Back to Journals » OncoTargets and Therapy » Volume 13
Methylcrotonoyl-CoA Carboxylase 2 Promotes Proliferation, Migration and Invasion and Inhibits Apoptosis of Prostate Cancer Cells Through Regulating GLUD1-P38 MAPK Signaling Pathway
Authors He J , Mao Y, Huang W, Li M, Zhang H, Qing Y, Lu S, Xiao H , Li K
Received 15 February 2020
Accepted for publication 8 July 2020
Published 28 July 2020 Volume 2020:13 Pages 7317—7327
DOI https://doi.org/10.2147/OTT.S249906
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Arseniy Yuzhalin
Jianwen He,* Yunhua Mao,* Wentao Huang,* Mingzhao Li, Huimin Zhang, Yunhao Qing, Shuo Lu, Hengjun Xiao, Ke Li
Department of Urology, The Third Affiliated Hospital of Sun Yat-sen University, Guangzhou 510630, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Ke Li Email [email protected]
Hengjun Xiao Email [email protected]
Purpose: Prostate cancer (PCa) is the most common cancer in American men, and the mechanisms of development and progression are still not completely clear. Methylcrotonoyl-CoA carboxylase 2 (MCCC2) was previously identified overexpressed in PCa with lymph node metastasis, but its specific role and mechanisms need further investigation. This study aimed to investigate the role of MCCC2 in PCa cells and its underlying mechanisms.
Materials and Methods: Quantitative RT-PCR and Western blotting were used to detect MCCC2 mRNA and protein expression in normal prostate epithelium and cancerous cells. Upon manipulation of MCCC2 expression, cell proliferation was measured by CCK-8 assays and migration and invasion were determined by transwell assays. Changes of apoptosis, cell cycle and mitochondrial membrane potential were evaluated by flow cytometry. MCCC2-mediated signaling pathways were screened by bioinformatics and verified by RT-PCR and Western blotting. Finally, immunohistochemistry was performed to detect the expression of MCCC2 and glutamate dehydrogenase 1 (GLUD1) in PCa tissues to analyze their correlation.
Results: We demonstrated that MCCC2 promoted cell proliferation, migration and invasion but inhibited apoptosis in PCa cells. In addition, MCCC2 in 22Rv1 cells induced mitochondrial damage. In PCa tissues, MCCC2 overexpression associated with lymph node metastasis (P=0.001) and high Gleason scores (P< 0.001). MCCC2 positively correlated with GLUD1 expression in PCa tissues (r=0.435, P< 0.001). Ectopic overexpression of MCCC2 up-regulated GLUD1 and p38 MAPK expression, whereas inhibition of MCCC2 decreased GLUD1 and p38 MAPK expression.
Conclusion: MCCC2 exerts oncogenic function in PCa through regulating GLUD1-p38 MAPK signaling pathway, and it may be a potential treatment target.
Keywords: prostate cancer, MCCC2, GLUD1, p38 MAPK
Introduction
Prostate cancer (PCa) is one of the most common types of cancer in American men, accounting for nearly one-fifth of the newly diagnosed cancers in American men in 2019.1 The incidence of PCa in China is much lower, but the incidence and mortality is increasing with the population aging progress.2 Surgery, radiation, androgen deprivation and chemotherapy are currently the main treatments and achieve good results for early localized PCa.3 However, when the disease progresses to the lethal stage which is called metastatic castration-resistant prostate cancer (mCRPC), patients have a median survival of about two years regardless of multidisciplinary treatments. At present, the mechanisms of development and progression of PCa are still not completely clear; therefore, it is of great significance to explore a potential therapeutic target and the molecular mechanisms as to improve the treatment effect.
Methylcrotonoyl-CoA carboxylase 2 (MCCC2) is a subunit of 3-methylcrotonyl-CoA carboxylase (MCC).4 MCC deficiency can lead to the disorder of leucine metabolism, which can be manifested in a variety of phenotypes, including neurological disorders, infant death and asymptomatic adults.5–8 A mouse study has shown that MCCC2 is associated with obesity, insulin resistance and dyslipidemia.9 In addition, MCC deletion indicates changes in glycolysis, TCA cycle, OXPHOS, gluconeogenesis, β-oxidation and branched chain fatty acid metabolism.10 Previously, colleagues in our institute first identified that MCCC2 was highly expressed in PCa with lymph node metastasis by DIGE-based proteomic analysis.11 Another study showed that the expression of MCCC2 may be related to the progression from androgen-dependent to androgen-independent PCa.12 A recent study revealed that MCCC2 associated with unfavorable clinical features and predicts poor prognosis in breast cancer.13 However, the specific role of MCCC2 and the underlying mechanisms in the development and progression of PCa still remains unclear.
