Back to Journals » Infection and Drug Resistance » Volume 13
Molecular Epidemiology of Carbapenemase-Producing Pseudomonas aeruginosa Isolated from an Iranian University Hospital: Evidence for Spread of High-Risk Clones
Authors Ohadian Moghadam S , Afshar D, Nowroozi MR, Behnamfar A, Farzin A
Received 13 March 2020
Accepted for publication 17 May 2020
Published 3 June 2020 Volume 2020:13 Pages 1583—1592
DOI https://doi.org/10.2147/IDR.S253756
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Suresh Antony
Solmaz Ohadian Moghadam,1 Davoud Afshar,2 Mohammad Reza Nowroozi,1 Amir Behnamfar,1 Amirreza Farzin1
1Uro-Oncology Research Center, Tehran University of Medical Sciences, Tehran, Iran; 2Department of Microbiology and Virology, Zanjan University of Medical Sciences, Zanjan, Iran
Correspondence: Solmaz Ohadian Moghadam
Uro-Oncology Research Center, Tehran University of Medical Sciences, Tehran, Iran
Tel/ Fax +98 21 66 43 77 25
Email [email protected]
Purpose: Given the importance of treatment failure due to multidrug-resistant (MDR) strains, studies on population structure of these organisms are necessary to improve control strategies. Accordingly, the current study aimed to determine the prevalence of carbapenem-resistant P. aeruginosa (CRPA) at a teaching referral hospital in Iran and to analyze their molecular clonality by multilocus sequence typing (MLST) and pulsed-field gel electrophoresis (PFGE) for epidemiological purposes.
Methods: In this study, modified Hodge test (MHT) and double-disk synergy test (DDST) were used for carbapenemase production and metallo-β-lactamases (MBLs) screening, respectively. All P. aeruginosa isolates were tested for antimicrobial resistance. Moreover, MBL genes (blaIMP, blaVIM, blaSPM, blaNDM) were detected by multiplex PCR assay.
Results: Among 68 P. aeruginosa clinical isolates, 38 (55.88%) isolates were CRPA. Antibiotic susceptibility testing revealed that most of these isolates were MDR. PFGE analyses showed 5 common types and 27 single types among CRPA isolates. MLST analysis revealed three major clusters (MLST-sequence types (STs): 235, 357, and 861) among them. The 30 non-CRPA isolates corresponded mainly to MLST-STs 253, 360, and 446.
Conclusion: Our results showed that internationally distributed MLST-STs with widely genomic diversity have spread in our hospital, and clonal expansion of MDR strains of P. aeruginosa was described as well.
Keywords: carbapenem-resistant P. aeruginosa, CRPA, carbapenemase, metallo-β-lactamases, MBLs, multidrug-resistant, MDR, Pseudomonas aeruginosa, multilocus sequence typing, MLST, pulsed-field gel electrophoresis, PFGE
Introduction
Pseudomonas aeruginosa, as one of the major cause of nosocomial infections, is associated with significantly higher mortality and morbidity rates compared to the other members of Pseudomonadaceae family. It is responsible for numerous types of infections, including surgical site infections, urinary tract infections, pneumonia and bloodstream infections.1 Currently, multidrug-resistant (MDR) and extensively drug-resistant (XDR) P. aeruginosa strains have become a global concern.2 Due to intrinsic and acquired resistance of P. aeruginosa to a wide range of commonly used antimicrobial agents, effective treatment of infections caused by this bacterium is often challenging.3 Currently, the most effective drugs for clinical management of severe P. aeruginosa infections are carbapenems.3,4 However, due to the emergence of resistant strains producing carbapenemase which can hydrolyze carbapenems, penicillins, and cephalosporins, their effectiveness has been endangered.4,5 According to the World Health Organization in 2017, new antibiotics were promptly required against carbapenem-resistant P. aeruginosa (CRPA) strains.6
Carbapenem resistance in P. aeruginosa can occur due to combination of mechanisms including decreased activity of efflux pump, decreased permeability of outer membrane as a result of porin alterations following chromosomal mutations, and carbapenemases production as the main resistance mechanism.4 Carbapenemases are a kind of β-lactamase being assigned to the classes of A, D (referred to as serine carbapenemases), and B (referred to as metallo-β-lactamases).5 Carbapenemase genes are commonly harbored on mobile genetic elements and have the potential for rapid dissemination. As a class of beta-lactams, carbapenems can inhibit the synthesis of the bacterial cell wall by binding to penicillin-binding proteins (PBPs). Clinically available carbapenems include imipenem, meropenem, ertapenem, and doripenem.4 Metallo-β-lactamases (MBLs) confer resistance to all β-lactams, except the monobactams.7 The most frequent MBLs comprise New Delhi metallo-β-lactamases (NDMs), Verona integron-encoded metallo-β-lactamase (VIM), and IMP (so-called for its imipenem-resistant phenotype) enzymes.8 In addition, the most prevalent MBL genes in P. aeruginosa are the VIM and IMP types; particularly, VIM-2 has become the dominant MBL type worldwide.9 However, in a previous study conducted in Iran blaIMP was reported as the most prevalent carbapenemase gene in P. aeruginosa isolates.10
