Back to Journals » OncoTargets and Therapy » Volume 13
RETRACTED ARTICLE: Long Noncoding RNA LINC00525 Promotes the Aggressive Phenotype of Chordoma Through Acting as a microRNA-505-3p Sponge and Consequently Raising HMGB1 Expression
Authors Li L, Lv G, Wang B, Ma H
Received 20 June 2020
Accepted for publication 11 August 2020
Published 10 September 2020 Volume 2020:13 Pages 9015—9027
DOI https://doi.org/10.2147/OTT.S268678
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Tohru Yamada
This paper has been retracted.
Lei Li, Guohua Lv, Bing Wang, Hong Ma
Department of Spine Surgery, The Second Xiangya Hospital, Central South University, Changsha, Hunan 410011, People’s Republic of China
Correspondence: Hong Ma
Department of Spine Surgery, The Second Xiangya Hospital, Central South University, 139 Renmin Road, Changsha, Hunan 410011, People’s Republic of China
Email [email protected]
Purposes: Long intergenic non-protein coding RNA 525 (LINC00525), a long noncoding RNA, has been implicated in the carcinogenesis and progression of many human cancer types. However, the detailed roles of LINC00525 in chordoma and the underlying mechanisms are not fully understood. Here, we aimed to determine whether LINC00525 could modulate the oncogenicity of chordoma cells and to elucidate in detail the molecular events underlying these tumor-promoting activities.
Methods: Reverse-transcription quantitative polymerase chain reactions were performed to assess LINC00525 expression in chordoma. The effects of LINC00525 silencing on chordoma cell proliferation, apoptosis, migration, and invasiveness in vitro and tumor growth in vivo were respectively tested using CCK-8 assay, flow cytometry, migration and invasion assays, and xenograft experiments.
Results: High LINC00525 expression levels were detected in chordoma tissues. The proliferative, migratory, and invasive abilities of chordoma cells in vitro and their tumor growth in vivo were suppressed by the LINC00525 knockdown, whereas apoptosis was induced by it. Mechanistically, LINC00525 acted as a molecular sponge of microRNA-505-3p (miR-505-3p) and upregulated the expression of high mobility group box 1 (HMGB1), which is directly targeted by miR-505-3p. Rescue assays indicated that increasing the output of miR-505-3p–HMGB1 axis attenuated the effects of LINC00525 depletion on chordoma cells.
Conclusion: LINC00525, a pro-oncogenic long noncoding RNA, promotes chordoma progression by regulating the miR-505-3p–HMGB1 axis. The LINC00525–miR-505-3p–HMGB1 pathway may be a novel therapeutic target in chordoma.
Keywords: chordoma, LINC00525, miR-505-3p, HMGB1, oncogenicity
Expression of Concern for this paper has been published
Introduction
Chordoma is a type of rare malignant bone tumor associated with a morbidity rate of 0.08 per 100,000 persons.1 Chordomas, which develop from benign notochordal rests, account for approximately 1–4% of all bone tumors.2 These tumors are characteristically low-grade, highly invasive, and chemo-resistant.3 Currently, surgical excision and radiotherapy are the primary types of therapies applied to chordomas.4 Despite considerable advances in chordoma diagnostic and treatment strategies in the last decade, the overall survival duration remains approximately 5 years, and no noticeable improvements in the prognosis of patients have been reported.5,6 This unsatisfactory clinical outcome is largely attributable to a lack of understanding of the pathogenesis of chordoma.7 Furthermore, this malignancy is difficult to treat due to an appallingly high recurrent rate, which has been reported to range from 44% to 78%.8,9 Hence, a comprehensive study of the molecular biological processes underlying chordoma oncogenesis and progression may help to identify potential targets for cancer diagnosis and management.
Long noncoding RNAs (lncRNAs) are RNA molecules with lengths exceeding 200 nucleotides.10 Although these lncRNAs do not code proteins, they perform other crucial functions in both normal body development and disease pathology by directly or indirectly modulating protein expression.11 The regulatory actions of lncRNAs on physiological and pathological processes are mediated by different molecular mechanisms, including lncRNA–microRNA (miRNA), lncRNA–protein, and lncRNA–mRNA interactions.12,13 A growing number of studies have shown the aberrant expression of numerous lncRNAs in chordomas.14,15 LncRNAs can exert tumor-suppressive or cancer-promoting activities during chordoma initiation and progression and may contribute to the regulation of nearly all processes involved in chordoma malignancy.16 Therefore, an investigation of the correlation between lncRNAs and chordoma may uncover a useful target with respect to chordoma prevention, diagnosis, and management.
The expression of lncRNA LINC00525 is dysregulated in many types of human cancer, and this molecule has been implicated in carcinogenesis and cancer progression.17,18 Nonetheless, details of the involvement of LINC00525 in chordoma and the related mechanism of action are not well understood. In this study, we aimed to ascertain the expression profile of LINC00525 in chordoma and to identify potential associated regulatory factors that contribute in chordoma progression. The molecular events underlying the tumor-promoting activities of LINC00525 in chordoma cells were also illustrated in detail.