In this study, we aimed to investigate the biological effects of MCCC2 on tumor behavior in PCa, such as cell proliferation, migration, invasion and apoptosis. Further, we explored the potential mechanisms whereby MCCC2 affects cancer aggressiveness. We also determined the immunohistostaining of MCCC2 in PCa tissues and analyzed the association with clinical features.
Materials and Methods
Cell, siRNA, Plasmid and Transfection
Human prostate cancer cells 22Rv1, LNCaP, C4-2, DU145 and PC3 were all purchased from ATCC in the United States. Prostate cancer cells were cultured in RPMI1640 medium (GIBCO, USA) supplemented with 10% fetal bovine serum (GIBCO, USA). All cells in a humidified incubator at 37 °C with 5% CO2. SiRNA to MCCC2 (sense 5ʹ-GCCCAAGAUUUCUCUACAUUUTT-3ʹ and antisense 5ʹ-AAAUGUAGAGAAAUCUUGGGCTT-3ʹ) was purchased from Jidan Biotech (Guangzhou, China). MCCC2 expression vector pCDNA3.1 (+)-Flag-MCCC2 was purchased from OBiO Technology (Shanghai, China) Corp. Overexpression plasmids and siRNA were transfected with Lipofectamine 3000 (Invitrogen) according to the instruction. Transfection of siRNA in 22Rv1 cells at a final concentration of 50nM. The cells were collected for further investigation at 48 h after transfection.
RNA Isolation and qRT-PCR
Total RNA was extracted using Trizol reagent (Invitrogen Life Technologies, CA, USA) then were reverse transcribed into cDNA by PrimeScriptTM RT reagent kit (TAKARA, Japan). After mixing cDNA and TB GreenR® Premix Ex TaqTM and primers, qRT-PCR was performed on the 7500 Fast Real-Time PCR System (Applied Biosystem). Method 2−ΔΔCT was used to calculate the relative difference of gene expression level. The following primer sequences are used: GAPDH forward 5ʹ-TCCTCTGACTTCAACAGCGACACC-3ʹ and reverse 5ʹ-TCTCTCTTCCTCTTGTGCTCTTGG-3ʹ, MCCC2 forward 5ʹ-TAGAATCGTGGATGGAAGCAG-3ʹ and reverse 5ʹ-TTTGAAGGAACAGCAGAGGAA-3ʹ, GLUD1 forward 5ʹ-GGGATTCTAACTACCACTTGCTCA-3ʹ and reverse 5ʹ-AACTCTGCCGTGGGTACAAT-3ʹ, GLUD2 forward 5ʹ-CACTCTGCCTTGGCATACAC-3ʹ and reverse 5ʹ-CTCAGGTCCAATCCCAGGTT-3ʹ.
Western Blot Analysis
Cells were lysed by using RIPA buffer supplemented with 1% PMSF for preparing whole-cell extracts. Protein concentration was quantified with BCA reagent (KeyGen Biotech, Nanjing, China) and equal amounts of total proteins were separated on 10% SDS-PAGE and transferred to PVDF membranes (Millipore, USA). Then, the membranes were blotted with antibody against MCCC2 (ab197297, Abcam, 1:500), GLUD1 (ab168352, Abcam, 1:500), GLUD2 (39,038, SAB, 1:500) or p38 MAPK (9212, CST, 1:500). GAPDH (2118, CST, 1:1000) was used to normalize the protein level. Visualization was then carried out using their own secondary antibodies (Affinity Biosciences. OH. USA) and ECL reagents (Advansta, USA).
Cell Proliferation Assay
22Rv1 cells transfected with siRNA and siNC, LNCaP cells transfected with pCDNA3.1(+)-Flag-MCCC2 and pCDNA3.1 (+)-Flag-NC. They were transferred to 96-well plates at 3000 cells per well. Detection was performed at 24, 48, 72 and 96h after transfection, and 10 μL of CCK-8 (Dojindo, Japan) were added and incubated at 37°C for 1 h, the absorbance at 450nm was determined by Automatic microplate reader (BioTek, USA).
Cell Migration and Invasion Assays
Cells were transfected for 24 h and plated in the upper chamber of the transwell in 24-well plates (8.0 µm Transparent PET Membrane, Corning, NY, USA) at 1×105 cells in 200μL of serum-free medium per chamber and for invasion there was Matrigel (BD Pharmingen) added into the upper chamber, 500 μL RPMI-1640 medium with 15% FBS were added into the lower chamber as a chemoattractant. After 48h, cells in the upper membrane were removed completely and 4% paraformaldehyde fixed for 15 minutes, then 1% crystal violet stained for 10 minutes, pictures were taken under the inverted microscope (NIKON, Japan).
