Back to Archived Journals » Open Access Animal Physiology » Volume 7
Molecular characterization and phylogenetic analysis of an anti-lipopolysaccharide factor from the crucifix crab, Charybdis feriatus
Authors Sruthy K, Nair A, Cubelio SS, Bright Singh IS, Philip R
Received 1 May 2015
Accepted for publication 4 August 2015
Published 21 September 2015 Volume 2015:7 Pages 149—156
DOI https://doi.org/10.2147/OAAP.S84508
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Peter Koulen
KS Sruthy,1 Aishwarya Nair,1 Sherine Sonia Cubelio,2 IS Bright Singh,3 Rosamma Philip1
1Department of Marine Biology, Microbiology and Biochemistry, School of Marine Sciences, Cochin University of Science and Technology, 2Centre for Marine Living Resources and Ecology, Ministry of Earth Sciences, Government of India, 3National Centre for Aquatic Animal Health, Cochin University of Science and Technology, Kochi, Kerala, India
Abstract: Antimicrobial peptides are small cationic, gene-encoded, amphipathic, host defense peptides with a ubiquitous distribution in all living kingdoms. They are <10 kDa in size, with 15–100 amino acids having a net positive charge of +2 to +9. Anti-lipopolysaccharide factor (ALF) is a cationic antimicrobial peptide which constitutes one of the key effector molecules in the innate immune system of crustaceans, and is capable of binding and neutralizing lipopolysaccharides. In the present study, an ALF homolog (Charybdis feriatus [Cf]-ALF1)-encoding cDNA sequence from the hemocytes of the crucifix crab, C. feriatus, was cloned, identified, and characterized. The deduced peptide of Cf-ALF1 encoded for a 123 amino acid peptide with a 97 residue mature peptide (11.16 kDa) that had a net charge of +10. Two conserved cysteine residues and a putative lipopolysaccharide binding domain were observed in the Cf-ALF1 mature peptide. BLAST analysis of Cf-ALF1 nucleotides showed a 99% similarity to Scylla serrata. The spatial structure of Cf-ALF1 was composed of three α-helices packed against a four-strand β-sheet. Two of these helices were linked by a disulfide bond to form an amphipathic loop similar to the structure of anti-lipopolysaccharide factor isoform 3 from Penaeus monodon (ALF-Pm3). All these features suggest that Cf-ALF1 could play a significant role in the innate immune defense mechanism of C. feriatus.
Keywords: antimicrobial peptides, crab, innate immunity, Charybdis feriatus, anti-lipopolysaccharide factor
Introduction
Crustaceans represent one of the most abundant groups of animals, inhabiting both aquatic and terrestrial areas. By virtue of their diversity and abundance, they have been considered as a potential source of bioactive compounds. The lack of an adaptive immune system in crustaceans has led to the evolution of potent compounds, including antimicrobial peptides (AMPs) that are used as defense against pathogens. Anti-lipopolysaccharide factor (ALF) represents one of the evolutionarily-conserved cationic AMPs distributed among horseshoe crabs and crustaceans which can bind against lipopolysaccharides (LPS) by neutralizing the lipid-A moiety of LPS.1
The first ALF was isolated from the amebocytes of two kinds of horseshoe crab, Tachypleus tridentatus and Limulus polyphemus.2 Since then, numerous ALF homologs have been identified and characterized from decapod crustaceans, including penaeid shrimp,3–6 nonpenaeid shrimp,7 lobsters,8,9 crabs, and crayfish.10–13 ALFs belong to the group of single-domain AMPs with a signal peptide at the N-terminal region followed by a conserved LPS-binding domain.14 Different ALF isoforms and variants are either encoded by distinct genes or formed as a consequence of alternative mRNA splicing.15 Functional characterization of ALFs has been conducted by chemically synthesizing the LPS domain region,16–22 and by recombinant expression of mature peptides.9,13,23–28 Biological activity reported so far for ALFs include antibacterial, 6,9,20 antifungal,6,27 antiviral,29,30 immunomodulatory,31 antitumor,32 anti-inflammatory, and antiprotozoal activity.32,33 As per the previous studies, the structure of ALF was elucidated using nuclear magnetic resonance (NMR) spectroscopy, with the finding that it was composed of three α-helices packed against a four-stranded β-sheet.34,35
Thus far, there are several studies concerning the molecular and phylogenetic characterization, gene expression analysis, and structural and functional analysis of ALFs from decapod crustaceans and chelicerates. However, there is little research of AMPs from the crucifix crab, Charybdis feriatus. C. feriatus, a member of the family Portunidae, is one of the important species of commercial crab sold in markets across East Asia, where it commands substantially higher premium prices than Portunus spp. (typically US$8–$15 per kg). Also it is a promising aquaculture species because of its meat quality, taste and size.
