Back to Journals » International Journal of Nanomedicine » Volume 10 » Issue 1
Magnetic removal of Entamoeba cysts from water using chitosan oligosaccharide-coated iron oxide nanoparticles
Authors Shukla S, Arora V, Jadaun A, Kumar J, Singh N, Jain V
Received 18 November 2014
Accepted for publication 22 January 2015
Published 31 July 2015 Volume 2015:10(1) Pages 4901—4917
DOI https://doi.org/10.2147/IJN.S77675
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Thomas Webster
Sudeep Shukla,1 Vikas Arora,2 Alka Jadaun,3 Jitender Kumar,1 Nishant Singh,1 Vinod Kumar Jain1
1School of Environmental Sciences, Jawaharlal Nehru University, New Delhi, Delhi, India; 2Department of Chemistry, Indian Institute of Technology, New Delhi, Delhi, India; 3School of Biotechnology, Jawaharlal Nehru University, New Delhi, Delhi, India
Abstract: Amebiasis, a major health problem in developing countries, is the second most common cause of death due to parasitic infection. Amebiasis is usually transmitted by the ingestion of Entamoeba histolytica cysts through oral–fecal route. Herein, we report on the use of chitosan oligosaccharide-functionalized iron oxide nanoparticles for efficient capture and removal of pathogenic protozoan cysts under the influence of an external magnetic field. These nanoparticles were synthesized through a chemical synthesis process. The synthesized particles were characterized by transmission electron microscopy, Fourier transform infrared spectroscopy, X-ray diffraction, and zeta potential analysis. The particles were found to be well dispersed and uniform in size. The capture and removal of pathogenic cysts were demonstrated by fluorescent microscopy, transmission electron microscopy, and scanning electron microscopy (SEM). Three-dimensional modeling of various biochemical components of cyst walls, and thereafter, flexible docking studies demonstrate the probable interaction mechanism of nanoparticles with various components of E. histolytica cyst walls. Results of the present study suggest that E. histolytica cysts can be efficiently captured and removed from contaminated aqueous systems through the application of synthesized nanoparticles.
Keywords: amebiasis, water treatment, nanotechnology
Introduction
With the expansion of the human imprint on the planet, sustained availability of key environmental resources is becoming a challenge. Among these challenges, availability of clean water is one of the top issues being debated. It is not just availability of fresh water, but the quality of it that is a key concern. The diseases caused due to the presence of harmful pathogens in natural water bodies are a major cause of morbidity and mortality affecting millions of individuals worldwide each year, especially in developing countries such as India.1–3 The most common pathogens that cause waterborne diseases are viruses, bacteria, and protozoa.
Among the protozoa, Entamoeba histolytica, Giardia lamblia, and Cryptosporidium parvum are the major causes of waterborne diseases.2,4 Amebiasis, caused by the ingestion of E. histolytica cysts, is a major health problem in developing countries and is the second most common cause of death due to parasitic infection globally. The ingested cysts get converted into the motile trophozoites in the ileocecal region of the intestine.5,6 Cysts are the infective stage, but so far it has not been possible to induce encystment in axenic E. histolytica trophozoites in culture.5 Therefore, Entamoeba invadens, a reptilian parasite that can be encysted in culture is generally used for the model system.7,8 The Entamoeba spp. cyst cell wall is composed of a fibrillar network of different polysaccharides and lectins.8 The major component of the cyst cell wall is chitin, a β-(1,4)-linked homopolymer of N-acetyl-d-glucosamine. Cyst wall lectins include Jacob (three to seven chitin-binding domains, each of which contains six Cys), Jessie (single N-terminal 8-Cys chitin-binding domains), and chitinase (single N-terminal chitin-binding domains containing eight Cys residues) have already been identified7,9 (Figure 1).
