Back to Archived Journals » Nanobiosensors in Disease Diagnosis » Volume 3
Nanobiosensors: the future for diagnosis of disease?
Authors Prasad S
Received 10 February 2014
Accepted for publication 26 March 2014
Published 4 August 2014 Volume 2014:3 Pages 1—10
DOI https://doi.org/10.2147/NDD.S39421
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Shalini Prasad
Department of Bioengineering, University of Texas, Dallas, TX, USA
Abstract: Nanotechnology is having a profound impact on the development of a new class of biosensors known as nanobiosensors. Nanobiosensors commonly comprise a biological recognition molecule immobilized onto the surface of a signal transducer. The reaction between the biorecognition molecule and the analyte is a heterogeneous reaction and therefore the design of the biosensing interface is important in determining the performance of the nanobiosensor. Nanobiosensors are being widely used for molecular detection of biomarkers associated with diagnosis of disease. The application of new nanomaterials in biosensing has influenced biosensing research. The use of high surface area nanomaterials has been important in producing nanobiosensors with greater sensitivity and shorter response times. This review summarizes the advances in disease diagnostics, primarily through the detection of molecular biomarkers, such as proteins and nucleic acids mediated by use of nanobiosensors.
Keywords: nanomaterial, biosensors, disease diagnosis, biomimetic, macromolecular crowding, performance metrics
Need for use of nanomaterial biosensors in diagnosis of disease
A biosensor is a device designed to detect or quantify a biochemical molecule, such as a particular DNA sequence or a particular protein. Many biosensors are affinity-based, meaning they use an immobilized capture probe that selectively binds the molecule being sensed, ie, the target or analyte, thus transferring the challenge of detecting a target in solution to detecting a change at a localized surface. This change can then be measured using a variety of methods, including biosensors requiring light (eg, surface plasmon resonance [SPR] or fluorescence), mechanical motion (eg, quartz crystal microbalance or resonant cantilever), or magnetic particles.
From the palette of techniques available to the user, the electrical method of interrogating sensors is widely used in designing “label-free” biosensors, which do not require a label or a tag to report the detection of a specific molecule. Electrical biosensors rely solely on the measurement of currents and/or voltages to detect binding.1–3 Due to their low cost, low power, and ease of miniaturization, electrical biosensors hold great promise for applications where minimizing size and cost is crucial, such as “point-of-care” disease diagnostics.
Biosensors of nanoscale dimensions may overcome many of the stumbling blocks that prevent wide scale use of affinity biosensors, without any of their major drawbacks. This is a conclusion that can be drawn from a recent research in the area of affinity-based biosensors.2,3 Affinity biosensors use a biorecognition molecule, such as an antibody or DNA, that selectively binds to an analyte to form a complex. A signal transducer determines the extent of the binding reaction and outputs this information to the end user. The interest in such devices is great, because the vast majority of analytes in disease diagnosis can be best detected using an affinity reaction. Furthermore, the specificity and high affinity of antibodies for their target species potentially allows biosensors to perform analyses at low concentrations of analyte in complex samples.
There are two major thrust areas in designing nanoscale affinity biosensors. The first is to reduce the detection limit and the second is the ability to detect a number of analytes in the same sample, which is known as multiplexing. Multiplexing is important because reliable diagnosis of disease often requires identification of the levels of a number of molecular markers.4 The barrier to reducing detection limits is the thermodynamics of the affinity reaction. Typically, dissociation constants (Kd) for antibody-antigen reactions are in the order of 108–1012 M, with the Kd for the known strongest bioaffinity reaction between biotin and streptavidin being 1015 M.5 With conventional transduction techniques, the detection limits are therefore in the range of 109–1014 M. In nanobiosensors, lower detection limits are achieved by utilizing nanomaterials such as nanoparticles, nanotubes, nanowires, nanomembranes, or nanotextured surfaces.6 The drawback of amplification techniques is that they typically require labeling and sample manipulation steps, which restrict these technologies to the hands of experienced operators and preclude real-time analysis. Highly sensitive, label-free transduction of biorecognition reactions has been achieved with cantilevers7 and field-effect transistor configurations.8,9
Definition of a nanobiosensor
Nanobiosensors are devices that measure a biochemical or biological event using any electronic, optical, or magnetic technology through a compact probe.6–9 Current advances in nanotechnology and advanced fabrication technology in electronics intersect towards creating a new set of biosensors called nanobiosensors and have ushered in a new era of bionanotechnology for disease diagnosis.
