Back to Journals » OncoTargets and Therapy » Volume 13
miR-30d Induced Apoptosis by Targeting Sox4 to Inhibit the Proliferation, Invasion and Migration of Nephroblastoma
Received 28 February 2020
Accepted for publication 15 May 2020
Published 27 July 2020 Volume 2020:13 Pages 7177—7188
DOI https://doi.org/10.2147/OTT.S251714
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Nicola Silvestris
This paper has been retracted.
Shi Zong, 1,* Jia Zhao, 2,* Ling Liu 3
1Department of Urology Surgery, China-Japan Union Hospital of Jilin University, Changchun 130000, Jilin, People’s Republic of China; 2Department of Anesthesia, China-Japan Union Hospital of Jilin University, Changchun 130000, Jilin, People’s Republic of China; 3Department of Pediatrics, China-Japan Union Hospital of Jilin University, Changchun 130000, Jilin, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Ling Liu Email [email protected]
Background: Wilms tumor (WT) is an embryonic malignant tumor, and its related mechanism is still unclear. microRNA (miR), as a short-chain non-coding RNA, has low expression in various tumors. In this study, WT differential miR was screened by multi-chip in GEO database and its mechanism was explored to provide potential therapeutic targets and ideas for clinic.
Methods: We logged into GEO database and downloaded GSE57370 and GSE48137 chip matrix files to analyze potential differences in miR. TargetScan, miRDB, miRTarBase and starBase were applied to predict the target genes of miR with significant differences. qRT-PCR was applied to determine the expression of miR-30d and Sox4 in WT tissue and cell line (G401). The interaction of miR-30d with Sox4 was confirmed by qRT-PCR, Western blot and luciferase assay, respectively. CCK-8, Transwell and flow cytometry were applied to determine the proliferation, invasion, migration and apoptosis of cells.
Results: We found that miR-30d was low expressed in two chips. qRT-PCR showed that miR-30d was down-regulated and SOX4 was up-regulated in WT tissues and cells. The online target gene prediction software showed there was a targeted binding site between Sox4 and miR-30d. Sox4 was negatively controlled by miR-30d. Subsequent studies found that over-expression of miR-30d inhibited the proliferation, invasion, migration and induced apoptosis of C64 and WiT49 cells. In addition, Sox4 could reverse the proliferation, invasion and migration of C64 and WiT49 induced by miR-30d and induce apoptosis.
Conclusion: miR-30d is poorly expressed in WT and can induce apoptosis and inhibit proliferation, invasion and migration by mediating Sox4.
Keywords: GEO, miR-30d, Sox4, nephroblastoma, proliferation, invasion
Introduction
Wilms tumor (WT), also namely nephroblastoma, is the most common solid carcinoma in children.1 WT originates from embryo, grows in renal parenchyma, deforms during growth and invades surrounding renal tissue.2 The data show that there is one WT child in every 100,000 children, and the peak incidence of WT is among 1–5 years old.3,4 Although the incidence rate of WT is low, it is easy to metastasize after illness, which is harmful to children. At present, clinical combined therapy can improve the prognosis of most children, but data show that 25% of children still cannot be effectively treated, and some children are prone to relapse and metastasis.5 Therefore, we urgently need to understand the occurrence and development of WT and find potential therapeutic targets for WT, so as to provide ideas for clinical practice.
In recent years, non-coding RNA has been a hot research field, in which micro RNA (miRNA/miRs) is participated in the occurrence and growth of many diseases.6,7 miR is an endogenous short-chain non-coding RNA, which is highly conservative and has a length of 21–23 nucleotides. Studies have shown that miR does not encode proteins, but it can regulate the transcription of target genes and inhibit their expression through the 3-quality untranslated region (3ʹ-UTR).8,9 Now there are many studies which can confirm that miR is participated in many cell events, namely cell proliferation, differentiation and apoptosis.10,11 In addition, studies found that miR participated in WT progress, which is a potential target for WT treatment.12,13 For example, miR-200c-3p targeting FRS2 controlled the proliferation, migration and invasion of nephroblastoma cells.14 miR−429 targeting c-myc regulated the proliferation and apoptosis of nephroblastoma cells in studies by Wang et al.15 Therefore, we hope to further explore the mechanism of miR and WT by finding out the potential differential miR in WT through selection.