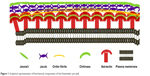  | Figure 1 Graphical representation of biochemical components of the Entamoeba cyst wall. |
With recent development in nanotechnology, various types of metal and metal oxide nanoparticles have been synthesized. Nanoscaled materials are now considered to be foremost among novel water purification materials.10 Magnetic nanoparticles are currently employed as a new tool for biomedical sciences in several ways such as imaging, sensing, and targeted drug delivery.11,12
Chitosan is a highly cationic biopolymer obtained through deacetylation of chitin. Chitin is isolated from crustaceans.13 Chitosan is a non-toxic, edible, and biocompatible polymer frequently used in biomedical and food applications.14,15 It is evident that molecular chitosan is difficult to dissolve in aqueous solutions at neutral pH.14,16 However, chitosan oligosaccharide (CSO) nanoparticles were found to be more stable, non-agglomerated, and well dispersed in aqueous systems at neutral pH and biological pH.17
In the current study, CSO-functionalized iron oxide nanoparticles (CSO-INPs) were synthesized and used for the magnetic separation of E. histolytica cysts from the water sample. Further studies were carried out to investigate the probable binding mechanism of synthesized nanoparticles with the various components of the cyst wall of E. histolytica cysts. The molecular affinity of nanoparticles with the cyst wall glycoprotein was also modeled and simulated.
Materials and methods
For present study, analytical grade chemicals, Iron (III) chloride (97%), oleic acid (90%), n-hexane (95%), anhydrous ethanol, 1-octadecene (90%), acetic acid, N-(3-Dimethylamino-propyl)-N-ethylcarbodiimide hydrochloride (EDC), N-hydroxy-succinimide (NHS), CSO (Mn 5,000 and >90% deacetylated) were purchased from Sigma-Aldrich Co, St Louis, MO, USA. N-[(3-Trimethoxysilyl)propyl] ethylenediaminetetraacetic acid trisodium salt (50% in water) was received from Gelest, Inc, Morrisville, PA, USA. The water used throughout this work was of reagent grade produced by a Milli-Q® water purification system.
Synthesis of nanoparticles
INPs ere synthesized as reported by Jana et al18 with slight modifications.19 In a typical synthesis of an iron–oleate complex, 2.55 g of iron chloride (FeCl3·6H2O) was dissolved in 100 mL of methanol and 11 mL of oleic acid under continuous stirring. Another solution prepared by dissolving 1.6 g of NaOH in 200 mL of methanol was added to the above solution in continuous stirring conditions. The observed brown precipitate of iron oleate was washed with methanol and dried under vacuum overnight to remove the solvent. The resultant synthesized solid mass (4.02 g) was dissolved in 30 mL of 1-octadecene at 70°C to make a stock solution. Thereafter, 10 mL of stock solution was mixed with 40 mL of 1-octadecene, and 0.1 equivalents of oleic acid and the solution were heated to 280°C for 30 minutes in an inert environment. When the reaction was complete, the mixture was precipitated twice with ethanol.
The oleic acid on the particle surface was replaced with a –COOH containing silane using a method reported by De Palma et al.20 Nanoparticles were further functionalized with CSO via carbodiimide activation, using EDC and NHS following a method developed by López-Cruz et al.17 In a typical synthesis, 150 g of silane-functionalized INPs were dissolved in 10 mL of MilliQ water (maintaining pH =8) with 0.04 g EDC and 0.05 g NHS for the carboiimide activation of the functional group, required for the covalent bonding of the amino group chitosan with the terminal carboxylic group of silane-functionalized INPs.17 Another mixture containing 0.2 g CSO lactate (chitosan derivative obtained from Sigma-Aldrich Co) was dissolved in 20 mL in MilliQ water. The two solutions were mixed in continuous stirring conditions, and were kept continuous stirring conditions for 24 hours. The observed milky solution was precipitated by ethanol. The resulting water-soluble precipitate of CSO-INPs was further re suspended in MilliQ water.
Characterization of nanoparticles
Transmission electron microscopy (TEM) images were recorded on a JEOL 2100F; the TEM operated at an accelerating voltage of 200 kV. Samples were prepared by adding 10 μL of the nanoparticles solution on 200-mesh carbon-coated Cu grids. For the rapid counting of nanoparticles, TEM images were further processed by NIH ImageJ software.21 Powder X-ray diffraction (XRD) studies were carried out through a Philips1820 advance diffractometer equipped with Ni-filtered Cu-Kα radiation, maintaining the scan rate of 0.24° per minute. Infrared (IR) spectroscopy studies were carried out by Varian 7000 Fourier transform infrared spectroscopy (FTIR) to determine the chemical functional groups in the sample.