The present study focused on identifying an antimicrobial peptide from the crucifix crab, C. feriatus.
Materials and methods
Animals and sample collection
Live and healthy adult crucifix crabs, C. feriatus, were collected from Cochin estuary along Vypeen, Kerala, India. To ensure survival of the crabs during their transport to the laboratory, adequate ventilation was provided. The hemolymph was extracted from the base of the crabs’ abdominal appendages using DEPC-treated, RNase-free capillary tubes that had been rinsed with precooled anticoagulant solution (RNase-free 10% sodium citrate, pH 7.0). The hemolymph was then homogenized in TRI reagent (Sigma-Aldrich Co, St Louis, MO, USA) and kept at −20°C until ready for total RNA isolation.
Total RNA isolation and first- strand cDNA synthesis
Total RNA was isolated from hemocytes using TRI reagent, as per the manufacturer’s instructions. Qualitative analysis of RNA was performed using agarose gel electrophoresis with a 0.8% agarose gel, as well as calculating the ratio of absorbance at 260 nm and 280 nm. RNA was quantified by spectrophotometry at 260 nm (Hitachi U-2900, Hitachi Ltd, Tokyo, Japan), and RNA samples with absorbance ratios (A260:A280) >1.8 were used for the present work. First-strand cDNA was generated in a 20 μL reaction volume containing 5 μg total RNA, 1× RT buffer, 2 mM dNTP, 2 mM oligo dT20, 20 U of RNase inhibitor, and 100 U of MMLV reverse transcriptase (New England Biolabs, Ipswich, MA, USA). The reaction was conducted at 42°C for 1 hour, followed by an inactivation step at 85°C for 15 minutes. Gene-specific primers β-actin-F (5′-CTTGTGGTTGACAATGGCTCCG-3′) and β-actin-R (5′-TGGTGAAGGAGTAGCCACGCTC-3′) were used to amplify the housekeeping gene β-actin (520 bp), which was used as an internal control to verify the reverse transcription polymerase chain reaction (RT-PCR).
PCR amplification and TA-Cloning
PCR amplification of 1 mL of cDNA was performed in a 25 μL reaction volume containing 1× standard Taq buffer (10 mM Tris HCl, 50 mM KCl, pH 8.3), 3.5 mM MgCl2, 200 mM dNTPs, 0.4 mM each primer, and 1 U Taq DNA polymerase (New England Biolabs, Ipswich, MA, USA). Amplification was performed using the primers, ALF-F (5′-GGACAGAAGAAACATTGAGGACGACGCA-3′) and ALF-R (5′-GGAAATCAAAAACATCCATTACAGGTCA-3′).11 The thermal profile used was 95°C for 2 minutes followed by 35 cycles of 94°C for 15 seconds, 60°C for 30 seconds, and 72°C for 30 seconds; final extension was at 72°C for 10 min. The PCR product was analyzed by electrophoresis in 1% agarose gels in TBE buffer, stained with ethidium bromide, and visualized using a gel documentation unit (Syngene, Synoptics Ltd, Cambridge, UK).
