Back to Journals » Infection and Drug Resistance » Volume 11
Multicenter prospective study on the prevalence of colistin resistance in Escherichia coli: relevance of mcr-1-positive clinical isolates in Lombardy, Northern Italy
Authors Principe L, Piazza A, Mauri C, Anesi A, Bracco S, Brigante G, Casari E , Agrappi C , Caltagirone M, Novazzi F, Migliavacca R, Pagani L, Luzzaro F
Received 21 December 2017
Accepted for publication 18 January 2018
Published 9 March 2018 Volume 2018:11 Pages 377—385
DOI https://doi.org/10.2147/IDR.S160489
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Luigi Principe,1 Aurora Piazza,2,3 Carola Mauri,1 Adriano Anesi,4 Silvia Bracco,5 Gioconda Brigante,6 Erminia Casari,7 Carlo Agrappi,8 Mariasofia Caltagirone,2 Federica Novazzi,2 Roberta Migliavacca,2 Laura Pagani,2 Francesco Luzzaro1
1Microbiology and Virology Unit, A. Manzoni Hospital, Lecco, Italy; 2Clinical-Surgical, Diagnostic and Pediatric Sciences Department, Unit of Microbiology and Clinical Microbiology, University of Pavia, Pavia, Italy; 3Romeo and Enrica Invernizzi Pediatric Research Center, Department of Biomedical and Clinical Sciences, University of Milan, Milan, Italy; 4Clinical Pathology Laboratory, ASST Lodi, Lodi, Italy; 5Clinical Pathology Laboratory, ASST Vimercate, Vimercate, Italy; 6Clinical Pathology Laboratory, ASST Valle Olona, Busto Arsizio, Italy; 7Clinical Pathology Laboratory, IRCCS “Humanitas,” Rozzano, Italy; 8Microbiology and Virology Unit, ASST Ovest Milanese, Legnano, Italy
Background: The emergence of the plasmid-mediated colistin resistance mechanism in Escherichia coli has raised concern among public health experts as colistin is a last-line antimicrobial resort. The primary aim of the study was to investigate the prevalence of this resistance trait in E. coli isolates circulating in the Lombardy region, Northern Italy. The presence of mcr-type genes and their genetic relationship were also studied.
Materials and methods: A prospective study was performed during a 4-month period (May to August, 2016) in six acute care Hospitals. Consecutive nonduplicate clinical isolates of E. coli from any type of clinical specimen, with the exception of rectal swabs, were included in the study. Isolates that exhibited MIC values for colistin >2 mg/L were further investigated. Bacterial identification was obtained by matrix-assisted laser desorption ionization-time of flight mass spectrometry. Amplification of mcr-type genes (−1 to −5 variants) and microarray analysis were accomplished. Repetitive sequence-based PCR (Rep-PCR) and multilocus sequence typing (MLST) analysis were used for genotyping.
Results: Overall, 3,902 consecutive E. coli isolates (2,342 from outpatients, 1,560 from inpatients) were evaluated during the study period. Of them, 18/3,902 (0.5%), collected from 4/6 centers, showed resistance to colistin. These isolates were mostly obtained from urine of both outpatients (n=12) and inpatients (n=6). Colistin MIC values ranged from 4 to 8 mg/L. The mcr-1 gene was detected in 10/18 isolates (7 from outpatients, 3 from inpatients). Rep-PCR and MLST analysis revealed the presence of nine different clusters. Further mcr-type genes were not detected.
Conclusion: Resistance to colistin in E. coli clinical isolates appears low in our geographic area. With regard to mcr-1-positive isolates, they accounted for approximately 50% of colistin-resistant E. coli isolates, thus representing a relevant resistance mechanism in this context. Although overall limited, the presence of mcr-1 determinant in our region should not be ignored and great concern should be given to the continuous surveillance.