Need for real-time measurements in disease diagnosis
Monitoring human health for early detection of disease conditions or health disorders is vital for maintaining a healthy life. Many biomolecules help determine the physiological state of a disease condition.10–12 These biomolecules are typically called biomarkers for diagnosing disease. In addition, analysis of food and the environment for perturbants such as pesticides and river water contaminants with harmful biomolecules have also become invaluable agents for health diagnosis. Thus, there is an ongoing need for rapid analysis, active continuous time monitoring systems with substantial accuracy for detecting biomolecules. A “real-time” biosensor that detects the analytes of interest in a near continuous time manner plays an important role in accomplishing effective data generation and data processing, supporting real-time decision-making and rapid manipulation.13,14 In order to accomplish these multiple requirements from multiple environments, one of the standard approaches adopted has been to develop hybrid biochemical analysis systems. These multiscale biosensors are versatile because they can monitor specific analytes from a wide range of environments at ultralow concentrations. They comprise a combination of nanomaterials integrated with microfluidic capabilities. This approach is similar to that followed by the semiconductor industry in integrated circuits. Microfluidic research involves the study of several fluid manipulation, detection, and separation techniques. Often these different components are integrated with essential electronics to develop a complete “on-chip” analysis system. Well established fabrication techniques are adapted from the semiconductor industry, such as micromachining, injection and replica molding, soft lithography, wet etching, and photolithography. These techniques enable miniaturization of fluid handling systems to palm-held “micro-total analysis systems” or “lab on a chip” devices which can perform the myriad of diagnostics and analysis tasks associated with a standard clinical laboratory assay. Hence, a lot of the current research in this area focuses on the integration of these complex requirements of real-time measurements with on-chip detection capabilities to build multifunctional nanobiosensors.15 The following section explains the basic concepts of nanobiosensors.
Basic concept of nanobiosensors
A nanobiosensor is a means of detecting biological agents such as antibodies, nucleic acids, pathogens, and metabolites. The working principle consists of binding bioanalytes of interest onto bioreceptors, which in turn modulate the physiochemical signal associated with the binding. Later, a transducer captures and converts the physiochemical signal into an electrical signal. The variation in signal such as electric potential, current, conductance, impedance, intensity and phase of electromagnetic radiation, mass, temperature, and viscosity is monitored. Analysis of the variation in one or more of these parameters quantifies the presence or absence of bioagents. The nanostructures in nanobiosensors act as an intermediate layer between biological agents and physicochemical detector components or biological agents, and the transducer is combined with nanomaterials to construct a biosensor.16
Existing biomolecule transduction methodologies
Biomolecule transduction can be broadly divided into two classes, ie, label-based and label-free detection.
Label-based detection
Most label-based detection technologies are based on immunoassays. These immunoassays are based on antigen–antibody reactions. Current immunoassays use enzyme-linked immunosorbent assay (ELISA) tests to detect the amount of protein present in blood. ELISA uses a biochemical technique to detect the presence of an antibody or antigen present in the sample. ELISA has been the standard diagnostic tool for the last 30 years. It is a very sensitive technique with low detection limits, but there are a few downsides to this technology. ELISA uses fluorophore-tagged linker molecules to improve detection, and these fluorophores might denature the proteins. Fluorescence detection techniques are based on fluorescent markers that emit light at specific wavelengths and the presence and the enhancement or reduction, as in fluorescence resonance energy transfer, the optical signal determines if the binding reaction has taken place.17 There are multiple wash steps, which increases the time involved. ELISA is a laboratory-based technique that needs trained professional technicians to run the tests and this leads to long delays. These drawbacks make ELISA unsuitable for rapid point-of-care biosensing applications.
Label-free detection methods
Rapid point-of-care diagnostics have been developed with the help of advances in microfabrication and nanotechnology. The following are a list of techniques that enable label-free detection of biomolecules. Figure 1 summarizes the label-free techniques available for designing nanobiosensors.
