Back to Journals » OncoTargets and Therapy » Volume 12
MTF2 Induces Epithelial-Mesenchymal Transition and Progression of Hepatocellular Carcinoma by Transcriptionally Activating Snail
Authors Wu TT, Cai J, Tian YH, Chen JF, Cheng ZL, Pu CS, Shi WZ, Suo XP, Wu XJ, Dou XW, Zhang KM
Received 6 August 2019
Accepted for publication 31 October 2019
Published 17 December 2019 Volume 2019:12 Pages 11207—11220
DOI https://doi.org/10.2147/OTT.S226119
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjay Singh
Tian-Tian Wu,* Jun Cai,* Yuan-Hu Tian, Jian-Fei Chen, Zhi-Lei Cheng, Chang-Sheng Pu, Wen-Zai Shi, Xiao-Peng Suo, Xian-Jia Wu, Xiao-Wei Dou, Ke-Ming Zhang
Department of Hepatobiliary Surgery, Peking University International Hospital, Beijing 102206, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Ke-Ming Zhang
Department of Hepatobiliary Surgery, Peking University International Hospital, No 1 Shengmingyuan Road, Zhongguancun Life Science Park, Changping District, Beijing 102206, People’s Republic of China
Email [email protected]
Background: Metal regulatory transcription factor 2 (MTF2) has been previously reported as a protein binding to the metal response element of the mouse metallothionein promoter, which is involved in chromosome inactivation and pluripotency. However, the function of MTF2 in tumor formation and progression has not yet been completely elucidated.
Methods: The expression of MTF2 and clinicopathological characteristics were evaluated by hepatocellular carcinoma (HCC) tissue microarray of 240 specimens. The role of MTF2 on HCC progression was determined using MTT, crystal violet, and transwell assays. Tumor growth was monitored in a xenograft model, and intrahepatic metastasis models were established.
Results: The expression of MTF2 was increased in HCC and strongly associated with the clinical characteristics and prognosis. Forced expression of MTF2 in HCC cells significantly promoted cell growth, migration, and invasion in vitro. In contrast, downregulation of MTF2 inhibited cell growth, migration, and invasion in vitro. Moreover, knock down of MTF2 suppressed tumorigenesis and intrahepatic metastasis of HCC cells in vivo. Mechanistically, MTF2 overexpression may promote growth and epithelial-mesenchymal transition processes of HCC cells by facilitating Snail transcription.
Conclusion: MTF2 promotes the proliferation, migration, and invasion of HCC cells by regulating Snail transcription, providing a potential therapeutic candidate for patients with HCC.
Keywords: MTF2, progression, hepatocellular carcinoma, EMT, Snail
Introduction
Hepatocellular carcinoma (HCC) is one of the most common malignant tumors, ranked the fifth most common cancer, and is the second largest cause of cancer-related death worldwide.1,2 China has a higher incidence and mortality of HCC than other countries, and the frequent intrahepatic and extrahepatic metastases of HCC are responsible for poor prognosis of patients with HCC.3,4
Recently, emerging evidences revealed that multi-gene mutation is involved in the progression of HCC.5,6 Epithelial-mesenchymal transition (EMT) confers metastatic properties upon cancer cells by enhancing mobility, invasion, and resistance.7,8 EMT is orchestrated by a restricted number of transcription factors including Snail, Twist, and the Zeb family.9,10 Therefore, it is important to understand the mechanism underlying HCC progression and metastasis, which may provide new therapeutic targets for the treatment of patients with HCC.
Metal regulatory transcription factor 2 (MTF2), also named polycomb-like 2 (Pcl2), is a member of the Pcl family of proteins.11 MTF2 has been previously reported as a protein binding to the metal response element of the mouse metallothionein promoter.12,13 MTF2 has been subsequently demonstrated to regulate the PRC2 activity at specific developmental genes, resulting in chromosome inactivation and pluripotency.14,15 In embryonic stem cells, MTF2 knockdown gives rise to reduced H3K27me3 levels at the MTF2 target loci without affecting global H3K27me3 levels.11,16 MTF2 plays its role as a PRC2 accessory protein in embryonic stem cells. However, previous studies have not explored the potential role of MTF2 in the progression of common cancers such as HCC, despite MTF2 being distinctively related to the prognosis and tumor stages of patients with HCC according to The Cancer Genome Atlas (TCGA) database.