Materials and Methods
Tissue Samples
Thirty-three chordoma tissue samples were collected from patients with chordoma who were admitted to the Second Xiangya Hospital (Xiangya, China). Seventeen nucleus pulposus tissue samples were obtained from patients who underwent total sacrectomy. All tissue samples were stored in liquid nitrogen. The Ethics Committee of The Second Xiangya Hospital approved the study protocol (#20160911), which was performed in accordance with the Declaration of Helsinki. Written informed consent was obtained from all the participating patients. The exclusion criteria for our study included a history of radiotherapy, chemotherapy, or other therapies or a second primary cancer.
Cell Culture
The human chordoma cell lines U-CH1 and U-CH2 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). Both lines were grown in a mixture of Iscove’s Modified Dulbecco’s Medium (ATCC) and RPMI-1640 Medium (Gibco; Thermo Fisher Scientific, Waltham, MA, USA) at a ratio of 1:4. This basal medium was supplemented with 10% fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc), 100 U/mL penicillin (Gibco/Thermo Fisher Scientific, Inc), 100 mg/mL streptomycin (Gibco; Thermo Fisher Scientific, Inc), and 1% L-glutamine (Gibco; Thermo Fisher Scientific, Inc). Both cell lines were cultured at 37 °C in a humidified atmosphere containing 5% CO2.
Cell Transfection
The small interfering RNAs (siRNAs) designed to target LINC00525 (si-LINC00525) specifically and negative control siRNA (si-NC) were purchased from RiboBio (Guangzhou, China). The oligonucleotides miR-505-3p mimic and miR-505-3p inhibitor were chemically synthesized by GenePharma (Shanghai, China). Negative control miRNA mimic (miR-NC) and NC inhibitor served as the controls for the miR-505-3p mimic and miR-505-3p inhibitor, respectively. The empty pcDNA3.1 vector and pcDNA3.1 plasmid harboring the full-length HMGB1 cDNA sequence (pcDNA3.1-HMGB1) were acquired from GeneChem Co., Ltd. (Shanghai, China). Cell transfection was performed using the Lipofectamine 2000 reagent (Invitrogen; Thermo Fisher Scientific).
Isolation of Cytoplasmic and Nuclear RNA Fractions
The nuclear and cytoplasmic fractions of chordoma cells were separated using a Cytoplasmic and Nuclear RNA Purification Kit (Norgen, Thorold, ON, Canada). RNA was extracted from both fractions and analyzed using the reverse-transcription quantitative polymerase chain reaction (RT-qPCR) method.
RT-qPCR
Total RNA was isolated using the TRIzol reagent (Invitrogen; Thermo Fisher Scientific) and reverse-transcribed into cDNA using the PrimeScript™ RT Reagent Kit (Takara Biotechnology Co., Ltd., Dalian China). Next, qPCR was performed to quantify HMGB1 mRNA and LINC00525 using SYBR Premix Ex Taq (Takara Biotechnology Co., Ltd.). GAPDH was chosen as an internal reference for the expression of HMGB1 mRNA and LINC00525.
To evaluate miR-505-3p expression, the miRNeasy Mini Kit (Qiagen GmbH, Hilden, Germany) was used to isolate miRNA from tissue samples or cells. Reverse transcription was conducted using the miScript Reverse Transcription Kit (Qiagen GmbH), while PCR amplification was performed using the miScript SYBR Green PCR Kit (Qiagen GmbH).
The expression of miR-505-3p was normalized to that of U6 small nuclear RNA. Relative gene expression levels were analyzed using the 2−ΔΔCq method.
Cell Counting Kit 8 (CCK-8) Assay
The CCK-8 reagent (Beyotime Institute of Biotechnology; Shanghai, China) was used to determine the proliferative capacities of chordoma cells. One hundred microliters of suspension containing 2 × 103 cells were seeded into each well of a 96-well plate. The CCK-8 assay was performed at 0, 1, 2, and 3 days after cell seeding. Specifically, 10 µL of the CCK-8 reagent was added to each well, and the cells were incubated at 37 °C for another 2 h. The optical density of each well at a wavelength of 450 nm was recorded and used to plot the growth curves.
Migration and Invasion Assays
The invasion capacity was assessed using 24-well Transwell chambers with inserts (Corning Incorporated, Corning, NY, USA). Prior to the assay, Matrigel (BD Biosciences, San Jose, CA, USA) was added to coat the membrane (8 µm pore size) in each well, and the chambers were incubated at 37 °C for 2 h. Next, the upper chambers were loaded with 5 × 104 cells resuspended in 200 µL of FBS-free culture medium. The lower chambers were covered with 500 µL of culture medium supplemented with 20% FBS. After a 24-h incubation at 37 °C with 5% CO2, the noninvasive cells were gently removed with a cotton swab. The invasive cells were fixed with 4% paraformaldehyde, stained with 0.1% crystal violet, washed extensively with phosphate-buffered saline, and air dried. The migration assay procedures were identical to the invasion assay procedures except that the membranes were not precoated with Matrigel. Finally, the cells that had traversed the membranes were imaged and counted under an inverted microscope (Olympus, Tokyo, Japan).