The potential of high-risk clones of P. aeruginosa for being selected and disseminating antimicrobial resistance has made them a serious issue for public health worldwide.2 There are controversies about population structure and epidemic clonality of P. aeruginosa.11 Multilocus sequence typing (MLST) is a typing method based on conserved housekeeping genes and is capable of tracking the global clonal history of the species with the highest accuracy.12
Hence, the aim of this study was to determine the prevalence of CRPA isolates at a teaching referral hospital in Iran and analyzing their molecular clonality by MLST as well as studying the clonal relationship among the isolates by pulsed-field gel electrophoresis (PFGE), in order to gain a better insight into the dynamics of MDR P. aeruginosa strains as it might help in the global comparisons.
Materials and Methods
Bacterial Isolation
From January 2018 to January of 2020, a total of 157 urine samples were collected from pathologically confirmed prostate cancer and bladder cancer patients hospitalized in the urology ward of Imam Khomeini hospital affiliated to Tehran University of Medical Sciences. The sampling was a part of routine hospital laboratory procedure and at the time of admission an informed consent was obtained from patients. All urine samples received in the microbiology laboratory were processed for identification of P. aeruginosa. Isolated bacteria were recovered on blood agar after an overnight incubation at 37°C. Sub-cultures were done on MacConkey agar for single colonies. Then the isolates were identified based upon standard microbiological procedures (Gram staining, colony morphology, and biochemical tests such as catalase, oxidase, lactose fermentation, Methyl red/Voges–Proskauer, Simmons’ citrate, triple sugar iron, oxidation/fermentation, urease, nitrate reduction, H2S production, and motility tests).13 Subsequently, strains were confirmed by polymerase chain reaction (PCR) for oprL gene.12
Antimicrobial Susceptibility Testing
Susceptibility testing against 10 antibiotics including piperacillin (100μg), piperacillin-tazobactam (100/10 μg), ceftazidime (30 μg), cefepime (30 μg), imipenem, meropenem (10 mg), aztreonam (30 μg), gentamicin (10 μg), tobramycin (10 μg), amikacin (30 μg), and ciprofloxacin (10 μg) (Oxoid; Basingstoke, UK) was performed for all non-duplicate P. aeruginosa isolates by Kirby-Bauer disk method on Mueller-Hinton agar (MHA) according to the Clinical & Laboratory Standards Institute (CLSI) guidelines.14 The reference strain was P. aeruginosa PAO1.12 In addition, E-test (AB Biodisk, Sweden) was used for determination of minimum inhibitory concentration (MIC) for ceftazidime, imipenem and meropenem according to the manufacturer’s instructions and CLSI guidelines. Carbapenem-resistant isolates were defined as imipenem or/and meropenem MIC ≥8mg/L.14 In addition, MDR was defined as resistance to at least one agent in more than three antimicrobial categories.15
Carbapenemase Assay
Carbapenemase assay was performed with the modified Hodge test (MHT) to identify carbapenemase activity in carbapenem-resistant isolates as described previously.16 Briefly, a turbidity of 0.5 McFarland standard of the E.coli ATCC 25922 as an indicator organism was prepared in 5 mL of saline and was subsequently diluted by sterile saline (1:10). This was streaked to a MHA plate. Then, a 10-μg IPM disk (Becton Dickinson) was located in the center of the MHA plate. The study isolate was heavily streaked in a straight line from the edge of the disk to the edge of the plate. The plate was incubated overnight at 37°C. Interpretation was performed according to the CLSI guidelines. Briefly, observation of clover leaf-like indentation inhibition zone was defined as positive result (carbapenemase production by the test isolate). In addition, lack of E.coli 25922 growth along the test isolate growth streak within the disc diffusion zone was defined as MHT negative.