In this study, we found that MTF2 expression was upregulated in human HCC tissues and was significantly related to prognosis and clinicopathologic parameters. We further demonstrated that MTF2 promoted proliferation, migration, and invasion abilities of HCC cells both in vitro and in vivo. In addition, we determined that MTF2 could accelerate the EMT of HCC cells by promoting Snail transcription. Finally, this study elucidated the function and mechanism of MTF2 in HCC growth and metastasis, providing a prognostic marker and a potential therapeutic candidate for patients with HCC.
Materials and Methods
HCC Tissue Samples and Tissue Microarray Analysis
After obtaining written informed consent, HCC tissues were obtained from the Peking University International Hospital, Beijing, China. Forty-three pairs of HCC tissues and the corresponding adjacent normal tissues were subjected to RNA extraction for quantitative real-time PCR (qPCR) analysis and protein extraction for western-blot detection was applied to 8 pairs of tissues. To evaluate the expression of MTF2, we examined the tissues array (TMA) which was constructed from 240 paraffin-embedded primary HCC tissues and paired non-tumor liver tissues and analyzed the correlation between MTF2 and clinical features. The protocol for tissue collection was approved by the Ethics Committee of the Peking University International Hospital, Beijing, China. The study was performed in accordance with the Declaration of Helsinki and the guidelines of the committee.
Immunohistochemistry (IHC)
Paraffin-embedded tissue sections were deparaffinized, rehydrated, subjected to antigen retrieval, and the endogenous peroxidases were blocked. After washing the section thrice with 0.01 mol/L PBS (8 mmol/L Na2HPO4, 2 mmol/L NaH2PO4, and 150 mmol/L NaCl), the sections were blocked in 0.01 mol/L PBS containing 0.3% Triton X-100 and 5% BSA. Next, sections were incubated with anti-MTF2 (1:200) antibody (Abcam) at 4 °C overnight. The sections were washed thrice with 0.01 M PBS and incubated with secondary antibody (1:500) for 2 h. After washing with 0.01 mol/L PBS and 0.05 M Tris-HCl (pH 7.6) solution twice, sections were visualized with 0.03% 3,3′-diaminobenzidine in 0.05 M Tris-HCl (pH 7.6). The extent and staining intensity of protein were scored automatically using the Vectra 2 system (Perkin-Elmer, USA). The outcome of staining was determined using the H-score, defined by the equation: H-score = 100ΣPi × i. The staining intensity (i) of the decorated tumor cells was graded from “0 to 3”, and Pi is the percentage of the stained cells at various intensities.
RNA Preparation and qPCR
The total RNA of 43 pairs of tumor tissues and adjacent noncancerous tissues and HCC cells was extracted using TRIzol reagent (Invitrogen). Total RNA (2 μg) was reverse transcribed to cDNA using the PrimeScript RT reagent Kit (Takara) according to the manufacturer’s instructions. SYBR Premix Ex Taq (Takara) was used for performing qPCR. The MTF2-specific primers were 5′-GTCCACCTGGCCCATATACAA-3′ (forward) and 5′-CCGTGAAATCCACATCTGAGG-3′ (reverse), the Snail-specific primers were 5′-CGCCTGACTGAGCAACTGG-3′ (forward) and 5′-GGAAGCTGGCGGTACTTGT-3′ (reverse), and the GAPDH primers were 5′-ATGACCCCTTCATTGACCTCA-3′ (forward) and 5′-GAGATGATCACCCTT TTGGCT-3′ (reverse). GAPDH was used as the internal control.
Cell Culture
HCC cell lines YY-8103, HepG2, Hep3B, MHCC97-H, Huh7, and LM3 were purchased from the Cell Bank of Type Culture Collection of the Chinese Academy of Sciences. All cell lines were maintained at 37 °C in a humidified atmosphere containing 5% CO2. All these cell lines were grown in DMEM medium (Gibco) supplemented with 10% fetal bovine serum (FBS; Invitrogen), and 1% penicillin/streptomycin (Sangong Biotech).
