Back to Journals » OncoTargets and Therapy » Volume 13
Long Non-Coding RNA PART1 Exerts Tumor Suppressive Functions in Glioma via Sponging miR-190a-3p and Inactivation of PTEN/AKT Pathway
Authors Jin Z, Piao L, Sun G, Lv C, Jing Y, Jin R
Received 29 September 2019
Accepted for publication 11 December 2019
Published 4 February 2020 Volume 2020:13 Pages 1073—1086
DOI https://doi.org/10.2147/OTT.S232848
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjeev K. Srivastava
Zheng Jin,1 Lianhua Piao,2 Guangchao Sun,1 Chuanxiang Lv,1 Yi Jing,1 Rihua Jin1
1Department of Neurosurgery, The First Hospital of Jilin University, Changchun 130021, People’s Republic of China; 2Department of Physiology, College of Basic Medical Sciences, Jilin University, Changchun 130021, People’s Republic of China
Correspondence: Rihua Jin
Department of Neurosurgery, The First Hospital of Jilin University, Changchun 130021, People’s Republic of China
Email [email protected]
Background: Glioma is the most commonly diagnosed primary brain tumor. Dysregulation of long non-coding RNA (lncRNA) is associated with initiation and development of various cancer types including glioma.
Methods: The relative expression of lncRNA was analyzed by real time-quantitative polymerase chain reaction (RT-qPCR). Cell counting kit (CCK-8) and flow cytometry analysis were applied to explore the role of prostate androgen-regulated transcript 1 (PART1) in glioma cell lines. Luciferase reporter assay, Western blotting and RT-qPCR were used to investigate the association between PART1, miR-190a-3p and phosphatase and tensin homolog deleted on chromosome ten (PTEN) in glioma cell lines.
Results: In the present study, we elucidated a pivotal role and molecular mechanism of lncRNA PART1 in glioma cell lines. It was found that PART1 was significantly downregulated in glioma tissues compared to normal tissues according to TCGA data and our RT-qPCR results. The cell-based assays showed that PART1 suppressed cell proliferation and triggered cell apoptosis in glioma cell lines. PART1 inactivated PI3K/AKT cascade in glioma cell lines. Transfection of constitutively activated AKT (Myr-AKT) reversed PART1 induced cell apoptosis and cell growth arrest. The bioinformatic analysis suggested that miR-190a-3p might bind to PART1. In the dual luciferase reporter assay, we validated that PART1 directly bound to miR-190a-3p in glioma cell lines. Furthermore, there was a reciprocal repression between PART1 and miR-190-3p. In addition, PART1 upregulated PTEN and inactivated PI3K/AKT pathway in glioma cell lines. Moreover, silencing of PTEN reversed PART1 overexpression induced cell growth arrest and apoptosis. In glioma tissues, the Pearson Correlation analysis showed that there was a strong-positive correlation between PART1 level and PTEN mRNA level.
Conclusion: Taken together, the current study revealed a PART1/miR-190a-3p/PTEN/PI3K/AKT axis in glioma and provided novel insights for understanding the complex lncRNA-miRNA network in glioma.
Keywords: prostate androgen-regulated transcript 1, microRNA-190a-3p, PI3K/AKT signaling, glioma
Introduction
Globally, nearly 300,000 patients were diagnosed with brain and nervous system tumors, in addition, approximately 240,000 patients lose their lives due to cancer which were developed from the brain and nervous system in 2018.1 Glioma is the most common primary malignant tumor which was originated from brain, accounting for 50% of all brain malignancies.2–5 Glioma can be classified into several major types including oligodendroglioma, astrocytomas, glioblastoma and mixed gliomas.6 Although the incidence of glioma is relatively lower compared to other major cancer types such as lung cancer, the prognosis of patients with glioma is very poor due to the aggressive nature of glioma cells.7,8 Glioblastoma (GBM), a high-grade glioma, is the most lethal type of glioma and the overall survival of patients with GBM was less than 12 months.9 Therefore, it is currently urgent to investigate the molecular mechanism of glioma.
Long non-coding RNAs (lncRNAs) are single-stranded RNA longer than 200 nucleotides with no protein coding potential.10 LncRNAs regulate gene expression and cellular distribution of proteins via a variety of mechanisms including but not limited to direct interaction with mRNA, miRNA, cirRNA and protein, binding to promoter region or regulatory elements of target genes and mediation of chromatin remodeling.11,12 Through control of gene expression and protein function, lncRNAs are essential for numerous normal physiology processes. Dysregulation of lncRNAs is commonly observed in human diseases such as cancer.13 For instance, it was found that lncRNA NEAT1 controlled promoter methylation and promoted invasion of glioblastoma cells by binding to EZH2;14 in glioblastoma stem cells, lncRNA SOX2OT sponged miR-194-5p and miR-122 to facilitate proliferation and survival of GBM cells;15 expression of PART1 was relatively higher in non-small cell lung cancer (NSCLC) and could serve as a biomarker for the prognosis of patients with NSCLC.16 Whereas, the role of PART1 in glioma remains unknown.
