Back to Journals » OncoTargets and Therapy » Volume 11
Minichromosome maintenance protein 2 correlates with the malignant status and regulates proliferation and cell cycle in lung squamous cell carcinoma
Authors Wu W, Wang X, Shan C, Li Y, Li F
Received 22 March 2018
Accepted for publication 8 May 2018
Published 20 August 2018 Volume 2018:11 Pages 5025—5034
DOI https://doi.org/10.2147/OTT.S169002
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Samir Farghaly
Wei Wu,1,* Xianwei Wang,1,* Changting Shan,1 Yong Li,2 Fengzhu Li3
1Department of Respiratory Medicine, Affiliated Hospital of Jining Medical University, Jining, Shandong 272029, People’s Republic of China; 2Department of Emergency, Affiliated Hospital of Jining Medical University, Jining, Shandong 272029, People’s Republic of China; 3Department of Paediatric Surgery, Jining No 1 People’s Hospital, Jining, Shandong 272011, People’s Republic of China
*These authors contributed equally to this work
Background: Minichromosome maintenance protein 2 (MCM2), which is a member of MCM family, has been found to be a relevant marker for progression and prognosis in a variety of human cancers. Due to lack of effective therapeutic target in lung squamous cell carcinoma (LUSC) patients, the aim of our study was to screen and identify biomarkers which are associated to clinicopathological characteristics including prognosis in LUSC patients.
Methods: The expression status of MCM2 in lung cancer was analyzed using the publicly available Gene Expression Omnibus databases (GSE3268 and GSE10245). The mRNA and protein expression of MCM2 in lung cancer tissues and cell lines was detected by quantitative real-time PCR and Western blot, and the association between MCM2 expression and clinicopathological factors was analyzed by immunohistochemistry. The loss-of-function study of MCM2 was conducted in LUSC cell lines.
Results: In our study, we found MCM2 expression was increased in LUSC tissues compared with paired adjacent normal lung tissues or lung adenocarcinoma tissues through analyzing microarray data sets (GSE3268 and GSE10245), which confirmed that MCM2 mRNA and protein were overexpressed in LUSC tissues and cell lines. Meanwhile, we analyzed the association between MCM2 protein expression and clinicopathological characteristics of LUSC patients, and found high expression of MCM2 protein was obviously associated with malign differentiated degree, advanced clinical stage, large tumor size, more lymph node metastasis and present distant metastasis. The survival analysis showed MCM2 overexpression was an independent unfavorable prognostic factor for LUSC patients.
Conclusion: MCM2 is involved in the development and progression of LUSC as an oncogene, and thereby may act as a potential therapeutic target for LUSC patients.
Keywords: MCM2, lung cancer, lung squamous cell carcinoma, biomarker, proliferation, cell cycle
Introduction
Lung cancer ranks first among all cancers in morbidity and mortality worldwide, and the incidence and mortality of lung cancer are increasing every year.1 According to the GLOBOCAN 2012 data, lung cancer is the most frequently diagnosed cancer worldwide with an estimated 1,824,700 cases and 1,589,900 deaths in 2012.1 In People’s Republic of China, about 733,300 newly diagnosed lung cancer cases and 610,200 lung cancer deaths were reported in 2015.2 In the USA, it is estimated that 234,030 new lung cancer cases and 154,050 lung cancer deaths will be recorded in 2018.3 Based on histological classification, lung cancer can be divided into small-cell lung cancer, lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), large-cell lung cancer and other rare types.4,5 LUAD and LUSC are the two major histological types of lung cancer, accounting for 80% of lung cancer cases.6 In recent decades, molecular targeted therapy has seen a rapid growth including the development of epidermal growth factor (EGF) receptor tyrosine kinase inhibitors and anaplastic lymphoma tyrosine kinase inhibitors, which are effective for the treatment of LUAD carrying activating mutations.7,8 Due to lack of effective therapeutic target in LUSC patients, the overall survival of LUSC patients is significantly shorter compared with LUAD patients.9,10 Hence, it is urgently necessary to identify novel and credible biomarkers for developing molecular targeted therapy for LUSC patients.