Analysis of Apoptosis Rate
Cells were collected and suspended in Binding Buffer and stained with Propidium Iodide (10 μL/mL), Annexin V-FITC conjugate (10 μL/mL) (KeyGEN Biotech, Nanjing, China) for 15 min according to the product instructions, and then detected by flow cytometry.
Cell Cycle Analysis
Cells were collected and suspended in PBS and fixed in 70% cold ethanol for 4 h at 4 °C and stained with Propidium Iodide and RNase A (KeyGen Biotech, Nanjing, China) in the dark for 30 min at room temperature according to the product instructions, and then detected by flow cytometry.
Mitochondrial Membrane Potential Analysis
Cells were collected and suspended in JC-1 working solution (for every 1 mL JC-1 working solution there was 2 μL 500× JC-1, 900 μL sterilized deionized water and 100 μL 10× Incubation Buffer) and incubated in a humidified incubator at 37 °C with 5% CO2 for 15min. Then, the cells were washed twice with 1× Incubation Buffer. Finally, resuscitate cells with 1 × Incubation Buffer according to the product instructions of JC-1 Apoptosis Detection Kit (KeyGen Biotech, Nanjing, China), and then detected by flow cytometry.
Functional Enrichment Analysis
We searched the mRNA sequencing data of PCa patients in the TCGA database and analyzed the genes that positively correlated with MCCC2 mRNA expression (Spearman correlation analysis, correlation coefficient <0.3 means low correlation, 0.3–0.6 means moderate correlation, >0.6 means high correlation). KEGG enrichment analysis was carried out by DAVID (https://david.ncifcrf.gov/), and the R language ggplot2 package was used to draw the bubble chart according to the results.
Immunohistochemistry (IHC)
The tissue was achieved from patients who underwent radical prostatectomy at the Third Affiliated Hospital of Sun Yat-sen University. The study involving human tissues was approved by the Ethics Committee of our Institutional. All patients signed informed consent forms and agreed to use the tissues which were surgically removed. The prostate tissue was embedded in paraffin, then paraffin was removed with xylene and hydrated with ethanol. And, 0.3% hydrogen peroxide (H2O2) and citrate buffer (pH6.0) were used to block the endogenous peroxidase and repair the antigen. Slides were incubated with primary antibodies against MCCC2 (ab197297, Abcam, 1:500) or GLUD1 (ab168352, Abcam, 1:500) overnight at 4 °C. All immunostaining scores were independently confirmed by two pathologists who were blinded with patients’ clinical information. Three pictures were taken by each pathologist; therefore, each sample had 6 pictures taken. Then, 3 pictures were randomly selected in the final analysis. The staining intensity of cells was divided into negative, weak positive, moderate positive and strong positive, which were recorded as 0, 1, 2 and 3 points, respectively. The proportion of positive cells <5%, ≥5% ~ <25%, 25% ~ <50%, ≥50% ~ <75% and ≥75% were recorded as 0, 1, 2, 3 and 4 points, respectively. Total immunohistochemical score for each picture was calculated by multiplying the above two scores. The average immunohistochemical score was generated from scores of three pictures from each sample. The score <8 was regarded as low expression, whereas ≥8 as high expression. Expression data of MCCC2 and GLUD1 were achieved in the cancer genome atlas (https://portal.gdc.cancer.gov/projects).
Statistical Analysis
Results were repeated independently in triplicates. Data are expressed by mean ± standard deviation. The statistical differences were analyzed by appropriate tests. Two-tailed P<0.05 indicated statistical significance. All analyses were conducted with SPSS version 20.0 (SPSS Company, Chicago, Illinois, USA).
Results
The Expression of MCCC2 in PCa Cell Lines
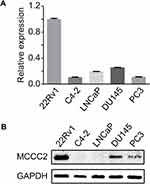
We first analyzed the mRNA expression of MCCC2 in PCa cell lines by qRT-PCR and found that it was highly expressed in 22Rv1 cells and significantly decreased in C4-2, LNCaP (Figure 1A). Similar results in protein expression were confirmed by Western blot (Figure 1B). In order to further investigate the cancer-related biological effect of MCCC2 on PCa, 22Rv1 and LNCaP cells were selected as research models.
