Back to Journals » International Journal of General Medicine » Volume 15
Multi-Omics Approaches Identify Necroptosis‑Related Prognostic Signature and Associated Regulatory Axis in Cervical Cancer
Authors Zhan J, Yang F, Ge C, Yu X
Received 18 March 2022
Accepted for publication 26 April 2022
Published 13 May 2022 Volume 2022:15 Pages 4937—4948
DOI https://doi.org/10.2147/IJGM.S366925
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Scott Fraser
JuanMei Zhan, Fenfang Yang, Cenhong Ge, Xiaojia Yu
Department of Medical Examination Center, Affiliated Hangzhou First People’s Hospital, Zhejiang University School of Medicine, Hangzhou City, Zhejiang Province, 310006, People’s Republic of China
Correspondence: Fenfang Yang, Department of Medical Examination Center, Affiliated Hangzhou First People’s Hospital, Zhejiang University School of Medicine, Hangzhou City, Zhejiang Province, 310006, People’s Republic of China, Email [email protected]
Background: Cervical cancer is the fourth most frequent malignancy among women globally, with approximately 604,000 new cases and 341,000 deaths per year. Necroptosis is a newly discovered mechanism of cell death involved in biological behaviors of cancer.
Methods: LASSO Cox regression analysis was conducted to construct a prognostic necroptosis-related signature. lncRNA–miRNA–mRNA regulatory axis was constructed with a ceRNA network. qRT-PCR was performed to verify our result.
Results: A total of 54 necroptosis-related genes were differentially expressed in cervical cancer (all p < 0.05). We also summarized genetic mutation landscape of necroptosis-related genes in cervical cancer. We then developed a necroptosis-related prognostic signature including 13 necroptosis-related genes (ATRX, AXL, DDX58, IDH1, ITPK1, MAP3K7, SLC39A7, TARDBP, TNF, TNFRSF1A, TNFRSF1B, TNFSF10, TRIM11) for cervical cancer. Cervical cancer patients with high riskscore had a poor overall survival (HR = 2.128, p = 0.00194) with an AUC of 0.725, 0.763 and 0.637 in 3-year, 5-year, and 10-year ROC curve. Consensus clustering analysis revealed that all cervical cancer cohort could be divided into three subtypes, which was correlated with different prognosis and immune infiltration (p < 0.05). A PPI network revealed TNF as the hub gene and TNF expression was correlated with immune infiltration (all p < 0.05), microsatellite instability (p < 0.012) and drug sensitivity (p < 0.05). The ceRNA network was performed and identified a lncRNA NUTM2B-AS1/miR-361-5p/TNF regulatory axis for cervical cancer. qRT-PCR result also suggested that TNF was upregulated in cervical cancer (p < 0.001) and associated with a poor overall survival (p = 0.007). Univariate and multivariate analysis demonstrated TNF expression, lymph node metastasis and clinical stage were prognosis factors of cervical cancer patients (p < 0.05).
Conclusion: We developed a necroptosis-related prognostic signature including 13 necroptosis-related genes for cervical cancer. Moreover, we also identified a lncRNA NUTM2B-AS1/miR-361-5p/TNF regulatory axis, which may play a vital role in the progression of cervical cancer. Further studies should be conducted to verify these results.
Keywords: cervical cancer, necroptosis, prognostic signature, NUTM2B-AS1, miR-361-5p
Introduction
Cervical cancer is the fourth most frequent malignancy among women globally, with approximately 604,000 new cases and 341,000 deaths per year.1 Cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC) were the most common two sub-types of cervical cancer. When diagnosed in early stage, cervical cancer could be easily treated.2 The incidence, recurrence and mortality in low-middle-income country were high in the absence of preventative and treatment programs.3 Until now, medical prognostic approaches have not provided a satisfactory biomarker to predict the prognosis of cervical cancer patients accurately enough. Despite human papillomavirus had been identified to be involved in the tumorigenesis and progression of cervical cancer, the specific mechanism was still not clarified. Therefore, it is of great significance to identify prognostic signature and associated potential mechanism of cervical cancer.