Keywords: MCR-1, colistin, Escherichia coli, prevalence, surveillance, epidemiology
Introduction
The increasing role of colistin in humans as a last antimicrobial resort in the treatment of infections caused by carbapenem-resistant Enterobacteriaceae has prompted more accurate and careful monitoring of resistance to this polypeptide.1 To this regard, the recent emergence of the plasmid-mediated colistin resistance encoded by mcr-1 in Escherichia coli has raised concern among public health experts worldwide.2 Due to its ability to transfer itself among bacterial strains and species by mobile genetic elements, the mcr-1 determinant could make real the nightmare of bacterial isolates resistant to all classes of antibiotics. In line with this worrisome prospect, mcr-1 gene has been also detected in Klebsiella pneumoniae, Salmonella spp., Enterobacter spp., Citrobacter spp., and Shigella spp., and sometimes associated with carbapenemase or extended-spectrum beta-lactamase (ESBL) producers.1,3–5
After its first description in People’s Republic of China (2015) during the routine surveillance of food animals, the mcr-1 gene has been reported (often retrospectively) across a wide geographic area, comprising 39 countries, in human, animal, and food-related samples.4,6 The first known MCR-1-producing isolate was from the 1980s and was detected in E. coli in People’s Republic of China from animal source, while in humans it was isolated in 2012 from the blood.7,8 In Europe, the first MCR-1-producing strain was an E. coli isolated in France from animal sources in 2005.9 Subsequently, the mcr-2 plasmid-mediated colistin resistance gene was detected from porcine and bovine E. coli in Belgium,10 and a variant of the mcr-1 determinant (named mcr-1.2) was isolated from the rectal swab of an Italian child in K. pneumoniae.11 A third mobile colistin resistance gene, mcr-3, has been reported in E. coli, Aeromonas spp., and Salmonella spp. isolates from human and animal samples,12–16 whereas the mcr-4 and mcr-5 genes were detected in Salmonella spp. and E. coli isolates, but only from animal sources.17–19 In summary, plasmid-mediated resistance to colistin had been around for more than 25 years, but without being detected until 2015.
The history of plasmid-mediated resistance to colistin had a very important veterinary component. Although colistin has been used in clinical settings in a limited manner in the past, due to its nephrotoxicity, its use in veterinary medicine has been carried on for decades (as it was so far a cheap antibiotic).1,20 The main indications for colistin use in veterinary setting are the prevention and treatment of infections caused by Enterobacteriaceae (especially gastrointestinal disorders), but it has been used as growth promoter in terrestrial and aquatic animals.20,21 Data regarding colistin resistance in bacteria from animals and food of animal origin are relatively scarce. Prevalence of colistin resistance in E. coli from animals (pigs, ruminants, poultries, and companion animals) shows wide differences ranging from 0% to 52.4%, with highest resistance percentages reported from Asia.20 It has been reported in several studies that the E. coli colistin resistance rate is higher in pigs compared with other animal productions.6,21–23 Not all studies recognized the colistin resistance mechanism, and so the real prevalence of mcr-1 determinant in the veterinary setting remains still largely underestimated.20 In this scenario, due to the high rate of colistin-resistant (CR) E. coli carrying the mcr-1 gene isolated from food animals compared with humans, livestock production was pinpointed as the greatest cause of colistin resistance amplification and spread, also in humans.6,21 This source of infection led to consider MCR-1-producing E. coli mostly as a community-associated microorganism, being isolated especially in outpatient samples. In this context, several publications have reported the detection of CR E. coli from healthy individuals without prior colistin usage.21,24–27 The observation of colistin resistance in humans without prior colistin exposure is of particular clinical importance and concern, because an antimicrobial stewardship program based on preservation of colistin in the hospital context could not be enough. However, mcr-1-positive E. coli has been almost never associated to hospital epidemic events, giving the reason to think to multi-variegate source of infection outside the hospital setting.
In Italy, data regarding the diffusion of E. coli clinical isolates harboring plasmid-mediated resistance to colistin are very scarce. The mcr-1 determinant was firstly described in 2016 in eight E. coli isolates collected from clinical specimens during the period 2013–2015 in two hospitals.28 Later, another study reported the presence of three E. coli isolates producing both MCR-1 and CTX-M-type ESBL enzymes as intestinal carriage in long-term care facilities residents, during a point prevalence survey on ESBL-producing Enterobacteriaceae.29 More recently, three cases of bloodstream infections caused by MCR-1-producing E. coli were reported among oncologic patients,30 whereas 37 out of 51 (72.5%) CR E. coli isolates from pigs were positive for mcr-1 gene.31 Finally, the mcr-1 determinant was detected in S. enterica isolates obtained from human and animals in the period 2012–2015,32 and the more recent mcr-4 gene was detected in S. enterica serovar Typhimurium (collected in 2013 and retrospectively studied) from an animal source.17 The aim of our study was to investigate 1) the prevalence of this resistance trait in E. coli isolates from clinical samples, 2) the presence of mcr-type genes, and 3) their genetic relationship. Our work represents the first evaluation of the diffusion of clinical mcr-1-positive E. coli in a specific defined area in our country.