With the continuous improvement of second-generation sequencing, more and more databases have been established. As one of the largest public databases in the world, GEO database contains gene chips for various diseases.16 In this study, we screened miR differential gene chip in WT through GEO database, and explored the related mechanisms to provide targets for clinical treatment.
Methods and Data
GEO Database Data Analysis
We logged into GEO database (https://www.ncbi.nlm.nih.gov/gds) to search Wilms tumor and microRNA related chips. Finally, we selected GSE57370 and GSE48137 chips, downloaded matrix file Series Matrix File (s) and then performed log2 conversion on the data. The processed data were analyzed for differences using limma package, and thermography and volcano plot were visualized. The screening criteria were log Fold Change=1, P < 0.05. miRs with differences were collected and jvenn visualizing Venn diagram was used to screen common differential miRs.
Clinical Data Collection
We collected 30 WT children admitted to our hospital from May 2014 to May 2018 and cancer tissues and adjacent tissues of the 30 children. The collected tissues were immediately sent to the laboratory for testing through liquid nitrogen transportation at −80°C. The survival time of the patients admitted to this study was expected to be more than 1 month. All the children have signed informed consent (signed by the children’s families). This study has been approved by the Ethics Committee of the China-Japan Union Hospital of Jilin University. All the tests conformed to the principles of Declaration of Helsinki of the World Medical Association.17
Cell Culture
Human WT cells HCM-BROD-0051-C64 (C64), G-401, SK-NEP-1, WiT49 and HEK-293T were collected from ATCC (Rockville, Maryland, USA) and then placed in Dulbecco’s Modified Eagle Medium (DMEM, Gibco, Rockville, MD, USA) including 10% FBS (Gibco, Rockville, Maryland, USA). A total of 100U/mL penicillin and 50 mg/mL streptomycin (Corning, New York, USA) were incubated at 37°C in a humid atmosphere with 5%CO2.
Cell Transfection
miR-30d mimetics, inhibitors (miR-30d-inhibit) and reference material (miR-NC) were designed and synthesized by GenePharma. Sox4 specific siRNA (si-Sox4) and siRNA negative pair (si-NC) were collected from Santa Cruz (USA). For over-expression of Sox4, the full-length Sox4 sequence was transfected into pcDNA-3.1 vector (ThermoFisher), called pcDNA-Sox4. pcDNA-3.1 was used as a blank control. According to the manufacturer’s plan, all the above steps were performed using Lipofectamine 3000 reagent (Life Technologies, Carlsbad, California, USA). Stable transfected C64 and WiT49 applied a medium including 0.5mg/mL G418 (Sigma-Aldrich, St. Louis, Missouri, USA) for subsequent experiments.
Cell Proliferation Detection
In this study, the proliferation was determined by Cell Counting Kit (CCK-8, Dojindo Laboratories, Japan). Transfected cells were collected and resuspended. The cells were adjusted to 1×104 cells per well, inoculated in 96-well plate, and incubated at 37°C for 24, 48, 72 and 96 h. After the incubation, CCK-8 solution was put into each well, and then the cells were placed at 37°C for 1.5h. Subsequently, the absorbance (450 nm) of each well was evaluated using an microplate reader.
Cell Invasion and Migration Detection
Transwell method was used to detect cell invasion and migration in this study. The cell invasion experiment was as follows: Matrigel was added to serum-free medium at 4°C to adjust its concentration to 1 mg/mL, and 50μL diluted media was put into the upper chamber and cultured at 37°C for 30min. Complete media including 15% FBS was put into lower chamber. The cells (5×104 cells/well) were put into the upper chamber, placed at 37°C. The 95% absolute ethyl alcohol was taken out from the upper chamber for fixation after incubation for 24h. The 0.1% crystal violet was used for dyeing at room temperature for 20min, and the number of cell invasions was observed using optical microscope. The cell migration experiment was as follows: Dulbecco’s modified Eagle medium (DMEM) with 15%FBS was put into the lower chamber of Transwell as induction. Cells were adjusted to 5×104 cells/well, put into the upper chamber, and cultured at 37°C. After 24h, the upper chamber was taken out, fixed with 95% anhydrous ethanol, dyed with 0.1% crystal violet for 20min, and the cell migration number was observed under optical microscope.