Plasmids and Cell Transfection
The coding sequence for the full length of MTF2 was cloned into the p23-ZsGreen plasmid (Invitrogen) to generate the MTF2 expression vector. Lentiviral short hairpin RNA (shRNA) targeting MTF2 and Snail were designed using software provided by Qiagen (Valencia, CA, USA). HepG2 and YY-8103 cells were infected with p23-ZsGreen-BUB1 lentiviral and Huh7 and LM3 cells were infected with pLKO.1-shRNA. Overexpressed and silenced cells were sorted using flow cytometry or selected using puromycin (4 ug/mL) for 4 days, respectively.
The sequences of shMTF2 were as follows: sh1 5′-CCGGCCACCAAATGTGGCTTTCAAACTCGAGTTTGAAAGCCACATTTGGTGGTTTTTG-3′ (forward) and 5′-AATTCAAAAACCACCAAATGTGGCTTTCAAACTCGAGTTTGAAAGCCACATTTGGTGG −3′ (reverse); sh2 5′-CCGGGCACTTAAGAAAGGACCAAATCTCGAGATTTGGTCCTTTCTTAAGTGCTTTTTG-3′ (forward) and 5′-AATTCAAAAAGCACTTAAGAAAGGACCAAATCTCGAGATTTGGTCCTTTCTTAAGTGC-3′ (reverse). The sequence of shSnail was: 5′-TATAGGTACCTGGGCTGGACAACAGTTCTTTAC-3′ (forward) and 5′-ATATAAGCTTTCGCTGTAGTTAGGCTTCC-3′ (reverse).
Western Blot Analysis
Cells and HCC samples were extracted using RIPA Lysis Buffer and PMSF (Thermo Scientific) according to the manufacturer’s instructions. The lysates were centrifuged at 12,000 rpm for 15 min at 4 °C, and the supernatant was extracted for further study. Before incubation with antibodies, proteins were separated using SDS-PAGE and transferred onto PVDF membranes. Antibodies against MTF2 (1:1000), Snail (1:1000), and Flag (1:4000) were purchased from Abcam; antibodies against Twist (1:1000) and ZEB1 (1:1000) were purchased from ProteinTech; antibodies against E-cadherin (1:1000), N-cadherin (1:1000), Vimentin (1:1000), and GAPDH (1:1000) were purchased from Cell Signaling Technology. Horseradish peroxidase-conjugated secondary antibodies (1:2000) were purchased from Sigma.
MTT Assay
The cell growth in vitro was measured using MTT assay. In the MTT assay, 1000 cells per well were seeded into 96-well plates. Twenty microliters of a 5 mg/mL MTT solution was added into the medium, and the plate was cultured at 37 °C for 4 h. Thereafter, the medium was removed and 200 μL DMSO was added to dissolve the formazan. The absorbance value was measured at a wavelength of 540 nm (OD 540) using an automatic microplate reader. The experimental process was performed every 24 h for 7 days to generate a cell growth curve.
Crystal Violet Assay
The cell growth in vitro was also measured by crystal violet assay. One thousand cells per well were seeded into 6-well plates, and the cells were cultured in a medium with 10% FBS. The medium was changed every 3 days, and cells were stained with 1 mL 0.5% crystal violet solution in 20% methanol after 2 weeks. Next, the fixed cells were washed with PBS and photographed. Glacial acetic acid (1 mL) was added to dissolve the cells, and the absorbance value was detected at 570 nm (OD 570) using an automatic microplate reader.
Transwell Assay
Migration and invasion of HCC cells was examined using the transwell assay with polyethylene terephthalate membranes (24-well inserts, 8.0 μm, Corning). For the migration assay, 150 μL cell suspension containing 2 × 105 cells was loaded into the upper well coated without Matrigel (BD Biosciences). For the invasion assay, the Matrigel (BD Biosciences) was added. Next, 500 μL DMEM medium with 10% FBS was placed into the bottom of the well. Seventy-two hours later, invasive cells were stained with 0.1% crystal violet. Four fields were randomly selected and the cells counted. The experiments were repeated thrice and the data are presented as the mean ± SD.