Sustained cell proliferation signaling is one of the hallmarks in cancer cells. As one of the most well-known tumor suppressor, PTEN shut down PI3K/AKT signaling, leading to cell growth arrest and cell apoptosis of cancer cells.17,18 Loss of PTEN is a frequent event in glioblastoma and is associated with the prognosis of patients with GBM.19 Recent studies have revealed that the downregulation of PTEN was the consequence of gene mutation, promoter methylation and deregulation of miRNA.20–22 LncRNAs were also involved in the regulation of PTEN expression in cancer cells.23
In the current study, we identified PART1 as a significantly downregulated lncRNA in glioma cells. In addition, PART1 reduced cell proliferation and induced cell apoptosis in glioma cells via inactivation of PI3K/AKT signaling. Mechanistically, PART1 sponged miR-190a-3p and upregulated PTEN expression. The association among PART1, miR-190a-3p and PTEN expression was also analyzed in glioma. Our findings indicated that PART1 was pivotal for glioma progression.
Materials and Methods
Cell Culture
Human glioma cell lines (U87MG, LN-18 and LN-428) were purchased from American Type Culture Collection (Manassas, VA). These cells were cultured in DMEM (Invitrogen, Carlsbad, CA) supplemented with 10% FBS (Gibco; Thermo Fisher Scientific, Waltham, MA) in a 37°C incubator with 5% CO2. These cells were passaged for less than 6 months after recipient from ATCC.
Clinical Samples
In total, 50 glioma tissues were collected from patients with glioma underwent surgery. Six normal brains were obtained from patients with severe traumatic brain injury undergoing internal decompression surgery. These tissues were collected in the first hospital of Jilin University from June, 2013 to July 2018. Written informed consents were obtained from all participants. The Ethic Committee of the first hospital of Jilin University approved and supervised the protocol and the whole procedures of experiments. Tissues were immediately stored in liquid nitrogen before subjection to RNA extraction or protein extraction.
Overexpression of PART1 in Glioblastoma Cells
The full length of PART1 sequence was ligated into pcDNA3.1 vector (Invitrogen) to obtain pcDNA3.1-PART1 reconstruction plasmid. In brief, 2μg pcDNA3.1 or pcDNA3.1-PART1 plasmid were transfected into 1X106 U87MG cells or LN-18 cells with Lipofectamine 3000 (Invitrogen) following manufacturer’s protocol. The cells were harvested and subjected to RT-qPCR for validation of overexpression efficiency at 48 h after transfection.
Overexpression and Inhibition of miR-190a-3p in Glioma Cells
miR-190a-3p mimic/miR-NC mimic and miR-190a-3p inhibitor/miR-NC inhibitor were purchased from Shanghai GenePharma (Shanghai, China). Their sequences were listed as follow: miR-190a-3p mimic: 5ʹ-CUAUAUAUCAAACAUAUUCCU-3ʹ; miR-NC mimic: 5ʹ-UCGCUUGGUGCAGGUCGGGAA-3ʹ; miR-190a-3p inhibitor: 5ʹ-AGGAAUAUGUUUGAUAUAUAG-3ʹ; miR-NC inhibitor: 5ʹ- UUCUCCGAACGUGUCACGUTT-3ʹ. miR-190a-3p mimic/miR-NC mimic and miR-190a-3p inhibitor/miR-NC inhibitor were transfected into U87MG cells and LN-18 cells at the work concentration of 80 nM with Lipofectamine 3000 reagent (Invitrogen) following the manufacturer’s protocol. After 48 h, the cells were harvested and subjected to RT-qPCR or Western blotting.
Small Interference RNA Mediated Silencing of PTEN in Glioma Cells
The siRNA target PTEN and the negative control siRNA were synthesized and purchased from GenePharma (Shanghai, China). The PTEN siRNA or control siRNA was transfected into U87MG cells or LN-18 cells with Lipofectamine RNAiMAX (Invitrogen) according to the manufacturer’s protocol. Seventy-two-hour later, the knockdown efficiency was confirmed via Western blotting.