Flow Cytometric Analysis
The Annexin V–Fluorescein Isothiocyanate (FITC) Apoptosis Detection Kit (Biolegend, San Diego, CA, USA) was used to detect apoptotic cells. Transfected cells were collected by digestion with ethylene diamine tetraacetic acid (EDTA)-free solution of 0.25% trypsin. Briefly, the harvested cells were resuspended in 100 µL of staining solution. Next, 5 µL of Annexin V–FITC and 5 µL of propidium iodide solution were added to each cell suspension. After a 15-min incubation at room temperature in the dark, the proportions of apoptotic cells were determined by flow cytometry (FACScan; BD Biosciences).
In vivo Xenograft Experiments
Lentiviral vectors expressing either short hairpin RNA (shRNA) specific for LINC00525 (sh-LINC00525) or negative control nonsensical sequence (sh-NC) were constructed by GenePharma. To obtain cells in which LINC00525 was stably silenced, U-CH1 cells were transduced with lentiviral vector expressing either sh-LINC00525 or sh-NC and subjected to puromycin selection for 2 weeks.
The animal procedures were approved by the Institutional Animal Care and Use Committee of the Second Xiangya Hospital (#20180428), and were executed in accordance with NIH guidelines for the care and use of laboratory animals. BALB/c nude mice (male; age 4–6 weeks; Shanghai Laboratory Animal Research Center, Shanghai, China) were inoculated subcutaneously with U-CH1 cells engineered to stably express either sh-LINC00525 or sh-NC. The lengths and widths of the tumors were measured every 5 days, and the tumor volume was calculated using the formula: volume (mm3) = 0.5 × length (mm) × width2 (mm2). On day 30 post-injection, all the mice were euthanized, and the subcutaneous xenograft tumors were resected, weighed, and stored for subsequent RT-qPCR and Western blotting analysis.
Bioinformatics Analysis
StarBase 3.0 (http://starbase.sysu.edu.cn/) was used to predict the potential miRNA(s) that would bind to LINC00525. The potential target mRNA(s) of miR-505-3p were identified using the miRDB tool (http://mirdb.org/miRDB/index.html), starBase 3.0, and TargetScan (http://www.targetscan.org/vert_60/).
Luciferase Reporter Assay
Fragments of LINC00525 containing either the predicted wild-type (wt) miR-505-3p–binding site or mutant (mut) site were constructed and inserted (separately) into the psiCHECK™-2 luciferase reporter vector (Promega Corporation, Madison, WI, USA). The recombinant luciferase reporter plasmids were respectively designated as wt-LINC00525 and mut-LINC00525. The wt-HMGB1 and mut-HMGB1 plasmids were constructed using a similar experimental process. Chordoma cells were co-transfected with either wt or mut reporter plasmid and either miR-505-3p mimic or miR-NC using Lipofectamine 2000 reagent. After a 48-h culture period, the luciferase activity was quantified using the Dual-Luciferase® Reporter Assay Kit (Promega). Renilla luciferase activity was normalized to firefly luciferase activity.
RNA Immunoprecipitation (RIP) Assay
A RIP assay to assess the interaction between LINC00525 and miR-505-3p was performed using the Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Billerica, MA). Chordoma cells were incubated in RIP lysis buffer, and the obtained cell lysates were incubated overnight at 4 °C in RIP buffer containing magnetic beads conjugated with either human anti-Argonaute 2 (Ago2) antibody (Millipore) or control IgG (Millipore). Prior to isolating the immunoprecipitated RNA, the harvested magnetic beads were treated with Proteinase K buffer at 4 °C for another 3 h. Target RNA enrichment was detected via RT-qPCR.
Western Blotting
Precooled RIPA lysis buffer (Beyotime; Shanghai, China) supplemented with protease inhibitors (Beyotime) was used to isolate total proteins from the cells. The total-protein concentrations were determined using the Bicinchoninic Acid Protein Assay Kit (Nanjing KeyGen Biotech Co., Ltd.). Equal amounts of protein were subjected to sodium dodecyl sulfate 10% polyacrylamide gel electrophoresis and subsequent electrophoretic transferred onto polyvinylidene fluoride membranes. Next, the membranes were blocked in 5% nonfat milk diluted in Tris-buffered saline containing 0.1% Tween 20 (TBST). The membranes were then incubated with primary antibodies specific for HMGB1 (1:1000 dilution in TBST; cat. No. ab79823; Abcam, Cambridge, UK) or GAPDH (1:1000 dilution in TBST; cat. No. ab128915; Abcam). Following an overnight incubation at 4 °C, the membranes were rinsed thrice with TBST and probed with horseradish peroxidase-conjugated secondary antibody (1:5000 dilution in TBST; cat. No. ab205718; Abcam) at room temperature for 1 h. The protein immunoblots were visualized using the Immobilon Western Chemilum HRP substrate (Millipore). GAPDH acted as the loading control.
Statistical Analysis
All results are presented as means ± standard deviations. Student’s t-test was performed to analyze differences between two groups. A one-way analysis of variance and subsequent Tukey’s post hoc test were used to characterize differences among multiple groups. The Pearson correlation coefficient was applied to examine the association between LINC00525 and miR-505-3p expression levels. Differences with a P value < 0.05 were considered statistically significant.