Detection of Apoptosis
The transfected cells were detected for apoptosis by Annexin V-PE cell apoptosis detection kit (South Biotech, USA). Cells in the media were rinsed with PBS, fixed with 70% ethanol, then washed twice and dyed with Annexin V labeled with propidium iodide/fluorescein isothiocyanate at 50 μ g/MLRNSEA. The samples were placed in dark at room temperature for 1 hour. Apoptosis (Annexin-V positive and PI negative) were determined using FACSCan (BD Biosciences, USA).
Double Fluorescein Report
The following two pmiR-RB-REPORT TM vectors were synthesized: Sox4 3ʹ-UTR with the putative target site of miR-30d (sox4 wt-3ʹ-utr); Sox4 3ʹ-UTR with mutation binding site (sox4 mut-3ʹ-UTR); 100ng of carrier (Sox4 WT-3ʹ-UTR, Sox4 Mut-3ʹ-UTR), miR-30d-mimics/inhibit and miR-NC (50 nM/well) were used together with Lipofectamine PTEN® 2000 reagent (Invitrogen; Thermo Fisher Scientific) and transfected into 293T and SW1353. Analog control and Sox4 Mut-3ʹUTR were applied as NC. Fluorescein activity was tested by a dual luciferase reporting kit (Promega, Madison, WI, USA).
RNA Extraction and Real-Time qPCR (RT-qPCR)
Total RNA was extracted with TRIzol reagent (Takara Bio, Inc.). The concentration and integrity of the extracted total RNA were tested. cDNA was used in accordance with the manufacturer’s plan. The production of Premix Ex Taq™II (TaKaRa) was carried out by executing SYBR PrimeScript analysis RT Kit (TaKaRa) synthesis ® referring to the manufacturer’s agreement. The reaction system and conditions were configured according to the kit instructions. A total of 40 cycles were performed. GAPDH (for mRNA) and U6 (for miRNA) were applied as internal references. The relative expression level was determined by 2− ΔΔCq method.18 All the above steps were repeated three times. miR-30d upstream primer: 5ʹ-GCCTGTAAACATCCCCGAC-3ʹ,downstream primer: 5ʹ-GTGCGTGTCGTGGAGTCG-3ʹ; Sox4 upstream primer: 5ʹ-GACCTGCTCGACCTGAACC-3ʹ, downstream primer: 5ʹ-CCGGGCTCGAAGTTAAAATCC-3ʹ; GAPDH upstream primer: 5ʹ-CTGCCAACG TGTCAGTGGTG-3ʹ, downstream primer: 5ʹ-TCAGTGTAGCCCAGGATGCC-3ʹ; U6 upstream primer: 5ʹ-GCTTCGGCAGCACATATACTAAAAT-3ʹ, downstream primer: 5ʹ-CGCTTCACGAATTTGCGTGTCAT-3ʹ
Western Blot Detection
RIPA buffer was used to extract the total protein of cells, and BCA (Thermo, PA, USA) was used to prepare and measure the total protein of cells. The total protein was separated on 12% SDS-PAGE, moved to PVDF membrane, and then sealed with dry milk and immunostained with primary antibody Sox4 and GAPDH at 4°C for a night at −4°C. After incubation with secondary antibody, the signal was visualized by chemiluminescence detection system (Pierce, Thermo, PA, USA).
Subcutaneous Tumor Model
Ten BALB/c-nu nude mice (5–6 weeks old, 18–20 g) were collected from Charles River (Beijing, China). All mice were subcutaneously inoculated with WiT49 cells transfected with miR-30d-mimics and miR-NC (5×106 cells per mouse) on the left dorsal side, and cultured for 28 days. The tumor volume of nude mice was detected every 7 days. The formula was (length× width2)/2(mm3). Animals were euthanized 28 days later, and the tumor was removed and weighed. This study was approved by the Animal Ethics Committee and performed referring to the Guide for the Care and Use of Laboratory Animals of USA National Institutes of health.