Tumorigenesis in vivo
Male nude mice were raised under standard conditions. Cell suspensions (1 × 106) of Huh7/iCon or Huh7/sh1 cells in a total volume of 100 µL were injected into the 5-week-old male nude mice (4 mice per group). Each nude mouse was injected subcutaneously in their flanks at two sites. Tumor volume (mm3) was measured each week. At the end of the experiment, the tumors were harvested, photographed, and weighed. All animal experiments were performed under the approval of the Animal Ethical and Welfare Committee of the Peking University International Hospital, Beijing, China. The study was performed in accordance with NC3Rs ARRIVE guidelines.17
In vivo Metastasis Assay
Huh7/iCon or Huh7/sh1 cells labelled with luciferase (2 × 106 cells in 50 µL Matrigel) were injected into the left livers of the nude mice (7 mice per group). Before the mice were anaesthetized with isoflurane, an aqueous solution of luciferin (150 mg/kg intraperitoneally) was injected 10 min before imaging. The metastatic lesions were monitored weekly by luciferase expression, which was determined by an in vivo Living Image System (Xenogen). Metastases in the injection lobe and non-injection lobes were counted. All experiments were approved by the ethics commitment of the Peking University International Hospital, Beijing, China.
Luciferase Reporter Assay
Cells were placed in a 24-well plate and 100 ng MTF2 expression vector or empty vector, 50 ng Snail reporter plasmid, and 20 ng Renilla luciferase were co-transfected into the cells for 48 h. The cells were then lysed 8 h after treatment and reporter activities were investigated using a dual-luciferase reporter assay system (Promega).
Statistical Analysis
All the results are shown as mean ± standard errors of the mean. Statistical evaluations were performed with GraphPad Prism (version 5). The Student’s t-test and chi-square test were used for comparing measurable variants of the two groups. Survival curves were calculated using the Kaplan–Meier method, and differences were assessed using a log rank test. Statistical significance was set at a p-value of < 0.05 and marked with an asterisk.
Results
Clinical Significance of MTF2 Expression in HCC
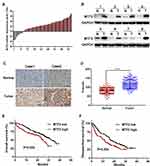
First, we quantified the mRNA level of MTF2 in 43 pairs of HCC tissues and their matched normal counterparts using qPCR, and we found that upregulation of MTF2 was observed in 35 pairs, accounting for 81% of the total specimens examined (Figure 1A). Consistent with the mRNA expression, we selected 8 pairs of HCC tissues and their matched normal tissues to measure the protein expression of MTF2 using Western blotting, and we observed elevated protein expression level of MTF2 in 7 of the 8 paired samples (Figure 1B). Next, to further explore the clinical significance of MTF2 in patients with HCC, we examined MTF2 expression in an HCC tissue microarray containing 240 specimens by immunohistochemical staining and staining intensity was scored in a standard manner as described previously.18 Higher protein expression of MTF2 in HCC tissues than in normal tissues was further confirmed by immunohistochemical staining and the MTF2 H-score (Figure 1C and D). Based on the intensity score, the patients with HCC were divided into the following two groups: MTF2 low (score < 102, 120 patients) and MTF2 high (score ≥ 102, 120 patients). In addition, we found that patients with HCC with higher MTF2 expression levels manifested a shorter overall survival and disease-free survival (Figure 1E and F; P = 0.002 and 0.003, respectively). Collectively, these results revealed that the expression of MTF2 was increased in HCC cells.