Materials and methods
Study design and participating centers
Bacterial isolates were obtained during a multicenter prospective study that involved six clinical microbiology laboratories located in the Lombardy region (Northern Italy). The following centers were included: Busto Arsizio, Lecco, Legnano, Lodi, Rozzano, and Vimercate (Figure 1). Participating hospitals had approximately 4,000 beds and served 2,400,000 people. The survey was conducted over a 4-month period, starting in May 2016. Consecutive nonduplicate clinical isolates of E. coli from any type of clinical specimen, with the exception of rectal swabs, were included in the study. Isolates that exhibited MIC values for colistin >2 mg/L were further investigated. Bacterial identification and antimicrobial susceptibility testing were routinely carried out by the collecting laboratories using either the Phoenix automated system (Becton Dickinson Diagnostic Systems, Sparks, MD, USA) or the Vitek2 system (bioMérieux, Marcy l’Etoile, France). Both inpatients and outpatients were included in the study. Outpatients were defined as patients not hospitalized at the time of specimen collection. For each isolate, information on the clinical specimen and type of ward (in the case of isolates from inpatients) was included. Moreover, each participating laboratory provided information on the total number of consecutive nonduplicate clinical isolates of E. coli observed during the collection period. The collected isolates were sent to reference laboratories for confirmation of both species identification and antimicrobial resistance. Characterization of the colistin resistance mechanism(s) and analysis of clonal relatedness were also carried out.
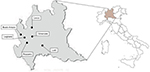  | Figure 1 Participating centers in the Lombardy region, Italy. |
Characterization of bacterial isolates
Bacterial identification of collected isolates was assessed by matrix-assisted laser desorption ionization-time of flight mass spectrometry (Vitek MS; bioMérieux). Antimicrobial susceptibility for colistin was evaluated by the reference broth microdilution method using a dedicated TREK panel (DKMGN; Thermo Fisher Diagnostics, Milan, Italy). This panel also provided MIC values for amoxicillin-clavulanate, piperacillin-tazobactam, cefotaxime, ceftazidime, aztreonam, ertapenem, imipenem, meropenem, ciprofloxacin, amikacin, gentamicin, tobramycin, trimethoprim-sulfamethoxazole, ceftolozane-tazobactam, ceftazidime-avibactam, and tigecycline. All collected isolates confirmed to be resistant to colistin according to EUCAST breakpoints33 (MIC value >2 mg/L) were evaluated for the presence of mcr-type genes.
Characterization of antimicrobial resistance determinants
The presence of the mcr-type determinants (−1 to −5 variants) was investigated by PCR using specific primers and conditions, as previously described.6,10,12,17,18
The content of the entire beta-lactamase resistance determinants of the mcr-type-positive isolates was tested by the Check-MDR CT103XL array (Check-Points Health B.V., Wageningen, The Netherlands).
Molecular typing
Repetitive sequence-based PCR (rep-PCR) was performed with the Diversilab (DL) System (bioMérieux), according to the manufacturer’s instructions. DNA extraction was performed with the UltraClean Microbial DNA isolation kit (Mo Bio Laboratories Inc). Analysis of the PCR amplicons was performed using a 2100 Bioanalyzer (Agilent Technologies , Santa Clara, CA, USA). DL fingerprints were analyzed with the DL software 3.4, using the Pearson correlation statistical method to determine clonal relationships.
Multilocus sequence typing (MLST) of mcr-1-positive E. coli isolates was carried out according to the protocol of Wirth et al. (2006).34 Allelic profiling and sequence-type (ST) determination were performed using the E. coli MLST scheme from the website of the University of Warwick (http://mlst.warwick.ac.uk/mlst/dbs/Ecoli). Phylogenetic groups were determined by a 2-step triplex PCR as described by Clermont et al.35
Plasmid incompatibility groups of mcr-1-positive strains were determined by the PCR-based replicon typing (PBRT) method using the commercially available PBRT kit (Diatheva),36 according to the manufacturer’s instructions. Specific primers and protocol were used for the amplification of IncX4 replicon.37
Ethics approval and consent to participate
Ethics approval and consent to participate were not required. Samples were taken from six different institutions as part of the standard patient care and used anonymously.