High Expression of MCCC2 Promotes Proliferation, Migration and Invasion of Prostate Cancer Cells
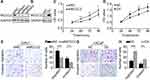
MCCC2-specific siRNA was transfected into 22Rv1 cells and recombinant plasmids that overexpressing Flag-MCCC2 was transfected into LNCaP cells. The efficiency of MCCC2-specific siRNA and overexpressing plasmids were confirmed by Western blot (Figure 2A and B). CCK-8 assays showed that cell proliferation of 22Rv1 cells was significantly inhibited when MCCC2 expression was suppressed by siRNA. Conversely, the proliferation of LNCaP cells was promoted after MCCC2 overexpression (P<0.05, Figure 2C and D). For cell migration, the number of transmembrane 22Rv1 cells in siRNA group was significantly less than that in the control group (32.86±3.72 versus 20.14±3.34, P<0.001), while the number of transmembrane LNCaP cells in the high expression group was significantly more than that in the control group (13.60±2.70 versus 31.80±4.50, P<0.001). For invasion, the invasiveness of 22Rv1 cells decreased significantly after inhibiting the expression of MCCC2 (P < 0.001). After overexpression of MCCC2, the invasiveness of LNCaP cells was significantly enhanced (Prun0.003) (Figure 2E–H). In general, MCCC2 can promote cell proliferation, migration and invasion of PCa cells.
MCCC2 Inhibits Apoptosis, Promote Cell Division and Reduce Mitochondrial Damage
We next explore the influence of MCCC2 on apoptosis, cell cycle and mitochondrial membrane potential of PCa cells. As shown in Figure 3A–D, annexin-V assays showed siRNA-induced inhibition of MCCC2 in 22Rv1 cells increased the apoptosis (P=0.0083), whereas ectopic overexpression of MCCC2 decreased apoptosis in LNCaP (P=0.003). For cell cycle, down-regulation of MCCC2 in 22Rv1 cells led to G0/G1 arrest (P = 0.035, Figure 3E and F). In contrast, up-regulation of MCCC2 in LNCaP cells resulted in the decrease of G0/G1 (P=0.016) phase arrest and the increase of S (P=0.035) phase cell percentage (Figure 3G and H). Since MCCC2 is located in mitochondria, and maintaining normal mitochondrial membrane potential is essential for cell survival. We used JC-1 Apoptosis Detection Kit to detect the changes in mitochondrial membrane potential. The results of JC-1 staining showed that down-regulation of MCCC2 in 22Rv1 cells induced mitochondrial damage (P<0.001), while up-regulation of MCCC2 in LNCaP cells reduced mitochondrial damage (P=0.004, Figure 3I–L). In general, MCCC2 can inhibit PCa cells’ apoptosis, promote cell division and reduce mitochondrial damage.
MCCC2 Led to PCa Progression via GLUD1-p38 MAPK Axis
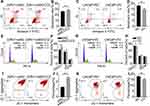
We analyzed the sequencing data of 499 patients with PCa in TCGA database, and the results showed that a total of 84 genes were moderately associated with MCCC2 (Spearman correlation analysis, correlation coefficient >0.3, P<0.05). KEGG enrichment analysis showed that metabolic pathway is one of the related pathways ((Figure 4A). The metabolic pathway enriches a total of 15 genes including MCCC2 (Figure 4B). Among the 15 genes, MCCC2, GLUD1 and GLUD2 are all located in mitochondria matrix. Further, we analyzed mRNA and protein expression of GLUD1 and GLUD2 in 22Rv1 cells and LNCaP cells upon manipulation of MCCC2 expression. There was no significant relationship between MCCC2 and GLUD1 or GLUD2 in the mRNA expression (Figure 4C and D). However, GLUD1 protein expression varied with MCCC2 while GLUD2 did not (Figure 4E). These results indicated that MCCC2 may regulate the expression of GLUD1 at the protein level rather than mRNA. Moreover, the expression of p38 MAPK was down-regulated in 22RV1 cells upon MCCC2 inhibition whereas it was up-regulated in LNCaP cells upon overexpression of MCCC2 (Figure 4E). Based on these results, MCCC2 may promote the PCa progression via GLUD1-p38 MAPK axis.
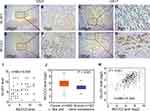
Relationship Between MCCC2 and GLUD1 Expression in PCa Tissue
The expression of MCCC2 and GLUD1 was detected by immunohistochemistry in 68 patients with PCa. Table 1 summarizes the basic clinical features of these 68 patients. The expression of MCCC2 was not related to age of onset (P=0.594), preoperative prostate-specific antigen (PSA) (P=0.454), tumor stage (P=0.969) and seminal vesicle invasion (P=0.523). Interestingly, increased expression of MCCC2 in the PCa tissues was associated with higher Gleason score (P<0.001) and lymph node metastasis (P = 0.001319, Table 1), which are important measurements of aggressiveness. The results also showed that there was a positive correlation between MCCC2 and GLUD1, with the correlation coefficient of 0.435 (P<0.001, Figure 5A–I and Table 1). We also analyzed the expression data of MCCC2 and GLUD1 in 499 cancer and 52 normal samples. TCGA data validated MCCC2 was increased in PCa compared to normal samples, with the fold change of 2.39, P<0.001, Figure 5J). Coexpression analysis showed a positive correlation of MCCC2 and GLUD1 expression in PCa, with correlation coefficient of 0.444 (P <0.001, Figure 5K). This provided clinical evidence that MCCC2 positively regulated GLUD1 thus promoting PCa progression.