Constitutive Activation of AKT Activity in Glioma Cells
pLNCX-Myr-AKT containing a constitutive activated mutant AKT was bought from Addgene (Watertown, MA). For transient transfection, 2 μg pLNCX-Myr-AKT or pLNCX was transfected into 1X106 U87MG cells or LN-18 cells with Lipofectamine 3000 (Invitrogen) following manufacturer’s protocol. The cells were harvested for Western blotting to validate the elevation of activated AKT at 48 h after transfection.
RNA Extraction and RT-qPCR
Total RNA from cultured cells or tissues was extracted with TRIzol reagent (Invitrogen). The concentration and quality of RNA were determined with the NanoDrop 2000 (Thermo Fisher Scientific). After that, RNA was reverse transcribed into first-strand cDNA with All‐in‐One First‐Strand cDNA Synthesis kit (GeneCopoeia, Rockville, MD). qPCR was performed with the TB Green Premix Ex Taq kit (Takara, Toyoko, Japan) on a CFX96 system (Bio-Rad, Hercules, CA). 2−ΔΔCT method was used to calculate the relative gene expression. GAPHD and U6 were used as internal controls for detection of mRNA level in PART1/PTEN and miR-190a-3p, respectively. PART1-forward: 5ʹ-AAGGCCGTGTCAGAACTCAA-3ʹ; PART1-reverse: 5ʹ-GTTTTCCATCTCAGCCTGGA-3ʹ; PTEN-forward: 5ʹ-TGGATTCGACTTAGACTTGACCT-3ʹ; PTEN-reverse: 5ʹ-GGTGGGTTATGGTCTTCAAAAGG-3ʹ. GAPDH-forward: 5ʹ-GTCTCCTCTGACTTCAACAGCG-3ʹ; GAPDH-reverse: 5ʹ-ACCACCCTGTTGCTGTAGCCAA-3ʹ; Stem-loop primer: 5ʹ-CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGGAAT-3ʹ; miR-190a-3p-forward: 5ʹ-TCGGCAGGCTATATATCAAAC-3ʹ; miR-190a-3p-reverse: 5ʹ-CTCAACTGGTGTCGTGGA-3ʹ; U6-forward: 5ʹ-CTCGCTTCGGCAGCACATATACT-3ʹ; U6-reverse: 5ʹ-ACGCTTCACGAATTTGCGTGTC-3ʹ.
Protein Extraction and Western Blotting
Primary antibodies against PI3K (#4255, 1:2000), p-PI3K (#17366, 1:2000), AKT (#2920, 1:2000), p-AKT (#4060, 1:2000) were bought from Cell Signaling Technology (Carlsbad, CA). Primary antibodies for PTEN (ab32199, 1:2000), Bax (ab32503, 1:2000), Bcl2 (ab32124, 1:2000) and GAPDH (ab8245, 1:5000) were obtained from Abcam (Cambridge, UK). HRP-conjugated secondary antibodies against mouse (AP308P, 1:10,000) and rabbit (AP132, 1:10,000) were purchased from Sigma Aldrich (St. Louis, MI). Protein lysates were prepared with RIPA lysis buffer (Thermo Fisher Scientific, Bremen, Germany). The concentration of protein lysates was determined via a BCA Protein Assay kit (Thermo Fisher Scientific). In brief, 20 μg proteins were loaded on each well of 8% SDS-PAGE gel. The PVDF membrane was used to transfer proteins from SDS-PAGE gel. The PVDF membrane was blocked with 5% non-fat milk at room temperature for 1 h. After that, the PVDF membrane was incubated with the representative primary antibody overnight at 4°C. On the next day, the PVDF membrane was incubated with the secondary antibody for 1 h at room temperature. The blots were developed with ECL reagent (Santa Cruz Biotechnology). The intensity of bands was normalized to GAPDH. The relative-protein expression was analyzed by Image J software 1.8.0 (NIH, Bethesda, MD).
Cell Proliferation Assay
Cell proliferation ability was measured by a CCK-8 kit (Dojindo, Toyoko, Japan). At 48 h after plasmid transfection, equal number of U87MG cells and LN-18 cells were plated in 96-well plates and maintained for 1, 2 or 3 days, respectively. At the corresponding timepoints of 0, 1, 2, 3 days, 10 μL of CCK-8 solution was added into indicated wells and sustained for 2 h. The medium containing CCK-8 solution was removed to new 96-well plates and the absorbance at 450 nM was detected to reflect cell number.
Cell Apoptosis Assay
Flow cytometry was used to detect the percentage of apoptotic cells by a Dead Cell Apoptosis Detection Kit with Annexin V and PI (Invitrogen). Briefly, at 48 h after transfection of plasmid, cells were harvested and suspended in Annexin Binding buffer. After that, PI and Annexin V were added into the above cell suspension and incubated at room temperature for 30 min. Afterwards, cells were subjected to flow cytometry for analysis of apoptosis. The cells positive for Annexin V was at early apoptosis and cells positive for both Annexin V and PI were at later apoptosis.