Electrochemical and electrical detection
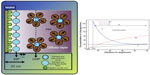
Electrical biosensors hold the greatest promise for applications where minimizing size with ease of use and cost are crucial, such as point-of-care diagnostics or patient care at the bedside. These biosensors primarily base measurement of the binding event on change in currents and/or voltages. Biosensors using electrochemical methods can be further classified according to how the electrical measurement is made, which includes voltammetric, amperometric/coulometric, and impedance measurements.18 Electrical biosensors border very closely with the electroanalytical techniques that have been developed in the field of chemistry. Figure 2 shows the charge distribution that occurs at the surface of the biosensor, which enables electrical measurements.
Optical detection
Chemiluminescence is the generation of energy in the form of light when a chemical reaction takes place. In synthetic compounds, such chemiluminescence takes place when a highly oxidized species such as peroxide emits energy during a chemical reaction. When some synthetic compounds are added to a biomolecule to form a conjugate, energy is released in the form of light, which could be used as a detection technique. Bioluminescence is another type of luminescent technique that has been reported using firefly luciferase/luciferin as the synthetic compound.19
SPR is a technique that measures surface activity. In this technique, a longitudinal wave of a certain charge density is propagated along the surface of the metal and the dielectric interface.20 Due to the total internal reflection of light against materials such as gold or silver, the evanescent wave or field created at the surface penetrates the interface into the dense medium (metal); this evanescent light is able to couple with free electrons known as plasmons on the surface, and this creates a resonance wave, which is recorded. Hence, if there is any biomolecule layer located on the surface of the metal, an SPR adsorption profile is obtained, and due to the properties of biomolecules, each SPR spectrum is different. The angle of the SPR is the factor that changes with each molecule. Use of metal nanoparticles instead of thin films produces significant amplification of the SPR signal, thus enabling ultrasensitive detection. SPR has already been commercialized, but the biosensors using this technique are yet to come. The issue with SPR-based biodetection systems is specificity and its very low signal-to-noise ratio.
Mass-based detection
Mechanical detection for biochemical entities is generally achieved by use of microscaled or nanoscaled cantilever sensors. The cantilever is excited mechanically to resonate at its resonant frequency, and then the resonant frequency is compared to the one with biomolecules bound to the surface of the cantilever, and the shift in resonant frequency determines the mass change on the surface. The advantage of using cantilever sensors is the ability to detect interacting compounds without attaching labels to the biomolecules. The biggest drawback is the complexity of the device and also the complexity of the detection mechanism. These problems, coupled with the ultrahigh sensitivity to temperature and ambient conditions, prevent this very accurate measurement technique from being popular for portable devices.
Nanomaterials for biosensing
In a biosensor, the sensing element that responds to the substance being measured is biological in nature. It has to be connected to a transducer of some sort, such that a visually observable signal can be recorded. At present, transducers that can be incorporated into the sensors are nanomaterials. The widespread interest in nanomaterials is driven by their many desirable properties; in particular, the ability to tailor the size and structure and hence the properties of nanomaterials offers excellent prospects for designing novel sensing systems and enhancing the performance of the biosensor.21,22
Nanoparticles
Conventional (optical) sandwich bioaffinity assays have the disadvantage of capturing only a small number of labels per binding event. Nanoparticles can be custom-made for specific bioassays, and because of their small size, their properties are strongly influenced by the binding of target biomolecules.23 Using nanoparticles significantly enhances the signal from the assays. The driving force behind the use of nanoparticle-quantifying tags in optical bioassays has been addressing the limitations of organic fluorophores. Mirkin’s group demonstrated that aggregation of gold nanoparticles induced by DNA hybridization leads to materials with remarkable optical properties.24 High fluorescence intensity of semiconductor quantum dots can also lead to remarkably sensitive bioassays. Hahn et al reported highly sensitive detection of the single bacterial pathogen Escherichia coli 0157 using CdSE/AnS core-shell quantum dots conjugated to streptavidin.25
Nanowires
Nanowires offer the best chance of creating robust, sensitive, and selective electrical detectors of biological binding events. Current flow in any one-dimensional system is extremely sensitive to minor perturbations, and in nanowires the current flows extremely close to the surface.26,27 Biological macromolecules are the same size as the diameter of these nanowires. The combination of the tunable conducting properties of semiconducting nanowires and the ability to bind analytes on their surface yields a direct, label-free electrical readout.28 These sensors work on the principle of ion-selective field effect transistors and rely on the interaction of external charges with carriers in the nearby semiconductor, which results in enhanced sensitivity at low ionic strength.