The purified PCR product was ligated into the pGEM®-T Easy Vector and transformed using DH5α-competent Escherichia coli cells, as per the manufacturer’s protocol (pGEM-T Easy TA Cloning Kit, Promega Corporation, Fitchburg, WI, USA). Transformed bacteria were cultured in Luria Bertani agar containing ampicillin, IPTG, and X-gal, at 37°C for 16 hours, and the recombinant clones with the inserts were selected by blue-white screening. Recombinant white colonies were selected and streaked onto fresh ampicillin plates, and screened by colony PCR using vector-specific primers (T7 F and Sp6 R) and the ALF sequence-specific primers. For vector-specific primers, the thermal profile used was 95°C for 3 minutes followed by 35 cycles of 94°C for 15 seconds, 57°C for 30 seconds, and 72°C for 30 seconds; final extension was at 72°C for 10 minutes. Amplicons were examined on 1% agarose gels in TBE buffer, stained with ethidium bromide, and visualized using a gel documentation unit. The potentially positive recombinant clones were selected for plasmid isolation as per the manufacturer’s protocol (GeneluteHp Plasmid Miniprep Kit Sigma-Aldrich Co, St Louis, MO, USA). Isolated plasmids were analyzed on a 0.8% agarose gel, and gene insertion was confirmed by PCR, using vector-specific and insert-specific primers. Selected recombinant plasmids were sequenced using T7 F and Sp6 R primers with the ABI PRISM® BigDye™ Terminator Cycle Sequencing Ready Reaction Kit on an ABI PRISM 377 DNA Sequencer (Thermo Fisher Scientific, Waltham, MA, USA) at SciGenom Sequencing Facility, India.
Characterization of nucleotide and amino acid sequences
GeneTool software (Syngene, Synoptics Ltd, Cambridge, UK) was used for the nucleotide sequence analysis and assembly. The cDNA sequences acquired were translated using the Expert Protein Analysis System (ExPASy) (http://web.expasy.org/tools/translate/) to obtain their amino acid sequences. Homology searches of nucleotide and deduced amino acid sequences were accomplished using the BLASTn and BLASTp algorithms of the NCBI (http://www.ncbi.nlm.nih.gov/blast). Presumed signal peptide region was identified using the online program SignalP (http://www.cbs.dtu.dk/services/SignalP). The transmembrane region within the mature peptide was predicted using the TMpred tool of ExPASy (http://embnet.vital-it.ch/software/TMPRED_form.html). Physicochemical parameters of the protein sequence were predicted using the online protein analysis tool, ExPASy ProtParam (http://web.expasy.org/protparam).
Phylogenetic analysis of C. feriatus-ALF1
Multiple sequence alignment of C. feriatus (Cf)-ALF1’s amino acid sequence was performed on ClustalW in BioEdit software (http://www.mbio.ncsu.edu/bioedit/bioedit.html) for the comparison and alignment with other crustacean ALF-encoding sequences. The phylogenetic tree was constructed based on the neighbor-joining method, using Molecular Evolutionary Genetics Analysis (http://www.megasoftware.net/) version 6.
Structure analysis of Cf-ALF1
The spatial structure of MnALF was established with PyMOL (https://www.pymol.org/), using the SWISS-MODE (http://swissmodel.expasy.org/) prediction algorithm, which is based on homology modeling. The helical property of Cf-ALF1 was predicted using HeliQuest (http://heliquest.ipmc.cnrs.fr/cgi-bin/ComputParamsV2.py).36 The cDNA sequence of Cf-ALF1 was converted to the corresponding RNA sequence (Attotron Biosensor Corporation, http://www.attotron.com/) and submitted to RNAStructure WebServer (http://rna.urmc.rochester.edu/RNAstructureWeb/Servers/Predict1/Predict1.html) to envisage the RNA structure with minimum free energy.
Prediction of Cf-ALF1 antimicrobial peptide properties
The antimicrobial activity of Cf-ALF1 was predicted using the Antimicrobial Peptide Database’s (APD2) calculation and prediction tool (http://aps.unmc.edu/AP/main.php).37 The amphipathicity and antimicrobial activity of the LPS domain was interpreted using HeliQuest.
Results and discussion
In the present study, an AMP belonging to the ALF family (herein referred to as Cf-ALF1) was identified and characterized from the hemocytes of C. feriatus. A 524 bp fragment of cDNA encoding an ORF of 123 amino acids was obtained from the hemocytes of C. feriatus (Figure 1). The Cf-ALF1 nucleotide and deduced amino acid sequences were deposited in the GenBank sequence database (GenBank ID: KP688577). BLAST analysis of the 372 bp of cDNA sequence, as well as the homology search by BLASTn and BLASTp of the deduced amino acid sequence of Cf-ALF1, revealed that it belongs to the DUF3254 superfamily. This superfamily includes all the family of ALFs. BLASTn analysis of the Cf-ALF1 nucleotide sequence showed that it exhibited a 99% similarity to an ALF isoform from Scylla serrata (SsALF) (GenBank ID: ACH87655.1), a 97% similarity to an S. paramamosain ALF (GenBank ID: EF207786.1), a 96% similarity to a P. pelagicus ALF (GenBank ID: JQ899452.1), an 87% similarity to a P. trituberculatus ALF (GenBank ID: GQ165621.2), and a 72% similarity to an Eriocheir sinensis ALF (GenBank ID: HQ850572.1). SignalP analysis of the peptides determined the existence of a signal peptide composed of 26 amino acids and containing an exceedingly hydrophobic core region. The presence of a putative cleavage site after the 26th amino acid (CEA26-Q27Y) demarcated the highly cationic mature peptide of 97 amino acid residues.