Necroptosis is a type of cell death-related mechanism triggering by RIP1, RIP3, and MLKL.4,5 As a crucial pathogenic mediator of human disease, necroptosis played a vital role in Parkinson, Alzheimer, vascular atherosclerosis, and infectious diseases.6 Accumulating studies also suggested that necroptosis was involved in the many biological behaviors in cancer.7 It was suggested that necroptosis could overcome apoptosis resistance and may trigger and amplify antitumor immunity in malignancy treatment.4 Moreover, necroptosis‑related signature could serve as prognosis biomarker in stomach adenocarcinoma.8 However, the role of necroptosis-related genes in the progression and prognosis in cervical cancer was not still clarified.
The Cancer Genome Atlas (TCGA, https://cancergenome.nih.gov/) platform is an open accessible public database containing a mass of clinical and bioinformatics data of over 30 types of human cancer.9 More and more studies suggested database mining as one of the promising approaches to explore the molecular mechanism and associated prognosis biomarkers in human cancer. In our study, we aimed to study prognosis significance of necroptosis-related genes and associated regulatory axis in cervical cancer.
Materials and Methods
Dataset and Preprocessing
We obtained 67 necroptosis-related genes from previous necroptosis-related studies (Supplementary Table 1). We then downloaded the gene expression profile of necroptosis-related genes of CESC from TCGA (n = 303). The gene expression profile of healthy control was obtained from TCGA and the Genotype-Tissue Expression (GTEx). Student’s t-test was conducted to detect the mRNA level of necroptosis-related genes in CESC and the result was visualized by R (version 4.0.3) with packages ggplot 2.
Mutation Landscape and Functional Enrichment Analysis
After obtaining the different expressed necroptosis-related genes, we then explored their mutation landscape in CESC using R software with a “maftools” package. A “cluster profiler” package was conducted to perform gene ontology (GO) analysis and Kyoto encyclopedia of genes and genomes (KEGG) pathways analysis.
Necroptosis-Related Prognostic Signature Analysis
In order to identify prognostic necroptosis-related genes, we performed Kaplan–Meier (KM) analysis with Log rank test calculating the p-values, hazard ratio (HR) and 95% confidence interval (CI). This was followed by construction of necroptosis-related prognostic signature with LASSO cox regression analysis using above identified prognostic biomarkers. The risk score of each patient was the sum of coefficients x necroptosis-related gene expression. The overall survival curve was drawn with KM analysis. Moreover, in order to evaluate predictive performance of this signature, we conducted a Time ROC analysis. The correlation between risk score and immune infiltration was evaluated with Pearson correlation analysis.
Consensus Cluster Analysis
Using above necroptosis-related prognostic signature gene, we performed consensus cluster analysis with “ConsensusClusterPlus” package. And “survival” package was used to explore the difference of each cluster in CESC. The difference of clinical characters of each cluster in CESC was calculated with “pheatmap” package. Moreover, the abundance of immune cells of each cluster in CESC was calculated with ESTIMATE algorithm and the result was visualized with “pheatmap” package.
LncRNA–miRNA–mRNA Regulatory Axis Analysis
To identify the hub gene, we performed a PPI network with STRING (https://string-db.org/). The miRNA targets of hub gene were detected with miRDB (http://mirdb.org/), StarBase (http://starbase.sysu.edu.cn/) and miRWalk (http://mirwalk.umm.uni-heidelberg.de/). The lncRNA targets interacting with miRNA were detected with LncBase (http://carolina.imis.athena-innovation.gr/) and StarBase (http://starbase.sysu.edu.cn/). Furthermore, Student’s t-test and KM test were performed to evaluate the expression and prognosis of miRNA and lncRNA in CESC.
Immune Infiltration and Drug Sensitivity Analysis
The abundance of immune cells was isolated from TIMER (https://cistrome.shinyapps.io/timer/). Pearson correlation test was performed to analyze the correlation of TNF expression and the abundance of immune cells. We collected the IC50 of 265 small molecules in 860 cell lines, and its corresponding TNF expression from Genomics of Drug Sensitivity in Cancer (GDSC). The TNF expression data and drug sensitivity data were merged. Pearson correlation analysis was performed to get the correlation between TNF expression and drug IC50. P-value was adjusted by FDR.