Carbon nanotubes
Carbon nanotubes are particularly exciting one-dimensional nanomaterials that have attracted considerable interest owing to their unique structure-dependent electronic and mechanical properties. Because of their high surface-to-volume ratio and novel electron transport properties, the electronic conductance of these nanostructures is strongly influenced by minor surface perturbations, such as those associated with binding of macromolecules. Such one-dimensional materials thus offer the prospect of rapid (real-time) and sensitive label-free bioelectronic detection, and massive redundancy in nanosensor arrays.29
Nanomembranes
Membranes of various pore size, length, morphology, and density have been synthesized from diverse materials for size exclusion-based separation.30,31 Specific bioagent immobilization and detection remains a great technical challenge in many bioanalytical assays. To achieve this, materials with controllable pore diameter, length, and surface chemistry are needed. Selective capture requires two steps, ie, collection and immobilization. Membranes are well suited for this because of their enhanced surface interactions with the liquid being analyzed.
The overall sensitivity of any sensor depends on signal transduction and the mass transport effect. Miniaturization of a sensor increases the signal-to-noise ratio, an inherent advantage for signal transduction. It has been reported that mass transport of analytical solution through the sensor surface plays an important role in determining sensitivity.32 The detection limit for bioassays depends on the amount of biomolecular interaction with the sensor surface. Whitman’s group at the Naval Research Laboratory, Washington, DC, reported that femtomolar detection limits for bioassays are likely.32 They predicted that the limit of detection would be due to limitations in analyte transport not limitations in signal transduction, and without directed transport of biomolecules, individual nanoscale sensors will be limited to picomolar-order sensitivity for practical time scales. Total flux to the sensor was studied as a function of sensor geometry and volumetric flow. Enhancing mass transport by conventional methods of decreasing height decreased the volumetric flow rate, which in turn decreased the total flux of the sensor. On the other hand, injecting the analyte directly into the sensor rather than merely streaming it past the sensing surface increased the mass transport effect, which in turn increased the total flux of the sensor. It was found that the flux of the sensor could be increased by using a nanoporous membrane. Thus, the sensitivity of sensors could be enhanced by incorporating nanoporous membranes into microfluidic-based biosensors.
Biomimetic nature of nanobiosensors
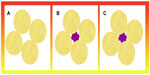
Nanomaterial structures enable spatial confinement. In support of this observation, there is a compelling argument in the realm of biophysical understanding of proteins and their structures. A crowded environment exists inside a cell structure, in which the cytoplasm is typically different from the dilute solutions generally used in in vitro studies of proteins, and there is a school of thought which states that this may significantly affect the behavior of proteins. The high concentration of proteins, nucleic acids, and complex sugars in cells have various energetic consequences and size constraints for smaller molecules; the effect of these bigger structures is known as “macromolecular crowding”.33,34 The concentration of macromolecules is up to 400 g/L, meaning that 5%–40% of the total volume is physically occupied by these molecules.35 An even larger fraction of the total volume is not available for molecules of similar sizes, as shown in Figure 3.
How important is confinement?