Bioinformatic Analysis
Via the Gene Expression Profiling Interactive Analysis database (GEPIA, http://gepia.cancer-pku.cn/), the expression of PART1 in normal brains and glioma tissues were compared based on TCGA-GBM and TCGA-LGG. MiRDB database (http://mirdb.org/) was used to predict potential miRNA targets of PART1. The correlation between PART1 and PTEN mRNA expression was analyzed with ENCORI database (http://starbase.sysu.edu.cn/) based on expression data 526 brain low-grade glioma.
Dual Luciferase Reporter Assay
Briefly, 2X105 U87MG or LN-18 cells were seeded in each well of 24-well plates and transfected with the indicated plasmids, pmirGLO-PART1-WT or pmirGLO-PART1-Mut or pmirGLO-PTEN 3ʹUTR-WT or pmirGLO-PTEN 3ʹUTR-Mut. Simultaneously, cells were transfected with miR-190a-3p mimic or miR-NC mimic using Lipofectamine 3000 (Invitrogen). Forty-eight-hour later, the relative luciferase activity in each group was determined by a Dual Luciferase Reporter Assay System kit (Promega, Madison, WI) on a Centro LB960 XS3 (Berthold, German) following manufacturer’s guideline.
Statistical Analysis
All data were analyzed with Graphpad 6.0 software and presented as mean±SD. Two groups were compared with Student’s t test. The experiments were repeated for three times. The differences among three groups were compared with one-way ANOVA followed by Newman Keuls test. The difference was considered as statistically significant when p value was less than 0.05.
Results
LncRNA PART1 Was Downregulated in Glioma
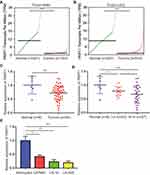
We used GEPIA to explore the expression pattern of PART1 in normal brains, GBM (high-grade glioma) and low grade glioma (LGG) from TCGA-GBM and TCGA-LGG datasets. The results suggested a significant downregulation of PART1 in GBM compared with normal brains (Figure 1A). In addition, PART1 was also significantly decreased in LGG compared with normal brains (Figure 1B). For validation, six-normal brain tissues and 50 glioma samples were collected for the study. The RT-qPCR showed that there was a significant downregulation of PART1 in glioma tissues compared with normal brains (Figure 1C). In addition, relatively lower levels of PART1 were observed in high-grade glioma (grade III-IV, n=27) compared with low-grade glioma (grade I-II, n=23) (Figure 1D). We also detected PART1 expression in glioma cell lines (U87MG, LN-18, LN-428) and primary astrocytes. PART1 was found to be decreased in glioma cells compared with astrocytes (Figure 1E). The data suggested that PART1 was decreased in glioma.
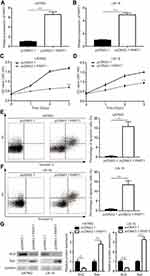
Overexpression of PART1 Inhibited Cell Proliferation and Induced Cell Apoptosis in Glioma
To investigate the role of PART1 in glioma, recombinant PART1 was transfected into U87MG cells and LN-18 cells. Transfection of recombinant PART1 significantly increased PART1 in U87MG cells and LN-18 cells (Figure 2A and B). The cell proliferation assay revealed that overexpression of PART1 greatly reduced cell proliferation of U87MG cells (Figure 2C). Consistently, PART1 overexpression decreased cell proliferation of LN-18 cells (Figure 2D). We proposed that the decreased cell proliferation might be the consequence of cell death. Consequently, the flow cytometry was used to detect the percentage of apoptotic cells after PART1 overexpression. As we expected, PART1 overexpression predominantly elevated the percentage of apoptotic cells to approximate 15% in U87MG cells and LN-18 cells (Figure 2E and F). As acknowledged, the apoptotic process is controlled by the Bcl-2 family member, including pro-apoptotic protein (Bax) and anti-apoptotic protein (Bcl2).24 Moreover, Western blotting showed that Bax protein level was increased while Bcl2 protein level was decreased in cells transfected with recombinant PART1 (Figure 2G). These data manifested that PART1 promoted cell apoptosis to inhibited cell proliferation in glioma.
