSCC develops via a prolonged multistage process that involves an accumulation of genetic and epigenetic alterations. Several investigations revealed several genes and pathways that play an important role in HNSCC tumorigenesis, including TP53, CDKN2A, Cyclin D1, PIK3CA, HRAS, and EGFR.17–20 However, these molecular alterations do not fully explain the pathogenesis of HNSCC.
Agrawal et al and Stransky et al confirmed the genes previously reported to play a role in HNSCC, and they also reported novel mutations in NOTCH1.21,22 The inactivating mutations of NOTCH1 were found in 10–15% of HNSCC tumors in both of those studies, which suggests NOTCH1 as the second most frequently mutated gene after TP53. Notch signaling is an attractive target of investigation among several molecular targets due to its involvement in a variety of cellular physiological processes, such as proliferation, differentiation, and cell survival. Notch proteins are a family of heterodimeric transmembrane receptors that are composed of an extracellular domain that is responsible for ligand recognition, a transmembrane domain, and an intracellular domain in signal transduction. To date, four Notch receptors (Notch 1–4) and five ligands (Jagged-1, Jagged-2, Delta-1, Delta-3, and Delta-4) have been described in mammals. The pathway is initiated when one cell expresses the appropriate ligand (Jagged or Delta) and interacts with another cell expression of Notch receptors (Notch 1–4). Upon ligand binding, the transmembrane Notch receptor is subsequently cleaved by ADAM metalloprotease and gamma-secretase complex. The cleaved product (NICD) translocates into the nucleus where it interacts with the nuclear DNA-binding factors. The most prominent targets of the Notch pathway comprise a set of basic-helix-loop factors of the Hes and Hey families.4
Notch signaling is tissue specific. Several studies suggested that NOTCH1 mutation can have pleiotropic effects. It was reported that gain-of-function NOTCH1 mutations are observed in leukemia’s cluster of the negative regulatory region and the C-terminal PEST domain.23 In contrast, loss-of-function NOTCH1 mutations were observed in cutaneous and lung squamous cell carcinoma.24 The cancer genome atlas of HNSCC reveals loss-of-function mutations of NOTCH1, and concludes that NOTCH1 acts as a tumor suppressor.25
Regarding the frequency of transcriptional alterations of Notch1 in HNSCC, we investigated NICD expression using IHC technique to identify the intranuclear expression of activated domain of Notch1. Our findings revealed an absence of NICD expression in HNSCC with and without SPT in 52.2% and 53.3% of cases, respectively. Despite the fact that no statistically significant difference was observed between groups, the high rate of NICD inactivation in this study supports its role as a tumor suppressor gene. Since the NOTCH gene would be unlikely to present in a significant proportion of normal cells, negative IHC staining may not be solely caused by loss of function due to mutation of NOTCH1. In hematologic malignancy, the perturbation of Notch signaling is mediated by aberration of Notch transcriptional coactivators.26
Discrepancy between NOTCH inactivation and NOTCH upregulation has been reported in HNSCC. Meta-analysis of data collected from many Asian studies found a high percentage of NOTCH activation.6 Gokulan et al reported that the intensity of NICD expression gradually progressed from dysplasia to carcinoma, and was significantly overexpressed in locally advanced HNSCC.27 Moreover, the expression of Notch1 might be used as an independent prognostic indicator of oral cancer. On the contrary, our primary tumors in both groups had less than 50% NICD expression. Furthermore, the percentage and intensity of expression did not indicate the development of SPT.
This study has some mentionable limitations. First, this was a retrospective, single-center study. Second, we decided not to calculate the survival rate in the non-SPT group because these patients required at least 3 years of disease-free follow-up. Additionally, more information was required to affirm the relationship between Notch1 expression and SPT development. Further study of other regulators, such as ligand binders and downstream activators, might provide more information about NOTCH1 mutation in HNSCC. In order to conclude the absence of protein expression caused by the NOTCH1 mutation, whole exome sequencing (WES) would be necessary.
Conclusion
Active smoking and alcohol use were found to be significantly associated with SPT development. A high prevalence of NICD inactivation was noted in HNSCC with no statistically significant difference between groups. The Notch signaling pathway is involved in HNSCC tumorigenesis, but may not be a suitable molecular marker for SPT development.
Acknowledgments
The authors gratefully acknowledge Mrs. Sunattee Kessung and Ms. Jeerapa Kerdnoppakhun for their assistance with manuscript development, and Miss Julaporn Pooliam of the Division of Clinical Epidemiology, Research Department, Faculty of Medicine Siriraj Hospital, Mahidol University for assistance with statistical analysis.
Disclosure
All authors declare no personal or professional conflicts of interest, and no financial support from the companies that produce and/or distribute the drugs, devices, or materials described in this report. This study project was supported by a grant from Siriraj Research Fund (grant no. [IO] R016032009), Faculty of Medicine Siriraj Hospital, Mahidol University.