Quantitative Real Time-Polymerase Chain Reaction Analysis
Approved by the Ethics Committee of Affiliated Hangzhou First People’s Hospital and obtained written informed consent from patient, we obtained serum from 60 cases of CESC. Our study was performed followed the guidelines outlined in the Declaration of Helsinki. The clinicopathological characteristics of these CESC patients is shown in Supplementary Table 2. All cases were diagnosed and assessed pathologically. Total RNA was extracted from CESC serum and healthy control serum using TRIzol kit (Vazyme, Nanjing, China). qRT-PCR experiments were used to verify TNF expression in CESC. Firstly, TNF primers were designed using the NCBI website with upstream sequence: 5’—3’ CCCGCATCCCAGGACCTCTCT and downstream sequence 5’—3’ CGGGGGACTGGCGA, and internal reference GAPDH primers were designed with upstream sequence 5’—3’ GGAGCGAGATCCCTCCAAAAT and downstream sequence 5’—3ʹGGCTGTTGTCATACTTCTCATGG, annealed at 60°C.
Results
The Expression of Necroptosis-Related Genes in Cervical Cancer
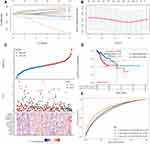
As shown in Figure 1A and B, a total of 54 necroptosis-related genes were differentially expressed in cervical cancer among these 63 necroptosis-related genes. More specifically, 34 necroptosis-related genes were upregulated while 20 necroptosis-related genes were downregulated in cervical cancer (p < 0.05).
Mutation Landscape and Functional Enrichment Analysis
The mutation landscape of necroptosis-related genes in cervical cancer is shown in Figure 2A and B, which revealed that 64% of cervical cancer samples had genetic mutation. ATRX was the gene with the highest frequency of mutation followed by CASP8 and DNMT1 (Figure 2B). Interestingly, missense mutation ranked the top variant classification and C>T was the most common SNV class (Figure 2A). And SNP was the most common variant type (Figure 2A). As shown in Figure 2C, these differentially expressed necroptosis-related genes were involved in necrotic cell death, programmed necrotic cell death, necroptosis process, positive regulation of proteolysis, and tumor necrosis factor receptor superfamily binding in GO analysis. As for KEGG pathway analysis, the data suggested the involvement in necroptosis, apoptosis, TNF signaling pathway, NF-kappa B signaling pathway, NOD-like receptor and IL-17 signaling pathway (Figure 2D).
The Prognostic Significance of Necroptosis-Related Genes in Cervical Cancer
In order to identify potential biomarker among necroptosis-related genes for cervical cancer, we performed overall survival (OS) analysis, disease-specific survival (DSS) analysis, progression-free survival (PFS) analysis and disease-free survival (DFS) analysis. As we could see in Figure 3A, a total of three necroptosis genes, including TRIM11, TNFRSF1A, and TARDBP, had a prognostic significance in OS analysis. In PFS analysis, cervical cancer patients with high expression of TRIM11, IDH1, as well as BNIP3 and low expression of TNFSF10, TNFRSF1B, as well as DDX58 had a poor prognosis (Figure 3B, all p < 0.05). Moreover, cervical cancer patients with high expression of TNF and low expression of DDX58, TNFSF10 as well as TNFRSF1B had a poor disease-specific survival (Figure 3C, all p < 0.05). As shown in Figure 3D, 6 necroptosis genes, including ATRX, AXL, ITPK1, MAP3K7, SLC39A7, and SQSTM1, had a prognostic significance in DFS analysis.
