Back to Journals » OncoTargets and Therapy » Volume 12
Long non-coding RNA CASC2 serves as a ceRNA of microRNA-21 to promote PDCD4 expression in oral squamous cell carcinoma
Authors Pan L, Chen H, Bai Y, Wang Q, Chen L
Received 20 December 2018
Accepted for publication 15 February 2019
Published 6 May 2019 Volume 2019:12 Pages 3377—3385
DOI https://doi.org/10.2147/OTT.S198970
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Prof. Dr. Geoffrey Pietersz
Lina Pan,1* Hui Chen,2* Yang Bai,1 Qibao Wang,2 Liang Chen1
1Shungeng Campus, Jinan Stomatological Hospital, Jinan, Shandong 250001, People’s Republic of China; 2Department of Endodontics, Jinan Stomatological Hospital, Jinan, Shandong 250001, People’s Republic of China
* These authors contributed equally to this work
Background: Oral squamous cell carcinoma (OSCC) is a common oral disease with high morbidity and mortality. Recently, long non-coding RNAs (lncRNAs) were identified as critical regulators in OSCC tumorigenesis. The present study aimed to work out the functions and the possible molecular mechanisms of lncRNA CASC2 in human OSCC.
Methods: The expression levels of CASC2 in clinical OSCC tissue samples and cultured OSCC cell lines were detected by RT-qPCR analysis. MTT assay was performed to detect the proliferation ability of OSCC cells, whereas the apoptosis rate and cell cycle distribution of OSCC cells were determined by flow cytometric analysis. The expression levels of relevant proteins were detected by Western blot assay. Dual-luciferase reporter assay was performed to validate the predicted relationship between CASC2, miR-21 amd PDCD4. The role of CASC2 in OSCC tumorigenesis in vivo was evaluated using a nude mouse tumor model.
Results: The results demonstrated that CASC2 was significantly downregulated in clinical OSCC tissue samples and cultured OSCC cell lines. Low CASC2 expression was closely correlated with adverse clinicopathological characteristics of OSCC patients. Functionally, overexpression of CASC2 remarkably inhibited cell proliferation partly through inducing cell cycle arrest and cell apoptosis. Furthermore, bioinformatics analysis and dual-luciferase reporter assay showed that CASC2 might act as a competing endogenous RNA of miR-21 to promote the expression of PDCD4. Rescue experiments also showed that miR-21 blocked the tumor-suppressive role that CASC2 exerted in OSCC cells. Finally, in vivo study indicated that overexpression of CASC2 restrained OSCC tumor growth in volume and weight.
Conclusion: In conclusion, these findings indicate that CASC2/miR-21/PDCD4 axis might be a potential regulator of OSCC tumorigenesis, and shed new light on lncRNA-directed diagnostics and therapeutics in OSCC.
Keywords: oral squamous cell carcinoma, long non-coding RNA, CASC2, microRNA-21, PDCD4
Corrigendum for this paper has been published
Introduction
Oral cancer is one of the 10 most common cancers worldwide, and oral squamous cell carcinoma (OSCC) is the predominant histological type of oral cancer.1 Despite the improved diagnostic and therapeutic methods in recent years, the 5-year survival rate of OSCC patients remains approximately 50%.2 Consequently, there is a compelling need for better understanding of the potential molecular mechanisms underlying OSCC progression.
Long non-coding RNAs (lncRNAs) belong to a subgroup of non-coding RNAs, whose length is more than 200 nucleotides. LncRNAs are deeply implicated in diverse biological courses, including cellular differentiation, cell lineage choice, organogenesis and tissue homeostasis,3 and recently, many lncRNAs are also known to play a key regulatory role in cancer biology.4 The lncRNA Cancer Susceptibility Candidate 2 (CASC2), that maps to human chromosome 10q26, has been frequently featured as a tumor suppressor in a wide variety of human cancers, such as bladder cancer,5 papillary thyroid cancer6 and hepatocellular carcinoma.7 However, the expression and function of CASC2 in OSCC remain to be further elucidated.