Imipenem (IPM)-EDTA Double-Disk Synergy Test (DDST)
To identify MBL activity, DDST was performed. Overnight liquid cultures of the test isolates were adjusted to the McFarland 0.5 standard turbidity and were inoculated to MHA plates. Two discs with 10 μg imipenem were placed on the plate. Then, 10 μL of 0.5 M solution of EDTA (as chelating agent) was added to one of the imipenem disc. After overnight incubation at 37 °C, the presence of increased inhibition zone (≥5 mm) in the EDTA-supplemented disc was considered as positive for MBL production. Additionally, carbapenem-susceptible isolates (disc inhibition zone diameters of ≤25 mm) were tested for carbapenemase activity as well. MBL production by the DDST-positive isolates were confirmed by PCR for carbapenemase genes.17
Detection of Resistance Genes
Chromosomal DNA was extracted using a DNAasy kit (Qiagen, Tokyo, Japan) as recommended by the manufacturer. Purified genomic DNA was used as the template for PCR. All carbapenem-resistant isolates were tested for MBL genes (blaIMP, blaVIM, blaSPM, blaNDM) using multiplex PCR assay described previously.18
MLST
MLST was accomplished for both CRPA and non-CRPA isolates, as described by Curran et al.19 Briefly, seven genes including acsA, aroE, guaA, mutL, nuoD, ppsA, and trpE were amplified, and the PCR products were purified with the purification kit (QIAGEN). Sequencing was performed with the primers as corrected at the http://pubmlst.org website using the Applied Biosystems (ABI) 3730XL DNA analyzer. Then their allelic profile and MLST-STs were determined via the P. aeruginosa MLST website (http://pubmlst.org/paeruginosa/).
Genotyping with PFGE
The P. aeruginosa isolates were genotyped by PFGE as described previously.20 DNA digestion was performed using SpeI restriction enzyme (New England BioLabs-IZASA, Barcelona, Spain) according to the manufacturer instructions (18 hours at 36 °C). The DNA macrorestriction fragments were separated in the agarose gel by PFGE using a contour-clamped homogeneous electric field (CHEF) electrophoresis System (Bio-Rad, Hercules, California, USA) on a 1.2% agarose. The genomic DNA of Salmonella enterica serotype Braenderup H9812 (CDC), digested with 20 IU XbaI (Roche Diagnostics) for 20 h at 37 °C was used as the size marker. The images of stained gels with ethidium bromide were visualized by ultraviolet transillumination and photographed. Digital images were stored electronically as TIFF files and were analyzed by GelComparII software version 4 (Applied Maths, Sint-Martens-Latem, Belgium). The UPGMA method (unweighted pair group method using arithmetic mean Algorithm) was utilized as the clustering method (1% band tolerance, 1% optimization). Interpretation of results was performed according to the criteria of relatedness suggested by Tenover et al.21 Clonal relatedness was defined according to a similarity (Dice) coefficient greater than 0.80; otherwise, the isolates were not considered to be related and were defined as single type or distinct PFGE profile.
Results
Clinical Isolates
In the current study, a total of 68 P. aeruginosa isolates were recovered from urine samples of urology cancer patients, among which 38 (55.88%) isolates were carbapenem resistant (Figure 1).
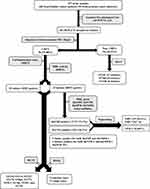 |
Figure 1 Flow of sample collection and study design through the study and detection of carbapenemase-producing P. aeruginosa isolate. |
Antibiotics Susceptibility Pattern
The susceptibility of CRPA isolates to the different antimicrobial agents tested is shown in Table 1. According to Etest method, 38 (100%) and 36 (92.1%) isolates were resistant to imipenem (MIC≥16mg/mL) and ceftazidime (MIC≥32 mg/mL), respectively (Table 2).
 |
Table 1 The Susceptibility of Carbapenem-Resistant P. Aeruginosa Isolates to the Different Antimicrobial Agents |
 |
Table 2 MIC90 and MIC50 Values for Meropenem, Ceftazidime, and Cefepime Among CRPA and CSPS Isolates |
Carbapenemase-Producing and Carbapenemase Genes
MHT and DDST were positive in 35 and 32 isolates, indicated carbapenemase and MBL production, respectively. PCR assay indicated that 23 and 9 of the isolates were positive for blaIMP and blaVIM genes, respectively. Sequencing of PCR products showed that 19 (82.60%) IMP products belonged to the IMP-1 group and 4 (17.40%) IMP products belonged to the IMP-7 group. All VIM products belonged to the VIM-1 group. Four isolates were positive for both blaIMP and blaNDM-1 genes and 2 isolates were found to be harboring with both blaVIM and blaNDM-1 genes. No blaSPM isolates were found. Three (7.9 %, 3/38) carbapenem-resistant isolates were found to be negative for both phenotypically carbapenemase tests and for the carbapenemase genes studied.