References
1. Leon X, Quer M, Diez S, Orus C, Lopez-Pousa A, Burgues J. Second neoplasm in patients with head and neck cancer. Head Neck. 1999;21(3):204–210. doi:10.1002/(SICI)1097-0347(199905)21:3<204::AID-HED4>3.0.CO;2-7
2. Chuang SC, Scelo G, Tonita JM, et al. Risk of second primary cancer among patients with head and neck cancers: a pooled analysis of 13 cancer registries. Int J Cancer. 2008;123(10):2390–2396. doi:10.1002/ijc.23798
3. Schwartz LH, Ozsahin M, Zhang GN, et al. Synchronous and metachronous head and neck carcinomas. Cancer. 1994;74(7):1933–1938. doi:10.1002/(ISSN)1097-0142
4. Sun W, Gaykalova DA, Ochs MF, et al. Activation of the NOTCH pathway in head and neck cancer. Cancer Res. 2014;74(4):1091–1104. doi:10.1158/0008-5472.CAN-13-1259
5. Lee SH, Do SI, Lee HJ, Kang HJ, Koo BS, Lim YC. Notch1 signaling contributes to stemness in head and neck squamous cell carcinoma. Lab Invest. 2016;96(5):508–516. doi:10.1038/labinvest.2015.163
6. Zhao YY, Yu GT, Xiao T, Hu J. The Notch signaling pathway in head and neck squamous cell carcinoma: a meta-analysis. Adv Clin Exp Med. 2017;26(5):881–887. doi:10.17219/acem/64000
7. Miele L. Notch signaling. Clin Cancer Res. 2006;12(4):1074–1079. doi:10.1158/1078-0432.CCR-05-2570
8. Zeng Q, Li S, Chepeha DB, et al. Crosstalk between tumor and endothelial cells promotes tumor angiogenesis by MAPK activation of Notch signaling. Cancer Cell. 2005;8(1):13–23. doi:10.1016/j.ccr.2005.06.004
9. Warren S, Gates O. Multiple primary malignant tumors; survey of literature and statistical study. Am J Cancer. 1932;16:1358–1414.
10. Hosokawa S, Takahashi G, Okamura J, et al. Risk and prognostic factors for multiple primary carcinomas in patients with head and neck cancer. Jpn J Clin Oncol. 2018;48(2):124–129. doi:10.1093/jjco/hyx178
11. Tripathi SC, Matta A, Kaur J, et al. Nuclear S100A7 is associated with poor prognosis in head and neck cancer. PLoS ONE. 2010;5(8):e11939. doi:10.1371/journal.pone.0011939
12. Ryser MD, Lee WT, Ready NE, Leder KZ, Foo J. Quantifying the dynamics of field cancerization in tobacco-related head and neck cancer: a multiscale modeling approach. Cancer Res. 2016;76(24):7078–7088. doi:10.1158/0008-5472.CAN-16-1054
13. Feng Z, Xu QS, Niu QF, et al. Risk factors for patients with multiple synchronous primary cancers involving oral and oropharyngeal subsites. Oral Surg Oral Med Oral Pathol Oral Radiol. 2016;121(4):360–366. doi:10.1016/j.oooo.2015.10.031
14. Irimie AI, Braicu C, Cojocneanu R, et al. Differential effect of smoking on gene expression in head and neck cancer patients. Int J Environ Res Public Health. 2018;15:7. doi:10.3390/ijerph15071558
15. Pongsapich W, Jotikaprasardhna P, Lianbanchong C, Phumchan A, Siritantikorn S, Chongkolwatana C. Human papillomavirus infection in oral cavity and oropharyngeal cancers: are they the same story? J Med Assoc Thai. 2016;99(6):684–690.
16. Pongsapich W, Eakkasem N, Siritantikorn S, Pithuksurachai P, Bongsabhikul K, Chongkolwatana C. Prevalence of HPV infection in hypopharyngeal and laryngeal squamous cell carcinoma at Thailand’s largest tertiary referral center. Infect Agent Cancer. 2017;12:58. doi:10.1186/s13027-017-0167-0
17. Poeta ML, Manola J, Goldenberg D, et al. The Ligamp TP53 assay for detection of minimal residual disease in head and neck squamous cell carcinoma surgical margins. Clin Cancer Res. 2009;15(24):7658–7665. doi:10.1158/1078-0432.CCR-09-1433
18. Demokan S, Chuang A, Suoglu Y, et al. Promoter methylation and loss of p16(INK4a) gene expression in head and neck cancer. Head Neck. 2012;34(10):1470–1475. doi:10.1002/hed.21949
19. Murugan AK, Hong NT, Fukui Y, Munirajan AK, Tsuchida N. Oncogenic mutations of the PIK3CA gene in head and neck squamous cell carcinomas. Int J Oncol. 2008;32(1):101–111.
20. Bonner JA, Harari PM, Giralt J, et al. Radiotherapy plus cetuximab for locoregionally advanced head and neck cancer: 5-year survival data from a Phase 3 randomised trial, and relation between cetuximab-induced rash and survival. Lancet Oncol. 2010;11(1):21–28. doi:10.1016/S1470-2045(09)70311-0
21. Agrawal N, Frederick MJ, Pickering CR, et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science. 2011;333(6046):1154–1157. doi:10.1126/science.1206923
22. Stransky N, Egloff AM, Tward AD, et al. The mutational landscape of head and neck squamous cell carcinoma. Science. 2011;333(6046):1157–1160. doi:10.1126/science.1208130
23. Weng AP, Ferrando AA, Lee W, et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science. 2004;306(5694):269–271. doi:10.1126/science.1102160
24. Wang NJ, Sanborn Z, Arnett KL, et al. Loss-of-function mutations in Notch receptors in cutaneous and lung squamous cell carcinoma. Proc Natl Acad Sci U S A. 2011;108(43):17761–17766. doi:10.1073/pnas.1114669108
25. Cancer Genome Atlas N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517(7536):576–582.
26. Kalaitzidis D, Armstrong SA. Cancer: the flipside of Notch. Nature. 2011;473(7346):159–160. doi:10.1038/473159a
27. Gokulan R, Halagowder D. Expression pattern of Notch intracellular domain (NICD) and Hes-1 in preneoplastic and neoplastic human oral squamous epithelium: their correlation with c-Myc, clinicopathological factors and prognosis in Oral cancer. Med Oncol. 2014;31(8):126. doi:10.1007/s12032-014-0374-0
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