Minichromosome maintenance protein (MCM) family is composed of six related proteins (MCM2–MCM7), which were originally identified in yeast to activate DNA replication.11,12 Closure of the MCM2/5 gate is required to activate DNA unwinding and elongation.13 Upon S-phase entry, several regulatory cyclin-dependent kinases and DBF4-dependent kinase activate MCM2–MCM7 by enabling the loading of the key accessory factors CDC45 and GINS that in combination with MCM2–MCM7 form the CMG complex.13 MCM2 has been found to be a relevant marker for progression and prognosis in a variety of human cancers.13 The aim of our study was to screen and identify biomarkers which are associated to clinicopathological characteristics including prognosis in LUSC patients. In the early stage of our study, we analyzed microarray data sets (GSE3268 and GSE10245), and found MCM2 expression was increased in LUSC tissues compared with paired adjacent normal lung tissues from microarray data (GSE3268) and MCM2 was overexpressed in LUSC tissue samples compared with LUAD tissue samples based on microarray data sets (GSE10245). Thus, we further explored the clinical value and biological function of MCM2 in LUSC. We measured MCM2 expression in LUSC tissues to observe the correlation between MCM2 expression and clinicopathological characteristics, and performed loss-of-function study to explore the effect of MCM2 on LUSC cell lines.
Methods
Ethics statement
The Research Ethics Committees of Affiliated Hospital of Jining Medical University and Jining No 1 People’s Hospital approved this protocol, and written informed consent was obtained from each patient. The entire study was performed based on the Declaration of Helsinki.
Database analysis
Microarray data sets were retrieved from the Gene Expression Omnibus database. The microarray data set GSE3268 comprised five pairs of LUSC tissues and adjacent normal lung tissues, and was submitted by Shinichiro Wachi et al (email: [email protected]). The microarray data set GSE10245 included LUSC tissue samples and LUAD tissue samples, and was submitted by Ruprecht Kuner et al (email: [email protected]). Raw data were processed, and differentially expressed genes were screened.
Patients and samples
Fresh and paraffin-embedded clinical samples (including LUAD tissue, LUSC tissue and normal lung tissue) were collected, and the pathological information (age, gender, histological type, clinical stage, tumor depth, lymph node metastasis, distant metastasis, differentiation and family history) was retrieved from the archives of the Department of Pathology of Affiliated Hospital of Jining Medical University and Jining No 1 People’s Hospital. None of the patients had received antitumor therapy before diagnosis. Clinical staging and system treatment were performed according to the 7th edition of the American Joint Committee on Cancer (AJCC) Cancer Staging Manual and National Comprehensive Cancer Network (NCCN) guideline, respectively.
RNA extraction and quantitative real-time PCR (qRT-PCR)
The total RNA from cell or tissue was extracted with RNAiso Plus (TaKaRa, Otsu, Japan). Total RNA was reverse-transcribed using PrimeScript® RT reagent kit (TaKaRa). The qRT-PCR was performed with SYBR® Premix Ex Taq™ II (TaKaRa) on a LightCycler® (Roche, Basel, Switzerland). The following primers were used for MCM2 and GAPDH: MCM2 forward, 5′-TGTCACCTGCTCTGCCACTAA-3′ and reverse, 5′-GCAGCATGCGCAAGACTTT-3′; GAPDH forward, 5′-CCATCAATGACCCCTTCATTG-3′ and reverse, 5′-CATGGGTGGAATCATATTGGAAC-3′. Relative expression was calculated via the comparative cycle threshold method and was normalized to the expression of GAPDH.
Immunohistochemistry
Immunohistochemical analysis was performed to detect MCM2 protein expression in LUAD tissues, LUSC tissues and normal lung tissues. In brief, paraffin-embedded sections were deparaffinized in xylene for 20 min and rehydrated in graded concentrations of ethanol (100%, 95%, 90%, 80% and 70%). The sections were submerged in ethylene diamine tetraacetic acid antigenic retrieval buffer and microwaved for antigen retrieval. They were then treated with 3% hydrogen peroxide in methanol to quench endogenous peroxidase activity, followed by incubation with 5% bovine serum albumin to block nonspecific binding. Sections were incubated with anti-MCM2 (Abcam, Cambridge, UK) overnight at 4°C. After washing, tissue sections were treated with secondary antibody, followed by incubation with conjugated horseradish peroxidase–streptavidin. Tissue sections were then counterstained with hematoxylin, dehydrated and mounted.