PFGE Typing
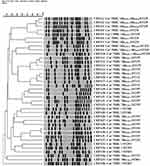
All CRPA isolates were analyzed by PFGE. Among these isolates, 5 common types (CTs; pulsotypes with a minimum similarity of 80%) and 27 single types (dissimilar pulsotypes) were observed. The constructed dendrogram for relative relatedness among CRPA isolates is shown in Figure 2.
MLST
MLST analysis revealed different STs for P. aeruginosa isolates (ST235, ST111, ST357, ST175, ST861, ST773, ST823, ST360, ST207, and ST253). ST235 and ST253 were the most prevalent STs among the carbapenem-resistant and carbapenem-susceptible isolates, respectively. Molecular characteristics of P. aeruginosa isolates are summarized in Table 1. The 30 non-CRPA isolates corresponded mainly to ST253, ST360, and ST446 (Figure 1, Table 1).
Discussion
The hospital environment is the most important source of MDR bacterial infections, especially infections involving P. aeruginosa.22 MDR P. aeruginosa strains are common causes of nosocomial infections worldwide23 and this can be particularly severe in patients with impaired immune systems, such as cancer patients.24
There are limited information on the CRPA population structure in Iran. In the present study, we observed that 55.88% (n =38) isolated P. aeruginosa were carbapenem resistant. The potential source of colonization could be both hospital and community. Resistance to imipenem has been reported to vary from 12.5% to 100% in different previous studies in Iran.25–27 Additionally, the prevalence of MDR and XDR P. aeruginosa strains has been reported to 15–30% in different geographical regions.28–30 Antimicrobial susceptibility profiles of CRPA isolates indicated a high level of resistance to most antibiotics studied (Table 1). CRPA isolates were resistant to the majority of antibiotics studied. The non-CRPA isolates showed lower MICs of ceftazidime, cefepime, and meropenem than CRPA isolates (Table 2). In this study, MHT and DDST indicated that carbapenem-resistance among P. aeruginosa isolates studied was mainly mediated by MBLs (32/35 isolates).
Among several molecular typing methods proposed for P. aeruginosa, PFGE seems to be the most discriminatory method for short-term epidemiological studies.31 However, due to its extensive genomic diversities, MLST is more suitable for evaluating long-term evolutionary relationships.19 In the present study, two typing methods were used to determine epidemic clones of P. aeruginosa isolates. Our results have shown high genetic diversity among them. The carbapenem-resistant isolates comprised 11 STs. Totally 26 of the isolates were distributed in STs 235 (n =14), 357 (n=8), and 111 (n=4), which have been considered as high-risk clones.32 However, the 30 non-resistant isolates comprised 3 MLST clones, with STs 360 (n=9), 446 (n=8) and 253 (n =13). Most of the STs have been reported previously.25,33
In this study, ST235 emerged as the most prevalent MLST-ST accounting for 14 CRPA isolates. This sequence type is widely disseminated in the world and has been reported from other countries including Hungary, Italy, Greece, Poland, Sweden, Japan, and Norway.34–37 Moreover, ST235 is an international clone associated with poor clinical outcomes.38 It has been reported as MBL producing strains responsible for outbreaks throughout Europe, Asia, and South America.35,39-44
In our study, all of the ST235 isolates were MBL positive and majority of them harbored blaIMP (n=11) while only 3 isolates produced VIM type MBLs. Four of carbapenem-resistant isolates belonged to ST235 were positive for both blaIMP and blaNDM genes. All were MDR as well, suggesting the key role of the strain ST235 in dissemination of the MDR strains at the studied hospital. Our results indicated that ST357 was the second most frequently isolated sequence type, with 8 isolates all were MDR. This ST was previously reported from both Iran and other Asian countries such as Japan and Korea.42,45 In addition, ST111 and ST175 were the other sequence types reported in the current study. ST111 as well as ST175 have been also recognized as MBL-producing epidemic lineages.46,47 ST175 is also a globally distributed clone and has been documented as a contaminant of the hospital environment.48 The nine remaining carbapenem-resistant isolates belonged to different STs including ST861, ST773, ST823, ST360, ST207, and ST253.