Cell culture and maintenance
E. invadens strain IP-1 was obtained from the American Type Culture Collection and maintained at 25°C in TYI-S-33 growth medium22 containing 15% heat-inactivated adult bovine serum, 125 μL/100 mL streptomycin (Nicholas Piramel India Limited, Maharashtra, India) and 2.5% vitamin mix (Sigma-Aldrich Co).
Cyst induction
Trophozoites were grown and kept on chilled ice for 10 minutes to collect the cells from the wall. Trophozoites (5×105/mL), at their log phase, were grown and centrifuged at 500 g for 5 minutes at 4°C, and thereafter were moved into induction medium (low glucose [LG]). Trypticase-Yeast Extract-Iron-Serum (TYI) medium (without glucose) was introduced and diluted to 2.12 times in MilliQ.5 Thereafter, the medium was added to 5% heat-inactivated adult bovine serum, 2.6% vitamin mix, and streptomycin (125 μL/100 mL).22 Cysts were incubated in LG medium for 3 days. They were harvested and treated with 0.05% sarkosyl.7 The spherical refractile cysts were identified using 0.01% calcofluor staining (Sigma-Aldrich Co).5
Removal of cysts in simulated conditions
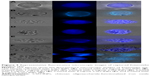
After incubating synthesized nanoparticles with cyst-spiked water, a magnetic field powered by permanent magnet was applied to separate out the nanoparticle/cyst aggregates. The supernatant was removed, and the remaining aggregates were washed thoroughly, stained with fluorescent dye calcofluor, transferred to a glass slide, and imaged.
Capture efficiency of nanoparticles was calculated by dividing the number of Entamoeba spp. in CSO-INP aggregates over the total number of cysts in both the supernatant and the aggregate. To evaluate the optimum capture efficiency of the synthesized nanoparticles, varied concentration of nanoparticles (0.5 mg/mL, 1 mg/mL, 2 mg/mL, 3 mg/mL, 4 mg/mL, and 5 mg/mL) were mixed with Entamoeba cyst-containing solutions (105 cells/mL) in 1 mL of Milli-Q (pH 7). This mixture was incubated for different time intervals (5, 10, 15, 20, 25, 35, and 45 minutes) and then analyzed for capture efficiency at room temperature while continuously shaking in a 2 mL microfuge tube. After incubation, a constant magnetic field was applied for separating nanoparticles and Entamoeba cyst aggregates. The supernatant was carefully removed, and the remaining aggregates were washed thoroughly with fresh 1× phosphate-buffered saline (PBS) buffer (pH 7.4). The aggregate was re-suspended in PBS buffer and stained with 0.01% calcofluor fluorescent dye. It has already been reported that calcofluor binds the cysts’ membrane.8 The cysts were subsequently transferred to a glass slide and imaged under fluorescent microscope. For negative controls, one tube was taken without nanoparticles but retaining the cysts.
A TEM (JEOL 2100F) study was also carried out for investigation of linkage of Entamoeba cysts with synthesized CSO-INPs. CSO-INPs (4 mg/mL) were added to Entamoeba cyst-containing solution diluted in 1 mL of 1× PBS buffer (pH 7.4) and visualized under TEM after fixation. The presence of CSO-INPs on the surface of cyst walls was further confirmed by the scanning electron microscopy coupled with energy dispersive X-ray (SEM-EDS) elemental analysis (Zeiss EVO40).
Modeling for binding of CSO with different structural components of Entamoeba cyst walls
In the present study, existing cyst wall structural models9 were used for the investigation of the interaction of various components of cyst cell wall (Figure 1) with synthesized CSO-INPs.