Results
LINC00525 Downregulation Promotes Cell Apoptosis but Suppresses Cell Proliferation, Migration and Invasion in Chordoma in vitro
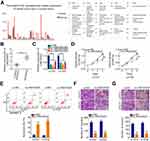
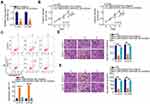
First, GEPIA (http://gepia.cancer-pku.cn/) was used to identify the expression of LINC00525 in human cancers. Notably, LINC00525 was upregulated in the majority of human cancer types (Figure 1A). To address the dysregulation of LINC00525 in chordoma, RT-qPCR was conducted to detect the expression of this lncRNA in 33 chordoma tissues and 17 nucleus pulposus tissues. LINC00525 was shown to be overexpressed in chordoma tissues relative to pulposus tissues (Figure 1B).
To examine whether LINC00525 exerts critical roles in cellular process, LINC00525 interference was induced by transfecting in U-CH1 and U-CH2 cells with si-LINC00525. RT-qPCR analysis presented that all of the three siRNAs was effective (Figure 1C). Here, si-LINC00525#2 was identified as the most potent silencer of LINC00525 and was used in subsequent experiments. CCK-8 assay was performed to test the effect of LINC00525 knockdown on the proliferation of chordoma cells. The results indicated that downregulation of LINC00525 strikingly suppressed the proliferative capacities of U-CH1 and U-CH2 cells (Figure 1D). In addition, transfection with si-LINC00525 led to an increased frequency of apoptosis in U-CH1 and U-CH2 cells, as demonstrated by the flow cytometric analysis (Figure 1E). Furthermore, migration and invasion assays revealed that the loss of LINC00525 restricted both the migration (Figure 1F) and invasion (Figure 1G) of U-CH1 and U-CH2 cells. These results suggest that LINC00525 is upregulated and executes cancer-promoting roles in chordomas.
LINC00525 Acts as a miR-505-3p Sponge in Chordoma Cells
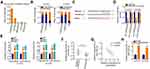
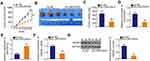
To unveil the molecular events underlying the tumor-promoting activities of LINC00525 in chordoma, the subcellular localization of LINC00525 was first predicted by lncLocator (http://www.csbio.sjtu.edu.cn/bioinf/lncLocator/). This analysis predicted that LINC00525 was mainly distributed in the cytoplasm (Figure 2A). To confirm this prediction, the cytoplasmic and nuclear RNA fractions of U-CH1 and U-CH2 cells were isolated, and the data further corroborated the largely cytoplasmic location of LINC00525 (Figure 2B). Accumulated studies have declared that cytoplasmic lncRNAs could exert their regulatory roles by acting as competing endogenous RNAs (ceRNAs) or molecular sponges for miRNAs in human cancers.19–21 Hence, StarBase 3.0 was used to predict potential miRNAs that may be sponged by LINC00525. Notably, miR-505-3p, which has considerable functions during cancer genesis and progression,22–24 was predicted to possess a putative binding site for LINC00525 (Figure 2C) and was therefore chosen for experimental reconfirmation.
Luciferase reporter assay was performed to explore the binding between LINC00525 and miR-505-3p in chordoma cells. Notably, the luciferase activity of wt-LINC00525 was clearly decreased in U-CH1 and U-CH2 cells after the introduction of miR-505-3p mimic. However, the luciferase activity of mut-LINC00525 was not altered significantly in response to co-transfection with miR-505-3p mimic (Figure 2D). RIP assay demonstrated the increased enrichment of both LINC00525 and miR-505-3p in the Ago2-immunoprecipitated pellet (Figure 2E), which further supported a direct interaction between LINC00525 and miR-505-3p in chordoma. Next, RT-qPCR analysis illustrated that miR-505-3p was weakly expressed in chordoma tissues (Figure 2F) and was negatively associated with LINC00525 expression (Figure 2G; r = −0.6844, P < 0.0001). To further verify the potential involvement of LINC00525 in modulating miR-505-3p expression, RT-qPCR analysis was performed to detect the expression of this miRNA in LINC00525-depleted U-CH1 and U-CH2 cells. Here, the downregulation of LINC00525 obviously promoted the expression of miR-505-3p in U-CH1 and U-CH2 cells (Figure 2H). In summary, these results proved that LINC00525 could act as a ceRNA in chordoma by directly sponging miR-505-3p.
miR-505-3p Overexpression Exerts Inhibitory Activities on Chordoma Cell Behaviors
We next investigated the specific roles of miR-505-3p in chordoma cells. Here, gain-of-functional assays were performed using the miR-505-3p mimic. Transfection with this mimic induced a remarkable upregulation of miR-505-3p in U-CH1 and U-CH2 cells (Figure 3A). CCK-8 assays presented an impaired proliferative ability in miR-505-3p overexpressing-U-CH1 and U-CH2 cells (Figure 3B). In addition, the ratios of apoptotic U-CH1 and U-CH2 cells were increased noticeably after upregulation of miR-505-3p, as demonstrated by flow cytometric analysis (Figure 3C). The migration and invasion assays illustrated that ectopic miR-505-3p expression reduced U-CH1 and U-CH2 cell migration (Figure 3D) and invasion (Figure 3E). Therefore, miR-505-3p is an anti-oncogenic miRNA in chordoma cells.