Several studies reported blaVIM gene as the most common MBL found in CRPA.49,50 However, in our study blaIMP gene was the most frequent detected carbapenemase gene in P. aeruginosa isolates, which is in consistent with a previous study in Iran.10
PFGE was used to investigate the epidemiological characteristics of P. aeruginosa isolates. This method is valuable for characterization of outbreaks and their sources.51 In the present study, 5 CTs were found among carbapenem-resistant isolates, which may represent five outbreaks. All the isolates belonged to the same PFGE-CTs had both similar MLST-ST and similar antibiotic resistance pattern. The chromosomal digestion patterns revealed a high diversity among CRPA isolates. This could be explained by acquisition of these strains from various sources.
This study is subjected to some limitations. First, the lack of enough previous information about the genetic composition of strains for comparison purposes. Second, it was a single-center study; therefore, additional studies are required to reach a broader insight on the clonal dynamics of CRPA contaminating hospitals in Iran. In summary, the population structure of P. aeruginosa present in our hospital shows that the MDR isolates studied are grouped in the prevalent STs reported in other Iranian hospitals and at the international level. In addition, IMP-1, producing P. aeruginosa are the carbapenemase-producing strains mainly circulating at our hospital environment.
Conclusions
The present study shows the prevalence of CRPA with limited susceptibility to antibiotics in hospital environment. Therefore, proper antibiotic strategies should be formulated in order to overcome these problems. Moreover, our results indicated that internationally distributed MLST-STs with widely genomic diversity have spread in our hospital. The PFGE analysis also showed the possibility of localized outbreak during the period of study. In conclusion, clonal expansion of MDR strains of P. aeruginosa was described in a urology ward in Iran. The finding highlights the significance of continuingly following the epidemiological relatedness of MDR P. aeruginosa spreading in hospital environment to improve infection control and prevention strategies.
Abbreviations
CRPA, Carbapenem-resistant P. aeruginosa; CHEF, Contour-clamped homogeneous electric field; DDST, Double-disk synergy test; XDR, Extensively drug-resistant; MBLs, Metallo-β-lactamases; MIC, Minimum inhibitory concentration; MHT, Modified Hodge test; MHA, Mueller-Hinton agar; MDR, Multidrug-resistant; MLST, Multilocus sequence typing; NDMs, New Delhi metallo-β-lactamases; PCR, Polymerase chain reaction; PFGE, Pulsed-field gel electrophoresis; PFGE-CT, PFGE-Common type; PFGE-ST, PFGE-Single type; MLST-ST, MLST-Sequence type; UPGMA, Unweighted pair group method using arithmetic mean Algorithm; VIM, Verona integron-encoded metallo-β-lactamase.
Acknowledgments
We give special thanks to all members of Uro‐Oncology Research Center for helpful discussions and friendly support.
Author Contributions
All authors have contributed in the acquisition of data, data analysis and interpretation, drafting of the manuscript, and critical revision of the manuscript for scientific and factual content. All authors have approved the final content of the manuscript and agree to be accountable for all aspects of the work.
Disclosure
The authors have no conflict of interest to declare.
References
1. Thaden JT, Park LP, Maskarinec SA, Ruffin F, Fowler VG
2. Horcajada JP, Montero M, Oliver A, et al. Epidemiology and treatment of multidrug-resistant and extensively drug-resistant pseudomonas aeruginosa infections. Clin Microbiol Rev. 2019;32(4). doi:10.1128/cmr.00031-19
3. Walters MS, Grass JE, Bulens SN, et al. Carbapenem-resistant Pseudomonas aeruginosa at US emerging infections program sites, 2015. Emerg Infect Dis. 2019;25(7):1281–1288. doi:10.3201/eid2507.181200
4. Papp-Wallace KM, Endimiani A, Taracila MA, Bonomo RA. Carbapenems: past, present, and future. Antimicrob Agents Chemother. 2011;55(11):4943–4960. doi:10.1128/aac.00296-11
5. Queenan AM, Bush K. Carbapenemases: the versatile beta-lactamases. Clin Microbiol Rev. 2007;20(3):440–458. doi:10.1128/cmr.00001-07.