Furthermore, we explored the relationship between MTF2 expression and clinicopathologic parameters in 240 patients with HCC. As shown in Table 1, high MTF2 expression was associated with a higher level of alpha-fetoprotein (AFP) (P = 0.001), larger tumor diameter (P < 0.001), satellite nodules (P = 0.006), and the histological presence of microvascular invasion (P = 0.016). Cox’s multivariate proportional hazards model demonstrated some risk factors for overall survival (Table 2): AFP ≥ 20 ng/mL (hazard ratio (HR): 1.113, 95% confidence interval (CI): 0.865–1.436, P = 0.013); tumor diameter ≥ 5 cm (HR: 1.866, 95% CI: 1.677–1.956, P = 0.022); presence of microvascular invasion (HR: 1675, 95% CI: 1.543–1.768, P = 0.006); and MTF2 ≥ 102 (HR: 1645, 95% CI: 0.098–1.965, P = 0.011). The risk factors for disease-free survival were as follows (Table 3): HBsAg positive (HR: 1.321, 95% CI: 0.991–1.987, P = 0.043); AFP ≥ 20 ng/mL (HR: 1.211, 95% CI: 0.793–1.532, P = 0.001); Tumor diameter ≥ 5 cm (HR: 1.732, 95% CI: 1.611–1.987, P = 0.038); presence of microvascular invasion (HR: 1.533, 95% CI: 1.345–1.699, P = 0.012); and MTF2 ≥ 102 (HR: 1.523, 95% CI: 0.087–1.822, P = 0.009).
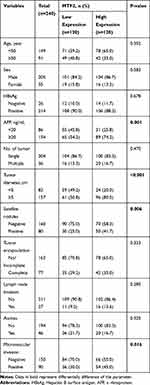 |
Table 1 Correlation Between the MTF2 Expression and the Clinicopathologic Features of HCC |
 |
Table 2 Univariate and Multivariate Analysis Associated with Overall Survival |
 |
Table 3 Univariate and Multivariate Analysis Associated with Disease-Free Survival |
MTF2 Overexpression Promoted the Growth, Migration, and Invasion of HCC Cells in vitro
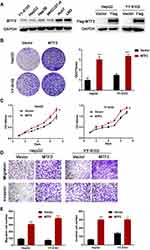
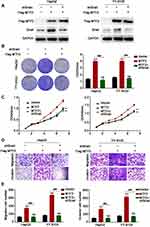
The above clinical observations encouraged us to investigate the function of MTF2 in HCC growth and metastasis in vitro. We constructed MTF2-overexpressing cells using lentiviruses in HepG2 and YY-8103 cells that have relatively low expression of MTF2 (Figure 2A). MTF2 expression was further confirmed by Western blot analysis (Figure 2B). Next, the effects of MTF2 on proliferation of HCC cells were assessed by crystal violet and MTT assays. We found that MTF2 overexpression markedly promoted the growth and colony formation of HepG2 and YY-8103 cells according to MTT (Figure 2B) and crystal violet assays (Figure 2C). In addition, the effects of MTF2 on migration and invasion of HCC cells were assessed by transwell assays, which showed that overexpression of MTF2 notably promoted the migration and invasion of HepG2 and YY-8103 cells (Figure 2D and E). Therefore, these results showed that the overexpression of MTF2 can promote the growth, migration, and invasion of HCC cells in vitro.
Knockdown of MTF2 Inhibited the Growth, Migration, and Invasion of HCC Cells in vitro
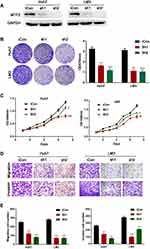
To further evaluate the function of MTF2 in HCC cells, we knocked down the expression of MTF2 in Huh7 and LM3 HCC cells which have relatively high expression of MTF2 (Figure 3A). In contrast to the results of the overexpression experiments, we found that knock down of MTF2 notably suppressed the growth and colony formation of Huh7 and LM3 cells according to MTT (Figure 3B) and crystal violet assays (Figure 3C). In addition, the transwell assay indicated that MTF2 downregulation can inhibit the migration and invasion of Huh7 and LM3 cells (Figure 3D and E). In conclusion, these results demonstrated that the knock down of MTF2 might play a protective role in HCC by decreasing the growth, migration, and invasion of HCC cells in vitro.