The binding affinities and rates of self-assembly can change by orders of magnitude as a result of confinement. Crowding is therefore a very important factor when performing an in vitro study. To study cells and their mechanisms in vivo, techniques known as cryoelectron tomography and fluorescent tags are used. Evidence has been presented that the reaction rates in cytoplasm filled with large ensembles of macromolecules are very similar to the ones predicted by the molecular crowding theory. In short, what this means is that the biochemistry that would take place in a test tube is not exactly the same as that taking place in the body. Thus, it is imperative to use this idea in a device that measures the accurate concentration of proteins and other biomolecules, as it is measured from physiological fluids.35,36
Protein folding and binding in confined spaces
Simple theoretical models presented by various groups to study and understand the essence of folding and binding of proteins in confined places was also taken into consideration during the conception of the device.37–40
Protein folding is the process by which the linear information contained in the amino acid sequence of a polypeptide gives rise to the well defined three-dimensional conformation of the functional protein. Because unfolded proteins can reach their native state spontaneously in vitro, it was assumed that folding (acquisition of the tertiary structure) and assembly (formation of protein oligomers) of newly synthesized polypeptides in vivo occurs essentially uncatalyzed and without the input of metabolic energy. This central idea has been revised in recent years after the discovery that correct folding of many proteins inside the cell depends on pre-existing protein machinery, known as molecular chaperones.41 This enables a certain level of spatial confinement, providing the former with a “cage” structure and a confined volume. Crowding prevents self-assembly of partly folded polypeptide chains, which is characterized as aggregation in in vitro studies.42 Confinement appears to stabilize proteins and accelerate their folding significantly,43 and would slow down the unfolding of proteins which is known to cause denaturation of protein structures and loss of their functionality.35–37
From the above discussion, it would seem useful to provide a very small confined area in which the proteins could be detected. To achieve this purpose, nanobiosensors comprising nanomaterial embedded on the surface of measurement electrodes have been designed for detecting disease biomarkers.1–3
Parameters indicating nanobiosensor performance
During the development of nanobiosensor devices, each of these individual elements must be individually optimized; however, for the device to meet requirements, each of the elements must work in unison.
Selectivity refers to the response of the nanobiosensor only to the target biomolecules and not to other similar molecules. It is extremely unlikely that a readout method would be able to distinguish between various specific and nonspecific interactions.
In real-world blood samples, the target biomolecules are present in lower concentrations than those of nonspecific biomolecules. Certain trade-offs exist between selectivity and preparation of real-world samples to be made suitable for detection. The most daunting challenge in the field of biosensors is meeting the selectivity requirements when tested with complex real-world samples. The concept of nonspecific binding further complicates the task at hand. This refers to the nonspecific, nontarget biomolecules that attach themselves to the biological probes located on the transducer probe layer. These nonspecific molecules reduce the number of binding locations for the corresponding target biomolecules, not only preventing the binding event but also creating the possibility of a false-positive signal. To overcome this problem, blocking agents such as bovine serum albumin which nonspecifically adsorbs, is readsorbed onto the surface of the immobilized biomolecule and this prevents any further nonspecific binding reactions during the exposure to the complex real-world samples.44
The limit of detection is the smallest or lowest amount of target biomolecule concentration that can be reliably detected by the nanobiosensor. This is the figure of merit of the sensor. The term “sensitivity of the nanobiosensor” could be used for some descriptions to refer to the same parameter.
The dose-response curve is the graph from which the limit of detection and range of concentrations that can be measured using the nanobiosensor are plotted. The dose-response curve is also known as the calibration curve of the device. Reproducibility is another important parameter associated with the device which can be seen from the error bars on a graph. Dynamic range is the ratio of the largest measurable target concentration and the limit of detection. Nanobiosensors can also be used to quantify the presence of the target biomolecule rather than just detect it. The resolution of the nanobiosensor is the smallest detectable change in target biomolecule concentration that can be detected. Dynamic range and limit of detection are interrelated, and the design of binding agents could be used to alter one parameter while reducing other parameters.
Multiplexing refers to detection of multiple target biomolecules from a single biological sample. This is one of the most important requirements of a nanobiosensor, and could be achieved by localization of different biological probes at different regions of the nanobiosensor. The biggest issue with multiplex detection is the problem of cross-reactivity. A certain biological probe may react with more than one of the target molecules, or certain target molecules might foul the probes. Using creative design techniques and highly specific probe molecules, these reactions could be reduced, and improved multiplexed detection could be achieved.
From the above description of the performance parameters, it appears that the limitations of the nanobiosensor lie in the biological affinity probes rather than in the actual electrical detection mechanism. With improved understanding of the immobilization chemistry and minimizing nonspecific binding issues, the limits of detection using biological probes could be enhanced further.
Integration of nanotechnology into biochip assay formats
Nanotechnology on a chip is the new paradigm for designing diagnostic assays for diagnosis of disease.25 The ability to achieve analytical quantification of chemical and biological information in a rapid and relatively inexpensive way will impact molecular diagnostics and health care. Some examples of devices that incorporate nanotechnology-based biochips and microarrays are nanofluidic arrays and protein nanobiochips. These devices can be adapted for point-of-care use.