Based on the ExPASy ProtParam analyses, Cf-ALF1 had an estimated molecular weight of 14.025 kDa, a net charge of +11, and a theoretical isoelectric point of 9.78. Comparatively, the mature peptide region measured 11.172 kDa, +10, and 10.01, respectively. Cationicity of the mature peptide was mainly attributed to the 18 positive amino acid residues (twelve lysine residues and six arginine residues), against eight negative residues (four aspartic acid residues and four glutamic acid residues). The estimated half-life of the mature peptide of Cf-ALF1 was found to be 0.8 hours (mammalian reticulocytes, in vitro), 10 minutes (yeast, in vivo), and 10 hours (E. coli, in vivo). The predicted instability index was computed to be 34.37, and thus classified the protein as stable. The presence of a transmembrane helical region was determined in the mature peptide region, starting from L43SGLWHSDTVDFMGHT59, which may assist in the peptide and bacterial membrane interaction.
The multiple protein sequence alignment of Cf-ALF1 (with representatives of each crustacean’s ALFs) revealed the existence of conserved sequence features (Figure 2), ie, cysteine residues (C55– C76) that established a disulfide bridge with a highly conserved sequence. This sequence contained a cluster of seven positively charged amino acid residues within the disulfide loop, forming the functional LPS domain of ALF, which is mandatory for LPS binding and neutralization.34 Sequence alignment also revealed that the amino acid sequence of the signal peptide was not conserved across different groups, but that of the mature peptide region showed a conserved pattern of the amino acid motif K71GR/KM/FWCPGW79 straddling the LPS domain; this domain interacts with microbial polysaccharides. The variation of sequences in the mature peptide leads to the functional divergence of ALF among different crustacean groups, and accounts for the differences in sequence characteristics such as theoretical isoelectric point, hydrophobicity, and net charge. In the case of Cf-ALF1, it differs from SsALF by only one amino acid residue in the signal peptide, whereas the mature peptide region was found to be highly conserved.
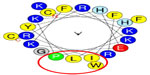
The level of antimicrobial activity of Cf-ALF1 predicted by Antimicrobial Peptide Database 2 revealed Cf-ALF1 as a potent AMP with a protein-binding potential (Boman index) and Wimley–White whole-residue hydrophobicity of 2.1 kcal/mol and 15.8 kcal/mol, respectively. In silico analysis of the anticipated LPS binding domain of Cf-ALF1 (C55HIRRKPKFRKFKLYHEGKFWC76) based on amphipathicity, using HeliQuest, showed that Cf-ALF1 is composed of 22 amino acids. Polar residues comprised 45.45% of these amino acids, with the remaining amino acids being nonpolar residues, thus establishing a hydrophobic face with IWLP residues and a hydrophobicity of 0.342 H with a +7 net charge. Such properties could assist in the binding of the LPS domain with the highly negatively-charged lipid A moiety of LPS, and therefore confirm the antimicrobial activity (Figure 3).
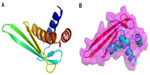
Recombinant expression of SsALF has previously been carried out using a bacterial expression system, and it showed a stronger inhibition of Gram-negative bacterial growth than that of Gram-positive bacteria.21 Using homology modeling with the template recombinant-ALF-Pm3 (SMTL id: 2job.1) NMR structure (chosen for its shared similarity of 38.61%), prediction of the spatial 3D structure of Cf-ALF1 demonstrated the presence of three α-helices packed against four strands of β-sheets (Figure 4). Somboonwiwat et al studied and elucidated the functional role of recombinant-ALF-Pm3 expressed in the Pichia pastoris system, and demonstrated a comprehensive spectrum of activity against the filamentous fungi, and both Gram-positive and Gram-negative bacteria.25 In the current study, the predicted RNA sequence based on the Cf-ALF1 cDNA sequence, as well as its ideal secondary structure with minimum free energy, was found to consist of both double-stranded and looped regions (Figure 5).