In the present study, we aimed to investigate the expression levels of CASC2 in human OSCC tissues and cell lines. Further experiments were performed to explore the biological functions of CASC2 with respect to OSCC cell phenotypes in vitro and in vivo.
Materials and methods
Patients and tissue samples
Paired tumor specimens and their adjacent non-tumor tissues were derived from 69 OSCC patients who underwent surgical resection at Jinan Stomatological Hospital (Jinan, China) between Jan 2016 and May 2018. None of the patients recruited in this study received any pre-operative chemotherapy or radiotherapy. All these tissue samples were immediately frozen in liquid nitrogen and stored at −80 °C until further use. This study was conducted in accordance with the Declaration of Helsinki, and was approved by the Ethics Committees of Jinan Stomatological Hospital. Written informed consent was obtained from all participants.
Cell culture
OSCC cell lines (Tca8113, SCC-9, TSCCa, CAL-27) and normal oral keratinocyte (NOK) cell line, obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China), were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM; Invitrogen, Carlsbad, CA, USA) containing 10% fetal bovine serum (FBS; Biowest, Loire, France), 100 U/ml penicillin and 100 µg/ml streptomycin in culture flasks at 37 °C with 5% CO2.
Cell transfection
miR-21 mimics and its negative control were purchased from RiboBio Co., Ltd. (Guangzhou, China). The full-length CASC2 cDNA was amplified and cloned into the pcDNA3.1 vector (Invitrogen). Cells were seeded in 24-well plates and transfected with the vectors or oligonucleotides using Lipofectamine 2000 Transfection Reagent (Invitrogen). Transfection efficiency was validated at 48 h after transfection.
RNA extraction and RT-qPCR analysis
Total RNA was extracted from tissues/cells using Trizol Reagent (Invitrogen). 1–3 μg RNA was reverse transcribed to cDNA using the PrimeScript RT reagent Kit (TaKaRa, Dalian, China) or commercial miRNA reverse transcription PCR kit (RiboBio). qPCR assay was then performed using the SYBR Premix Ex Taq™ II kit (TaKaRa), primers and cDNA templates on the ABI PRISM 7900 Sequence Detection System (Applied Biosystems, Foster City, CA, USA). The 2−ΔΔCt method was used for data quantification,8 and GAPDH or U6 was used as the internal control.
MTT assay
Cell proliferation was measured using 3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyl tetrazolium bromide (MTT) assay. 48 h after transfection, the cells were seeded at a density of 3×103 cells/well in 96-well plates. At indicated time points, 20 μl MTT regents (5 mg/ml; Sigma-Aldrich, St. Louis, MO, USA) was supplemented into each well, and the plates were incubated for another 4 h. Then the supernatant was removed before the addition of 150 μl DMSO (Sigma-Aldrich). A microplate reader (Bio-Rad Laboratories, Hercules, CA, USA) was used to measure the absorbance at 570 nm.
Cell apoptosis analysis
Cell apoptosis was evaluated using the Annexin-V-FITC Apoptosis Detection Kit (BD Biosciences, Franklin lakes, NJ, USA). 48 h after transfection, the cells were collected, washed with PBS, resuspended in binding buffer and stained with 5 μl of Annexin V-FITC and 2 μl of propidium iodide (PI) in dark at room temperature for 30 min. The samples were then analyzed by FACScan flow cytometer (BD Biosciences).
Cell cycle analysis
48 h after transfection, the cells were harvested, washed with PBS and fixed with 70% ethanol for 4 h. The fixed cells were resuspended in PBS containing RNase A (10 mg/ml; BD Biosciences) and stained with PI in dark at room temperature for 30 min. Cell cycle distribution was detected and analyzed by FACScan flow cytometer.