One of the more promising uses of nanofluidic devices is isolation and analysis of individual biomolecules that have diagnostic relevance, such as DNA and protein detection. This capability could lead to new detection schemes for a number of chronic diseases. Nanofluidic technology is expected to have broad applications in systems biology, personalized medicine, detection of pathogens, drug development, and clinical research.
Protein microarrays for the study of protein function are not widely used, in part because of the challenges in producing proteins to spot on the arrays. Protein microarrays can be generated by printing complementary DNAs onto glass slides and then translating target proteins with mammalian reticulocyte lysate.45 Epitope tags fused to the proteins allow them to be immobilized in situ. This procedure obviates the need to purify proteins, avoids protein stability problems during storage, and captures sufficient protein for functional studies.
Roadmap toward implementation of nanobiosensors for diagnosis of disease
Within the next decade, diagnostic devices based on nanotechnology will become available, and be able to perform thousands of measurements very rapidly and inexpensively. Future trends in diagnostics will continue in miniaturization of biochip technology to the nanoscale range. The most common clinical diagnostic application will be analysis of blood proteins. Blood in the systemic circulation reflects the state of health or disease of most organs. Therefore, detection of blood molecular fingerprints will provide a sensitive assessment of health and disease.
Molecular electronics and nanoscale chemical sensors will enable construction of microscopic sensors capable of detecting patterns of chemicals in a fluid. Information from a large number of such devices flowing passively in the bloodstream allows estimation of the properties of tiny chemical sources in a macroscopic tissue volume. Estimates of plausible device capabilities have been used to evaluate their performance for typical chemicals released into the blood by tissues in response to localized injury or infection.46 These observations indicate that the devices can readily enable differentiation of a single cell-sized chemical source from the background chemical concentration in vivo, providing high-resolution sensing in both time and space. With the methods currently used for blood analysis, such a chemical source would be difficult to distinguish from background when diluted throughout the blood volume and withdrawn as a blood sample.
The trend will be to build diagnostic devices from the bottom up, starting with the most fundamental building blocks. Unless there are early successes that translate into large-volume sales and early adoption, the long range forecast for nanobiosensors in disease diagnosis is promising. A factor that may support the implementation of nanobiosensors is the trend of moving away from fluorescent labeling as miniaturization reduces the signal intensity, but there have been some improvements making fluorescent labeling methods viable with nanoparticles. Nanobiosensors will also facilitate the development of non-polymerase chain reaction diagnostic technologies. As a further refinement, nanotechnology can potentially be used for analysis of a single cell to enable a genetic diagnosis.
In the near future, use of nanodiagnostics could reduce waiting times for test results. For example, patients with contagious diseases could provide urine samples when they first arrive at the clinic, and the results could be ready by the time they see the physician. Patients could then be given a prescription immediately, reducing the length of time that the patient has to wait for results, thereby decreasing anxiety, improving compliance, and making the whole process less costly.
In the next decade, nanobiotechnology-based biosensors will play an important role not only in diagnosis but also in linking diagnosis with treatment and development of personalized medicine. Because of the integration and interrelationship of several technologies involved in nanodiagnostics, those who conduct these tests or devise new tests will be taking a more active part in decision-making in future health care systems.
Another important area of application will be cancer diagnostics. Methods for molecular diagnosis of cancer, including genetic profiling, are currently commercially available. However, by the time a cancer is detected by currently available methods, it is often too late for curative treatment. Nanobiosensors can offer ultrasensitivity in detection of biomarkers for cancer, which may be applied in the future for early detection as well as treatment of cancer. Nanodevices for this purpose are now in the feasibility stage. A nanodevice for use in both cancer diagnostics and therapeutics, known as a nanotheranostic device, could be implanted as a prophylactic measure in individuals who do not have any obvious manifestations of cancer, and cancer surveillance could be conducted by remote monitoring. Such monitoring could detect cancer at the earliest stages and enable appropriate therapeutic intervention. These monitoring devices should be biodegradable, and safety must be established before implantation. Such a surveillance system would be the ultimate in personalized prevention of cancer. Early detection would increase the chances of a cure. Such a device would have advantages over detection of biomarkers in specimens of body fluids, because such examinations can be performed only periodically and would be less accurate than analyses conducted continuously in vivo.46
In conclusion, nanotechnologies promise to extend the limits of current molecular diagnostics and enable point-of-care diagnosis, integration of diagnostics with therapeutics, and development of personalized medicine. The most important clinical applications of the currently available nanotechnology are in the areas of biomarker discovery, cancer diagnosis, and detection of infectious micro-organisms. Nanomedicine promises to play an important role in the future development of diagnostics and therapeutics.