Knockdown of MTF2 Suppressed Tumorigenesis and Intrahepatic Metastasis of HCC Cells in vivo
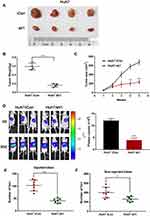
Based on the results of in vitro experiments, control and MTF2 knockdown Huh7 cells were injected subcutaneously into the flanks of nude mice, and the tumors were harvested and photographed. Consistent with the in vitro study, knock down of MTF2 in Huh7 cells markedly suppressed tumor growth compared with that in the control cells in xenograft mouse model (Figure 4A). The tumors were lighter and grew slower in the MTF2 downregulation group than those generated by the control cells (n = 4, P < 0.001) (Figure 4B and C). These results indicated that MTF2 may act as an oncogene that promotes tumorigenesis of HCC cells in vivo.
Based on the preliminary migration and invasion results, we speculated that MTF2 might promote the metastasis of HCC cells in vivo. The intrahepatic metastasis mouse model was applied to verify the function of MTF2 in HCC metastasis. Luciferase-labelled Huh7 con or Huh7 knockdown cells were injected into the left lobes of the nude mice. As shown in Figure 4D, luciferase signals from the livers of the MTF2 knockdown group were significantly lower than those from the control group at 5 weeks after injection (n = 7, P < 0.001). In addition, less foci were detected on injected and non-injected lobes in the MTF2 knockdown group, compared with that in the control group (Figure 4E and F; P < 0.001 and P < 0.05, respectively). Therefore, these results revealed that knock down of MTF2 suppressed intrahepatic metastasis of HCC cells in vivo.
MTF2 Promoted EMT and Snail Transcription in HCC Cells
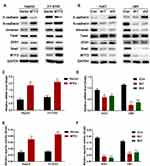
It has been reported that constitutive EMT activation was common in the development and progression of HCC. We hypothesized that MTF2 might promote growth and metastasis of HCC cells by activating the EMT, and Western blot analysis was performed to investigate the expression levels of EMT related genes E-cadherin, N-cadherin, and Vimentin in MTF2 overexpression and knockdown cell lines. We found that MTF2 overexpression in HepG2 and YY-8103 cells increased the expression of the mesenchymal markers, N-cadherin and Vimentin, and repressed the expression of the epithelial marker, E-cadherin (Figure 5A). In contrast, downregulation of MTF2 in Huh7 and LM3 cells reduced the expression of N-cadherin and Vimentin, while the expression of E-cadherin was increased in HCC cells (Figure 5B).
Molecular reprogramming occurring during EMT is triggered and orchestrated by EMT-activating transcriptional factors, including the Snail family, Twist family, and ZEB proteins. Among them, we found that only the expression of Snail was associated with MTF2 according to the Western blot analysis (Figure 5A and B). To further confirm their relationship, qPCR was conducted to examine the expression of Snail in MTF2 overexpression and knockdown cell lines, which revealed that MTF2 overexpression increased the mRNA expression of Snail, while MTF2 knockdown decreased the mRNA expression of Snail (Figure 5C and D). In addition, luciferase reporter assay revealed that the activity of the Snail promoter was promoted by MTF2 overexpression and inhibited in MTF2 knockdown HCC cells (Figure 5E and F). Taken together, these results showed that MTF2 promoted EMT and Snail transcription by facilitating its promoter activity in HCC cells.
Next, we examined whether the effects of MTF2 on the progression of HCC were Snail-dependent. We knocked down the expression of Snail in HepG2 and YY-8103 cell lines after MTF2 overexpression (Figure 6A). As shown in Figure 6B and E, the MTF2-induced promoting effects on cell growth, migration, and invasion were rescued by knockdown of Snail. These observations indicated that the promoting effects of MTF2 were Snail-dependent.
Discussion
As a member of the Pcl family of proteins, it is reported that MTF2 is actively involved in chromosome inactivation and pluripotency. Perino et al demonstrated that MTF2 directly binds to DNA and is a methylation-sensitive PRC2 recruiter in mouse embryonic stem cells.14 Rothberg et al revealed that MTF2 is an important epigenetic regulator of the Wnt signaling pathway during erythropoiesis and recast the role of polycomb accessory proteins in a tissue-specific context.16 However, the potential function of MTF2 in the progression of common cancers like hepatocellular carcinoma has not been elucidated to date.