HMGB1 is a Direct Target of miR-505-3p and LINC00525 Positively Regulates HMGB1 Expression in Chordoma Cells
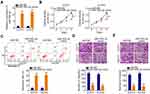
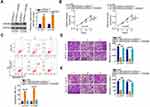
The putative targets of miR-505-3p were predicted automatically using bioinformatics analysis tools (TargetScan, miRDB and StarBase), based on the assumption that miR-505-3p inhibits the aggressiveness of chordoma cells. The 3′-untranslated region (UTR) of HMGB1 was found to harbor a complementary binding site for miR-505-3p (Figure 4A). Luciferase reporter assay indicated that the luciferase activity of wt-HMGB1 was suppressed in U-CH1 and U-CH2 cells following the introduction of miR-505-3p mimic. However, this inhibitory action was abolished when the binding sequences were mutated (Figure 4B). Additionally, RT-qPCR confirmed the strong expression of HMGB1 mRNA in chordoma tissues (Figure 4C). Pearson correlation coefficient analysis verified the inverse correlation between miR-505-3p and HMGB1 mRNA expression in the chordoma tissues (Figure 4D; r = −0.7263, P < 0.0001). Additionally, the HMGB1 mRNA (Figure 4E) and protein (Figure 4F) levels were noticeably reduced when miR-505-3p was overexpressed in U-CH1 and U-CH2 cells.
Next, we attempted to address whether LINC00525 could control HMGB1 expression in chordoma cells. U-CH1 and U-CH2 cells were transfected with si-LINC00525 or si-NC, and HMGB1 expression was measured. Notably, the levels of both HMGB1 mRNA (Figure 4G) and protein (Figure 4H) were decreased in U-CH1 and U-CH2 cells after the knockdown of LINC00525. A correlation analysis revealed a positive correlation between HMGB1 mRNA and LINC00525 expression in chordoma tissues (Figure 4I; r = 0.6394, P < 0.0001). To illustrate the interaction of miR-505-3p with LINC00525 and HMGB1 in chordoma cells, U-CH1 and U-CH2 cells were co-transfected with si-LINC00525 and either the miR-505-3p inhibitor or NC inhibitor. RT-qPCR and Western blotting analyses verified that si-LINC00525 downregulated the expression of HMGB1 mRNA (Figure 4J) and protein (Figure 4K) in U-CH1 and U-CH2 cells, whereas this expression was partly recovered by co-transfection with the miR-505-3p inhibitor. Collectively, these data illustrate that LINC00525 positively regulates HMGB1 expression in chordoma cells by binding competitively to miR-505-3p.
LINC00525 Exerts Its Functions in Chordoma Cells via the miR-505-3p/HMGB1 Axis
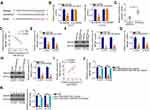
Rescue assays were designed and conducted to confirm that LINC00525 exerted its cancer-promoting functions in chordoma cells via the miR-505-3p/HMGB1 axis. The follow-up assays were performed using the miR-505-3p inhibitor, and the transfection efficiency data are presented in Figure 5A. U-CH1 and U-CH2 cells transfected with si-LINC00525 were further transfected with miR-505-3p inhibitor or NC inhibitor. CCK-8 assay and flow cytometric analysis data respectively verified that si-LINC00525 inhibited cell proliferation (Figure 5B) and improved cell apoptosis (Figure 5C) in U-CH1 and U-CH2 cells. However, these effects were neutralized in response to miR-505-3p inhibitor co-transfection. Additionally, the use of miR-505-3p inhibitor partially abolished the suppressive effects of si-LINC00525 on the migratory (Figure 5D) and invasive (Figure 5E) capacities of U-CH1 and U-CH2 cells.
Meanwhile, Western blotting was employed to evaluate the transfection efficiency of HMGB1 overexpression plasmid pcDNA3.1-HMGB1, and the results indicated that transfection with pcDNA3.1-HMGB1 resulted in an obvious increase of HMGB1 protein expression in U-CH1 and U-CH2 cells (Figure 6A). The si-LINC00525 in parallel with pcDNA3.1-HMGB1 or pcDNA3.1 was introduced into U-CH1 and U-CH2 cells. The LINC00525 knockdown-induced suppression of U-CH1 and U-CH2 cell proliferation (Figure 6B) and enhancement of cell apoptosis (Figure 6C) were reversed by HMGB1 reintroduction. Furthermore, the hindered migration (Figure 6D) and invasion (Figure 6E) of U-CH1 and U-CH2 cells after LINC00525 silencing was recovered with the co-transfection of pcDNA3.1-HMGB1. Taken together, these results suggest that LINC00525 promotes chordoma progression through the miR-505-3p/HMGB1 axis.