Western blot analysis
Total protein was extracted from tissues/cells using radioimmunoprecipitation-assay buffer (Beyotime, Shanghai, China) and quantified using a BCA kit (Beyotime). Identical quantities of proteins were separated on SDS-polyacrylamide gel electrophoresis, and then transferred onto the polyvinylidene difluoride (PVDF) membranes (Millipore, Bedford, MA, USA). The membranes were blocked in 5% non-fat milk at room temperature for 2 h, and then incubated with specific primary antibodies against PDCD4 (1:1000; Abcam, Cambridge, MA, USA), Bax (1:1000; Abcam), Bcl-2 (1:1000; Abcam) and GAPDH (1:1000; Santa Cruz Biotechnology, Inc., Dallas, TX, USA) at 4 °C for one night. The membranes were then incubated with HRP-conjugated secondary antibody at room temperature for 2 h. The signals were visualized by the enhanced chemiluminescence detection kit (Millipore). GAPDH was used as the internal control.
Dual-luciferase reporter assay
Partial CASC2 sequences or PDCD4 3′-UTR fragments containing WT or MUT miR-21 binding site were amplified by PCR and inserted into psiCHECK-2 luciferase reporter vector (Promega, Madison, WI, USA). HEK293T cells were seeded in 24-well plates and co-transfected with the luciferase report vectors and miR-21 mimics or its negative control using Lipofectamine 2000 Transfection Reagent. After 48 h of transfection, cells were harvested, and the luciferase activities were measured by using the Dual-Luciferase Reporter Assay System (Promega).
Xenografts in nude mice
The male athymic BALB/c nude mice (4–5 weeks of age) were obtained from Shanghai SLAC Laboratory Animal, Co., Ltd. (Shanghai, China) and housed under SPF condition. Tca8113 cells (5×106 cells) were suspended in 150 μl PBS and then subcutaneously injected into the left side of the posterior flank of nude mice. The width (W) and length (L) of the tumors were measured using a caliper every 3 days, and the volume (V) was calculated using the formula: V =0.5× W2× L. After 21 days, the mice euthanized, and the tumors were excised and weighted. All experimental procedures involving the use of animals were approved by the Ethics Committees of Jinan Stomatological Hospital, and great efforts were given to reduce animal suffering, according to the NIH Guide for the Care and Use of Laboratory Animals.
Statistical analysis
All statistical analyses were performed using GraphPad Prism 6.0 software (GraphPad Software, San Diego, CA, USA) and SPSS 23.0 software (SPSS, Chicago, IL, USA). The differences between groups were analyzed using Student’s t-test or one-way ANOVA test. The associations between CASC2 expression and the clinicopathological characteristics of OSCC patients were examined using the χ2 test. A value of P<0.05 was considered to indicate a statistically significant difference.
Results
CASC2 is underexpressed in OSCC
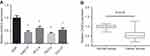
Firstly, RT-qPCR analysis was performed to detect the CASC2 expression in OSCC cell lines, and the result indicated that CASC2 was underexpressed in four OSCC cell lines (Tca8113, SCC-9, TSCCa, CAL-27) compared with that in NOK cells (Figure 1A). In addition, we found that the expression levels of CASC2 were also significantly decreased in human OSCC tissues compared with the adjacent non-tumor tissues (Figure 1B). The patients were then divided into two groups: CASC2 high expression group (n=30) and CASC2 low expression group (n=39), according to the median CASC2 expression in OSCC tissues. As shown in Table 1, the expression level of CASC2 in OSCC patients was significantly correlated with lymph node metastasis (P=0.024) and TNM stage (P=0.039).