6. Tacconelli E, Carrara E, Savoldi A, et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis. 2018;18(3):318–327. doi:10.1016/s1473-3099(17)30753-3
7. Bush K, Bush K. New beta-lactamases in gram-negative bacteria: diversity and impact on the selection of antimicrobial therapy. Clin Infect Dis. 2001;32(7):1085–1089. doi:10.1086/319610
8. Tamma PD, Simner PJ, Kraft CS. Phenotypic detection of carbapenemase-producing organisms from clinical isolates. J Clin Microbiol. 2018;56(11). doi:10.1128/jcm.01140-18
9. Walsh TR. Clinically significant carbapenemases: an update. Curr Opin Infect Dis. 2008;21(4):367–371. doi:10.1097/QCO.0b013e328303670b
10. Dogonchi AA, Ghaemi EA, Ardebili A, Yazdansetad S, Pournajaf A. Metallo-β-lactamase-mediated resistance among clinical carbapenem-resistant Pseudomonas aeruginosa isolates in northern Iran: a potential threat to clinical therapeutics. Ci Ji Yi Xue Za Zhi. 2018;30(2):90–96. doi:10.4103/tcmj.tcmj_101_17
11. Pirnay JP, Bilocq F, Pot B, et al. Pseudomonas aeruginosa population structure revisited. PLoS One. 2009;4(11):e7740. doi:10.1371/journal.pone.0007740
12. Fazeli H, Sadighian H, Esfahani BN, Pourmand MR. Molecular epidemiology and mechanisms of antimicrobial resistance in Pseudomonas aeruginosa isolates causing burn wound infection in Iran. J Chemother (Florence, Italy). 2014;26(4):222–228. doi:10.1179/1973947813y.0000000132
13. LaBauve AE, Wargo MJ. Growth and laboratory maintenance of Pseudomonas aeruginosa. Curr Protoc Microbiol. 2012;Chapter 6:Unit6E 1. doi:10.1002/9780471729259.mc06e01s25
14. Institute CaLS. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically. Wayne, PA: CLSI Document M07– A9; 2012.
15. Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. doi:10.1111/j.1469-0691.2011.03570.x
16. Okoche D, Asiimwe BB, Katabazi FA, Kato L, Najjuka CF, Zhang Q. Prevalence and characterization of carbapenem-resistant enterobacteriaceae isolated from mulago national referral hospital, Uganda. PLoS One. 2015;10(8):e0135745. doi:10.1371/journal.pone.0135745
17. Lee K, Lim YS, Yong D, Yum JH, Chong Y. Evaluation of the hodge test and the imipenem-EDTA double-disk synergy test for differentiating metallo-beta-lactamase-producing isolates of Pseudomonas spp. and Acinetobacter spp. J Clin Microbiol. 2003;41(10):4623–4629. doi:10.1128/jcm.41.10.4623-4629.2003
18. Woodford N. Rapid characterization of beta-lactamases by multiplex PCR. Methods Mol Biol (Clifton, N J). 2010;642:181–192. doi:10.1007/978-1-60327-279-7_14
19. Curran B, Jonas D, Grundmann H, Pitt T, Dowson CG. Development of a multilocus sequence typing scheme for the opportunistic pathogen Pseudomonas aeruginosa. J Clin Microbiol. 2004;42(12):5644–5649. doi:10.1128/jcm.42.12.5644-5649.2004
20. Yetkin G, Otlu B, Cicek A, Kuzucu C, Durmaz R. Clinical, microbiologic, and epidemiologic characteristics of Pseudomonas aeruginosa infections in a University Hospital, Malatya, Turkey. Am J Infect Control. 2006;34(4):188–192. doi:10.1016/j.ajic.2005.11.010
21. Tenover FC, Arbeit RD, Goering RV, et al. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33(9):2233–2239. doi:10.1128/JCM.33.9.2233-2239.1995
22. Aruhomukama D, Najjuka CF, Kajumbula H, et al. bla(VIM)- and bla(OXA)-mediated carbapenem resistance among Acinetobacter baumannii and Pseudomonas aeruginosa isolates from the mulago hospital intensive care unit in Kampala, Uganda. BMC Infect Dis. 2019;19(1):853. doi:10.1186/s12879-019-4510-5
23. Kateete DP, Nakanjako R, Namugenyi J, Erume J, Joloba ML, Najjuka CF. Carbapenem resistant Pseudomonas aeruginosa and Acinetobacter baumannii at Mulago Hospital in Kampala, Uganda (2007–2009). SpringerPlus. 2016;5(1):1308. doi:10.1186/s40064-016-2986-7
24. Pagani L, Mantengoli E, Migliavacca R, et al. Multifocal detection of multidrug-resistant Pseudomonas aeruginosa producing the PER-1 extended-spectrum beta-lactamase in Northern Italy. J Clin Microbiol. 2004;42(6):2523–2529. doi:10.1128/jcm.42.6.2523-2529.2004
25. Fazeli H, Sadighian H, Esfahani BN, Pourmand MR. Genetic characterization of Pseudomonas aeruginosa-resistant isolates at the university teaching hospital in Iran. Adv Biomed Res. 2015;4(1):156. doi:10.4103/2277-9175.161583
26. Yousefi S, Farajnia S, Nahaei MR, et al. Detection of metallo-β-lactamase-encoding genes among clinical isolates of Pseudomonas aeruginosa in northwest of Iran. Diagn Microbiol Infect Dis. 2010;68(3):322–325. doi:10.1016/j.diagmicrobio.2010.06.018
27. Ranjbar R, Owlia P, Saderi H, et al. Characterization of Pseudomonas aeruginosa strains isolated from burned patients hospitalized in a major burn center in Tehran, Iran. Acta Med Iran. 2011;49(10):675–679.