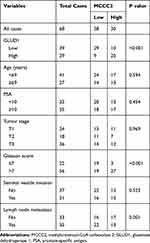 |
Table 1 Clinicopathological Characteristics and MCCC2 Expression of 68 Prostate Cancer Patients |
Discussion
In our present study, we demonstrated that MCCC2 promoted cell proliferation, migration and invasion but inhibited apoptosis in PCa cells. In addition, MCCC2 overexpression associated with unfavorable clinical features such as lymph node metastasis and high Gleason scores. We performed KEGG enrichment analysis and found that MCCC2 was mainly related to metabolic pathway in PCa. What is more, we demonstrated that MCCC2 positively correlated with GLUD1 protein expression in PCa tissues and regulated GLUD1-p38 MAPK axis in PCa cells. Our results suggest that MCCC2 may promote the progression and metastasis of prostate cancer via regulating GLUD1-p38 MAPK axis.
Cancer cells require constant energy supply to keep rapid growth and division. Glucose is the most abundant nutrient utilized by cancer cells. The vast majority of solid cancers accommodate altered energy requirement via increasing glucose uptake and glycolysis, which is called Warburg effect.14 Although aerobic glycolysis is inefficient in ATP production per unit of glucose compared to mitochondrial respiration, the rate of glucose metabolism is very high, such that the amount of ATP biosynthesized are comparable in either means. In the tumor microenvironment, tumor cells compete with stromal cells and the immune compartment for limited resources such as nutrients and oxygen, thus cells may gain advantages of aerobic glycolysis.15 However, PCa does not seem to get energy through Warburg effect, but relies heavily on amino acids and lipids to generate energy.16,17
Human methylcrotonyl-CoA carboxylase (MCC) which is formed by subunits MCCCα and MCCCβ encoded by MCCC1 and MCCC2 is a biotin-dependent mitochondrial enzyme that participates in the fourth step of leucine metabolism.18,19 Leucine is validated as a crucial activator of mammalian target of rapamycin complex 1 (mTORC1) by triggering the Rag Guanosine triphosphate (GTP) binding proteins which in turn stimulates mTORC1 translocation to the lysosome surface where it is activated. Aberrant activation of mTORC1 signaling is common in many cancers, and consequently, it promotes cancer progression through transactivating pathways that support cell proliferation, survival and invasiveness.20 L-type amino acid transporters (LAT) mediate the entry of leucine into cancer cells and promote mTORC1 activity, reinforces MYC and EZH2 signaling.21 Elegant studies revealed that LAT1 and LAT3 expression were highly associated with cell cycle progression, E2F transcription factors and high-grade metastatic and hormone-resistant PCa.22,23 Inhibiting leucine transporters could lead to significant cell growth inhibition in PCa cells as well as tumor growth and metastasis in xenografts, suggesting anti-leucine transporter therapy could be employed as an important option against PCa.23,24 Our study found high expression of MCCC2 in highly proliferative PCa cell line 22RV1 whereas low expression in low-speed proliferative cell line LNCaP, the difference in MCCC2 expression reflects energy supply which is important for cancer cell proliferation. We also confirmed MCCC2 associated with unfavorable clinical features and promoted aggressiveness of PCa cells in vitro, adding evidence that MCCC2 may modulate leucine metabolism to exhibit an oncogenic role in PCa.