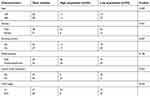 | Table 1 Correlation between the CASC2 expression and clinicopathological characteristics of OSCC patients |
CASC2 inhibits cell proliferation and induces cell apoptosis in OSCC
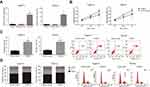
To explore the functions of CASC2 in OSCC tumorigenesis, CASC2 overexpression plasmid pcDNA3.1-CASC2 was transfected into Tca8113 and TSCCa cells, which expressed lowest CASC2 levels among four OSCC cell lines. 48 h after transfection, RT-qPCR analysis confirmed that CASC2 was overexpressed in Tca8113 and TSCCa cells (Figure 2A). MTT assay showed that overexpression of CASC2 notably inhibited the proliferative abilities of Tca8113 and TSCCa cells (Figure 2B). Next, we determined the effects of CASC2 on the cell cycle and apoptosis of OSCC cells by flow cytometric analysis. We found that CASC2 overexpression significantly increased the apoptosis levels of Tca8113 and TSCCa cells (Figure 2C). We also observed that CASC2 overexpression induced a cell cycle arrest at G0/G1 phase, and the cells distributed at S and G2/M phases in both Tca8113 and TSCCa cells were deceased (Figure 2D).
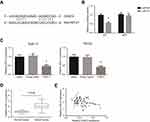
CASC2 serves as a ceRNA of miR-21 in OSCC
Through the online database (
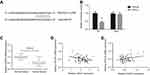
MiR-21 targets PDCD4 in OSCC
We also predicted the direct interaction between miR-21 and PDCD4 3′-UTR through TargetScan website (
MiR-21 blocks the tumor suppressive role of CASC2 in OSCC
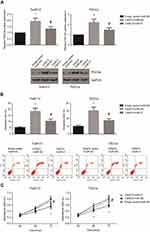
Next, we aimed to investigate whether miR-21/PDCD4 axis was partly implicated in the CASC2-induced OSCC apoptosis. We found that CASC2 overexpression significantly increased the PDCD4 protein expression levels in Tca8113 and TSCCa cells, and these effects were blocked by co-transfection with miR-21 mimics (Figure 5A). In addition, as demonstrated in Figure 5B, the apoptosis levels of CASC2-overexpressing Tca8113 and TSCCa cells were significantly decreased by miR-21 mimics. Besides, we also found that miR-21 mimics rescued the impaired proliferative abilities of CASC2-overexpressing Tca8113 and TSCCa cells (Figure 5C).
CASC2 inhibits OSCC tumor growth in vivo
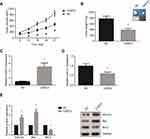
To further analyze the effect of CASC2 on OSCC tumor growth in vivo, we established xenograft tumors in nude mice using Tca8113 cells. The animals were closely monitored for three weeks, and we found that CASC2 overexpression notably suppressed the growth of OSCC xenografts in nude mice (Figure 6A). At the end of the assay, all of the tumors were excised and weighed, and we noticed that the average tumor weight of CASC2 group was significantly lower than NC group (Figure 6B). RT-qPCR and western blot analysis further validated that CASC2, PDCD4 and Bax expression levels were markedly increased, whereas miR-21 and Bcl-2 were underexpressed in the tumors of CASC2 group (Figure 6C-E).
Discussion
OSCC development is a multistep process driven by the accumulation of genetic and epigenetic changes.9 Among these risk factors, plentiful lncRNAs have been universally investigated and acknowledged as oncogenes or tumor suppressors in OSCC. For example, Lu et al reported that lncRNA C5orf66-AS1 may prevent OSCC progression by inhibiting cell growth and metastasis.10 Therefore, identification of OSCC-related lncRNAs may provide valuable therapeutic targets for OSCC patients.
In this study, we took lncRNA CASC2 as a study target. We found that CASC2 expression was significantly decreased in OSCC tissues compared with adjacent normal tissues and was negatively associated with adverse clinicopathological characteristics of OSCC patients. To further elucidate the biological function of CASC2 in OSCC, we performed gain-of-function experiments and showed that the proliferation and cell cycle progression of OSCC cells were inhibited, whereas OSCC cell apoptosis was enhanced by CASC2 overexpression. In vivo study also indicated that CASC2 overexpression restrained the volume and weight of OSCC xenografts in nude mice. All these findings suggested that CASC2 may have a tumor suppressive role in OSCC.