28. Peña C, Cabot G, Gómez-Zorrilla S, et al. Influence of virulence genotype and resistance profile in the mortality of Pseudomonas aeruginosa bloodstream infections. Clin Infect Dis. 2015;60(4):539–548. doi:10.1093/cid/ciu866
29. Walkty A, Lagace-Wiens P, Adam H, et al. Antimicrobial susceptibility of 2906 Pseudomonasaeruginosa clinical isolates obtained from patients in Canadian hospitals over a period of 8 years: results of the Canadian ward surveillance study (CANWARD), 2008–2015. Diagn Microbiol Infect Dis. 2017;87(1):60–63. doi:10.1016/j.diagmicrobio.2016.10.003
30. Sader HS, Castanheira M, Duncan LR, Flamm RK. Antimicrobial susceptibility of enterobacteriaceae and Pseudomonas aeruginosa isolates from United States medical centers stratified by infection type: results from the International Network for Optimal Resistance Monitoring (INFORM) surveillance program, 2015–2016. Diagn Microbiol Infect Dis. 2018;92(1):69–74. doi:10.1016/j.diagmicrobio.2018.04.012
31. Johnson JK, Arduino SM, Stine OC, Johnson JA, Harris AD. Multilocus sequence typing compared to pulsed-field gel electrophoresis for molecular typing of Pseudomonas aeruginosa. J Clin Microbiol. 2007;45(11):3707–3712. doi:10.1128/jcm.00560-07
32. Woodford N, Turton JF, Livermore DM. Multiresistant gram-negative bacteria: the role of high-risk clones in the dissemination of antibiotic resistance. FEMS Microbiol Rev. 2011;35(5):736–755. doi:10.1111/j.1574-6976.2011.00268.x
33. Yousefi S, Nahaei MR, Farajnia S, et al. A multiresistant clone of Pseudomonas aeruginosa sequence type 773 spreading in a burn unit in Orumieh, Iran. APMIS. 2013;121(2):146–152. doi:10.1111/j.1600-0463.2012.02948.x
34. Giske CG, Libisch B, Colinon C, et al. Establishing clonal relationships between VIM-1-like metallo-beta-lactamase-producing Pseudomonas aeruginosa strains from four European countries by multilocus sequence typing. J Clin Microbiol. 2006;44(12):4309–4315. doi:10.1128/jcm.00817-06
35. Glupczynski Y, Bogaerts P, Deplano A, et al. Detection and characterization of class A extended-spectrum-beta-lactamase-producing Pseudomonas aeruginosa isolates in Belgian hospitals. J Antimicrob Chemother. 2010;65(5):866–871. doi:10.1093/jac/dkq048
36. Samuelsen O, Buarø L, Toleman MA, et al. The first metallo-beta-lactamase identified in Norway is associated with a TniC-like transposon in a Pseudomonas aeruginosa isolate of sequence type 233 imported from Ghana. Antimicrob Agents Chemother. 2009;53(1):331–332. doi:10.1128/aac.00785-08
37. Maatallah M, Cheriaa J, Backhrouf A, et al. Population structure of Pseudomonas aeruginosa from five Mediterranean countries: evidence for frequent recombination and epidemic occurrence of CC235. PLoS One. 2011;6(10):e25617. doi:10.1371/journal.pone.0025617
38. Treepong P, Kos VN, Guyeux C, et al. Global emergence of the widespread Pseudomonas aeruginosa ST235 clone. Clin Microbiol Infect. 2018;24(3):258–266. doi:10.1016/j.cmi.2017.06.018
39. Empel J, Filczak K, Mrówka A, Hryniewicz W, Livermore DM, Gniadkowski M. Outbreak of Pseudomonas aeruginosa infections with PER-1 extended-spectrum beta-lactamase in Warsaw, Poland: further evidence for an international clonal complex. J Clin Microbiol. 2007;45(9):2829–2834. doi:10.1128/jcm.00997-07