Glutamine is another essential nutrient for energy production, as it provides not only carbon for biosynthesis but also nitrogen for anabolic reactions.25 Glutamate dehydrogenase 1 (GLUD1), as a key mitochondrial enzyme in glutaminolysis, is transactivated by the direct binding of leucine and makes glutamine-derived carbon enter the tricarboxylic acid (TCA) cycle by converting glutamate to α-ketoglutarate (α-KG).26,27 GLUD1 is commonly elevated in human cancers enabling the cancer cells to produce ATP and synthesize amino biomacromolecules (amino acids, nucleotides and lipids, etc) for rapid growth and proliferation.28 It has been reported that GLUD1 is overexpressed in childhood acute myeloid leukemia, glioma, colorectal or lung cancer,29–31 while GLUD1 expression differs in molecular subtypes of breast cancer.32 Mechanistically, GLUD1 plays a predominant role in controlling intracellular levels of α-KG and consequently fumarate, which activates glutathione peroxidase 1 (GPx) to increase reactive oxygen species (ROS), regulating redox homeostasis to promote cancer cell proliferation and tumor growth.30 Anoikis is a programmed cell death resulting from loss of cell and extracellular membrane, which is a known physiological barrier for cancer metastasis.33,34 GLUD1 activates CamKK2-AMPK1 signaling to contribute to bioenergetics that confers anoikis resistance, making cancer cells survive during dissemination and metastatic colonization in distant organs.34 In the cancer cells, GLUD1 also has a role in the activation of mTORC1, which can be stimulated by leucine.32 Interestingly, GLUD1 catalyzes reductive amination to maximize nitrogen utilization via metabolic recycling of ammonia, thus accelerating cell proliferation and supporting tumor biomass of breast cancer.35 In PCa, GLUD1 expression is positively correlated with Gleason score and significantly associated with shorter survival.36 Gleason score is a commonly used scoring system to evaluate the aggressiveness of PCa. Consistently, our results showed MCCC2 expression positively correlated with GLUD1. The qRT-PCR results show that MCCC2 cannot affect the mRNA level of GLUD1, but Western blot confirmed MCCC2 up-regulated protein expression of GLUD1. This may suggest that MCCC2 positively regulates GLUD1 in the post-translational modification stage.
It is well known that p38 MAPK participates in the proliferation and cycle of many kinds of tumor cells. Although most studies of p38 MAPK show that it is mainly anti-proliferation in cancer, this effect is the opposite in prostate cancer. For example, in an in vitro study of PCa, it has been reported that TBL-12 can inhibit the proliferation, migration and invasion of PCa cells due to its down-regulation of MMP-2/MMP-9 by inhibiting p38 MAPK signaling.37 In addition, COPS3 gene knockout inhibits the progression of PCa by reducing the expression of phosphorylated p38 MAPK.38 A study of LNCaP cells showed that apoptosis and proliferation inhibition induced by exposure to high concentrations of leptin were associated with reduced p38 MAPK expression.39 Nontoxic EEAC can reduce the expression of GLUD1 in hepatoma cells, and it can inhibit the migration of hepatoma cells by the down-regulation of MAPK.40 These studies remind us that p38 may be a link in which GLUD1 changes the behavior of prostate cancer cells. We further confirm this conjecture that MCCC2 may regulate the progression and metastasis of prostate cancer through the GLUD1-p38 MAPK axis.
Little is known about MCCC2 function in cancers, here we uncovered a new underlying mechanism that MCCC2 promoted GLUD1 expression and subsequent activation of p38 MAPK signaling, which shed insights into the understanding of MCCC2 function in PCa. In terms of therapeutic implication, inhibiting GLUD1 achieved tumor-suppressing effect in many kinds of cancer cells,30 thus targeting MCCC2 may be a treatment strategy in PCa.
This study also has some limitations. Although statistic differences were confirmed, the number of PCa patients whose cancerous tissues were used to determine MCCC2 and GLUD1 immunostaining is relatively small. In addition, the predictive values were not evaluated due to the limited data of follow-up. Thirdly, the tumor-promoting role of MCCC2 needs further verification in the animal models.
Conclusion
MCCC2 can promote cell proliferation, migration, invasion and survival via regulating GLUD1-p38 MAPK axis, suggesting MCCC2 exerts oncogenic function and may be a potential treatment target in PCa.
Acknowledgments
This work was supported by Pearl River S&T Nova Program of Guangzhou (201710010039), Basic Service Charge Young Teachers Cultivation Project of Sun Yat-sen University (17ykpy48), and National Natural Science Foundation (81771573).