Screening and modeling of various cyst wall components and CSO
CSO was used as a ligand for this analysis. The structure of CSO was retrieved by CORINA software.23 The structure of one of the receptors, chitin, was adopted from the Pubchem database (https://pubchem.ncbi.nlm.nih.gov/substance/599272#section=Top). As the structure of other receptor, namely chitinase, Jacob and Jessie lectins were not available in the Pubchem database, so their amino acid sequences were obtained from the National Center for Biotechnology Information (NCBI) database. A Basic Local Alignment Search Tool (BLAST) search was performed against the Protein Data Bank (PDB) for homology modeling with similar sequence and known three-dimensional (3D) structures (http://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastp&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome). PDB-BLAST was repeated four times for the best template selection (in the current study, we could not model Jessie lectin, as we did not find any available database for it). These PDB templates were selected on the basis of high score, maximum percentage identity, and low E-value. Chitinase and Jacob were modeled through the ESyPred3D program. The ESyPred3D program is an automated homology modeling program based on an artificial neural networks computational model.24 Final models were visualized by PyMOL viewer.25
Validation of 3D homology model structures
After homology modeling of chitinase and Jacob, their structures were validated through three structure verification programs, PROCHECK,26 VERIFY_3D,27 and ProSA.28 These verification programs validate the predicted model on the basis of various parameters. The specific position of each amino acid residue in each model was validated by the PROCHECK program that relies on Ramachandran plots.29 The VERIFY_3D program determines the compatibility of an atomic model (3D) with its own amino acid sequence (1D), based on its position and surroundings. The ProSA program generates an energy plot. The energy plot provides z scores on the basis of total interaction energy. The z score was further used for the validation of accuracy and stability of the model.
Automated flexible docking of chitinase, Jacob, and chitin with CSO
Final verified models of the biochemical components of cyst wall were docked against CSO by Autodock software.25 For adequate simulation salvation parameters, Kollman charges and polar hydrogens were added onto the protein structures. Gasteiger charge and non-polar hydrogens were assigned to CSO.23 This docking process was based on Lamarckian genetic algorithm with a population size of 2,000; maximum number of energy evaluation was 10 million and maximum number of generations was 300,000.30 The best confirmation of docking was received after 100 simulations. Interaction of ligand (CSO) and receptors (chitin, Jacob, and chitinase) was visualized through the PyMOL viewer.
Results and discussion
Characterization of nanoparticles
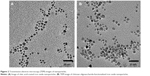
The typical TEM images of INPs and CSO-INPs are shown in Figure 2A and B, respectively. Processing of these TEM images by US National Institutes of Health (NIH) ImageJ software provided mean INP and CSO-INP diameter values of 6.5±1.2 and 9.1±2.3 nm, respectively. Further, it was observed that the INPs and CSO-functionalized nanoparticles were well dispersed and non-aggregated.31–33 XRD patterns of synthesized INPs exhibited peaks at 2θ at 30.1, 35.5, 42.6, 53.6, 57.0, and 62.8, which were respectively assignable to diffraction of the (220), (311), (400), (422), (511), (440) planes of magnetite nanoparticles (JCPDS card number 82-1533).34 However, the broad reflection planes are perhaps due to the nanosize of the synthesized particles (black line in Figure 3). The XRD pattern for CSO-INPs exhibited its two characteristic peaks at 2θ =20.1, 30.1, 35.5, and 62.8. Presence of characteristic peaks at 2θ =20.1 for CSO, along with peaks at 2θ =30.1, 35.5, and 62.8 associated with the iron nanoparticles, confirm the coating of CSO on iron nanoparticles (red line in Figure 3).32,35
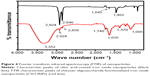
The FTIR graph (Figure 4) indicates the characteristic peaks of oleic acid-coated INPs. The –CH2 stretch is seen at 2,846 cm−1 and again at 2,924 cm−1, C=C at 1,646 cm−1, and the –CH3 stretch can be seen at 1,459 cm−1 (black line in Figure 4), indicating the presence of oleic acid on nanoparticle surfaces. Successful amide formation between amine groups in CSO and the carboxylic group of silane (red line in Figure 4) is confirmed by the appearance of characteristic bands such as the band representing the –OH group at 3,355 cm−1, and the band representing C=O (secondary amide formation) at 1,635 cm−1 for CSO.17,31,32
Zeta potential data in Figure S1A indicates that CSO-INPs were positively charged, with a surface potential greater than +37 mV at pH 3. This confirms the presence of amino groups on the nanoparticle surfaces in their protonated form, thus establishing the presence of CSO on the particle surface. Results indicate that with an increase of pH, the surface charge of the particle decreased, which was probably due to the deprotonation tendency of the surface-exposed amino groups at higher pH values.17 Figure S1B shows that particles possess positive zeta potential of +11 mV at pH 7, which corresponds to the pH of natural water. However, at pH 9, particles show a negative zeta potential of −2.7 mV (Figure S1C). These results confirm that the nanoparticles have sufficient colloidal stability that is necessary for biological and environmental applications.32
Magnetic removal of Entamoeba cysts from spiked water samples
Capture efficiency for the cysts cells treated with CSO-INPs was calculated. Different concentrations of nanoparticles, from 0.5 mg/mL to 5 mg/mL were used along with 105 cysts/mL spiked aqueous solution (Figure 5). Initially, all the experiments were performed at pH 7.0 and at 37°C temperature. The highest capture efficiency was found to be at 4 mg/mL concentration of nanoparticles under the same experimental conditions. By keeping the concentration constant (4 mg/mL) and varying the incubation time, highest capture efficiency (approximately 86%) was observed at 35 minutes (Figures S2 and S3).