Results
Bacterial isolates and epidemiological data
A total of 3,902 consecutive nonduplicate E. coli clinical isolates (outpatients, n=2,342; inpatients, n=1,560) were evaluated during the collection period. Of note, E. coli isolates obtained from patients admitted to nursing homes (included in the outpatients group) accounted for 5.8% (n=135) of study isolates. Clinical isolates were mostly obtained from urine samples (n=3070, 78.7%), followed by skin and soft tissues (n=316, 8.1%), and blood cultures (n=301, 7.7%).
Overall, 18 out of 3,902 (0.5%) isolates, collected from 4/6 centers, were confirmed as CR (MIC>2 mg/L). In particular, 6/18 were from inpatients and 12/18 from outpatients (no one from nursing homes). Thus, the prevalence of colistin resistance was 0.5% (6/1560) and 0.4% (12/2342) among inpatients and outpatients, respectively. Particularly, CR isolates recovered from hospitalized patients came from medical (n=3), rehabilitation (n=2), and surgical (n=1) wards. Overall, CR isolates were obtained from patients (male, n=8; female, n=10) aging from 52 to 94 years, mostly from urine samples (n=16), while the remaining isolates were from blood cultures (n=2).
Molecular characterization and genetic relationship among mcr-1-positive E. coli clinical isolates
PCR analysis detected the mcr-1 gene in 10/18 CR isolates, all of which were from urine samples (seven from outpatients and three from inpatients). Isolates were uniformly negative for other mcr-type genes. Genetic relationship among mcr-1-positive isolates was investigated using different methods. The Rep-PCR technique showed the presence of nine different clusters (data not shown). These data agreed with MLST analysis that revealed nine different STs, with a new one consisting of the following allelic profile: 6–23–5–8–24–18–6. The phylogenetic group analysis showed high heterogeneity among isolates: four belonged to groups A and D, respectively, whereas the remaining two strains were from groups B1 and B2. Seven mcr-1-positive isolates harbored a plasmid of IncX4 group; in three cases, the IncHI2 incompatibility group was found. Details are reported in Table 1.
Antimicrobial susceptibility of CR isolates and associated resistance mechanisms
As shown in Table 2, mcr-1-positive isolates showed MIC values for colistin ranging from 4 to 8 mg/L. These isolates were frequently resistant to co-trimoxazole (8/10) and ciprofloxacin (8/10), and sometimes also to gentamicin (3/10) and tobramycin (3/10). Notably, two of them (both positive for the SHV-12 determinant) were not susceptible to third-generation cephalosporins (cefotaxime and ceftazidime). In all cases, however, carbapenems (ertapenem, imipenem, and meropenem), amikacin, ceftazidime/avibactam, ceftolozane/tazobactam, and tigecycline maintained their activity. As assessed by microarray analysis, six isolates co-harbored the blaTEM-1 gene (Table 1).
Similarly to mcr-1-positive isolates, mcr-type-negative isolates had MIC values for colistin ranging from 4 to 8 mg/L (Table 3). With the exception of ciprofloxacin (4/8 isolates), resistance to other antimicrobials was overall rare, even though three of them were not susceptible to third-generation cephalosporins (cefotaxime and ceftazidime) due to ESBL production.
Discussion
Colistin is increasingly used as one of the last available treatment options for patients with severe infections caused by carbapenem-resistant Enterobacteriaceae, Acinetobacter baumannii, and Pseudomonas aeruginosa.2,38 Colistin resistance follows the increasing trend in consumption of colistin in human medicine, especially in countries with high rates of carbapenem-resistant gram-negative bacilli, including Italy.39 Chromosomally mediated resistance, often generated by mutations in the mgrB gene and upregulation of PhoP/PhoQ system, seems to be related to this trend and mostly associated with K. pneumoniae in the hospital setting.40–43 A mutation in the pmrB gene has been also recently described in E. coli.44 On the contrary, the plasmid-mediated colistin resistance (mostly due to the mcr-1 determinant) has been especially found in E. coli, a common cause of urinary tract infections in healthy individuals in the community setting without prior colistin usage.21,24–27
Prevalence data on colistin resistance are overall scarce. In particular, data regarding the plasmid-mediated resistance to colistin among clinical isolates of E. coli are lacking in Italy. This prospective multicenter study represents the first evaluation on the dissemination of clinical isolates of mcr-1-positive E. coli in Lombardy, the most inhabited Italian region, accounting for about 10 million of residents. Our results show that resistance to colistin in E. coli clinical isolates is almost low in this area (0.5%), with similar percentages among both inpatients and outpatients (0.5% and 0.4%, respectively). Notably, considering only outpatients, resistance to colistin was not detected in nursing home patients, thus enforcing the theory of a major risk source outside the health-care setting.1,3,4,20,21,24,38
As a limitation of the study, however, it should be taken into account that some methodological difficulties affect automated systems in determining the correct MIC value for colistin, especially when it ranges from 1 to 2 mg/L. This issue could lead to a possible underestimation of colistin resistance.