Development of a Necroptosis-Related Prognostic Signature
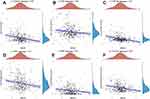
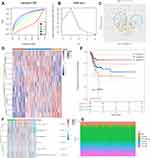
Based on above potential prognosis biomarkers, we then performed LASSO cox regression analysis to develop a necroptosis-related prognostic signature. As a result, a total of 13 necroptosis-related genes (ATRX, AXL, DDX58, IDH1, ITPK1, MAP3K7, SLC39A7, TARDBP, TNF, TNFRSF1A, TNFRSF1B, TNFSF10, TRIM11) were included in the prognostic signature. Figure 4A and B revealed the coefficient and partial likelihood deviance of prognostic signature. Cervical cancer cases were divided into high- and low-risk group according to the riskscore = (0.0285)*ATRX+(0.0073)*AXL+(−0.0665)*DDX58+(0.0514)*IDH1+(−0.0296)*ITPK1+(0.2065)*MAP3K7+(0.0956)*SLC39A7+(−0.4932)*TARDBP+(0.2339)*TNF+(0.3095)*TNFRSF1A+(−0.2588)*TNFRSF1B+(−0.0228)*TNFSF10+(0.2128)*TRIM11. The riskscore, survival status, and gene expression of prognostic signature are shown in Figure 4C. As expected, cervical cancer patients with high riskscore had a poor overall survival (Figure 4D, HR = 2.128, p = 0.00194), with an AUC of 0.725, 0.763 and 0.637 in 3-year, 5-year, and 10-year ROC curve (Figure 4E), demonstrating a good performance of this signature in predicting the prognosis of cervical cancer patients. We also analyzed the correlation between riskscore and immune infiltration, which indicated a negative correlation between the riskscore and the immune infiltration level of B cells (p = 5.21e−6, Figure 5A), CD4+ T cells (p = 0.01, Figure 5B), CD8+ T cells (p = 3.07e−8, Figure 5C), Neutrophils (p = 0.007, Figure 5D), Macrophage (p = 0.003, Figure 5E) and Dendritic cells (p = 4.31e−5, Figure 5F).
Consensus Clustering of Necroptosis-Related Prognostic Genes in Three Clusters in Cervical Cancer
Based on above 13 necroptosis-related prognostic genes, consensus clustering was conducted to cluster cervical cancer patients. As displayed in Figure 6A and B, the k = 3 was identified with optimal clustering stability from k = 2 to 6 based on the similarity displayed by gene expression. As a result, TCGA cervical cancer cohort were classified into cluster 1, cluster 2 and cluster 3 (Figure 6C and D). Among these 3 clusters, cluster 1 patients had a best prognosis and cluster 3 patients had a worst prognosis (Figure 6E, p = 0.0072). Interestingly, these 3 clusters had distinct difference in the immune cell infiltration, including CD4+ T cells, Neutrophils, Macrophage, and Dendritic cells (Figure 6F and G, all p < 0.05).
Hub Gene Analysis
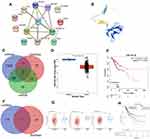
In order to identify the hub gene among the necroptosis-related prognostic signature, we performed a PPI network and the result suggested TNF as the hub gene among necroptosis-related prognostic signature (Figure 7A). The structure of TNF is shown in Figure 7B. Interestingly, further analysis revealed that TNF expression was significantly correlated with the abundance of CD4+ T cell, Macrophage, Neutrophils, and Dendritic cells (Supplementary Figure 1A, all p < 0.05). Increasing evidences suggested TMB and MSI as predictive markers for tumor immunotherapy efficacy in cancer.10,11 In our study, the result suggested a significant correlation between TNF expression and MSI score (Supplementary Figure 1B, p = 0.012) but not TMB score (Supplementary Figure 1C, p = 0.382). To explore the potential of TNF as a drug scanning biomarker for cervical cancer, we then conducted drug sensitivity analysis. The data showed that low expression of TNF were correlated with drug resistance of GDSC (Supplementary Figure 1D).