Based on the clinical sample results, this study revealed MTF2 was significantly upregulated in HCC tissues compared with that in their paired normal counterparts. Patients with HCC with high MTF2 expression had worse overall survival and higher recurrence rates than those with low MTF2 expression. In addition, the expression levels of MTF2 correlated with the clinical characteristics of the patients, such as AFP, tumor diameter, satellite nodules, and microvascular invasion. Moreover, MTF2 overexpression, as well as high level of AFP, large tumor diameter, and presence of microvascular invasion was an independent risk factor for overall survival and disease-free survival, suggesting that it might play a suppressive role during the progression of HCC. These clinical results indicated that MTF2 might play an incentive role during the progression of HCC.
The functional analysis for MTF2 in the progression of HCC cells revealed that overexpression of MTF2 could promote HCC cell proliferation, migration, and invasion in vitro. In contrast, knockdown of MTF2 could suppress proliferation, migration, and invasion of HCC cells. Furthermore, MTF2 knockdown also inhibited tumorigenesis and intrahepatic metastasis of HCC cells in nude mice.
During the EMT process, epithelial tumor cells lose their polarity and transform into motile mesenchymal cells, and this process plays a predominant role in the dissemination of malignant hepatocytes during HCC progression.19,20 EMT is characterized by the loss of epithelial cell junction markers, such as E-cadherin, and the gain of mesenchymal proteins, such as N-cadherin and Vimentin.21,22 In this study, we found that cells with overexpression of MTF2 expressed high levels of N-cadherin and Vimentin, and low levels of E-cadherin, suggesting that MTF2 may be a potent inducer of EMT, which may lead to more invasive and metastatic HCC cells. In addition, EMT is orchestrated by a restricted number of transcription factors including Snail, Twist, and Zeb families.9,10 Our study showed that only the expression of Snail was significantly associated with MTF2. Furthermore, MTF2 regulated Snail expression by directly binding to its promoter. The rescue experiment also indicated that the promoting effects of MTF2 were Snail-dependent. Therefore, we conclude that MTF2 overexpression may promote EMT processes of HCC cells through Snail-mediated E-cadherin expression. To the best of our knowledge, this is the first study to reveal how MTF2 is involved in the progression of HCC.
In summary, our current study demonstrated a novel role for MTF2 in the growth and EMT of HCC cells through transcriptional activation of Snail. Overexpression of MTF2 in HCC is a strong indicator of aggressive malignant behavior and poor clinical outcome. These results provide a better understanding of the mechanisms underlying the role of MTF2 in tumor development, which might provide a potential therapeutic target for HCC.
Ethics Approval and Consent
HCC tissue samples and paired cancer-adjacent normal tissues were obtained from the Peking University International Hospital, after written informed consent was obtained from the patients. Use of these tissue samples was approved by the Ethics Committee for the Peking University International Hospital, and the study was performed in accordance with the Declaration of Helsinki.
Availability of Data and Materials
All data generated or analyzed during this study are included in this published article.
Consent for Publication
All the authors are aware of and agree to the content of the paper.