Statistical Analysis
GraphPad 8 was used to analyze the data, and independent sample t-test was used for inter-group comparison. The counting data were expressed by percentage (%). Chi-square test was used and expressed by χ2. One-way ANOVA was applied for multiple groups comparison, which was expressed by F. LSD-t-test was used for post-event comparison, repeated measurement ANOVA was used for expression at multiple time points, which was expressed as F. Bonferroni was used for back testing and Pearson test for analysis of correlation of genes, and when P < 0.05, there were statistical differences.
Results
WT GEO Chip Differential miR Analysis
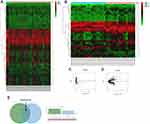
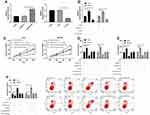
We analyzed the differential miR of GSE57370 and GSE48137 chips, respectively. As a result, we found that there were 50 differential miRs in GSE57370 chips, including 5 high expression, 45 low expression (See Table 1, Figure 1A and C). Among them, there were 31 differential miRs in GSE48137 chips, including 20 high expression and 11 low expression (See Table 2, Figure 1B and D). We found through Venn diagram analysis that there were two intersecting miRs in the two chips, namely miR-30d and miR-1973 (See Figure 1E). We searched the relevant literature and there is no research to prove that miR-1973 is expressed in any sample, so we chose miR-30d for further research in combination with the existing literature.
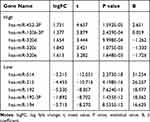 |
Table 1 miR with Top 5 Differences of High and Low Expression in GSE57370 Chip |
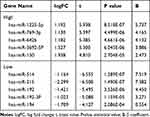 |
Table 2 miR with Top 5 Differences of High and Low Expression in GSE48137 Chip |
Clinical Value of miR-30d in WT Children
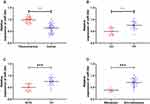
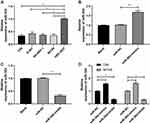
In order to determine the expression of miR-30d in WT children, miR-30d in tumor tissues of children with WT was tested by RT-qPCR. It was indicated that miR-30d was low expressed in tissues of WT children (See Figure 2A). Furthermore, we explored the correlation of miR-30d with clinical data of children and found the increasing probability of UH (See Figure 2B), NWTS high staging (See Figure 2C) and lymphatic metastasis (See Figure 2D) in children with miR-30d low expression. This suggested that miR-30d may be a potential clinical observation indicator of WT (See Table 3).
 |
Table 3 Relationship Between miR-30d and WT Clinical Data |
Up-Regulation of miR-30d Could Inhibit the Growth and Metastasis of WT Cells
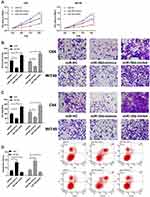
In this study, the expression of miR-30d in WT cells was tested by RT-qPCR, and it was found that miR-30d down-regulated in WT cells (Figure 3A). C64 and WiT49 cells were selected for transfection respectively, and miR-30d-mimics/inhibit cell lines were constructed to explore the role of miR-30d on WT cells (Figure 3B–D). Further observation of CCK-8 experiment found that the proliferation ability of WT cells transfected with miR-30d-mimics was significantly inhibited (Figure 4A). Transwell experiment found that the invasion ability and migration ability were significantly inhibited (Figure 4B and C). Flow cytometry found that it further induced apoptosis, but the above results of it were reversed (Figure 4D). In addition, we observed through subcutaneous tumor model that the tumor volume and mass of nude mice injected with miR-30d-mimics were significantly reduced compared with those transfected with miR-NC (Figure 5A and B). This showed that miR-30d is a potential target for WT treatment.