lncRNA–miRNA–mRNA Regulatory Axis Analysis
In order to further clarify TNF-associated mechanism in cervical cancer, we then performed a lncRNA–miRNA–mRNA regulatory axis analysis. As shown in Figure 7C, miR-361-5p and miR-3666 were identified as the most potential miRNA targets of TNF based on the results of miRDB, miRWalk and StarBase. Further analysis demonstrated that miR-361-5p expression was decreased in cervical cancer (Figure 7D, p = 4.77e−3) and patient with high miR-361-5p level had a better survival (Figure 7E, p = 0.0033). Thus, we suggested miR-361-5p as the miRNA target of TNF. In lncRNA target analysis, a total of four lncRNAs (NUTM2B-AS1, TRIM52-AS1, LINC01468, COX10-AS1) were suggested as the most potential targets of miR-361-5p based on the data of lncBase and Starbase (Figure 7F). Among these four genes, the expression of COX10-AS1, TRIM52-AS1, and NUTM2B-AS1 were downregulated in cervical cancer (Figure 7G, all p < 0.05). However, further prognosis analysis suggested a significant correlation between NUTM2B-AS1 expression and the overall survival of cervical cancer (Figure 7H, p = 0.0485). These results suggested lncRNA NUTM2B-AS1 as the lncRNA target of miR-361-5p. Thus, lncRNA NUTM2B-AS1/miR-361-5p/TNF regulatory axis may play a vital role in the progression in cervical cancer and further in vivo and in vitro studies should be conducted to verify this hypothesis.
The Expression and Prognostic Significance of TNF in Cervical Cancer
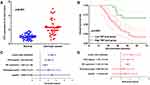
We finally verified the expression and prognostic significance of TNF in cervical cancer. And the data indicated an upregulation of TNF in cervical cancer (Figure 8A, p<0.001) and a poor overall survival in cervical cancer patients with high TNF level (Figure 8B, p=0.007). Considering age, histological grade, FIGO staging, lymph node metastasis and TNF expression, we then performed univariate and multivariate analysis, which demonstrated TNF expression, lymph node metastasis and FIGO stage were prognosis factors of cervical cancer patients (Figure 8C and D).
Discussion
Despite being largely preventable, cervical cancer is still one of most common cancers of women globally and a leading cause of cancer-related deaths in low-middle-income country.12 Worse still, the incidence and mortality of cervical cancer have increased significantly in developing country, especially among young women.13 Until now, medical prognostic approaches have not provided a satisfactory biomarker to predict the prognosis of cervical cancer patients accurately enough. Increasing evidences indicated the prognostic significance of necroptosis-related signature in cancers, including stomach adenocarcinoma.8 However, there is limited study about the role of necroptosis in cervical cancer. Thus, we explored the significance of necroptosis-related gene in the prognosis of cervical cancer.
Gene expression profile suggested upregulation of 34 necroptosis-related genes and downregulation of 20 necroptosis-related genes in cervical cancer. Further functional enrichment analysis indicated that these different expressed necroptosis-related genes were involved in necrotic cell death, programmed necrotic cell death, necroptosis process, positive regulation of proteolysis, TNF signaling pathway, NF-kappa B signaling pathway, NOD-like receptor and IL-17 signaling pathway. In fact, these pathways were proved to be correlated with the progression of cervical cancer. By regulating TNF signaling pathway, miR-195-5p/MMP14 could regulate tumor cell proliferation and invasion in cervical carcinoma.14 Another study revealed that IL-17 promoted the tumorigenesis of cervical cancer via JAK2/STAT3, PI3K/Akt and NF-κB signaling pathway.15
We then performed OS analysis, DSS analysis, PFS analysis and DFS analysis. As a result, a total of 15 potential prognostic biomarkers were identified. Based on these potential prognostic biomarkers, LASSO Cox regression analysis was performed and a necroptosis-related prognostic signature including 13 genes (ATRX, AXL, DDX58, IDH1, ITPK1, MAP3K7, SLC39A7, TARDBP, TNF, TNFRSF1A, TNFRSF1B, TNFSF10, TRIM11) was developed. As expected, this prognostic signature could predict the OS of cervical cancer patients with medium to high accuracy. Actually, several prognostic signatures had been identified for cervical cancer. Jie Mei et al. constructed an immune-related 4 gene signature, which could predict the prognosis of patients with cervical cancer.16 Another six-lncRNA immune prognostic signature had a good performance in prognosis prediction in cervical cancer.17 Based on previous evidences, only two necroptosis‑related prognostic signatures had been identified for human cancers.8,18 Thus, the current study was the first study to identify necroptosis‑related prognostic signature for cervical cancer.