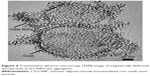
TEM images clearly indicate the successful trapping of Entamoeba cyst by magnetic CSO nanoparticles under the influence of an external magnetic field. TEM images of magnetically defected mass show many cysts entrapped in clusters of nanoparticles (Figures 6 and S4).
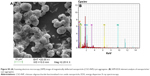
The scanning electron microscopy with energy dispersive X-ray spectroscopy (SEM-EDX) element analysis further confirmed presence of CSO-INPs on the surface of cyst walls. SEM image of magnetically deflected mass clearly indicate the successful trapping of Entamoeba cysts (Figure S5A). Spectral profile of Fe Kα and its corresponding X-ray mapping suggests that iron is almost uniformly distributed on cyst walls (Figure S5B).
Modeling for binding of CSO with different components of Entamoeba cyst walls
CORINA software generated a 3D structure of CSO that was used as a ligand for further modeling (Figure S6). After using BLAST, the best template for chitinase homology modeling was identified as human chitotriosidase (PDB entry 1GUV_A, resolution =2.35 Å) with 39% homology similarity (Figure S7),36 and diisopropyl fluorophosphatase of Loligo vulgaris (PDB entry 2IAX_A, resolution =1.10 Å) was identified as best template for Jacob with 18.2% homology similarity (Figure S8).37 Ramachandran plot of chitinase showed 91.1% amino acid residue in a favorable region, 8.0% residue in an additionally allowed region, and 0.9% residue in a disallowed region (Figure S9A). In contrast, the Ramachandran plot of Jacob showed 71.1% residue in a favorable region, 27.1% residue in an additionally allowed region, and 1.8% residue in a disallowed region (Figure S9B). It was confirmed that most of the residues were located in the phi-psi range, which made the plot reliable for further analysis.26 A ProSA energy plot represented interaction energy of each residue of protein that was identified by the z score. z scores for the chitinase model and the Jacob model were −9.39 and −1.85 kcal/mol, respectively (Figure S10A, B). Chitinase has a more negative z score, which confirms the stability and suitability of the model for further analysis compared to other models.28 The VERIFY_3D program also showed good score as 93.92% and 46.27% for chitinase and Jacob models, respectively (Table S1), which implied that surrounding side chains of the models are acceptable for docking purposes.27
Automated flexible docking analysis of chitinase, Jacob, and chitin with CSO
Docking of CSO against chitinase, Jacob, and chitin was performed via Autodock software. Homology models of chitinase, Jacob, and chitin were used as receptors. Receptor models were kept in non-flexible conditions, but the CSO ligand was kept flexible for superior binding.25 The ligand–receptor interaction analysis was based on Lamarckian genetic algorithm, with a higher degree of freedom for ligands. This degree of freedom accounts for better binding of ligands on receptors.30
Site-specific interaction of chitinase with CSO
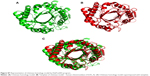
In the current study, docking studies of chitinase with CSO also revealed that amino acid residues such as Tyr167, Asn222, Asn246, Trp245, and Trp504 of chitinase participated in hydrogen bonding with CSO. Gly244 was found to be involved in a hydrophobic interaction with CSO (Figure 7).