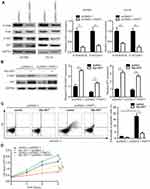
PART1 Evoked Cell Apoptosis via Inactivation of PI3K/AKT Pathway
Sustained activation of PI3K/AKT is essential for the cell survival of glioma cells. Western blotting revealed that overexpression of PART1 significantly decreased the phosphorylation levels of PI3K and AKT while the total PI3K and AKT protein levels were not altered in U87MG cells and LN-18 cells (Figure 3A), suggesting the inactivation of PI3K/AKT pathway. To support that PART1 repressed PI3K/AKT activity, a constitutively activated AKT (Myr-AKT) mutant was transfected into U87MG cells. Transfection of Myr-AKT elevated p-AKT in U87MG cells and partly restored p-AKT expression in U87MG cells transfected with pcDNA3.1-PART1 (Figure 3B). Furthermore, Myr-AKT reversed cell apoptosis induced by PART1 overexpression in U87MG cells (Figure 3C). The cell proliferation assay showed that Myr-AKT promoted cell proliferation and partly reversed PART1 overexpression mediated cell proliferation inhibition of U87MG cells (Figure 3D). The results collectively suggested that PART1 inhibited glioma proliferation via inactivation of PI3K/AKT pathway.
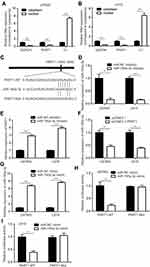
PART1 Sponged miR-190a-3p in Glioma
To explore the subcellular localization of PART1 in glioma cells, the cytoplasmatic and nuclear fractions of glioma were isolated. RT-qPCR suggested that PART1 was mainly localized in the cytoplasm (Figure 4A and B), which indicated that PART1 might function as a competing endogenous RNA (ceRNA) in glioma cells. We predicted the potential miRNAs that might be associated with PART1 by miRDB database. Among the predicted miRNAs, miR-190a-3p was an oncogenic miRNA and a negative regulator of PI3K/AKT pathway in glioma. The sequence of miR-190a-3p was complementary to PART1 at 1983–1989 (Figure 4C). miR-190a-3p inhibitor was transfected into glioma cells. Transfection of miR-190a-3p inhibitor significantly decreased miR-190a-3p levels in U87MG cells and LN-18 cells (Figure 4D). Downregulation of miR-190a-3p significantly increased PART1 expression in U87MG cells and LN-18 cells (Figure 4E). On the contrary, overexpression of PART1 predominantly decreased miR-190a-3p expression in U87MG cells and LN-18 cells (Figure 4F). Transfection of miR-190a-3p mimic predominantly elevated miR-190a-3p expression in U87MG cells and LN-18 cells (Figure 4G). The dual luciferase assay indicated that miR-190a-3p mimic significantly repressed the relative luciferase activity of PART1-WT instead of PART1-Mut in U87MG cells (Figure 4H). Meanwhile, the similar results were observed in LN-18 cells (Figure 4I).
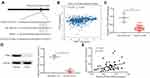
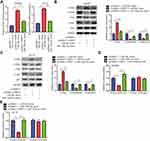
PART1 Upregulated PTEN and Inactivated AKT Signaling via Sponging miR-190a-3p
miR-190a-3p facilitated glioma cell proliferation via targeting the well-known tumor suppressor, PTEN.25 There were complementary binding sites among miR-190a-3p, PART1 and PTEN 3ʹUTR (Figure 5A). To investigate the clinical association between PART1 and PTEN in glioma, we analyzed the level of PART1 and PTEN in LGG by ENCORI database. There was a positive correlation between the level of PART1 and mRNA level of PTEN in LGG (n=526) (Figure 5B). In our collected samples, RT-qPCR suggested a significant downregulation of PTEN mRNA level in glioma compared with normal brains (Figure 5C). Additionally, Western blotting also showed a significant reduction of PTEN protein level in glioma (n=11) compared with normal brain (n=3) (Figure 5D). The Pearson correlation analysis further confirmed a positive correlation between the PART1 level and PTEN mRNA level in glioma tissues (Figure 5E). Overexpression of PART1 increased PTEN mRNA level which was reversed after transfection of miR-190a-3p mimic in U87MG cells and LN-18 cells (Figure 6A). At protein level, PART1 overexpression increased PTEN level while decreased P-PI3K and P-AKT level in U87MG cells which was reversed after overexpression of miR-190a-3p (Figure 6B). The consistent results were observed in LN-18 cells (Figure 6C). More importantly, in the dual luciferase assay, transfection of miR-190a-3p mimic repressed the relative luciferase activity of PTEN 3ʹUTR-WT which was reversed after PART1 overexpression in U87MG cells and LN-18 cells (Figure 6D and E).