Analysis of the phylogenetic tree constructed using the neighbor-joining method clearly showed that ALF sequences were clustered according to species (Figure 6). The phylogenetic tree revealed that ALFs including Cf-ALF1 and SsALF shared a common ancestral origin and could have subsequently diverged in the course of evolution. The ALF family of AMPs were found to be evolutionarily conserved, with a diversity of ALF isoforms exhibiting the same physicochemical properties that formed one of the main AMPs involved in the removal of pathogenic microorganisms. The phylogenetic tree could be divided into three groups: Group I consisted of shrimp ALFs, including both penaeid and nonpenaeid shrimp ALFs; Group II comprised all crustacean ALFs; and Group III included crab ALFs. Cf-ALF1 was grouped along with SsALF and other portunid crab ALFs in Group III.
Conclusion
The present study reports the identification, cloning, molecular, and phylogenetic characterization of an isoform of ALF from the crucifix crab, C. feriatus, exhibiting an 100% identity to SsALF. The occurrence of sequence variation mainly in the mature peptide could be an important factor that has led to the functional divergence of ALF isoforms from limulids to decapod crustaceans. C. feriatus is now an emerging species in the aquaculture market, due to the lower quantities in wild catch and the requirement to meet the demands of the market. Also, C. feriatus has been suggested as one of the appropriate species for stock enhancement and aquaculture in the international forum on the culture of portunid crabs.38
To date, there is little research regarding the physiological processes involved in the immune system or the host defense peptides from C. feriatus. As there are currently restrictions in antibiotic usage due to the emergence of multidrug resistant strains, the present study is of great significance, particularly in the aquaculture system because C. feriatus has shown its ability to combat pathogenic microorganisms. Since recombinant expression of SsALF has already been carried out using a bacterial expression system,27 further work with Cf-ALF1 could include heterologous expression with post-translational modification, using a eukaryotic system such as Baculovirus-mediated insect cell line expression, and characterization of biological activity. These approaches would shed light on the possibilities of the Cf-ALF1 peptide as a therapeutic candidate in aquaculture and medicine. Bioprospecting for AMPs and other host defense molecules from potential aquaculture species would contribute to the understanding of crab immunology, and their potential role in prevention of microbial diseases in wild and domestic animals.
Acknowledgments
The authors are grateful to the Director of the Centre for Marine Living Resources and Ecology (CMLRE), and the Ministry of Earth Sciences (MoES), Govt. of India for the research grant (MoES/CMLRE/10-MLR-TD/12/2012) and scientific support for the work. Authors would also like to thank the Coordinator of the National Centre for Aquatic and Animal Health (NCAAH) for scientific support. KS Sruthy gratefully acknowledges Kerala State Council for Science, Technology and Environment (KSCSTE) for the Award of Fellowship.
Disclosure
The authors report no conflicts of interest in this work.
References
Ried C, Wahl C, Miethke T, et al. High affinity endotoxin-binding and neutralizing peptides based on the crystal structure of recombinant Limulus anti-lipopolysaccharide factor. J Biol Chem. 1996;271(45):28120–28127. | |
Tanaka S, Nakamura T, Morita T, Iwanaga S. Limulus anti-LPS factor: an anticoagulant which inhibits the endotoxin mediated activation of Limulus coagulation system. Biochem Biophys Res Commun. 1982;105(2):717–723. | |