Site-specific interaction of Jacob with CSO
Docking studies of Jacob with CSO revealed that Phe183, Asn236, Glu171, Cys190, Cys389, and Pro388 residues participated in hydrogen bond interactions, whereas Trp391, Val191, Try189, and Asp387 residues were associated with hydrophobic interactions (Figure 8).
Site-specific interaction of chitin to CSO
Interaction analysis of chitin with CSO (Figure 9) clearly indicated that it possesses low binding energy (Table S2) and evidences lesser formation of hydrogen bonds as compared to chitinase and Jacob. Therefore, chitin cannot be considered as a good receptor for binding of CSO.
  | Figure 9 Representation of docked complex of chitin with chitosan oligosaccharide and its hydrogen and hydrophobic interaction (chitin is in blue color, chitosan in red color). |
Table S2 clearly indicates the higher negative value of intermolecular energy, hydrogen bonding energy, electrostatic energy, total internal energy, and torsional free energy for Jacob and Chitinase receptors with respect to chitin. These findings of differential binding affinity (additional outcome of final intermolecular energy, hydrogen bonding energy, electrostatic energy, total internal energy, and torsional free energy)30 revealed that CSO has a higher affinity for chitinase and Jacob lectins. Therefore, the 3D homology model of chitinase, Jacob can be used as a therapeutic target for the removal of Entamoeba histolytica from environmental samples.
In summary, CSO-INPs were synthesized for removal of Entamoeba cysts from the water under the influence of an external magnetic field. Synthesized particles were found to be well dispersed, non-agglomerative, and stable over a range of pH values, which makes them suitable for water treatment. Different lectins and chitin components of cyst walls were found to be interactive with CSO nanoparticles, making the latter very much suitable for pathogenic Entamoeba cyst removal from water.
Present work endorses integration of magnetic CSO nanomaterials into existing water purification systems. To fully utilize the potential of CSO-INPs, especially for future water treatment applications, various parameters such as size and shape of nanoparticles, temperature, and pH need to be optimized. Toxicity of synthesized nanomaterials is another challenging issue for upscaling this process. However, biocompatibility, nontoxic nature,16,19,38–42 and antimicrobial properties of chitosan16,43,44 makes the synthesized nanomaterial a suitable candidate for the treatment of the contaminated water systems.
Acknowledgments
Sudeep Shukla was a recipient of the Senior Research Fellowship from the Council of Scientific and Industrial Research (CSIR). We are grateful to Dr Sudha Bhattacharya for providing the E. invadens cysts used in this work. We sincerely acknowledge Dr A Dixit and her research group for their help in developing the bioinformatics part of this manuscript. We are also thankful to Advance Instrumentation Facility (AIRF), Jawaharlal Nehru University, New Delhi for various analytical instruments used in this work.
Disclosure
The authors report no conflicts of interest in this work.