The role of nanobiosensors is to continuously monitor biomarker concentration levels for active tracking and effective treatment of disease. It also needs to be a point-of-care device which in turn means simplistic human involvement towards detection. The best possible solution would be to be able to just drop the sample onto a sensing device and continuously monitor the concentration of biomarkers present in the sample.
Disclosure
The author reports no conflicts of interest in this work.
References
Zheng G, Patolsky F, Cui Y, Wang WU, Lieber CM. Multiplexed electrical detection of cancer markers with nanowire sensor arrays. Nat Biotechnol. 2005;23:1294–1301. | |
Ferrari M. Cancer nanotechnology: opportunities and challenges. Nat Rev Cancer. 2005;5:161–171. | |
Bayer EA, Wilchek M. Biotin-binding proteins, overview and prospects. Methods Enzymol. 1990;184:49–51. | |
Nam JM, Stoeva SI, Mirkin CA. Bio-bar-code-based DNA detection with PCR-like sensitivity. J Am Chem Soc. 2004;126:5932–5933. | |
Oh BK, Nam JM, Lee SW, Mirkin CA. A fluorophore-based bio-barcode amplification assay for proteins. Small. 2006;1:103–108. | |
Di Giusto DA, Wlassoff WA, Gooding JJ, Messerle BA, King GC. Proximity extension of circular DNA aptamers with real-time protein detection. Nucleic Acids Res. 2005;33:e64. | |
Fredriksson S, Gullberg M, Jarvius J, et al. Protein detection using proximity-dependent DNA ligation assays. Nat Biotechnol. 2002;20: 473–477. | |
Gullberg M, Gústafsdóttir SM, Schallmeiner E, et al. Cytokine detection by antibody-based proximity ligation. Proc Natl Acad Sci U S A. 2004;101:8420–8424. | |
Fritz J, Baller MH, Lang HP, et al. Biosensor-based label-free assays of amyloid growth. Science. 2000;288:316–318. | |
Wu G, Datar RH, Hansen KM, Thundat T, Cote RJ, Majumdar A. Bioassay of prostate-specific antigen (PSA) using microcantilevers. Nat Biotechnol. 2001;19:856–860. | |
Chen RJ, Bangsaruntip S, Drouvalakis KA, et al. Noncovalent functionalization of carbon nanotubes for highly specific electronic biosensors. Proc Natl Acad Sci U S A. 2003;100:4984–4989. | |
Prime KL, Whitesides GM. Self-assembled organic monolayers: model systems for studying adsorption of proteins at surfaces. J Am Chem Soc. 1993;115:10714–10721. | |
Bard AJ, Faulkner LR. Electrochemical Methods: Fundamentals and Applications. New York, NY, USA: Wiley Interscience; 2000. | |
Yogeswaran U, Chen SM. A Review on the electrochemical sensors and biosensors composed of nanowires as sensing material. Sensors. 2008;8(1):290–313. | |
Reddy RK. Nanomonitors: electrical immunoassay for clinical diagnostics implementation of a microfabricated biosensor for the detection of proteins. MSc thesis. Portland, OR, USA: Portland State University; 2007. | |
Adamson A, Gast A. Physical Chemistry of Surfaces. 6th ed. New York, NY, USA: John Wiley & Sons, Inc.; 1997. | |
Bard A. Electroanalytical Chemistry: A Series of Advances. Boca Raton, FL, USA: CRC Press; 1988. | |
Bard A, Parsons R, Jordan J, editors. Standard Potentials in Aqueous Solution. Boca Raton, FL, USA: CRC Press; 1985. | |
Hibbert DB. Introduction to Electrochemistry. London, UK: Macmillan; 1993. | |
Bondar VS, Puzyr AA. Nanodiamonds for biological investigations. Physics of the Solid State. 2004;46:761–763. | |
Guedon P, Livache T, Martin F, et al. Characterization and optimization of a real-time, parallel, label-free, polypyrrole-based DNA sensor by surface plasmon resonance imaging. Anal Chem. 2000;72:6003–6009. | |