Our study also identified lncRNA NUTM2B-AS1/miR-361-5p/TNF regulatory axis and this regulatory axis may play a significant role in the progression in cervical cancer. Previous study revealed that miR-361-5p could serve as prognostic biomarker and associated with biological processes in cervical cancer. miRNA-361-5p could promote tumor progression via ETM in cervical cancer.19 Moreover, miR-361 level served as an independent biomarker predicting a favorable survival in cervical cancer.20 Moreover, TNF was also involved in the progression of cervical cancer. By regulating TNF, miR-21 could regulate tumor cell proliferation and apoptosis in cervical cancer.21 However, there was limited study about the role of NUTM2B-AS1 in human cancer and lncRNA NUTM2B-AS1/miR-361-5p/TNF regulatory axis has not been studied before. Further in vivo and in vitro studies should be conducted to verify this hypothesis.
Cancer subtype analysis, as an extension of cancer diagnosis, can be regarded as a consensus clustering problem. Consensus clustering analysis and identification different cancer subtype is beneficial for providing patients with more accurate treatment.22 By consensus clustering analysis, we found that all cervical cancer cohort could be divided into three subtypes with different prognosis and immune infiltration, providing potential clinical therapeutic targets based on the specific molecular features. Another consensus clustering analysis identified several subtypes of colorectal cancer had a significant different prognosis and therapy response.23 Pengfei et al identified immune landscape and subtypes in oral squamous cell carcinoma facilitating development of tailed immunotherapeutic strategies.24
There is no doubt that our research has some limitations. Firstly, another dataset should be used to verify the necroptosis-related prognostic signature. Moreover, further in vivo and in vitro studies should be performed to verify lncRNA NUTM2B-AS1/miR-361-5p/TNF regulatory axis.
Conclusion
In conclusion, we developed a necroptosis-related prognostic signature and a lncRNA NUTM2B-AS1/miR-361-5p/TNF regulatory axis for cervical cancer. Further in vivo and in vitro studies should be conducted to verify these results.
Data Sharing Statement
The analyzed data sets generated during the study are available from the corresponding author on reasonable request.
Ethics Approval and Consent to Participate
Our study was approved by the Ethics Committee of Affiliated Hangzhou First People’s Hospital. Informed consent was signed by all participating patients.
Author Contributions
JuanMei Zhan performed data analysis work and aided in writing the manuscript. Fenfang Yang designed the study, assisted in writing the manuscript. Cenhong Ge and Xiaojia Yu edit the manuscript. All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
This study was funded by the research on the application of Internet-health management in medical examination (NO.2021WJCY086).
Disclosure
The authors declare that they have no competing interests in this work.
References
1. Buskwofie A, David-West G. Clare CA: a review of cervical cancer: incidence and disparities. J Natl Med Assoc. 2020;112(2):229–232. doi:10.1016/j.jnma.2020.03.002
2. Pimple SA, Mishra GA. Global strategies for cervical cancer prevention and screening. Minerva Ginecol. 2019;71(4):313–320. doi:10.23736/S0026-4784.19.04397-1
3. Vu M, Yu J, Awolude OA, Chuang L. Cervical cancer worldwide. Curr Probl Cancer. 2018;42(5):457–465. doi:10.1016/j.currproblcancer.2018.06.003
4. Gong Y, Fan Z, Luo G, et al. The role of necroptosis in cancer biology and therapy. Mol Cancer. 2019;18(1):100. doi:10.1186/s12943-019-1029-8
5. Zhe-Wei S, Li-Sha G, Yue-Chun L. The role of necroptosis in cardiovascular disease. Front Pharmacol. 2018;9:721. doi:10.3389/fphar.2018.00721