Evaluation of staining
The tissue sections stained immunohistochemically for MCM2 were reviewed, and scored separately by two pathologists blinded to the clinical parameters. Any disagreements were arbitrated by a third pathologist. For MCM2 assessment, staining intensity was scored as 0=negative, 1=weak, 2=moderate or 3=strong, and staining extent was scored as 0=0%, 1=1%–25%, 2=26%–50%, 3=51%–75% or 4=70%–100%, depending on the percentage of positively stained cells.14 The sum of staining intensity and staining extent scores was used as final staining score. Low expression of MCM2 was defined as scores 0–4, and high expression was defined as scores >4.
Western blot
Total protein was extracted using radioimmunoprecipitation assay buffer (Thermo Fisher Scientific, Waltham, MA, USA) for Western blot. Equal amounts of protein were denatured and then separated by 10% SDS-PAGE. The target proteins were incubated with the following primary antibodies: MCM2 antibody (Abcam) or GAPDH antibody (CWBIO, Beijing, People’s Republic of China). Then, the proteins were incubated with homologous secondary antibodies (CWBIO). For horseradish peroxidase detection, an enhanced chemiluminescent kit (CWBIO) was used. Intensity of blots was determined by Quantity One Software (Bio-Rad, Hercules, CA, USA).
Cell lines and cell cultures
Two human LUAD cell lines (A549 and H1299), two human LUSC cell lines (SK-MES-1 and H2170) and a human bronchial epithelial cell line (HBEC) were obtained from American Type Culture Collection and Cell Bank of Type Culture Collection of the Chinese Academy of Sciences. A549, H1299, SK-MES-1 and H2170 were cultured in Roswell Park Memorial Institute 1640 medium (Thermo Fisher Scientific) supplemented with 10% fetal bovine serum (Thermo Fisher Scientific). HBEC was cultured in complete Keratinocyte-SFM (Thermo Fisher Scientific) supplemented with 5 mg/L EGF and 50 mg/L bovine pituitary extract. All cells were maintained at 37°C in a humidified atmosphere with 5% CO2.
Cell transfection
The siRNA for downregulation of MCM2 was purchased from Guangzhou RiboBio Co., Ltd. (Guangzhou, People’s Republic of China). Cells were transfected using Lipofectamine™ 3000 reagent (Thermo Fisher Scientific) in Opti-MEM (Gibco), according to the manufacturer’s instructions. The transfection efficiency was tested by qRT-PCR and Western blot.
Cell proliferation and colony formation assays
Cell proliferation was analyzed using MTT assay and colony formation assay according to previous description.15 After transfected with plasmids and cultured in normal medium, cells were grown in a 96-well plate for 24, 48, 72, 96 and 120 h.
Cell cycle analysis
LUSC cells transfected with siRNA-MCM2 or siRNA-negative control (NC) were harvested after 48 h, rinsed with cold phosphate buffer saline and fixed with 70% ice-cold ethanol for 48 h at 4°C overnight. After incubation with PBS containing 10 mg/mL propidium iodide and 0.5 mg/mL RNase A for 15 min at 37°C, fixed cells were washed with cold PBS three times. FACSCalibur™ flow cytometry (BD Biosciences, Franklin Lakes, NJ, USA) was used to determine the DNA content of labeled cells.
Statistical analysis
All data were analyzed for statistical significance using SPSS 17.0 software and GraphPad Prism 5.0 software. The chi-square test was used to analyze the correlation between MCM2 expression and clinicopathological characteristics of lung cancer patients. Survival times were detected using the Kaplan–Meier survival curves, and compared by the log-rank test. The prognostic significance of various variables was analyzed by univariate and multivariate survival analysis using Cox’s regression model. Two-tailed Student’s t-test was used for comparisons of two independent groups. A P-value of <0.05 was considered statistically significant.