LINC00525 Depletion Inhibits Tumor Growth in vivo
Finally, in vivo xenograft experiments were performed to analyze the effect of LINC00525 on chordoma tumorigenicity in vivo. The volumes of tumor xenografts were smaller in sh-LINC00525-treated mice than in sh-NC-treated mice (Figure 7A and B). Simultaneously, the tumor weights were demonstrably lower in the mice inoculated with U-CH1 cells engineered to stably express sh-LINC00525 (Figure 7C). Furthermore, the expression of LINC00525, miR-505-3p and HMGB1 in the tumor xenografts was measured. A lower level of LINC00525 (Figure 7D) and higher level of miR-505-3p (Figure 7E) were detected in the tumors collected from the sh-LINC00525 group, compared to tumors from the sh-NC group. Furthermore, the RT-qPCR and Western blotting data affirmed that HMGB1 mRNA (Figure 7F) and protein (Figure 7G) levels were weakly expressed in tumor xenografts that originated from stably LINC00525-silenced U-CH1 cells. Our results suggest that LINC00525 promotes the tumorigenesis of chordoma cells in vivo by regulating the miR-505-3p/HMGB1 axis.
Discussion
LncRNAs have recently attracted considerable attention, and an increasing body of evidence indicates the dysregulation of numerous lncRNAs in chordoma.14,15 This dysregulation correlates significantly with the clinical outcomes of chordoma patients.16 Functionally, lncRNAs play key regulatory roles in the formation and evolution of chordoma.14,15 Hence, lncRNAs are likely to have the potential to be developed as effective targets for chordoma therapy. In this study, we focused on whether LINC00525 could modulate the malignancy of chordoma cells in vitro and in vivo by influencing the miR-505-3p/HMGB1 axis.
LINC00525 expression has been studied in several human cancer types. For example, LINC00525 is upregulated in non-small cell lung cancer and is closely correlated with the TNM stage and lymph node metastasis.17 The overall survival of patients with non-small cell lung cancer in the high-LINC00525 group was notably shorter relative to the patients in the low-LINC00525 group.17 High LINC00525 expression has been also identified in colorectal cancer.18 In contrast, the LINC00525 expression profile in chordoma remains under-investigated. In this study, the TCGA normal and GTEx databases were used to perform the initial analysis of LINC00525 expression in human cancers, and the data demonstrated the overexpression of LINC00525 in nearly all types of human cancer types. Our data further verified that LINC00525 is strongly expressed in chordoma tissues.
LINC00525 performs an oncogenic function in the progression of non-small cell lung cancer, where it influences cell proliferation, colony-formation, migration, and invasion.17 In colorectal cancer, LINC00525 inhibition suppresses cancer cell stemness properties and oxaliplatin sensitivity.18 Even though the roles of LINC00525 in multiple human cancers have been well studied, the functions of this lncRNA in chordoma were explored for the first time in this study. Chordoma cells were subjected to loss-of-function assays to determine whether LINC00525 contributes to chordoma progression. Interference with LINC00525 expression led to reductions in chordoma cell proliferation, migration, and invasion and an increase in apoptosis in vitro. Additionally, the knockdown of LINC00525 decelerated the growth of chordoma tumors in vivo.
The interactions between lncRNAs and miRNAs have been confirmed as the critical mechanisms underlying the lncRNA-mediated malignant phenotypes of tumor cells.25 LncRNAs can control gene expression by binding competitively to miRNAs and consequently abrogating miRNA-induced translational suppression and/or mRNA degradation.26 Herein, LINC00525 was largely concentrated in the cytoplasm of chordoma cells, suggesting that LINC00525 may act as a ceRNA in chordoma. A subsequent bioinformatics analysis identified miR-505-3p as a miRNA that may bind to sequences within LINC00525. Our luciferase reporter assay and RIP assay confirmed that LINC00525 is an upstream regulator of miR-505-3p activity and acts as a molecular sponge of miR-505-3p in chordoma. In addition, we demonstrated that miR-505-3p was downregulated in chordoma tissues, and observed an inverse correlation of this miRNA with LINC00525 expression. The depletion of LINC00525 led to an increase in miR-505-3p expression and decreased in HMGB1 expression. Further experiments revealed that the inhibition of miR-505-3p could reverse the suppressive action of LINC00525 knockdown on the expression of HMGB1 expression in chordoma cells. Collectively, these results suggest that LINC00525 acts as a ceRNA of miR-505-3p and thus increases HMGB1 expression.
MiR-505-3p acts as a tumor-suppressor during carcinogenesis and cancer progression and is downregulated in many tumor types.22–24 To the best of our knowledge, this study is the first to explore in detail the expression, roles, and mechanisms of action of miR-505-3p in chordoma. Our data indicate that miR-505-3p is downregulated in chordoma, whereas exogenous miR-505-3p expression attenuates the malignant chordoma cell phenotype. An in-depth mechanistic study identified HMGB1 mRNA as a direct miR-505-3p target in chordoma cells.