miR-30d Could Target Sox4
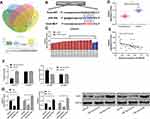
miR regulated downstream target genes, which has become a classic approach. In order to further explore the in-depth mechanism of miR-30d, we predicted its target genes through online miR prediction websites of TargetScan,19 miRDB,20 miRTarBase,21 and starBase,22 respectively (See Figure 6A). As a result, we found targeted binding sites between Sox4 and miR-30d (See Figure 6B). In order to explore the expression of Sox4 in WT tumor, we searched relevant chips through GEO database and found that Sox4 was highly expressed in WT tumor tissue in GSE66405 (See Figure 6C). For further determining the expression of Sox4 in WT children, we found that the expression of Sox4 in WT children’s tissues up-regulated significantly (See Figure 6D), and further found that the expression of Sox4 in WT children’s tissues was negatively correlated with miR-30d (See Figure 6E), which suggested that miR-30d and Sox4 had targeted relationship. For verifying the relationship between the two, we found that miR-30d-mimics could inhibit Sox4-WT fluorescence activity through double luciferase report detection, while Sox4-WT fluorescence activity was inhibited after co-transfection of miR-30d-inhibit (See Figure 6F), and Sox4 mRNA and protein in transfected cells increased and decreased in different degrees through detection of WT cells transfected with miR-30d-mimics/inhibit (See Figure 6G), which suggested that miR-30d could target Sox4 and might be participated in the occurrence and development of WT.
miR-30d Participated in WT Growth and Transfer by Regulating Sox4
At the end of the study, in order to determine the targeting of miR-30d to Sox4, we co-transfected miR-30d-mimics, pcDNA-Sox4 (See Figure 7A), miR-30d-inhibit, si-Sox4 (See Figure 7B), and further observed the growth and metastasis. After knocking-down, the proliferation (See Figure 7C), invasion (See Figure 7D), and migration (See Figure 7E) of Sox4 WT cells were significantly inhibited, and apoptosis was induced (See Figure 7F), while the above results of up-regulation of Sox4 were reversed. Moreover, after co-transfection of miR-30d-mimics, pcDNA-Sox4, miR-30d-inhibit, and si-Sox4 respectively, the cell proliferation, invasion, migration, and apoptosis were not different from those of miR-NC+si-NC, suggesting that miR-30d participated in WT growth and metastasis by regulating Sox4.
Discussion
WT is a multiple solid tumor in children. More and more studies have proved that miR are closely related to WT.23 Abnormal expression of miR has a close correlation with the occurrence of tumors, and many miRs play important regulatory roles in the biological functions, proliferation, invasion, migration and apoptosis.24,25 Therefore, we hope to provide potential therapeutic targets for clinic by exploring the mechanisms in miR and WT.
miR-30d belongs to the miR family and is located on human chromosome 8q24.22. More and more studies have found that miR-30d is low expressed in various tumors such as lung cancer,26 colon cancer27 and ovarian cancer.28 Moreover, the biological function can be reversed by up-regulating miR-30d. However, there is no research on miR-30d in WT at present. In this study, we searched the common differential miR through GEO database, and learned that miR-30d was decreased in GSE57370 and GSE48137 chips, which suggested that miR-30d had a close correlation with the occurrence and development of WT. In order to further verify the relationship between miR-30d and WT, miR-30d in tumor tissues of WT children was subsequently detected, and we found miR-30d down-regulated in both tumor tissues and cell lines, and the analysis also found the increasing probability of UH, NWTS high staging and lymphatic metastasis in children with miR-30d low expression. This showed that miR-30d could be a potential indicator to observe the condition of WT patients. We regulated the expression of miR-30d in WT cells and found that the cell proliferation, invasion, migration were inhibited and apoptosis was induced after increasing miR-30d. This showed that miR-30d could be a potential target for treating WT.
More and more studies have found that miR participates in tumor growth and metastasis through targeted regulation of downstream target genes.29 For further exploring whether miR-30d could participate in occurrence of WT by regulating downstream target genes, the downstream target genes of miR-30d were predicted, and 177 related target genes were predicted, among which we found Sox4, which belongs to Sox family. Like other members of Sox family, Sox4 participates in the regulation of embryo development and the determination of cell fate.30 Some studies have found that Sox4 may play a role in the apoptosis pathway leading to cell death and tumor occurrence.31 In order to verify the expression of Sox4 in WT, Sox4 was found to be highly expressed in WT through GEO database chip screening analysis. Further tests showed that Sox4 was also highly expressed in WT children’s tissues. In addition, it was found that Sox4 was negatively correlated to miR-30d in WT children’s tissues, which further confirmed the possible targeting relationship between miR-30d and Sox4. Subsequently, double luciferase report detection and Sox4 expression in WT cells transfected with miR-30d-mimics/inhibit were carried out, respectively, to verify the targeting correlation of miR-30d with Sox4. We found that miR-30d could directly target Sox4. So far, we could confirm that miR-30d could target Sox4.