With regard to mcr-1-positive isolates, they accounted for approximately 50% of CR E. coli isolates, thus representing a relevant mechanism in the context of colistin resistance. Overall, however, these isolates represented a low rate (10 isolates, 0.2%) of total isolates studied in the survey. These results are similar to the previously published prevalence data, ranging from 0.05% to 1%.6,45–52 The aforementioned studies included isolates from infected or colonized patients and showed higher prevalence rates in Asian countries compared with those reported from Europe, thus highlighting a major concern toward mcr-related colistin resistance in that geographic area. This issue is reinforced by a high prevalence value of 3.5% described in a report including colonized patients from People’s Republic of China.53
These data, showing a low prevalence of mcr-1-positive isolates, are mostly reassuring since mcr-1 appears as a transferable resistance determinant capable of limited propensity to spread so far. To date, it was never associated with epidemic events, even though association of mcr-type determinants with high-risk clones (e.g., E. coli ST131) capable of large diffusion has been described.16,28 Moreover, other resistance determinants (including those responsible for carbapenemase and ESBL production) have been already reported in association with mcr genes, mainly limiting therapeutic options really effective against these strains.54,55
As previously reported, mcr-1-positive isolates usually show a multi-susceptible profile.1,6,28 In our study, resistance to co-trimoxazole (8/10 isolates) and ciprofloxacin (8/10 isolates) was common. Interestingly, mcr-1-positive isolates were detected only in urine samples. Furthermore, 2/10 isolates were resistant to third-generation cephalosporins (cefotaxime and ceftazidime) due to ESBL production. This worrisome finding could essentially reflect Italian epidemiology for ESBL production in E. coli isolates circulating among both inpatients and outpatients.56
All mcr-1-positive isolates were genetically unrelated, as demonstrated by molecular typing. Both Rep-PCR and MLST revealed nine different clusters, giving the reason to assess a multi-variegate source of infection. Only one couple of isolates was genetically related despite these isolates had been collected from different centers and had no obvious epidemiological link.
In conclusion, we can speculate that the prevalence of CR E. coli isolates is low in our region, and the diffusion of mcr-1 determinant is very limited among clinical isolates. No epidemic events caused by CR E. coli are so far described in Italy in the hospital setting, thus highlighting the community origin of these isolates. Accordingly, in our experience, mcr-1-positive strains were not genetically related and were mostly isolated from outpatients, evidencing their different sources and the low-level diffusion in the community. Although limited, the presence of mcr-1 determinant in our region should not be ignored. Great concern should be given to continuous surveillance, improving prevalence data in both human and veterinary settings in our country.
Ethics approval and consent to participate
Ethics approval and consent to participate were not required. Samples were taken from six different institutions as part of the standard patient care and used anonymously.
Acknowledgments
The abstract of this paper was presented at the 27th ECCMID Congress 2017, Vienna (Austria), as a poster presentation, with interim findings. The poster’s abstract was published in “ESCMID eLibrary” (poster code P0697, year 2017), available from: https://www.escmid.org/escmid_publications/escmid_elibrary.
Disclosure
The authors report no conflicts of interest in this work.