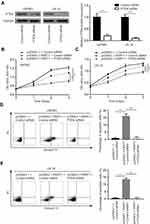
PART1 Reduced Cell Proliferation and Induced Cell Apoptosis via Upregulation of PTEN in Glioma
PTEN siRNA was transfected into U87MG cells and LN-18 cells. The transfection of PTEN siRNA significantly downregulated PTEN protein level in these cells (Figure 7A). In the cell proliferation assay, we observed that the cell proliferation inhibition induced by PART1 was partially reversed upon PTEN knockdown in U87MG cells and LN-18 cells (Figure 7B and C). In addition, the silencing of PTEN reduced percentage of apoptotic cells in U87MG cells and LN-18 cells transfected with PART1 (Figure 7D and E). The data suggested that PART1 inhibited glioma cell proliferation via elevation of PTEN.
Discussion
Studies on the signaling network have obtained satisfactory achievements for the development of target therapy for patients with glioma.26 Growing evidences have elaborated a pivotal role of lncRNA during cancer cell proliferation, metastasis and the development of drug resistance.27 LncRNA PART1 was firstly identified as a human prostate-specific, androgen-induced gene which was located in chromosome 5q12, a region that harbored tumor suppressor genes.28 The previous reports have suggested a controversial role of PART1 in cancers. For instance, Hu et al found that PART1 was overexpressed in bladder cancer specimens and silencing of PART1 led to cell proliferation inhibition and apoptosis in bladder cancer cells;29 data mining of TCGA dataset revealed that higher expression of PART1 was associated with a higher risk of recurrence for patients with hepatocellular carcinoma;30 in oral squamous cell carcinoma, however, patients with higher expression of PART1 exhibited longer overall survival compared with patients with lower expression of PART1.31 Although one study indicated that higher expression of PART1 was associated with longer survival in patients with GBM,32 the role and molecular mechanisms of PART1 in glioma are not known. We firstly discovered that PART1 was a downregulated lncRNA in glioma via bioinformatic analysis of TCGA-GBM dataset and RT-qPCR in glioma tissues and normal brain tissues. The cell proliferation and apoptosis assays suggested that overexpression of PART1 markedly inhibited cell proliferation and induced apoptosis in glioma cells.
PI3K/AKT signaling is a well-characterized drug target for its central role in mediating cancer cell survival and proliferation.33,34 Hyper-activation of PI3K/AKT cascade is essential for cell proliferation and survival of glioma cells.35 Inhibition of PI3K/AKT pathway led to cell apoptosis and cell growth arrest in glioma.36 It was observed that overexpression of PART1 decreased phosphorylation levels of PI3K and AKT in glioma cells, suggesting that PART1 inactivated PI3K/AKT pathway in glioma. More importantly, transfection of constitutively activated AKT mutant (Myr-AKT) reversed cell proliferation inhibition and cell apoptosis induced by PART1 in glioma cells. These data suggested that PART1 relied on inactivation of PI3K/AKT cascade to inhibit glioma cell proliferation and survival. The hyper-activation of PI3K/AKT cascade is due to the loss of several tumor suppressors such as PTEN, NHERF1 and PHLPP.37 The downregulation of PTEN in glioma is the consequence of mutation and epigenetic regulation.38,39 Dysregulation of miRNA was pivotal for the development of glioma.40 Previous studies have found that PTEN could be targeted by several miRNAs including miR-130a, miR-374b and miR-190a-3p.25,41,42 We found that miR-190a-3p targeted PTEN in glioma cells which was consistent with a recent report.25 PART1 directly sponged miR-190a-3p to promote PTEN expression and inactivated PI3K/AKT cascade in glioma cells. Furthermore, PART1 inhibited cell proliferation and induced cell apoptosis in glioma via upregulation of PTEN. In glioma specimens, PART1 level was positively correlated with PTEN mRNA level.