At the end of the study, in order to determine that miR-30d inhibits WT cell growth and metastasis by targeting Sox4, we transfected different expressions of Sox4 into WT cells, and found that the proliferation, invasion and migration were significantly accelerated after up-regulation of Sox4, and cell apoptosis was inhibited, while the result was reversed after knocking-down of Sox4. In addition, we also found that after miR-30d-mimics and miR-30d-inhibit were co-transfected with pcDNA-Sox4 and si-Sox4 respectively, the proliferation, invasion, migration and apoptosis were not different from the control group, which indicated that miR-30d participated in occurrence of WT and it was completed by regulating Sox4.
Through the above research, we could determine the relevant mechanism of miR-30d in WT, but this research still has certain limitations. Firstly, due to the low incidence of WT, the samples collected in this study are relatively small. Secondly, the diagnostic value of miR-30d and Sox4 in WT children is not evaluated. Therefore, we hope to further explore the relationship of miR-30d and Sox4 with clinical diagnosis in future research to supplement the results of this study.
Conclusion
mir-30d is low expression in WT, and participates in the occurrence of WT by regulating Sox4, which is a promising target for the treatment of WT.
Acknowledgment
Shi Zong and Jia Zhao are co-first authors for this study.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Friedman AD. Wilms tumor. Pediatr Rev. 2013;34(7):
2. Charlton J, Irtan S, Bergeron C, Pritchard-Jones K. Bilateral Wilms tumour: a review of clinical and molecular features. Expert Rev Mol Med. 2017;19:e8. doi:10.1017/erm.2017.8
3. Hohenstein P, Pritchard-Jones K, Charlton J. The yin and yang of kidney development and Wilms’ tumors. Genes Dev. 2015;29(5):467–482. doi:10.1101/gad.256396.114
4. Charles A. Phylogeny of Wilms tumor? EBioMedicine. 2016;9:11–12. doi:10.1016/j.ebiom.2016.06.044
5. Dome JS, Graf N, Geller JI, et al. Advances in Wilms tumor treatment and biology: progress through international collaboration. J Clin Oncol. 2015;33(27):2999–3007. doi:10.1200/JCO.2015.62.1888
6. Lu TX, Rothenberg ME. MicroRNA. J Allergy Clin Immunol. 2018;141(4):1202–1207. doi:10.1016/j.jaci.2017.08.034
7. Gebert LFR, MacRae IJ. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol. 2019;20(1):21–37. doi:10.1038/s41580-018-0045-7
8. Tutar Y. Editorial (Thematic Issue: “miRNA and Cancer; Computational and Experimental Approaches”). Curr Pharm Biotechnol. 2014;15(5):429. doi:10.2174/138920101505140828161335
9. Amirkhah R, Schmitz U, Linnebacher M, Wolkenhauer O, Farazmand A. MicroRNA-mRNA interactions in colorectal cancer and their role in tumor progression. Genes Chromosomes Cancer. 2015;54(3):129–141. doi:10.1002/gcc.22231
10. Li S, Geng J, Xu X, et al. miR-130b-3p modulates epithelial-mesenchymal crosstalk in lung fibrosis by targeting IGF-1. PLoS One. 2016;11(3):e0150418. doi:10.1371/journal.pone.0150418
11. Wu H, Huang T, Ying L, et al. MiR-155 is involved in renal ischemia-reperfusion injury via direct targeting of FoxO3a and regulating renal tubular cell pyroptosis. Cell Physiol Biochem. 2016;40(6):1692–1705. doi:10.1159/000453218
12. Wang HF, Zhang YY, Zhuang HW, Xu M. MicroRNA-613 attenuates the proliferation, migration and invasion of Wilms’ tumor via targeting FRS2. Eur Rev Med Pharmacol Sci. 2017;21(15):3360–3369.