References
Nordmann P, Poirel L. Plasmid-mediated colistin resistance: an additional antibiotic resistance menace. Clin Microbiol Infect. 2016;22(5):398–400. | ||
Poirel L, Jayol A, Nordmann P. Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin Microbiol Rev. 2017;30(2):557–596. | ||
Schwarz S, Johnson AP. Transferable resistance to colistin: a new but old threat. J Antimicrob Chemother. 2016;71(8):2066–2070. | ||
Al-Tawfiq JA, Laxminarayan R, Mendelson M. How should we respond to the emergence of plasmid-mediated colistin resistance in humans and animals? Int J Infect Dis. 2017;54:77–84. | ||
Sennati S, Di Pilato V, Riccobono E, et al. Citrobacter braakii carrying plasmid-borne mcr-1 colistin resistance gene from ready-to-eat food from a market in the Chaco region of Bolivia. J Antimicrob Chemother. 2017;72(7):2127–2129. | ||
Liu YY, Wang Y, Walsh TR, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16(2):161–168. | ||
Shen Z, Wang Y, Shen Y, Shen J, Wu C. Early emergence of mcr-1 in Escherichia coli from food-producing animals. Lancet Infect Dis. 2016;16(3):293. | ||
Rapoport M, Faccone D, Pasteran F, Ceriana P, Albornoz E, Petroni A; MCR Group, Corso A. First description of mcr-1-mediated colistin resistance in human infections caused by Escherichia coli in Latin America. Antimicrob Agents Chemother. 2016;60(7):4412–4413. | ||
Haenni M, Poirel L, Kieffer N, et al. Co-occurrence of extended spectrum β lactamase and MCR-1 encoding genes on plasmids. Lancet Infect Dis. 2016;16(3):281–282. | ||
Xavier BB, Lammens C, Ruhal R, et al. Identification of a novel plasmid-mediated colistin-resistance gene, mcr-2, in Escherichia coli, Belgium, June 2016. Euro Surveill. 2016;21(27):30280. | ||
Di Pilato V, Arena F, Tascini C, et al. mcr-1.2, a new mcr variant carried on a transferable plasmid from a colistin-resistant KPC carbapenemase-producing Klebsiella pneumoniae strain of sequence type 512. Antimicrob Agents Chemother. 2016;60(9):5612–5615. | ||
Yin W, Li H, Shen Y, et al. Novel plasmid-mediated colistin resistance gene mcr-3 in Escherichia coli. MBio. 2017;8(4):e00543-17. | ||
Ling Z, Yin W, Li H, et al. Chromosome-mediated mcr-3 variants in Aeromonas veronii from chicken meat. Antimicrob Agents Chemother. 2017;61(11):e01272-17. | ||
Hernández M, Iglesias MR, Rodríguez-Lázaro D, et al. Co-occurrence of colistin-resistance genes mcr-1 and mcr-3 among multidrug-resistant Escherichia coli isolated from cattle, Spain, September 2015. Euro Surveill. 2017;22(31):30586. | ||
Litrup E, Kiil K, Hammerum AM, Roer L, Nielsen EM, Torpdahl M. Plasmid-borne colistin resistance gene mcr-3 in Salmonella isolates from human infections, Denmark, 2009-2017. Euro Surveill. 2017;22(31):30587. | ||
Roer L, Hansen F, Stegger M, Sönksen UW, Hasman H, Hammerum AM. Novel mcr-3 variant, encoding mobile colistin resistance, in an ST131 Escherichia coli isolate from bloodstream infection, Denmark, 2014. Euro Surveill. 2017;22(31):22846. | ||
Carattoli A, Villa L, Feudi C, et al. Novel plasmid-mediated colistin resistance mcr-4 gene in Salmonella and Escherichia coli, Italy 2013, Spain and Belgium, 2015 to 2016. Euro Surveill. 2017;22(31):30589. | ||
Borowiak M, Fischer J, Hammerl JA, Hendriksen RS, Szabo I, Malorny B. Identification of a novel transposon-associated phosphoethanolamine transferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J Antimicrob Chemother. 2017;72(12):3317–3324. | ||
Fukuda A, Sato T, Shinagawa M, et al. High prevalence of mcr-1, mcr-3 and mcr-5 in Escherichia coli derived from diseased pigs in Japan. Int J Antimicrob Agents. 2018;51(1):163–164. | ||
Kempf I, Jouy E, Chauvin C. Colistin use and colistin resistance in bacteria from animals. Int J Antimicrob Agents. 2016;48(6):598–606. | ||
Rhouma M, Beaudry F, Letellier A. Resistance to colistin: what is the fate for this antibiotic in pig production? Int J Antimicrob Agents. 2016;48(2):119–126. | ||