HMGB1, which is located on human chromosome 13q12, encodes a highly conserved DNA-binding protein that can translocate from the cytoplasm to the nucleus and interact with transcription factors, nucleosomes, and histones.27 Previously, HMGB1 was proven to act as an oncogene by affecting a wide spectrum of biological activities in human cancers.28–30 HMGB1 was validated as a master driver of tumorigenesis and tumor progression, largely achieved via pro-inflammatory interactions with toll-like receptor 4 in particular, as well as the receptor for advanced glycation end products.31 Furthermore, several important signaling pathways can be activated by HMGB1, thereby promoting the hallmarks of cancer.32 Our results suggest that miR-505-3p directly targets HMGB1 mRNA and thus suppresses the aggressive behavior of chordoma cells. Judging by our findings, LINC00525 interacts directly with miR-505-3p and acts as a molecular sponge for miR-505-3p, thereby preventing miR-505-3p–mediated HMGB1 downregulation. This study provides a novel understanding of the participation of the LINC00525–miR-505-3p–HMGB1 ceRNA regulatory network in the oncogenesis and progression of chordoma.
Therapy targeting the dysregulated lncRNAs is a potential technique to treat chordoma. Exogenous expression of decreased lncRNAs or knocking down increased lncRNAs via siRNAs or antisense oligonucleotides have been explored to be applicable in the treatments of chordoma. Furthermore, the combination of radiochemotherapy and lncRNAs-based therapeutic interventions may be a promising way to manage chordoma patients at advanced stage. However, the lncRNAs-mediated treatments are not yet possible in clinical practice as a result of delivery problems, and this may be resolved in the future by ongoing nanotechnological approaches.
In this study, we did not analyze the association between LINC00525, miR-505-3p, or HMGB1 with prognosis in patients with chordoma. It was mainly due to the small sample size and inadequate follow-up time. It was a limitation of our study, and we will resolve it in the near future.
Conclusion
In summary, we validated a novel chordoma-related lncRNA, LINC00525, and revealed for the first time that this pro-oncogenic lncRNA promotes the progression of chordoma in vitro and in vivo. Our mechanistic investigation revealed that LINC00525 operates as a ceRNA by sponging miR-505-3p and consequently enhancing the expression of HMGB1. Therefore, the LINC00525–miR-505-3p–HMGB1 pathway may represent a novel treatment target in chordoma.
Disclosure
The authors declare that they have no competing interests for this work.
References
1. Kreshak J, Larousserie F, Picci P, et al. Difficulty distinguishing benign notochordal cell tumor from chordoma further suggests a link between them. Cancer Imaging. 2014;14(1):4. doi:10.1186/1470-7330-14-4
2. Bettegowda C, Yip S, Lo SL, et al. Spinal column chordoma: prognostic significance of clinical variables and T (brachyury) gene SNP rs2305089 for local recurrence and overall survival. Neuro-Oncology. 2017;19(3):405–413. doi:10.1093/neuonc/now156
3. Makhdoomi R, Ramzan A, Khursheed N, et al. Clinicopathological characteristics of chordoma: an institutional experience and a review of the literature. Turk Neurosurg. 2013;23(6):700–706. doi:10.5137/1019-5149.JTN.5941-12.3
4. Gagliardi F, Boari N, Riva P, Mortini P. Current therapeutic options and novel molecular markers in skull base chordomas. Neurosurg Rev. 2012;35(1):1–13. doi:10.1007/s10143-011-0354-1
5. Nguyen-Huynh A, Blevins NH, Jackler RK. The challenges of revision skull base surgery. Otolaryngol Clin North Am. 2006;39(4):783–799. doi:10.1016/j.otc.2006.04.006
6. Stacchiotti S, Casali PG, Lo Vullo S, et al. Chordoma of the mobile spine and sacrum: a retrospective analysis of a series of patients surgically treated at two referral centers. Ann Surg Oncol. 2010;17(1):211–219. doi:10.1245/s10434-009-0740-x
7. Whelan JS, Davis LE. Osteosarcoma, chondrosarcoma, and chordoma. J Clin Oncol. 2018;36(2):188–193. doi:10.1200/JCO.2017.75.1743
8. McPherson CM, Suki D, McCutcheon IE, Gokaslan ZL, Rhines LD, Mendel E. Metastatic disease from spinal chordoma: a 10-year experience. J Neurosurg Spine. 2006;5(4):277–280. doi:10.3171/spi.2006.5.4.277
9. Chen KW, Yang HL, Lu J, Liu JY, Chen XQ. Prognostic factors of sacral chordoma after surgical therapy: a study of 36 patients. Spinal Cord. 2010;48(2):166–171. doi:10.1038/sc.2009.95