Gross PS, Bartlett TC, Browdy CL, Chapman RW, Warr GW. Immune gene discovery by expressed sequence tag analysis of hemocytes and hepatopancreas in the Pacific White Shrimp, Litopenaeus vannamei, and the Atlantic White Shrimp, L. setiferus. Dev Comp Immunol. 2001; 25(7):565–577. | |
Supungul P, Klinbunga S, Pichyangkura R, Hirono I, Aoki T, Tassanakajon A. Antimicrobial peptides discovered in the black tiger shrimp Penaeus monodon using the EST approach. Dis Aquat Organ. 2004;61(1–2):123–135. | |
Liu F, Liu Y, Li F, Dong B, Xiang J. Molecular cloning and expression profile of putative antilipopolysaccharide factor in Chinese shrimp (Fenneropenaeus chinensis). Mar Biotechnol (N Y). 2005;7(6):600–608. | |
de la Vega E, O’Leary NA, Shockey JE, et al. Anti-lipopolysaccharide factor in Litopenaeus vannamei (LvALF): a broad spectrum antimicrobial peptide essential for shrimp immunity against bacterial and fungal infection. Mol Immunol. 2008;45(7):1916–1925. | |
Lu KY, Sung HJ, Liu CL, Sung HH. Differentially enhanced gene expression in hemocytes from Macrobrachium rosenbergii challenged in vivo with lipopolysaccharide. J Invertebr Pathol. 2009; 100(1):9–15. | |
Beale KM, Towle DW, Jayasundara N, et al. Anti-lipopolysaccharide factors in the American lobster Homarus americanus: molecular characterization and transcriptional response to Vibrio fluvialis challenge. Comp Biochem Physiol Part D Genomics Proteomics. 2008;3(4):263–269. | |
Zhang Y, Wang L, Wang L, et al. The second anti-lipopolysaccharide factor (EsALF-2) with antimicrobial activity from Eriocheir sinensis. Dev Comp Immunol. 2010;34(9):945–952. | |
Imjongjirak C, Amparyup P, Tassanakajon A, Sittipraneed S. Antilipopolysaccharide factor (ALF) of mud crab Scylla paramamosain: molecular cloning, genomic organization and the antimicrobial activity of its synthetic LPS binding domain. Mol Immunol. 2007;44(12):3195–3203. | |
Afsal VV, Antony SP, Sathyan N, Philip R. Molecular characterization and phylogenetic analysis of two antimicrobial peptides: Anti-lipopolysaccharide factor and crustin from the brown mud crab, Scylla serrata. Results Immunol. 2011;1(1):6–10. | |
Afsal VV, Antony SP, Sanjeevan VN, Anil Kumar PR, Singh ISB, Philip R. A new isoform of anti-lipopolysaccharide factor identified from the blue swimmer crab, Portunus pelagicus: Molecular characteristics and phylogeny. Aquaculture. 2012;356–357:119–122. | |
Sun C, Xu WT, Zhang HW, et al. An anti-lipopolysaccharide factor from red swamp crayfish, Procambarus clarkii, exhibited antimicrobial activities in vitro and in vivo. Fish Shellfish Immunol. 2011;30(1):295–303. | |
Rosa RD, Barracco MA. Antimicrobial peptides in crustaceans. Invertebr Surv J. 2010;7:262–284. | |
Tharntada S, Somboonwiwat K, Rimphanitchayakit V, Tassanakajon A. Anti-lipopolysaccharide factors from the black tiger shrimp, Penaeus monodon, are encoded by two genomic loci. Fish Shellfish Immunol. 2008;24(1):46–54. | |
Arockiaraj J, Kumaresan V, Bhatt P, et al. A novel single-domain peptide, anti-LPS factor from prawn: synthesis of peptide, antimicrobial properties and complete molecular characterization. Peptides. 2013;53:79–88. | |
Imjongjirak C, Amparyup P, Tassanakajon A. Molecular cloning, genomic organization and antibacterial activity of a second isoform of antilipopolysaccharide factor (ALF) from the mud crab, Scylla paramamosain. Fish Shellfish Immunol. 2011;30(1):58–66. | |
Li S, Guo S, Li F, Xiang J. Characterization and function analysis of an anti-lipopolysaccharide factor (ALF) from the Chinese shrimp Fenneropenaeus chinensis. Dev Comp Immunol. 2014;46(2):349–355. | |
Nagoshi H, Inagawa H, Morii K, et al. Cloning and characterization of a LPS-regulatory gene having an LPS binding domain in kuruma prawn Marsupenaeus japonicus. Mol Immunol. 2006;43(13); 2061–2069. | |
Pan C, Chao T, Chen J, et al. Shrimp (Penaeus monodon) anti-lipopolysaccharide factor reduces the lethality of Pseudomonas aeruginosa sepsis in mice. Int Immunopharmacol. 2007;7(5):687–700. | |