de Jong A, Thomas V, Simjee S, et al. Pan-European monitoring of susceptibility to human-use antimicrobial agents in enteric bacteria isolated from healthy food-producing animals. J Antimicrob Chemother. 2012;67(3):638–651. | ||
Malhotra-Kumar S, Xavier BB, Das AJ, Lammens C, Butaye P, Goossens H. Colistin resistance gene mcr-1 harboured on a multidrug resistant plasmid. Lancet Infect Dis. 2016;16(3):283–284. | ||
Olaitan AO, Morand S, Rolain JM. Emergence of colistin-resistant bacteria in humans without colistin usage: a new worry and cause for vigilance. Int J Antimicrob Agents. 2016;47(1):1–3. | ||
Prim N, Rivera A, Español M, Mirelis B, Coll P. In vivo adaptive resistance to colistin in Escherichia coli isolates. Clin Infect Dis. 2015;61(10):1628–1629. | ||
Olaitan AO, Thongmalayvong B, Akkhavong K, et al. Clonal transmission of a colistin-resistant Escherichia coli from a domesticated pig to a human in Laos. J Antimicrob Chemother. 2015;70(12):3402–3404. | ||
Urban C, Tiruvury H, Mariano N, Colon-Urban R, Rahal JJ. Polymyxin-resistant clinical isolates of Escherichia coli. Antimicrob Agents Chemother. 2011;55(1):388–389. | ||
Cannatelli A, Giani T, Antonelli A, Principe L, Luzzaro F, Rossolini GM. First detection of the mcr-1 colistin resistance gene in Escherichia coli in Italy. Antimicrob Agents Chemother. 2016;60(5):3257–3258. | ||
Giufrè M, Monaco M, Accogli M, Pantosti A, Cerquetti M; PAMURSA Study Group. Emergence of the colistin resistance mcr-1 determinant in commensal Escherichia coli from residents of long-term-care facilities in Italy. J Antimicrob Chemother. 2016;71(8):2329–2331. | ||
Corbella M, Mariani B, Ferrari C, et al. Three cases of mcr-1-positive colistin-resistant Escherichia coli bloodstream infections in Italy, August 2016 to January 2017. Euro Surveill. 2017;22(16):30517. | ||
Curcio L, Luppi A, Bonilauri P, et al. Detection of the colistin resistance gene mcr-1 in pathogenic Escherichia coli from pigs affected by post-weaning diarrhoea in Italy. J Glob Antimicrob Resist. 2017;10:80–83. | ||
Carnevali C, Morganti M, Scaltriti E, Bolzoni L, Pongolini S, Casadei G. Occurrence of mcr-1 in colistin-resistant Salmonella enterica isolates recovered from humans and animals in Italy, 2012 to 2015. Antimicrob Agents Chemother. 2016;60(12):7532–7534. | ||
The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 7.1, 2017. Available from: http://www.eucast.org. Accessed December 15, 2017. | ||
Wirth T, Falush D, Lan R, et al. Sex and virulence in Escherichia coli: an evolutionary perspective. Mol Microbiol. 2006;60(5):1136–1151. | ||
Clermont O, Bonacorsi S, Bingen E. Rapid and simple determination of the Escherichia coli phylogenetic group. Appl Environ Microbiol. 2000;66(10):4555–4558. | ||
Carattoli A, Bertini A, Villa L, Falbo V, Hopkins KL, Threlfall EJ. Identification of plasmids by PCR-based replicon typing. J Microbiol Methods. 2005;63(3):219–228. | ||
Johnson TJ, Bielak EM, Fortini D, et al. Expansion of the IncX plasmid family for improved identification and typing of novel plasmids in drug-resistant Enterobacteriaceae. Plasmid. 2012;68(1):43–50. | ||
Catry B, Cavaleri M, Baptiste K, et al. Use of colistin-containing products within the European Union and European Economic Area (EU/EEA): development of resistance in animals and possible impact on human and animal health. Int J Antimicrob Agents. 2015;46(3):297–306. | ||
European Centre for Disease Prevention and Control (ECDC). Plasmid-mediated colistin resistance in Enterobacteriaceae. Stockholm: ECDC; 2016. Available from https://ecdc.europa.eu/sites/portal/files/media/en/publications/Publications/enterobacteriaceae-risk-assessment-diseases-caused-by-antimicrobial-resistant-microorganisms-europe-june-2016.pdf. | ||