Results
Expression of MCM2 in LUSC tissue
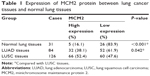
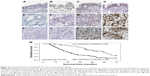
We analyzed the microarray data (GSE3268) to screen and identify valuable biomarkers for LUSC, and found MCM2 expression was increased in LUSC tissues compared with paired adjacent normal lung tissues (Figure 1A). Meanwhile, MCM2 was overexpressed in LUSC tissue samples compared with LUAD tissue samples based on microarray data sets (GSE10245; Figure 1B). Furthermore, qRT-PCR and immunohistochemistry were performed to detect mRNA and protein expressions of MCM2 in LUAD tissues, LUSC tissues and normal lung tissues. Compared with normal lung tissues or LUAD tissues, MCM2 mRNA expression was significantly elevated in LUSC tissues (both P<0.001; Figure 1C). Meanwhile, immunohistochemical analysis (Table 1 and Figure 2A–L) showed MCM2 protein expression was increased in LUSC tissues (52.4%, 66/126) in comparison to normal lung tissues (16.1%, 5/31, P<0.001) and LUAD tissues (38.1%, 32/84, P=0.042).
Expression of MCM2 in LUSC cells
MCM2 mRNA and protein levels were detected in two human LUAD cell lines (A549 and H1299), two human LUSC cell lines (SK-MES-1 and H2170) and a HBEC through qRT-PCR and Western blot. Levels of MCM2 mRNA were increased in LUSC cells compared with bronchial epithelial cells or LUAD cells (both P<0.001; Figure 1D). Meanwhile, we observed that MCM2 protein level was increased in LUSC cells compared with bronchial epithelial cells or LUAD cells (Figure 1E).
MCM2 overexpression correlates with the malignant status of LUSC patients
We measured MCM2 protein expression in LUAD tissue, LUSC tissue and normal lung tissue by using immunohistochemical staining (Figure 2A–L), and explored the association between MCM2 protein expression and clinicopathological characteristics of LUSC patients. As summarized in Table 2, MCM2 protein expression was obviously associated with differentiated degree (high and middle vs low; P=0.003), clinical stage (I–II vs III–IV; P<0.001), tumor size (≤5 cm vs >5 cm; P=0.034), lymph node metastasis (N0–N1 vs N2–N3; P<0.001) and distant metastasis (No vs Yes; P=0.001). However, MCM2 protein expression was not associated with gender, age and smoking.
MCM2 overexpression is an independent poor predictor for LUSC patients
According to LUSC patients’ prognosis information, we analyzed the relationship between MCM2 expression and patients’ survival. We observed that patients with higher levels of MCM2 expression had poorer survival than those with lower levels of MCM2 expression (P<0.001; Figure 2M), as MCM2 overexpression level predicted overall survival. Univariate and multivariate analyses suggested that MCM2 overexpression was an independent unfavorable prognostic factor for LUSC patients (HR=1.847, 95% CI: 1.080–3.158, P=0.025; Table 3).
  | Table 3 Summary of univariate and multivariate Cox regression analyses of overall survival duration |
Downregulation of MCM2 suppresses LUSC cell proliferation
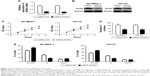
To explore the biological function of MCM2 in LUSC cells, we induced downregulation of MCM2 in SK-MES-1 and H2170 cells by transfecting siRNA-MCM2, and the results were confirmed by qRT-PCR and Western blot (Figure 3A and B). The results of MTT assay showed that downregulation of MCM2 obviously decreased SK-MES-1 and H2170 viability in comparison to control cells after transfection for 48, 72 and 96 h (all P<0.05; Figure 3C). The colony formation assay suggested that downregulation of MCM2 markedly reduced the number of colonies in SK-MES-1 and H2170 cells over a 12-day period (both P<0.05; Figure 3D). The analysis of cell cycle distribution showed that downregulation of MCM2 obviously decreased the percentage of S-phase cells and increased the percentage of G0/G1-phase cells, suggesting that downregulation of MCM2 might cause a G0/G1- to S-phase arrest in SK-MES-1 and H2170 cells (all P<0.05; Figure 3E).