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Disclosure
The authors declare no conflict of interest in this work.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69(1):7–34. doi:10.3322/caac.21551
2. Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. doi:10.3322/caac.21338
3. Litwin MS, Tan HJ, Diagnosis T. Treatment of prostate cancer: a review. JAMA. 2017;317(24):2532–2542. doi:10.1001/jama.2017.7248
4. Abe S, Chamnan C, Miyamoto K, Minamino Y, Nouda M. Isolation and identification of 3-methylcrotonyl coenzyme A carboxylase cDNAs and pyruvate carboxylase, and their expression in red seabream (Pagrus major) organs. Mar Biotechnol (NY). 2004;6(6):527–540. doi:10.1007/s10126-004-4203-x
5. Stadler SC, Polanetz R, Maier EM, et al. Newborn screening for 3-methylcrotonyl-CoA carboxylase deficiency: population heterogeneity of MCCA and MCCB mutations and impact on risk assessment. Hum Mutat. 2006;27(8):748–759. doi:10.1002/humu.20349
6. Morscher RJ, Grunert SC, Burer C, et al. A single mutation in MCCC1 or MCCC2 as a potential cause of positive screening for 3-methylcrotonyl-CoA carboxylase deficiency. Mol Genet Metab. 2012;105(4):602–606. doi:10.1016/j.ymgme.2011.12.018
7. Grunert SC, Stucki M, Morscher RJ, et al. 3-methylcrotonyl-CoA carboxylase deficiency: clinical, biochemical, enzymatic and molecular studies in 88 individuals. Orphanet J Rare Dis. 2012;7(1):31. doi:10.1186/1750-1172-7-31
8. Cozzolino C, Villani GR, Frisso G, et al. Biochemical and molecular characterization of 3-Methylcrotonylglycinuria in an Italian asymptomatic girl. Genet Mol Biol. 2018;41(2):379–385. doi:10.1590/1678-4685-gmb-2017-0093
9. Meierhofer D, Halbach M, Sen NE, Gispert S, Auburger G. Ataxin-2 (Atxn2)-knock-out mice show branched chain amino acids and fatty acids pathway alterations. Mol Cell Proteomics. 2016;15(5):1728–1739. doi:10.1074/mcp.M115.056770
10. Zandberg L, van Dyk HC, van der Westhuizen FH, van Dijk AA. A 3-methylcrotonyl-CoA carboxylase deficient human skin fibroblast transcriptome reveals underlying mitochondrial dysfunction and oxidative stress. Int J Biochem Cell Biol. 2016;78:116–129. doi:10.1016/j.biocel.2016.07.010
11. Pang J, Liu WP, Liu XP, et al. Profiling protein markers associated with lymph node metastasis in prostate cancer by DIGE-based proteomics analysis. J Proteome Res. 2010;9(1):216–226. doi:10.1021/pr900953s
12. Marques RB, Dits NF, Erkens-Schulze S, van Ijcken WF, van Weerden WM, Jenster G. Modulation of androgen receptor signaling in hormonal therapy-resistant prostate cancer cell lines. PLoS One. 2011;6(8):e23144. doi:10.1371/journal.pone.0023144
13. Liu Y, Yuan Z, Song C. Methylcrotonoyl-CoA carboxylase 2 overexpression predicts an unfavorable prognosis and promotes cell proliferation in breast cancer. Biomark Med. 2019;13(6):427–436. doi:10.2217/bmm-2018-0475
14. Pacini N, Borziani F. Cancer stem cell theory and the warburg effect, two sides of the same coin? Int J Mol Sci. 2014;15(5):8893–8930. doi:10.3390/ijms15058893
15. Liberti MV, Locasale JW. The warburg effect: how does it benefit cancer cells? Trends Biochem Sci. 2016;41(3):211–218. doi:10.1016/j.tibs.2015.12.001
16. Twum-Ampofo J, Fu DX, Passaniti A, Hussain A, Siddiqui MM. Metabolic targets for potential prostate cancer therapeutics. Curr Opin Oncol. 2016;28(3):241–247. doi:10.1097/CCO.0000000000000276
17. Zadra G, Photopoulos C, Loda M. The fat side of prostate cancer. Biochim Biophys Acta. 2013;1831(10):1518–1532. doi:10.1016/j.bbalip.2013.03.010
18. Baumgartner MR, Almashanu S, Suormala T, et al. The molecular basis of human 3-methylcrotonyl-CoA carboxylase deficiency. J Clin Invest. 2001;107(4):495–504. doi:10.1172/JCI11948
19. Fonseca H, Azevedo L, Serrano C, Sousa C, Marcao A, Vilarinho L. 3-Methylcrotonyl-CoA carboxylase deficiency: mutational spectrum derived from comprehensive newborn screening. Gene. 2016;594(2):203–210. doi:10.1016/j.gene.2016.09.003
20. Saxton RA, Sabatini DM. mTOR signaling in growth, metabolism, and disease. Cell. 2017;168(6):960–976. doi:10.1016/j.cell.2017.02.004
21. Salisbury TB, Arthur S. The regulation and function of the L-Type Amino Acid Transporter 1 (LAT1) in cancer. Int J Mol Sci. 2018;19(8):2373. doi:10.3390/ijms19082373
22. Wang Q, Bailey CG, Ng C, et al. Androgen receptor and nutrient signaling pathways coordinate the demand for increased amino acid transport during prostate cancer progression. Cancer Res. 2011;71(24):7525–7536. doi:10.1158/0008-5472.CAN-11-1821
23. Wang Q, Tiffen J, Bailey CG, et al. Targeting amino acid transport in metastatic castration-resistant prostate cancer: effects on cell cycle, cell growth, and tumor development. J Natl Cancer Inst. 2013;105(19):1463–1473. doi:10.1093/jnci/djt241
24. Otsuki H, Kimura T, Yamaga T, Kosaka T, Suehiro JI, Sakurai H. Prostate cancer cells in different androgen receptor status employ different leucine transporters. Prostate. 2017;77(2):222–233. doi:10.1002/pros.23263
25. Hensley CT, Wasti AT, DeBerardinis RJ. Glutamine and cancer: cell biology, physiology, and clinical opportunities. J Clin Invest. 2013;123(9):3678–3684. doi:10.1172/JCI69600
26. Coloff JL, Murphy JP, Braun CR, et al. Differential glutamate metabolism in proliferating and quiescent mammary epithelial cells. Cell Metab. 2016;23(5):867–880. doi:10.1016/j.cmet.2016.03.016
27. Tzimagiorgis G, Leversha MA, Chroniary K, et al. Structure and expression analysis of a member of the human glutamate dehydrogenase (GLUD) gene family mapped to chromosome 10p11.2. Hum Genet. 1993;91(5):433–438. doi:10.1007/BF00217767
28. Plaitakis A, Kalef-Ezra E, Kotzamani D, Zaganas I, Spanaki C. The glutamate dehydrogenase pathway and its roles in cell and tissue biology in health and disease. Biology. 2017;6(1):11.