Giacobbe DR, Del Bono V, Trecarichi EM, et al; ISGRI-SITA (Italian Study Group on Resistant Infections of the Società Italiana Terapia Antinfettiva). Risk factors for bloodstream infections due to colistin-resistant KPC-producing Klebsiella pneumoniae: results from a multicenter case-control-control study. Clin Microbiol Infect. 2015;21(12):1106. | ||
Giani T, Arena F, Vaggelli G, et al. Large nosocomial outbreak of colistin-resistant, carbapenemase-producing Klebsiella pneumoniae traced to clonal expansion of an mgrB deletion mutant. J Clin Microbiol. 2015;53(10):3341–3344. | ||
Monaco M, Giani T, Raffone M, et al. Colistin resistance superimposed to endemic carbapenem-resistant Klebsiella pneumoniae: a rapidly evolving problem in Italy, November 2013 to April 2014. Euro Surveill. 2014;19(42):20939. | ||
Poirel L, Jayol A, Bontron S, et al. The mgrB gene as a key target for acquired resistance to colistin in Klebsiella pneumoniae. J Antimicrob Chemother. 2015;70(1):75–80. | ||
Cannatelli A, Giani T, Aiezza N, et al. An allelic variant of the PmrB sensor kinase responsible for colistin resistance in an Escherichia coli strain of clinical origin. Sci Rep. 2017;7(1):5071. | ||
Terveer EM, Nijhuis RHT, Crobach MJT, et al. Prevalence of colistin resistance gene (mcr-1) containing Enterobacteriaceae in feces of patients attending a tertiary care hospital and detection of a mcr-1 containing, colistin susceptible E. coli. PLoS One. 2017;12(6):e0178598. | ||
Zhong LL, Phan HTT, Shen C, et al. High rates of human fecal carriage of mcr-1-positive multi-drug resistant Enterobacteriaceae isolates emerge in China in association with successful plasmid families. Clin Infect Dis. Epub 2017 Oct 10. | ||
Wang Y, Tian GB, Zhang R, et al. Prevalence, risk factors, outcomes, and molecular epidemiology of mcr-1-positive Enterobacteriaceae in patients and healthy adults from China: an epidemiological and clinical study. Lancet Infect Dis. 2017;17(4):390–399. | ||
Quan J, Li X, Chen Y, et al. Prevalence of mcr-1 in Escherichia coli and Klebsiella pneumoniae recovered from bloodstream infections in China: a multicentre longitudinal study. Lancet Infect Dis. 2017;17(4):400–410. | ||
Heras-Cañas VJ, López-Cerero L, Díaz de-Alba P, Pascual Á. Low prevalence of mcr-1 positive Enterobacteriaceae isolates in a health area. Enferm Infecc Microbiol Clin. 2017;35(7):467–468. | ||
Kuo SC, Huang WC, Wang HY, Shiau YR, Cheng MF, Lauderdale TL. Colistin resistance gene mcr-1 in Escherichia coli isolates from humans and retail meats, Taiwan. J Antimicrob Chemother. 2016;71(8):2327–2329. | ||
Liassine N, Assouvie L, Descombes MC, et al. Very low prevalence of MCR-1/MCR-2 plasmid-mediated colistin resistance in urinary tract Enterobacteriaceae in Switzerland. Int J Infect Dis. 2016;51:4–5. | ||
Prim N, Rivera A, Rodríguez-Navarro J, et al. Detection of mcr-1 colistin resistance gene in polyclonal Escherichia coli isolates in Barcelona, Spain, 2012 to 2015. Euro Surveill. 2016;21(13):30183. | ||
Bi Z, Berglund B, Sun Q, et al. Prevalence of the mcr-1 colistin resistance gene in extended-spectrum β-lactamase-producing Escherichia coli from human faecal samples collected in 2012 in rural villages in Shandong Province, China. Int J Antimicrob Agents. 2017;49(4):493–497. | ||
Beyrouthy R, Robin F, Lessene A, et al. MCR-1 and OXA-48 in vivo acquisition in KPC-producing Escherichia coli after colistin treatment. Antimicrob Agents Chemother. 2017;61(8):e02540-16. | ||
Savov E, Todorova I, Politi L, et al. Colistin resistance in KPC-2- and SHV-5-producing Klebsiella pneumoniae clinical isolates in Bulgaria. Chemotherapy. 2017;62(6):339–342. | ||
Giani T, Antonelli A, Caltagirone M, et al. Evolving beta-lactamase epidemiology in Enterobacteriaceae from Italian nationwide surveillance, October 2013: KPC-carbapenemase spreading among outpatients. Euro Surveill. 2017;22(31):30583. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.