Discussion
MCM2, which is a member of the MCM family, has been found to be a relevant marker for progression and prognosis in a variety of human cancers.13 Primarily, MCM2 has been considered as a sensitive marker for early detection of pulmonary malignant lesions. Tan et al reported that MCM2 was detectable in two to three times more proliferating premalignant lung cells than Ki-67 index, and acted as an easy-to-use marker for the early detection of lung cancer which may significantly enhance lung cancer survival.16 Furthermore, Muñoz-Antonia et al showed progression of premalignant lesions toward carcinoma in situ was accompanied by an increase in the expression of MCM2.17 Subsequently, Dehan et al and Kadara et al congruously suggested that MCM2 was significantly upregulated in non-small-cell lung cancer through microarrays.18,19 In our study, we analyzed microarray data sets (GSE3268 and GSE10245), and found MCM2 expression was increased in LUSC tissues compared with paired adjacent normal lung tissues and LUAD tissues. Furthermore, we confirmed that MCM2 mRNA and protein expressions were increased in LUSC tissues and cell lines compared with normal lung tissues and LUAD tissues, or bronchial epithelial cell line or LUAD cell lines. In addition, high expression of MCM2 was observed in several types of human cancers such as breast cancer,20,21 gastric cancer,22,23 colorectal cancer,24,25 esophageal cancer,26 liver cancer,27,28 renal cell carcinoma29 and so on.
Due to lack of effective therapeutic target in LUSC patients, we supposed MCM2 might act as an important biomarker for LUSC patients based on the expression of MCM2 in LUSC tissues. We analyzed the association between MCM2 protein expression and clinicopathological characteristics of LUSC patients, and found high expression of MCM2 protein was obviously associated with malign differentiated degree, advanced clinical stage, large tumor size, more lymph node metastasis and present distant metastasis. Similarly, Veena et al found MCM2 proteins have significant association with tumor stage in lung cancer patients,30 and Liu et al reported that high expression of MCM2 in non-small cell lung cancer was observed more frequently in aged patients and in patients at later stage.31 Moreover, MCM2 expression positivity significantly increased with increasing tumor grade, the presence of residual disease and advancing tumor stage in ovarian cancer patients.32 In esophageal squamous cell carcinoma patients, Kato et al demonstrated that there were significant associations between MCM2 expression and tumor status, lymph node status, metastatic status, pathological stage and histological grade.33 On the whole, high expression of MCM2 is associated with aggressive progression in most types of human cancers.
In the last 10 years, the prognostic significance of MCM2 has been explored in various types of human cancers. Generally, high expression of MCM2 predicts a poor prognosis in most types of human cancers such as gastric cancer,22 osteosarcoma,34 oral cavity squamocellular carcinoma,35 bladder cancer,36 prostate cancer,37 breast cancer,38 renal cell cancer39,40 and penile cancer.41 In non-small-cell lung cancer, Liu et al,31 Ramnath et al,42 Hashimoto et al43 and Cheung et al44 consistently revealed MCM2 overexpression was associated with unfavorable prognosis. However, Werynska et al reported no significant differences in overall survival related to the expression of MCM-2 in non-small-cell lung cancer patients.45 In our study, we presented more evidence suggesting that MCM2 expression was significantly associated with poor overall survival of LUSC patients, and MCM2 overexpression was an independent unfavorable prognostic factor for LUSC patients. However, Nishihara et al showed that no correlation was observed between the MCM2 expression and cumulative survival in colorectal cancer patients.46 Similarly, MCM2 expression has been indicated to lack prognostic significance in medulloblastomas and myxofibrosarcomas.47,48
In order to explore the biological function of MCM2 in LUSC cells, we performed loss-of-function study to explore the effect of MCM2 on LUSC cell line. In our study, we found downregulation of MCM2 obviously decreased LUSC cell proliferation, and also decreased the percentage of S-phase cells and increased the percentage of G0/G1-phase cells. Similarly, Cheung et al showed MCM2 promoted lung cancer cell proliferation via the regulation of HMGA1 phosphorylation.44 Moreover, Zhang et al confirmed that MCM2 was a therapeutic target of lovastatin in suppressing lung cancer cell viability.49 Meanwhile, Zhang et al found that downregulation of MCM2 expression increased Rb, cyclin D1 and CDK4 expression, and decreased p53 and p21 expression, which suggested that MCM2 triggered the arrest of G1/S or G2/M through the induction of cyclin-dependent kinase inhibitor p21cip14 and cyclin D/CDK4 cyclin D/CDK4.49 Besides, MCM2 has been found to act as a functional target for lncRNA and microRNA in regulating tumor cell proliferation, migration and apoptosis.50–52
Conclusion
In summary, MCM2 is significantly increased in LUSC tissues and cell lines, and correlates with the malignant status and prognosis of LUSC patients. MCM2 functions as an oncogene in regulating LUSC cell proliferation and cell cycle distribution.