29. Kanderova V, Kuzilkova D, Stuchly J, et al. High-resolution antibody array analysis of childhood acute leukemia cells. Mol Cell Proteomics. 2016;15(4):1246–1261. doi:10.1074/mcp.M115.054593
30. Jin L, Li D, Alesi GN, et al. Glutamate dehydrogenase 1 signals through antioxidant glutathione peroxidase 1 to regulate redox homeostasis and tumor growth. Cancer Cell. 2015;27(2):257–270. doi:10.1016/j.ccell.2014.12.006
31. Wang YQ, Wang HL, Xu J, et al. Sirtuin5 contributes to colorectal carcinogenesis by enhancing glutaminolysis in a deglutarylation-dependent manner. Nat Commun. 2018;9(1):545. doi:10.1038/s41467-018-02951-4
32. Craze ML, El-Ansari R, Aleskandarany MA, et al. Glutamate dehydrogenase (GLUD1) expression in breast cancer. Breast Cancer Res Treat. 2019;174(1):79–91. doi:10.1007/s10549-018-5060-z
33. Paoli P, Giannoni E, Chiarugi P. Anoikis molecular pathways and its role in cancer progression. Biochim Biophys Acta. 2013;1833(12):3481–3498. doi:10.1016/j.bbamcr.2013.06.026
34. Jin L, Chun J, Pan C, et al. The PLAG1-GDH1 axis promotes anoikis resistance and tumor metastasis through CamKK2-AMPK signaling in LKB1-deficient lung cancer. Mol Cell. 2018;69(1):87–99 e87. doi:10.1016/j.molcel.2017.11.025
35. Spinelli JB, Yoon H, Ringel AE, Jeanfavre S, Clish CB, Haigis MC. Metabolic recycling of ammonia via glutamate dehydrogenase supports breast cancer biomass. Science. 2017;358(6365):941–946. doi:10.1126/science.aam9305
36. Li Y, Li X, Li X, et al. PDHA1 gene knockout in prostate cancer cells results in metabolic reprogramming towards greater glutamine dependence. Oncotarget. 2016;7(33):53837–53852. doi:10.18632/oncotarget.10782
37. Yuan L, Huang X, Zhou K, et al. Sea cucumber extract TBL-12 inhibits the proliferation, migration, and invasion of human prostate cancer cells through the p38 mitogen-activated protein kinase and intrinsic caspase apoptosis pathway. Prostate. 2019;79(8):826–839. doi:10.1002/pros.23788
38. Zhu Z, Hong Y, Zhang F, et al. Knockdown of COPS3 inhibits the progress of prostate cancer through reducing phosphorylated p38 MAPK expression and impairs the epithelial-mesenchymal transition process. Prostate. 2019;79(16):1823–1831. doi:10.1002/pros.23907
39. Szyszka M, Paschke L, Tyczewska M, et al. Analysis of transcriptome, selected intracellular signaling pathways, proliferation and apoptosis of LNCaP cells exposed to high leptin concentrations. Int J Mol Sci. 2019;20(21):5412. doi:10.3390/ijms20215412
40. Chen YY, Liu FC, Wu TS, Sheu MJ. Antrodia cinnamomea inhibits migration in human hepatocellular carcinoma cells: the role of ERp57 and PGK-1. Am J Chin Med. 2015;43(8):1671–1696. doi:10.1142/S0192415X15500950
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.