References
World Health Organization [webpage on the Internet]. WHO Guidelines for Drinking-water Quality. 4th ed. Geneva, Switzerland: World Health Organization; 2011:1–564. Available from: http://whqlibdoc.who.int/publications/2011/9789241548151_eng.pdf?ua=1. Accessed April 13, 2013. | ||
Sharma S, Sachdeva P, Virdi JS. Emerging water-borne pathogens. Appl Microbiol Biotechnol. 2003;61(5–6):424–428. | ||
Gadgil A. Drinking water in developing countries. Annu Rev Energy Env. 1998;23:253–286. | ||
Marshall MM, Naumovitz D, Ortega Y, Sterling CR. Waterborne protozoan pathogens. Clin Microbiol Rev. 1997;10(1):67–85. | ||
Singh N, Bhattacharya S, Paul J. Entamoeba invadens: dynamics of DNA synthesis during differentiation from trophozoite to cyst. Exp Parasitol. 2011;127(2):329–333. | ||
Stanley S. Amoebiasis. Lancet. 2003;361:1025–1034. | ||
Dellen K Van, Ghosh SK, Robbins PW, Loftus B, Samuelson J. Entamoeba histolytica lectins contain unique 6-Cys or 8-Cys chitin-binding domains. Infect Immun. 2002;70(6):3259–3263. | ||
Arroyo-Begovich AA, Cárabez-Trejo A, Ruíz-Herrera J. Identification of the structural component in the cyst wall of Entamoeba invadens. J Parasitol. 1980;66(5):735–741. | ||
Chatterjee A, Ghosh SK, Jang K, et al. Evidence for a “wattle and daub” model of the cyst wall of Entamoeba. PLoS Pathog. 2009;5(7): e1000498. | ||
Pradeep T. Noble metal nanoparticles for water purification: a critical review. Thin Solid Films. 2009;517(24):6441–6478. | ||
Koo OM, Rubinstein I, Onyuksel H. Role of nanotechnology in targeted drug delivery and imaging: a concise review. Nanomedicine. 2005; 1(3):193–212. | ||
Farokhzad OC, Langer R. Impact of nanotechnology on drug delivery. ACS Nano. 2009;3(1):16–20. | ||
Rinaudo M. Chitin and chitosan: properties and applications. Prog Polym Sci. 2006;31(7):603–632. | ||
Larsen MU, Seward M, Tripathi A, Shapley NC. Biocompatible nanoparticles trigger rapid bacteria clustering. Biotechnol Prog. 2009;25(4):1094–1102. | ||
Alves NM, Mano JF. Chitosan derivatives obtained by chemical modifications for biomedical and environmental applications. Int J Biol Macromol. 2008;43(5):401–414. | ||
Qin C, Li H, Xiao Q, Liu Y, Zhu J, Du Y. Water-solubility of chitosan and its antimicrobial activity. Carbohydr Polym. 2006;63(3):367–374. | ||
López-Cruz A, Barrera C, Calero-DdelC VL, Rinaldi C. Water dispersible iron oxide nanoparticles coated with covalently linked chitosan. J Mater Chem. 2009;19(37):6870–6876. | ||
Jana NR, Chen Y, Peng X. Size- and shape-controlled magnetic (Cr, Mn, Fe, Co, Ni) oxide nanocrystals via a simple and general approach. Chem Mater. 2004;16(20):3931–3935. | ||
Shukla S, Jadaun A, Arora V, Sinha RK, Biyani N, Jain VK. In vitro toxicity assessment of chitosan oligosaccharide coated iron oxide nanoparticles. Toxicol Reports. 2015;2:27–39. | ||
De Palma R, Peeters S, Van Bael MJ, et al. Silane ligand exchange to make hydrophobic superparamagnetic nanoparticles water-dispersible. Chem Mater. 2007;(19):1821–1831. | ||
Woehrle GH, Hutchison JE, Ozkar S, Finke RG. Analysis of nanoparticle transmission electron microscopy data using a public-domain image-processing program, image. Turk J Chem. 2006;30(1):1–13. | ||
Diamond LS. Techniques of axenic cultivation of Entamoeba histolytica Schaudinn, 1903 and E. Histolytica-like amebae. J Parasitol. 1968;54(5):1047–1056. | ||
Gasteiger J, Rudolph C, Sadowski J. Automatic generation of 3D-atomic coordinates for organic molecules. Tetrahedron Comput Methodol. 1992; 3(6):537–547. | ||
Lambert C, Léonard N, De Bolle X, Depiereux E. ESyPred3D: prediction of proteins 3D structures. Bioinformatics. 2002;18(9):1250–1256. | ||
Seeliger D, de Groot BL. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput Aided Mol Des. 2010;24(5):417–422. | ||
Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr. 1993;26(2):283–291. | ||
Eisenberg D, Lüthy R, Bowie JU. VERIFY3D: assessment of protein models with three-dimensional profiles. Methods Enzymol. 1997; 277:396–406. | ||