Disclosure
The authors report no conflicts of interest in this work.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108. | ||
Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. | ||
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68(1):7–30. | ||
Tang Y, Qiao G, Xu E, Xuan Y, Liao M, Yin G. Biomarkers for early diagnosis, prognosis, prediction, and recurrence monitoring of non-small cell lung cancer. Onco Targets Ther. 2017;10:4527–4534. | ||
Zhang Y, Yang SH, Guo XL. New insights into Vinca alkaloids resistance mechanism and circumvention in lung cancer. Biomed Pharmacother. 2017;96:659–666. | ||
Phillips I, Ajaz M, Ezhil V, et al. Clinical applications of textural analysis in non-small cell lung cancer. Br J Radiol. 2018;91(1081):20170267. | ||
Sgambato A, Casaluce F, Maione P, Gridelli C. Targeted therapies in non-small cell lung cancer: a focus on ALK/ROS1 tyrosine kinase inhibitors. Expert Rev Anticancer Ther. 2018;18(1):71–80. | ||
Mayekar MK, Bivona TG. Current landscape of targeted therapy in lung cancer. Clin Pharmacol Ther. 2017;102(5):757–764. | ||
Sawabata N, Miyaoka E, Asamura H, et al; Japanese Joint Committee for Lung Cancer Registration. Japanese lung cancer registry study of 11,663 surgical cases in 2004: demographic and prognosis changes over decade. J Thorac Oncol. 2011;6(7):1229–1235. | ||
Mari-Alexandre J, Diaz-Lagares A, Villalba M, et al. Translating cancer epigenomics into the clinic: focus on lung cancer. Transl Res. 2017;189:76–92. | ||
Maine GT, Sinha P, Tye BK. Mutants of S. cerevisiae defective in the maintenance of minichromosomes. Genetics. 1984;106(3):365–385. | ||
Das SP, Rhind N. How and why multiple MCMs are loaded at origins of DNA replication. Bioessays. 2016;38(7):613–617. | ||
Simon NE, Schwacha A. The Mcm2-7 replicative helicase: a promising chemotherapeutic target. Biomed Res Int. 2014;2014:549719. | ||
Fathi A, Mosaad H, Hussein S, Roshdy M, Ismail EI. Prognostic significance of CD133 and ezrin expression in colorectal carcinoma. IUBMB Life. 2017;69(5):328–340. | ||
Shan C, Fei F, Li F, et al. miR-448 is a novel prognostic factor of lung squamous cell carcinoma and regulates cells growth and metastasis by targeting DCLK1. Biomed Pharmacother. 2017;89:1227–1234. | ||
Tan DF, Huberman JA, Hyland A, et al. MCM2 – a promising marker for premalignant lesions of the lung: a cohort study. BMC Cancer. 2001;1:6. | ||
Muñoz-Antonia T, Muro-Cacho C, Sharma S, Cantor A, Bepler G. Expression of TGFbeta type-II receptor in association with markers of proliferation and apoptosis in premalignant lung lesions. Cancer. 2007;110(7):1527–1531. | ||
Dehan E, Ben-Dor A, Liao W, et al. Chromosomal aberrations and gene expression profiles in non-small cell lung cancer. Lung Cancer. 2007;56(2):175–184. | ||
Kadara H, Lacroix L, Behrens C, et al. Identification of gene signatures and molecular markers for human lung cancer prognosis using an in vitro lung carcinogenesis system. Cancer Prev Res (Phila). 2009;2(8):702–711. | ||
Tőkés T, Tőkés AM, Szentmártoni G, et al. Expression of cell cycle markers is predictive of the response to primary systemic therapy of locally advanced breast cancer. Virchows Arch. 2016;468(6):675–686. | ||