6. Choi ME, Price DR, Ryter SW, Choi AMK. Necroptosis: a crucial pathogenic mediator of human disease. JCI Insight. 2019;4(15). doi:10.1172/jci.insight.128834
7. Su Z, Yang Z, Xu Y, Chen Y, Yu Q. Apoptosis, autophagy, necroptosis, and cancer metastasis. Mol Cancer. 2015;14:48. doi:10.1186/s12943-015-0321-5
8. Wang N, Liu D. Identification and validation a necroptosis‑related prognostic signature and associated regulatory axis in stomach adenocarcinoma. Onco Targets Ther. 2021;14:5373–5383. doi:10.2147/OTT.S342613
9. Tomczak K, Czerwińska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol. 2015;19(1a):A68–77. doi:10.5114/wo.2014.47136
10. Liu L, Bai X, Wang J, et al. Combination of TMB and CNA stratifies prognostic and predictive responses to immunotherapy across metastatic cancer. Clin Cancer Res. 2019;25(24):7413–7423. doi:10.1158/1078-0432.CCR-19-0558
11. Samstein RM, Lee CH, Shoushtari AN, et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet. 2019;51(2):202–206. doi:10.1038/s41588-018-0312-8
12. Mehrotra R, Yadav K. Cervical Cancer: formulation and Implementation of Govt of India guidelines for screening and management. Indian J Gynecol Oncol. 2022;20(1):4. doi:10.1007/s40944-021-00602-z
13. Feng Y, Wang Y, Xie Y, Wu S, Li Y, Li M. Nomograms predicting the overall survival and cancer-specific survival of patients with stage IIIC1 cervical cancer. BMC Cancer. 2021;21(1):450. doi:10.1186/s12885-021-08209-5
14. Li M, Ren CX, Zhang JM, et al. The effects of miR-195-5p/MMP14 on proliferation and invasion of cervical carcinoma cells through TNF signaling pathway based on bioinformatics analysis of microarray profiling. Cell Physiol Biochem. 2018;50(4):1398–1413. doi:10.1159/000494602
15. Bai Y, Li H, Lv R. Interleukin-17 activates JAK2/STAT3, PI3K/Akt and nuclear factor-κB signaling pathway to promote the tumorigenesis of cervical cancer. Exp Ther Med. 2021;22(5):1291. doi:10.3892/etm.2021.10726
16. Mei J, Xing Y, Lv J, et al. Construction of an immune-related gene signature for prediction of prognosis in patients with cervical cancer. Int Immunopharmacol. 2020;88:106882. doi:10.1016/j.intimp.2020.106882
17. Chen Q, Hu L, Huang D, Chen K, Qiu X, Qiu B. Six-lncRNA immune prognostic signature for cervical cancer. Front Genet. 2020;11:533628.
18. Zhao Z, Liu H, Zhou X, et al. Necroptosis-related lncRNAs: predicting prognosis and the distinction between the cold and hot tumors in gastric cancer. J Oncol. 2021;2021:6718443. doi:10.1155/2021/6718443
19. Wu X, Xi X, Yan Q, et al. MicroRNA-361-5p facilitates cervical cancer progression through mediation of epithelial-to-mesenchymal transition. Med Oncol. 2013;30(4):751. doi:10.1007/s12032-013-0751-0
20. Liu S, Song L, Yao H, et al. Preserved miR-361-3p expression is an independent prognostic indicator of favorable survival in cervical cancer. Dis Markers. 2018;2018:8949606. doi:10.1155/2018/8949606
21. Xu L, Xu Q, Li X, Zhang X. MicroRNA-21 regulates the proliferation and apoptosis of cervical cancer cells via tumor necrosis factor-α. Mol Med Rep. 2017;16(4):4659–4663. doi:10.3892/mmr.2017.7143
22. Li J, Xie L, Xie Y, Wang F. Bregmannian consensus clustering for cancer subtypes analysis. Comput Methods Programs Biomed. 2020;189:105337. doi:10.1016/j.cmpb.2020.105337
23. Sadanandam A, Lyssiotis CA, Homicsko K, et al. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat Med. 2013;19(5):619–625. doi:10.1038/nm.3175
24. Diao P, Jiang Y, Li Y, et al. Immune landscape and subtypes in primary resectable oral squamous cell carcinoma: prognostic significance and predictive of therapeutic response. J Immunother Cancer. 2021;9(6):e002434. doi:10.1136/jitc-2021-002434
 © 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.