13. Che G, Gao H, Tian J, Hu Q, Xie H, Zhang Y. MicroRNA-483-3p promotes proliferation, migration, and invasion and induces chemoresistance of Wilms’ tumor cells. Pediatr Dev Pathol. 2019;23:1093526619873491.
14. Li T, Zhao P, Li Z, Wang CC, Wang YL, Gu Q. miR-200c-3p suppresses the proliferative, migratory, and invasive capacities of nephroblastoma cells via targeting FRS2. Biopreserv Biobank. 2019;17(5):444–451. doi:10.1089/bio.2019.0009
15. Wang HF, Wang WH, Zhuang HW, Xu M. MiR-429 regulates the proliferation and apoptosis of nephroblastoma cells through targeting c-myc. Eur Rev Med Pharmacol Sci. 2018;22(16):5172–5179. doi:10.26355/eurrev_201808_15713
16. Barrett T, Wilhite SE, Ledoux P, et al. NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res. 2012;41(D1):D991–995. doi:10.1093/nar/gks1193
17. World Medical A. World Medical Association Declaration of Helsinki: ethical principles for medical research involving human subjects. JAMA. 2013;310(20):2191–2194. doi:10.1001/jama.2013.281053
18. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
19. Agarwal V, Bell GW, Nam JW, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs. Elife. 2015;4. doi:10.7554/eLife.05005
20. Chen Y, Wang X. miRDB: an online database for prediction of functional microRNA targets. Nucleic Acids Res. 2020;48(D1):D127–D131. doi:10.1093/nar/gkz757
21. Chou CH, Shrestha S, Yang CD, et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018;46(D1):D296–D302. doi:10.1093/nar/gkx1067
22. Li JH, S L, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42(Database issue):D92–97. doi:10.1093/nar/gkt1248
23. Tian F, Yourek G, Shi X, Yang Y. The development of Wilms tumor: from WT1 and microRNA to animal models. Biochim Biophys Acta. 2014;1846(1):180–187. doi:10.1016/j.bbcan.2014.07.003
24. Ren Y, Zhang H, Jiang P. MicroRNA-382 inhibits cell growth and migration in colorectal cancer by targeting SP1. Biol Res. 2018;51(1):51. doi:10.1186/s40659-018-0200-9
25. Wu D, Niu X, Tao J, et al. MicroRNA-379-5p plays a tumor-suppressive role in human bladder cancer growth and metastasis by directly targeting MDM2. Oncol Rep. 2017;37(6):3502–3508. doi:10.3892/or.2017.5607
26. Gao L, He RQ, Wu HY, et al. Expression signature and role of miR-30d-5p in non-small cell lung cancer: a comprehensive study based on in silico analysis of public databases and in vitro experiments. Cell Physiol Biochem. 2018;50(5):1964–1987. doi:10.1159/000494875
27. Zhang R, Xu J, Zhao J, Bai J. Mir-30d suppresses cell proliferation of colon cancer cells by inhibiting cell autophagy and promoting cell apoptosis. Tumour Biol. 2017;39(6):1010428317703984. doi:10.1177/1010428317703984
28. Ye Z, Zhao L, Li J, Chen W, Li X. miR-30d blocked transforming growth factor β1–induced epithelial-mesenchymal transition by targeting snail in ovarian cancer cells. Int J Gynecol Cancer. 2015;25(9):1574–1581. doi:10.1097/IGC.0000000000000546
29. Ludwig N, Werner TV, Backes C, et al. Combining miRNA and mRNA expression profiles in Wilms tumor subtypes. Int J Mol Sci. 2016;17(4):475. doi:10.3390/ijms17040475
30. Moreno CS. SOX4: the unappreciated oncogene. Semin Cancer Biol. 2019. doi:10.1016/j.semcancer.2019.08.027
31. Liu Y, Li F, Lai D, et al. MicroRNA-140 inhibits proliferation and promotes apoptosis and cell cycle arrest of prostate cancer via degrading SOX4. J BUON. 2019;24(1):249–255.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.