Yousef EM, Furrer D, Laperriere DL, et al. MCM2: an alternative to Ki-67 for measuring breast cancer cell proliferation. Mod Pathol. 2017;30(5):682–697. | ||
Yang C, Wen Y, Li H, et al. Overexpression of minichromosome maintenance 2 predicts poor prognosis in patients with gastric cancer. Oncol Rep. 2012;27(1):135–142. | ||
Giaginis C, Giagini A, Tsourouflis G, et al. MCM-2 and MCM-5 expression in gastric adenocarcinoma: clinical significance and comparison with Ki-67 proliferative marker. Dig Dis Sci. 2011;56(3):777–785. | ||
Hanna-Morris A, Badvie S, Cohen P, McCullough T, Andreyev HJ, Allen-Mersh TG. Minichromosome maintenance protein 2 (MCM2) is a stronger discriminator of increased proliferation in mucosa adjacent to colorectal cancer than Ki-67. J Clin Pathol. 2009;62(4):325–330. | ||
Wang Y, Zhou ZG, Xia QJ, Zhang WY, Li HG, Wang R. [Expression of minichromosome maintenance protein 2 in colonic adenocarcinoma, adenoma and normal colonic mucosa and its clinical significance]. Zhonghua Wei Chang Wai Ke Za Zhi. 2008;11(5):465–468. Chinese [with English abstract]. | ||
Huang B, Hu B, Su M, et al. Potential role of minichromosome maintenance protein 2 as a screening biomarker in esophageal cancer high-risk population in China. Hum Pathol. 2011;42(6):808–816. | ||
Sun M, Wu G, Li Y, et al. Expression profile reveals novel prognostic biomarkers in hepatocellular carcinoma. Front Biosci (Elite Ed). 2010;2:829–840. | ||
Qin LX, Tang ZY. The prognostic molecular markers in hepatocellular carcinoma. World J Gastroenterol. 2002;8(3):385–392. | ||
Zhong H, Chen B, Neves H, et al. Expression of minichromosome maintenance genes in renal cell carcinoma. Cancer Manag Res. 2017;9:637–647. | ||
Veena VS, Rajan K, Saritha VN, et al. DNA replication licensing proteins for early detection of lung cancer. Asian Pac J Cancer Prev. 2017;18(11):3041–3047. | ||
Liu YZ, Wang BS, Jiang YY, et al. MCMs expression in lung cancer: implication of prognostic significance. J Cancer. 2017;8(18):3641–3647. | ||
Gakiopoulou H, Korkolopoulou P, Levidou G, et al. Minichromosome maintenance proteins 2 and 5 in non-benign epithelial ovarian tumours: relationship with cell cycle regulators and prognostic implications. Br J Cancer. 2007;97(8):1124–1134. | ||
Kato H, Miyazaki T, Fukai Y, et al. A new proliferation marker, minichromosome maintenance protein 2, is associated with tumor aggressiveness in esophageal squamous cell carcinoma. J Surg Oncol. 2003;84(1):24–30. | ||
Cheng DD, Zhang HZ, Yuan JQ, Li SJ, Yang QC, Fan CY. Minichromosome maintenance protein 2 and 3 promote osteosarcoma progression via DHX9 and predict poor patient prognosis. Oncotarget. 2017;8(16):26380–26393. | ||
Szelachowska J, Dziegiel P, Jelen-Krzeszewska J, et al. Mcm-2 protein expression predicts prognosis better than Ki-67 antigen in oral cavity squamocellular carcinoma. Anticancer Res. 2006;26(3B):2473–2478. | ||
Burger M, Denzinger S, Hartmann A, Wieland WF, Stoehr R, Obermann EC. Mcm2 predicts recurrence hazard in stage Ta/T1 bladder cancer more accurately than CK20, Ki67 and histological grade. Br J Cancer. 2007;96(11):1711–1715. | ||