In conclusion, our data indicated that PART1 was downregulated in glioma. PART1 inhibited cell proliferation and induced apoptosis of glioma cells. Mechanistically, PART1 sponged miR-190a-3p to increase PTEN expression in glioma. The current study revealed that PART1 might represent a novel therapeutic target or biomarker for glioma. The molecular mechanism and role of PART1 in glioma will be further investigated in vivo in the future.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi:10.3322/caac.21492
2. Morgan LL. The epidemiology of glioma in adults: a “state of the science” review. Neuro Oncol. 2015;17:623–624. doi:10.1093/neuonc/nou358
3. Louis DN, Perry A, Reifenberger G, et al. The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 2016;131:803–820. doi:10.1007/s00401-016-1545-1
4. Chai RC, Li YM, Zhang KN, et al. RNA processing genes characterize RNA splicing and further stratify lower-grade glioma. JCI Insight. 2019;5. doi:10.1172/jci.insight.130591
5. Chai RC, Zhang KN, Chang YZ, et al. Systematically characterize the clinical and biological significances of 1p19q genes in 1p/19q non-codeletion glioma. Carcinogenesis. 2019;40:1229–1239. doi:10.1093/carcin/bgz102
6. Davis ME. Epidemiology and overview of gliomas. Semin Oncol Nurs. 2018;34:420–429. doi:10.1016/j.soncn.2018.10.001
7. Magnani I, Novielli C, Bellini M, Roversi G, Bello L, Larizza L. Multiple localization of endogenous MARK4L protein in human glioma. Cell Oncol. 2009;31:357–370. doi:10.3233/CLO-2009-0481
8. Jiang T, Mao Y, Ma W, et al. CGCG clinical practice guidelines for the management of adult diffuse gliomas. Cancer Lett. 2016;375:263–273. doi:10.1016/j.canlet.2016.01.024
9. Ostrom QT, Gittleman H, Stetson L, Virk SM, Barnholtz-Sloan JS. Epidemiology of gliomas. Cancer Treat Res. 2015;163:1–14. doi:10.1007/978-3-319-12048-5_1
10. Kung JT, Colognori D, Lee JT. Long noncoding RNAs: past, present, and future. Genetics. 2013;193:651–669. doi:10.1534/genetics.112.146704
11. Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77:3965–3981. doi:10.1158/0008-5472.CAN-16-2634
12. Akhade VS, Pal D, Kanduri C. Long noncoding RNA: genome organization and mechanism of action. Adv Exp Med Biol. 2017;1008:47–74. doi:10.1007/978-981-10-5203-3_2
13. Peng WX, Koirala P, Mo YY. LncRNA-mediated regulation of cell signaling in cancer. Oncogene. 2017;36:5661–5667. doi:10.1038/onc.2017.184
14. Chen Q, Cai J, Wang Q, et al. Long noncoding RNA NEAT1, regulated by the EGFR pathway, contributes to glioblastoma progression through the WNT/beta-catenin pathway by scaffolding EZH2. Clin Cancer Res. 2018;24:684–695. doi:10.1158/1078-0432.CCR-17-0605
15. Su R, Cao S, Ma J, et al. Knockdown of SOX2OT inhibits the malignant biological behaviors of glioblastoma stem cells via up-regulating the expression of miR-194-5p and miR-122. Mol Cancer. 2017;16:171. doi:10.1186/s12943-017-0737-1
16. Li M, Zhang W, Zhang S, Wang C, Lin Y. PART1 expression is associated with poor prognosis and tumor recurrence in stage I-III non-small cell lung cancer. J Cancer. 2017;8:1795–1800. doi:10.7150/jca.18848
17. Moon SH, Kim DK, Cha Y, Jeon I, Song J, Park KS. PI3K/Akt and Stat3 signaling regulated by PTEN control of the cancer stem cell population, proliferation and senescence in a glioblastoma cell line. Int J Oncol. 2013;42:921–928. doi:10.3892/ijo.2013.1765
18. Bao L, Li X, Lin Z. PTEN overexpression promotes glioblastoma death through triggering mitochondrial division and inactivating the Akt pathway. J Recept Signal Transduct Res. 2019;1–11. doi:10.1080/10799893.2019.1655051
19. Koul D. PTEN signaling pathways in glioblastoma. Cancer Biol Ther. 2008;7:1321–1325. doi:10.4161/cbt.7.9.6954
20. Li H, Yang BB. Stress response of glioblastoma cells mediated by miR-17-5p targeting PTEN and the passenger strand miR-17-3p targeting MDM2. Oncotarget. 2012;3:1653–1668. doi:10.18632/oncotarget.810
21. Xu J, Li Z, Wang J, Chen H, Fang JY. Combined PTEN mutation and protein expression associate with overall and disease-free survival of glioblastoma patients. Transl Oncol. 2014;7:196–205 e1. doi:10.1016/j.tranon.2014.02.004
22. Wiencke JK, Zheng S, Jelluma N, et al. Methylation of the PTEN promoter defines low-grade gliomas and secondary glioblastoma. Neuro Oncol. 2007;9:271–279. doi:10.1215/15228517-2007-003
23. Qiao Q, Li H. LncRNA FER1L4 suppresses cancer cell proliferation and cycle by regulating PTEN expression in endometrial carcinoma. Biochem Biophys Res Commun. 2016;478:507–512. doi:10.1016/j.bbrc.2016.06.160
24. Zhou L, Gao R, Wang Y, Zhou M, Ding Z. Loss of BAX by miR-365 promotes cutaneous squamous cell carcinoma progression by suppressing apoptosis. Int J Mol Sci. 2017;18. doi:10.3390/ijms18061157
25. Chu L, Yu L, Liu J, et al. Long intergenic non-coding LINC00657 regulates tumorigenesis of glioblastoma by acting as a molecular sponge of miR-190a-3p. Aging (Albany NY). 2019;11:1456–1470. doi:10.18632/aging.101845
26. Lin L, Cai J, Jiang C. Recent advances in targeted therapy for glioma. Curr Med Chem. 2017;24:1365–1381. doi:10.2174/0929867323666161223150242
27. Pecero ML, Salvador-Bofill J, Molina-Pinelo S. Long non-coding RNAs as monitoring tools and therapeutic targets in breast cancer. Cell Oncol (Dordr). 2019;42:1–12. doi:10.1007/s13402-018-0412-6
28. Lin B, White JT, Ferguson C, et al. PART-1: a novel human prostate-specific, androgen-regulated gene that maps to chromosome 5q12. Cancer Res. 2000;60:858–863.