Acknowledgment
This study was funded by the Capital Characteristic Clinic Project (No. Z141107002514108).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Forner A, Reig M, Bruix J. Hepatocellular carcinoma. Lancet. 2018;391(10127):1301–1314. doi:10.1016/S0140-6736(18)30010-2
3. Marrero JA, Kulik LM, Sirlin CB, et al. Diagnosis, staging, and management of hepatocellular carcinoma: 2018 practice guidance by the American Association for the study of liver diseases. Hepatology. 2018;68(2):723–750. doi:10.1002/hep.29913
4. Galle PR, Forner A, Llovet JM, European Association for the Study of the Liver, Electronic address eee, European Association for the Study of the L. EASL clinical practice guidelines: management of hepatocellular carcinoma. J Hepatol. 2018;69(1):182–236. doi:10.1016/j.jhep.2018.03.019
5. Hu J, Gao DZ. Distinction immune genes of hepatitis-induced heptatocellular carcinoma. Bioinformatics. 2012;28(24):3191–3194. doi:10.1093/bioinformatics/bts624
6. Tornesello ML, Buonaguro L, Izzo F, Buonaguro FM. Molecular alterations in hepatocellular carcinoma associated with hepatitis B and hepatitis C infections. Oncotarget. 2016;7(18):25087–25102. doi:10.18632/oncotarget.v7i18
7. Huang RY, Guilford P, Thiery JP. Early events in cell adhesion and polarity during epithelial-mesenchymal transition. J Cell Sci. 2012;125(Pt 19):4417–4422. doi:10.1242/jcs.099697
8. Hay ED. An overview of epithelio-mesenchymal transformation. Acta Anat. 1995;154(1):8–20. doi:10.1159/000147748
9. Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2014;15(3):178–196. doi:10.1038/nrm3758
10. Dongre A, Weinberg RA. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat Rev Mol Cell Biol. 2019;20(2):69–84. doi:10.1038/s41580-018-0080-4
11. Walker E, Chang WY, Hunkapiller J, et al. Polycomb-like 2 associates with PRC2 and regulates transcriptional networks during mouse embryonic stem cell self-renewal and differentiation. Cell Stem Cell. 2010;6(2):153–166. doi:10.1016/j.stem.2009.12.014
12. Casanova M, Preissner T, Cerase A, et al. Polycomblike 2 facilitates the recruitment of PRC2 Polycomb group complexes to the inactive X chromosome and to target loci in embryonic stem cells. Development. 2011;138(8):1471–1482. doi:10.1242/dev.053652
13. Inouye C, Remondelli P, Karin M, Elledge S. Isolation of a cDNA encoding a metal response element binding protein using a novel expression cloning procedure: the one hybrid system. DNA Cell Biol. 1994;13(7):731–742. doi:10.1089/dna.1994.13.731
14. Perino M, van Mierlo G. MTF2 recruits Polycomb Repressive Complex 2 by helical-shape-selective DNA binding. Nat Genet. 2018;50(7):1002–1010.
15. Walker E, Manias JL, Chang WY, Stanford WL. PCL2 modulates gene regulatory networks controlling self-renewal and commitment in embryonic stem cells. Cell Cycle. 2011;10(1):45–51. doi:10.4161/cc.10.1.14389
16. Rothberg JLM, Maganti HB, Jrade H, Porter CJ. Mtf2-PRC2 control of canonical Wnt signaling is required for definitive erythropoiesis. Cell Discovery. 2018;4:21. doi:10.1038/s41421-018-0022-5
17. Kilkenny C, Browne WJ, Cuthill IC, Emerson M, Altman DG. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol. 2010;8(6):e1000412. doi:10.1371/journal.pbio.1000412
18. Stack EC, Wang C, Roman KA, Hoyt CC. Multiplexed immunohistochemistry, imaging, and quantitation: a review, with an assessment of tyramide signal amplification, multispectral imaging and multiplex analysis. Methods. 2014;70(1):46–58. doi:10.1016/j.ymeth.2014.08.016
19. Thiery JP, Acloque H, Huang RY, Nieto MA. Epithelial-mesenchymal transitions in development and disease. Cell. 2009;139(5):871–890. doi:10.1016/j.cell.2009.11.007
20. Yang J, Weinberg RA. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell. 2008;14(6):818–829. doi:10.1016/j.devcel.2008.05.009
21. Manjunath Y, Upparahalli SV, Avella DM, Deroche CB. PD-L1 expression with epithelial mesenchymal transition of circulating tumor cells is associated with poor survival in curatively resected non-small cell lung cancer. Cancers. 2019;11(6):806.
22. Zhou ZJ, Dai Z, Zhou SL, et al. HNRNPAB induces epithelial-mesenchymal transition and promotes metastasis of hepatocellular carcinoma by transcriptionally activating SNAIL. Cancer Res. 2014;74(10):2750–2762. doi:10.1158/0008-5472.CAN-13-2509
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.