Toubaji A, Sutcliffe S, Chaux A, et al. Immunohistochemical expression of minichromosome maintenance complex protein 2 predicts biochemical recurrence in prostate cancer: a tissue microarray and digital imaging analysis-based study of 428 cases. Hum Pathol. 2012;43(11):1852–1865. | ||
Joshi S, Watkins J, Gazinska P, et al. Digital imaging in the immunohistochemical evaluation of the proliferation markers Ki67, MCM2 and Geminin, in early breast cancer, and their putative prognostic value. BMC Cancer. 2015;15:546. | ||
Dudderidge TJ, Stoeber K, Loddo M, et al. Mcm2, Geminin, and KI67 define proliferative state and are prognostic markers in renal cell carcinoma. Clin Cancer Res. 2005;11(7):2510–2517. | ||
Rodins K, Cheale M, Coleman N, Fox SB. Minichromosome maintenance protein 2 expression in normal kidney and renal cell carcinomas: relationship to tumor dormancy and potential clinical utility. Clin Cancer Res. 2002;8(4):1075–1081. | ||
May M, Burger M, Otto W, et al. Ki-67, mini-chromosome maintenance 2 protein (MCM2) and geminin have no independent prognostic relevance for cancer-specific survival in surgically treated squamous cell carcinoma of the penis. BJU Int. 2013;112(4):E383–E390. | ||
Ramnath N, Hernandez FJ, Tan DF, et al. MCM2 is an independent predictor of survival in patients with non-small-cell lung cancer. J Clin Oncol. 2001;19(22):4259–4266. | ||
Hashimoto K, Araki K, Osaki M, et al. MCM2 and Ki-67 expression in human lung adenocarcinoma: prognostic implications. Pathobiology. 2004;71(4):193–200. | ||
Cheung CHY, Hsu CL, Chen KP, et al. MCM2-regulated functional networks in lung cancer by multi-dimensional proteomic approach. Sci Rep. 2017;7(1):13302. | ||
Werynska B, Pula B, Muszczynska-Bernhard B, et al. Correlation between expression of metallothionein and expression of Ki-67 and MCM-2 proliferation markers in non-small cell lung cancer. Anticancer Res. 2011;31(9):2833–2839. | ||
Nishihara K, Shomori K, Fujioka S, et al. Minichromosome maintenance protein 7 in colorectal cancer: implication of prognostic significance. Int J Oncol. 2008;33(2):245–251. | ||
Lau KM, Chan QK, Pang JC, et al. Minichromosome maintenance proteins 2, 3 and 7 in medulloblastoma: overexpression and involvement in regulation of cell migration and invasion. Oncogene. 2010;29(40):5475–5489. | ||
Huang HY, Kang HY, Li CF, et al. Skp2 overexpression is highly representative of intrinsic biological aggressiveness and independently associated with poor prognosis in primary localized myxofibrosarcomas. Clin Cancer Res. 2006;12(2):487–498. | ||
Zhang X, Teng Y, Yang F, et al. MCM2 is a therapeutic target of lovastatin in human non-small cell lung carcinomas. Oncol Rep. 2015;33(5):2599–2605. | ||
Liu F, Yuan JH, Huang JF, et al. Long noncoding RNA FTX inhibits hepatocellular carcinoma proliferation and metastasis by binding MCM2 and miR-374a. Oncogene. 2016;35(41):5422–5434. | ||
Jin Y, Xiong A, Zhang Z, et al. MicroRNA-31 suppresses medulloblastoma cell growth by inhibiting DNA replication through minichromosome maintenance 2. Oncotarget. 2014;5(13):4821–4833. | ||
Cheung CC, Chung GT, Lun SW, et al. miR-31 is consistently inactivated in EBV-associated nasopharyngeal carcinoma and contributes to its tumorigenesis. Mol Cancer. 2014;13:184. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.