40. Juan C, Zamorano L, Mena A, Albertí S, Pérez JL, Oliver A. Metallo-beta-lactamase-producing Pseudomonas putida as a reservoir of multidrug resistance elements that can be transferred to successful Pseudomonas aeruginosa clones. J Antimicrob Chemother. 2010;65(3):474–478. doi:10.1093/jac/dkp491
41. Koh TH, Khoo CT, Tan TT, et al. Multilocus sequence types of carbapenem-resistant Pseudomonas aeruginosa in Singapore carrying metallo-beta-lactamase genes, including the novel bla(IMP-26) gene. J Clin Microbiol. 2010;48(7):2563–2564. doi:10.1128/jcm.01905-09
42. Kouda S, Ohara M, Onodera M, et al. Increased prevalence and clonal dissemination of multidrug-resistant Pseudomonas aeruginosa with the blaIMP-1 gene cassette in Hiroshima. J Antimicrob Chemother. 2009;64(1):46–51. doi:10.1093/jac/dkp142
43. Libisch B, Poirel L, Lepsanovic Z, et al. Identification of PER-1 extended-spectrum beta-lactamase producing Pseudomonas aeruginosa clinical isolates of the international clonal complex CC11 from Hungary and Serbia. FEMS Immunol Med Microbiol. 2008;54(3):330–338. doi:10.1111/j.1574-695X.2008.00483.x
44. Samuelsen O, Toleman MA, Sundsfjord A, et al. Molecular epidemiology of metallo-beta-lactamase-producing Pseudomonas aeruginosa isolates from Norway and Sweden shows import of international clones and local clonal expansion. Antimicrob Agents Chemother. 2010;54(1):346–352. doi:10.1128/aac.00824-09
45. Seok Y, Bae IK, Jeong SH, Kim SH, Lee H, Lee K. Dissemination of IMP-6 metallo-β-lactamase-producing Pseudomonas aeruginosa sequence type 235 in Korea. J Antimicrob Chemother. 2011;66(12):2791–2796. doi:10.1093/jac/dkr381
46. Edalucci E, Spinelli R, Dolzani L, et al. Acquisition of different carbapenem resistance mechanisms by an epidemic clonal lineage of Pseudomonas aeruginosa. Clin Microbiol Infect. 2008;14(1):88–90. doi:10.1111/j.1469-0691.2007.01874.x
47. Elias J, Schoen C, Heinze G, et al. Nosocomial outbreak of VIM-2 metallo-β-lactamase-producing Pseudomonas aeruginosa associated with retrograde urography. Clin Microbiol Infect. 2010;16(9):1494–1500. doi:10.1111/j.1469-0691.2009.03146.x
48. Gomila M, Del Carmen Gallegos M, Fernández-Baca V, et al. Genetic diversity of clinical Pseudomonas aeruginosa isolates in a public hospital in Spain. BMC Microbiol. 2013;13(1):138. doi:10.1186/1471-2180-13-138
49. Ellappan K, Belgode Narasimha H, Kumar S. Coexistence of multidrug resistance mechanisms and virulence genes in carbapenem-resistant Pseudomonas aeruginosa strains from a tertiary care hospital in South India. J Glob Antimicrob Resist. 2018;12:37–43. doi:10.1016/j.jgar.2017.08.018
50. Al Dawodeyah HY, Obeidat N, Abu-Qatouseh LF, Shehabi AA. Antimicrobial resistance and putative virulence genes of Pseudomonas aeruginosa isolates from patients with respiratory tract infection. Germs. 2018;8(1):31–40. doi:10.18683/germs.2018.1130
51. Ohadian Moghadam S, Pourmand MR, Douraghi M, Sabzi S, Ghaffari P. Utilization of PFGE as a powerful discriminative tool for the investigation of genetic diversity among MRSA strains. Iran J Public Health. 2017;46(3):351–356.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