29. Hu X, Feng H, Huang H, et al. Downregulated long noncoding RNA PART1 inhibits proliferation and promotes apoptosis in bladder cancer. Technol Cancer Res Treat. 2019;18:1533033819846638. doi:10.1177/1533033819846638
30. Ye J, Zhang J, Lv Y, et al. Integrated analysis of a competing endogenous RNA network reveals key long noncoding RNAs as potential prognostic biomarkers for hepatocellular carcinoma. J Cell Biochem. 2019;120:13810–13825. doi:10.1002/jcb.28655
31. Li S, Chen X, Liu X, et al. Complex integrated analysis of lncRNAs-miRNAs-mRNAs in oral squamous cell carcinoma. Oral Oncol. 2017;73:1–9. doi:10.1016/j.oraloncology.2017.07.026
32. Zhang XQ, Sun S, Lam KF, et al. A long non-coding RNA signature in glioblastoma multiforme predicts survival. Neurobiol Dis. 2013;58:123–131. doi:10.1016/j.nbd.2013.05.011
33. Starska K, Forma E, Lewy-Trenda I, Stasikowska-Kanicka O, Skora M, Brys M. Fibroblast growth factor receptor 1 and 3 expression is associated with regulatory PI3K/AKT kinase activity, as well as invasion and prognosis, in human laryngeal cancer. Cell Oncol (Dordr). 2018;41:253–268. doi:10.1007/s13402-017-0367-z
34. Zhou F, Geng J, Xu S, et al. FAM83A signaling induces epithelial-mesenchymal transition by the PI3K/AKT/Snail pathway in NSCLC. Aging (Albany NY). 2019;11:6069–6088. doi:10.18632/aging.102163
35. Zhao HF, Wang J, Shao W, et al. Recent advances in the use of PI3K inhibitors for glioblastoma multiforme: current preclinical and clinical development. Mol Cancer. 2017;16:100. doi:10.1186/s12943-017-0670-3
36. Foster KA, Jane EP, Premkumar DR, Morales A, Pollack IF. NVP-BKM120 potentiates apoptosis in tumor necrosis factor-related apoptosis-inducing ligand-resistant glioma cell lines via upregulation of Noxa and death receptor 5. Int J Oncol. 2015;47:506–516. doi:10.3892/ijo.2015.3035
37. Molina JR, Agarwal NK, Morales FC, et al. PTEN, NHERF1 and PHLPP form a tumor suppressor network that is disabled in glioblastoma. Oncogene. 2012;31:1264–1274. doi:10.1038/onc.2011.324
38. Janssen EA, Soiland H, Skaland I, et al. Comparing the prognostic value of PTEN and Akt expression with the mitotic activity index in adjuvant chemotherapy-treated node-negative breast cancer patients aged <55 years. Cell Oncol. 2007;29:25–35.
39. Elhag R, Mazzio EA, Soliman KF. The effect of silibinin in enhancing toxicity of temozolomide and etoposide in p53 and PTEN-mutated resistant glioma cell lines. Anticancer Res. 2015;35:1263–1269.
40. Han B, Meng X, Chen H, et al. Epigenetic silencing of miR-338 facilitates glioblastoma progression by de-repressing the pyruvate kinase M2-beta-catenin axis. Aging (Albany NY). 2017;9:1885–1897. doi:10.18632/aging.101271
41. Ye L, Wang Y, Nie L, Qian S, Xu M. MiR-130 exerts tumor suppressive function on the tumorigenesis of human non-small cell lung cancer by targeting PTEN. Am J Transl Res. 2017;9:1856–1865.
42. Noguchi S, Ogusu R, Wada Y, Matsuyama S, Mori T. PTEN, a target of Microrna-374b, contributes to the radiosensitivity of canine oral melanoma cells. Int J Mol Sci. 2019;20:4631. doi:10.3390/ijms20184631
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.