Ren J, Gao H, Tang M, Gu J, Xia P, Xiao G. Lipopolysaccharide (LPS) detoxification of analogue peptides derived from limulus anti-LPS factor. Peptides. 2010;31(10):1853–1859. | |
Vallespi MG, Glaria LA, Reyes O, Garay HE, Ferrero J, Araña MJ. A Limulus Antilipopolysaccharide factor-derived peptide exhibits a new immunological activity with potential applicability in infectious diseases. Clin Diagn Lab Immunol. 2000;7(4):669–675. | |
Liu H, Chen R, Zhang, Q, et al. Characterization of two isoforms of antilipopolysacchride factors (Sp-ALFs) from the mud crab Scylla paramamosain. Fish Shellfish Immunol. 2012;33(1):1–10. | |
Liu Y, Cui Z, Li X, Song C, Li Q, Wang S. Molecular cloning, expression pattern and antimicrobial activity of a new isoform of anti-lipopolysaccharide factor from the swimming crab Portunus trituberculatus. Fish Shellfish Immunol. 2012;33(1):85–91. | |
Somboonwiwat K, Marcos M, Tassanakajon A, et al. Recombinant expression and anti-microbial activity of anti-lipopolysaccharide factor (ALF) from the black tiger shrimp Penaeus monodon. Dev Comp Immunol. 2005;29(10):841–851. | |
Sun W, Wan W, Zhu S, et al. Characterization of a novel anti- lipopolysaccharide factor isoform (SpALF5) in mud crab, Scylla paramamosain. Mol Immunol. 2015;64(2):262–275. | |
Yedery RD, Reddy KV. Identification, cloning, characterization and recombinant expression of an anti-lipopolysaccharide factor from the hemocytes of Indian mud crab, Scylla serrata. Fish Shellfish Immunol. 2009;27(2):275–284. | |
Zhu L, Lan JF, Huang YQ, et al. SpALF4: a newly identified anti-lipopolysaccharide factor from the mud crab Scylla paramamosain with broad spectrum antimicrobial activity. Fish Shellfish Immunol. 2014;36(1):172–180. | |
Liu H, Jiravanichpaisal P, Söderhäll I, Cerenius L, Söderhäll K. Antilipopolysaccharide factor interferes with white spot syndrome virus replication in vitro and in vivo in the crayfish Pacifastacus leniusculus. J Virol. 2006;80(21):10365–10371. | |
Antony SP, Philip R, Joseph V, Singh ISB. Anti-lipopolysaccharide factor and Crustin-III, the anti-white spot virus peptides in Penaeus monodon: Control of viral infection by up-regulation. Aquaculture. 2011;319(1–2):11–17. | |
Montero T, Vallespi MG, Garay H, Reyes O, Aran MJ. A Limulus anti-LPS factor-derived peptide modulates cytokine gene expression and promotes resolution of bacterial acute infection in mice. Int Immunopharmacol. 2003;3(2):247–256. | |
Lin MC, Pan CY, Hui CF, Chen JY, Wu JL. Shrimp anti-lipopolysaccharide factor (SALF), an antimicrobial peptide, inhibits proinflammatory cytokine expressions through the MAPK and NF-κB pathways in LPS-induced HeLa cells. Peptides. 2013;40:42–48. | |
Pan CY, Chen JY, Lin TL, Lin CH. In vitro activities of three synthetic peptides derived from epinecidin-1 and an anti-lipopolysaccharide factor against Propionibacterium acnes, Candida albicans, and Trichomonas vaginalis. Peptides. 2009;30(6):1058–1068. | |
Hoess A, Watson S, Siber GR, Liddington R. Crystal structure of an endotoxin-neutralizing protein from the horseshoe crab, Limulus anti-LPS factor, at 1.5 A resolution. EMBO J. 1993;12(9):3351–3356. | |
Yang Y, Boze H, Chemardin P, et al. NMR structure of rALF-Pm3, an anti-lipopolysaccharide factor from shrimp: model of the possible lipid A-binding site. Biopolymers. 2009;91(3):207–220. | |
Gautier R, Douguet D, Antonny B, Drin G. HELIQUEST: a web server to screen sequences with specific alpha-helical properties. Bioinformatics. 2008;24(18):2101–2102. | |
Wang G, Li X, Wang Z. APD2: the updated antimicrobial peptide database and its application in peptide design. Nucleic Acids Res. 2009;37(Database issue):D933–D937. | |
Williams MJ, Primavera JH. Choosing tropical portunid species for culture, domestication and stock enhancement in the Indo-Pacific. Asian Fish Sci. 2001;14:121–142. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.