Homola J, Yee S, Gauglitz G. Surface plasmon resonance sensors: review. Sens Actuators B Chem. 1999;54:3–15. | |
van den Berg B, Wain R, Dobson CM, Ellis RJ. Macromolecular crowding perturbs protein refolding kinetics: implications for protein folding inside the cell. EMBO J. 2000;19:3870–3875. | |
Goluch ED, Stoeva SI, Lee JS, Shaikh KA, Mirkin CA, Liu C. A microfluidic detection system based upon a surface immobilized biobarcode assay. Biosens Bioelectron. 2009;24:2397–2403. | |
Hahn MA, Tabb JS, Krauss TD. Detection of single bacterial pathogens with semiconductor quantum dots. Anal Chem. 2005;77:4861–4869. | |
Gooding JJ. Nanoscale biosensors: significant advantages over larger devices? Small. 2006;2:313–315. | |
Ramanathan K, Bangar MA, Yun M, Chen W, Myung NV, Mulchandani A. Bioaffinity sensing using biologically functionalized conducting-polymer nanowire. J Am Chem Soc. 2005;127:496–497. | |
Cui Y, Lieber CM. Functional nanoscale electronic devices assembled using silicon nanowire building blocks. Science. 2001;291:851–853. | |
Desai T, Hansford D, Ferrari M. Characterization of micromachined silicon membranes for immunoisolation and bioseparation applications. J Memb Sci. 1999;159:221–231. | |
Desai TA, Hansford DJ, Ferrari M. Micromachined interfaces: new approaches in cell immunoisolation and biomolecular separation. Biomol Eng. 2000;17:23–36. | |
Desai T, Hansford D, Kulinsky L, et al. Nanopore technology for biomedical applications. Biomed Microdevices. 1999;2:11–40. | |
Sheehan PE, Whitman LJ. Detection limits for nanoscale biosensors. Nano Lett. 2005;5:803–807. | |
Zimmerman SB, Minton AP. Macromolecular crowding: biochemical, biophysical, and physiological consequences. Annu Rev Biophys Biomol Struct. 1993;22:27–65. | |
Zimmerman SB, Trach SO. Estimation of macromolecule concentrations and excluded volume effects for the cytoplasm of Escherichia coli. J Mol Biol. 1991;222:599–620. | |
Ellis RJ. Macromolecular crowding: obvious but underappreciated. Trends Biochem Sci. 2001;26:597–604. | |
Dobson CM. Chemical space and biology. Nature. 2004;432: 824–828. | |
Minton AP. Models for excluded volume interaction between an unfolded protein and rigid macromolecular cosolutes: macromolecular crowding and protein stability revisited. Biophys J. 2005;88: 971–985. | |
Ellis RJ. Molecular chaperones: assisting assembly in addition to folding. Trends Biochem Sci. 2006;31:395–401. | |
Ellis RJ, Minton AP. Cell biology: join the crowd. Nature. 2003;425: 27–28. | |
Ellis RJ, Pinheiro TJ. Medicine: Danger-misfolding proteins. Nature. 2002;416:483–484. | |
Hall D, Minton AP. Macromolecular crowding: qualitative and semiquantitative successes, quantitative challenges. Biochim Biophys Acta. 2003;1649:127–139. | |
Minton AP. Macromolecular crowding. Curr Biol. 2006;16: R269–R271. | |
van den Berg B, Ellis RJ, Dobson CM. Effects of macromolecular crowding on protein folding and aggregation. EMBO J. 1999;18: 6927–6933. | |
Baszkin A, Norde W. Physical Chemistry of Biological Interfaces. New York, NY, USA: MarcelDekker; 2000. | |
Jain KK. Role of nanodiagnostics in personalized cancer therapy. Clin Lab Med. 2012;32:15–31. | |
Jain KK. Synthetic biology and personalized medicine. Med Princ Pract. 2013;22:209–219. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.