Wiederstein M, Sippl MJ. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 2007;35(Web Server issue):W407–W410. | ||
Ramachandran GN, Venkatachalam CM. Stereochemical criteria for polypeptides and proteins. IV. Standard dimensions for the cis-peptide unit and conformation of cis-polypeptides. Biopolymers. 1968;6(9):1255–1262. | ||
Morris GM, Goodsell DS, Halliday RS, et al. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem. 1998;19(14):1639–1662. | ||
Li G, Jiang Y, Huang K, Ding P, Chen J. Preparation and properties of magnetic Fe3O4–chitosan nanoparticles. J Alloys Compd. 2008; 466(1–2):451–456. | ||
Unsoy G, Yalcin S, Khodadust R, Gunduz G, Gunduz U. Synthesis optimization and characterization of chitosan-coated iron oxide nanoparticles produced for biomedical applications. J Nanoparticle Res. 2012;14(11):1–13. | ||
Chang YC, Chen DH. Preparation and adsorption properties of monodisperse chitosan-bound Fe3O4 magnetic nanoparticles for removal of Cu(II) ions. J Colloid Interface Sci. 2005;283(2):446–451. | ||
Zhang L, He R, Gu H. Oleic acid coating on the monodisperse magnetite nanoparticles. Appl Surf Sci. 2006;253:2611–2617. | ||
Li G, Huang K, Jiang Y, Ding P, Yang D. Preparation and characterization of carboxyl functionalization of chitosan derivative magnetic nanoparticles. Biochem Eng J. 2008;40(3):408–414. | ||
Fusetti F, von Moeller H, Houston D, et al. Structure of human chitotriosidase. Implications for specific inhibitor design and function of mammalian chitinase-like lectins. J Biol Chem. 2002;277(28):25537–25544. | ||
Katsemi V, Lücke C, Koepke J, et al. Mutational and structural studies of the diisopropylfluorophosphatase from Loligo vulgaris shed new light on the catalytic mechanism of the enzyme. Biochemistry. 2005;44(25):9022–9033. | ||
Worthington KLS, Adamcakova-Dodd A, Wongrakpanich A, et al. Chitosan coating of copper nanoparticles reduces in vitro toxicity and increases inflammation in the lung. Nanotechnology. 2013; 24(39):395101. | ||
Shi SF, Jia JF, Guo XK, et al. Biocompatibility of chitosan-coated iron oxide nanoparticles with osteoblast cells. Int J Nanomedicine. 2012;7:5593–5602. | ||
Dung DTK, Hai TH, Long BD, Truc PN. Preparation and characterization of magnetic nanoparticles with chitosan coating. J Phys Conf Ser. 2009;187(1):012036. | ||
Kumar MNR. A review of chitin and chitosan applications. React Funct Polym. 2000;46(1):1–27. | ||
Qi L, Xu Z, Jiang X, Hu C, Zou X. Preparation and antibacterial activity of chitosan nanoparticles. Carbohydr Res. 2004;339(16): 2693–2700. | ||
Chávez de Paz LE, Resin A, Howard KA, Sutherland DS, Wejse PL. Antimicrobial effect of chitosan nanoparticles on streptococcus mutans biofilms. Appl Environ Microbiol. 2011;77(11):3892–3895. | ||
Zheng L, Zhu JF. Study on antimicrobial activity of chitosan with different molecular weights. Carbohydr Polym. 2003;54(4): 527–530. |
Supplementary materials
  | Figure S1 Zeta potential measurement of chitosan oligosaccharide-functionalized iron oxide nanoparticles (CSO-INPs) at pH 3 (A), pH 7 (B), and pH 9 (C). |
  | Figure S3 Capture efficiency of nanoparticles to Entamoeba cysts at different concentrations of nanoparticles at different times. |
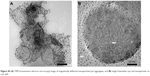  | Figure S4 (A) TEM (transmission electron microscopy) image of magnetically deflected nanoparticles/cyst aggregates, and (B) single Entamoeba cyst and nanoparticles on cyst wall. |
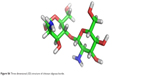  | Figure S6 Three dimensional (3D) structure of chitosan oligosaccharide. |
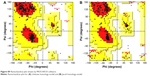  | Figure S9 Ramachandran plot drawn by PROCHECK software. |
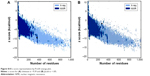  | Figure S10 z score representation by ProSA energy plot. |
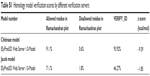  | Table S1 Homology model verification scores by different verification servers |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.