10. Djebali S, Davis CA, Merkel A, et al. Landscape of transcription in human cells. Nature. 2012;489(7414):101–108. doi:10.1038/nature11233
11. Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;136(4):629–641. doi:10.1016/j.cell.2009.02.006
12. Zhu R, Liu Z, Jiao R, et al. Updates on the pathogenesis of advanced lung cancer-induced cachexia. Thorac Cancer. 2019;10(1):8–16. doi:10.1111/1759-7714.12910
13. Zhu W, Liu H, Wang X, Lu J, Yang W. Long noncoding RNAs in bladder cancer prognosis: a meta-analysis. Pathol Res Pract. 2019;215(6):152429. doi:10.1016/j.prp.2019.04.021
14. Ma X, Qi S, Duan Z, et al. Long non-coding RNA LOC554202 modulates chordoma cell proliferation and invasion by recruiting EZH2 and regulating miR-31 expression. Cell Prolif. 2017;50(6):e12388. doi:10.1111/cpr.12388
15. Wang CB, Wang Y, Wang JJ, Guo XL. LINC00662 triggers malignant progression of chordoma by the activation of RNF144B via targeting miR-16-5p. Eur Rev Med Pharmacol Sci. 2020;24(3):1007–1022. doi:10.26355/eurrev_202002_20151
16. Chen H, Zhang K, Lu J, Wu G, Yang H, Chen K. Comprehensive analysis of mRNA-lncRNA co-expression profile revealing crucial role of imprinted gene cluster DLK1-MEG3 in chordoma. Oncotarget. 2017;8(68):112623–112635. doi:10.18632/oncotarget.22616
17. Yang Z, Lin X, Zhang P, et al. Long non-coding RNA LINC00525 promotes the non-small cell lung cancer progression by targeting miR-338-3p/IRS2 axis. Biomed Pharmacother. 2020;124:109858. doi:10.1016/j.biopha.2020.109858
18. Wang S, Li J, Yang X. Long non-coding RNA LINC00525 promotes the stemness and chemoresistance of colorectal cancer by targeting miR-507/ELK3 axis. Int J Stem Cells. 2019;12(2):347–359. doi:10.15283/ijsc19041
19. Qi X, Zhang DH, Wu N, Xiao JH, Wang X, Ma W. ceRNA in cancer: possible functions and clinical implications. J Med Genet. 2015;52(10):710–718. doi:10.1136/jmedgenet-2015-103334
20. Wang L, Cho KB, Li Y, Tao G, Xie Z, Guo B. Long noncoding RNA (lncRNA)-mediated competing endogenous RNA networks provide novel potential biomarkers and therapeutic targets for colorectal cancer. Int J Mol Sci. 2019;20(22).
21. Ye Y, Shen A, Liu A. Long non-coding RNA H19 and cancer: a competing endogenous RNA. Bull Cancer. 2019;106(12):1152–1159. doi:10.1016/j.bulcan.2019.08.011
22. Tian L, Wang ZY, Hao J, Zhang XY. miR-505 acts as a tumor suppressor in gastric cancer progression through targeting HMGB1. J Cell Biochem. 2018.
23. Ren L, Yao Y, Wang Y, Wang S. MiR-505 suppressed the growth of hepatocellular carcinoma cells via targeting IGF-1R. Biosci Rep. 2019;39(7). doi:10.1042/BSR20182442
24. Kapora E, Feng S, Liu W, Sakhautdinova I, Gao B, Tan W. MicroRNA-505-5p functions as a tumor suppressor by targeting cyclin-dependent kinase 5 in cervical cancer. Biosci Rep. 2019;39(7). doi:10.1042/BSR20191221
25. Yu Y, Chen X, Cang S. Cancer-related long noncoding RNAs show aberrant expression profiles and competing endogenous RNA potential in esophageal adenocarcinoma. Oncol Lett. 2019;18(5):4798–4808. doi:10.3892/ol.2019.10808
26. Zhou Y, Zheng X, Xu B, Hu W, Huang T, Jiang J. The Identification and analysis of mRNA-lncRNA-miRNA cliques from the integrative network of ovarian cancer. Front Genet. 2019;10:751. doi:10.3389/fgene.2019.00751
27. Bianchi ME, Agresti A. HMG proteins: dynamic players in gene regulation and differentiation. Curr Opin Genet Dev. 2005;15(5):496–506. doi:10.1016/j.gde.2005.08.007
28. Li Y, Qin C. MiR-1179 inhibits the proliferation of gastric cancer cells by targeting HMGB1. Hum Cell. 2019;32(3):352–359. doi:10.1007/s13577-019-00244-6
29. Chen J, Liu X, Zhang J, Zhao Y. Retraction: targeting HMGB1 inhibits ovarian cancer growth and metastasis by lentivirus-mediated RNA interference. J Cell Physiol. 2015;230(10):2579. doi:10.1002/jcp.25027
30. Dong YD, Cui L, Peng CH, Cheng DF, Han BS, Huang F. Expression and clinical significance of HMGB1 in human liver cancer: knockdown inhibits tumor growth and metastasis in vitro and in vivo. Oncol Rep. 2013;29(1):87–94. doi:10.3892/or.2012.2070
31. Rapoport BL, Steel HC, Theron AJ, et al. High mobility group box 1 in human cancer. Cells. 2020;9(7):1664. doi:10.3390/cells9071664
32. Cheng KJ, Alshawsh MA, Mejia Mohamed EH, Thavagnanam S, Sinniah A, Ibrahim ZA. HMGB1: an overview of its versatile roles in the pathogenesis of colorectal cancer. Cell Oncol. 2020;43(2):177–193. doi:10.1007/s13402-019-00477-5
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.