Among the working mechanisms of lncRNAs, competing endogenous RNA (ceRNA) theory hypothesizes that lncRNAs can serve as ceRNAs of miRNAs to prevent miRNAs from negatively regulating their target genes.11,12 We then focused on the role of CASC2 as a ceRNA by bioinformatics analysis, and found that miR-21, one of the most studied oncogenic miRNAs,13 contains the binding site of CASC2. In OSCC tissues, upregulation of miR-21 was inversely correlated with downregulation of CASC2, and rescue experiments further confirmed that miR-21 partly mediated the tumor suppressive role of CASC2 in OSCC. PDCD4 is a famous tumor suppressor protein involved in programmed cell death.14 Many study confirmed that PDCD4 is a target of miR-21, and the regulatory role of miR-21/PDCD4 axis has been well documented in OSCC.15,16 Our findings illustrated the potential role of CASC2 as a ceRNA of miR-21 to increase the expression of PDCD4 mRNA in OSCC.
In short, our study showed for the first time that CASC2 is downregulated in OSCC tissues and cell lines, and functional experiments confirmed that CASC2 inhibits the malignant phenotypes of OSCC cells through the regulation of miR-21/PDCD4 axis. Our findings suggested that CASC2 might potentially act as an effective therapeutic target for OSCC patients in the future.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Rivera C. Essentials of oral cancer. Int J Clin Exp Pathol. 2015;8(9):11884–11894.
2. Massano J, Regateiro FS, Januario G, Ferreira A. Oral squamous cell carcinoma: review of prognostic and predictive factors. Oral Surg Oral Med Oral Pathol Oral Radiol Endod. 2006;102(1):67–76. doi:10.1016/j.tripleo.2005.07.038
3. Schmitz SU, Grote P, Herrmann BG. Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci. 2016;73(13):2491–2509. doi:10.1007/s00018-016-2174-5
4. Qiu MT, Hu JW, Yin R, Xu L. Long noncoding RNA: an emerging paradigm of cancer research. Tumour Biol. 2013;34(2):613–620. doi:10.1007/s13277-013-0658-6
5. Pei Z, Du X, Song Y, et al. Down-regulation of lncRNA CASC2 promotes cell proliferation and metastasis of bladder cancer by activation of the Wnt/beta-catenin signaling pathway. Oncotarget. 2017;8(11):18145–18153. doi:10.18632/oncotarget.15210
6. Zhou T, Zhong M, Zhang S, et al. LncRNA CASC2 expression is down- regulated in papillary thyroid cancer and promotes cell invasion by affecting EMT pathway. Cancer Biomark. 2018;23(2):185–191. doi:10.3233/CBM-181198
7. Gao X, Du H, Zhang R, et al. Overexpression of cancer susceptibility candidate 2 inhibited progression of hepatocellular carcinoma cells. J Cell Physiol. 2018. doi:10.1002/jcp.27573
8. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta delta C(T)) method. Methods. 2001;25(4):402–408. Doi:10.1006/meth.2001.1262
9. Williams HK. Molecular pathogenesis of oral squamous carcinoma. Mol Pathol. 2000;53(4):165–172.
10. Lu T, Liu H, You G. Long non-coding RNA C5orf66-AS1 prevents oral squamous cell carcinoma through inhibiting cell growth and metastasis. Int J Mol Med. 2018;42(6):3291–3299. doi:10.3892/ijmm.2018.3913
11. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146(3):353–358. doi:10.1016/j.cell.2011.07.014
12. Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505(7483):344–352. doi:10.1038/nature12986
13. Pfeffer SR, Yang CH, Pfeffer LM. The role of miR-21 in cancer. Drug Dev Res. 2015;76(6):270–277. doi:10.1002/ddr.21257
14. Li JZ, Gao W, Ho WK, et al. The clinical association of programmed cell death protein 4 (PDCD4) with solid tumors and its prognostic significance: a meta-analysis. Chin J Cancer. 2016;35(1):95. doi:10.1186/s40880-016-0158-3
15. Reis PP, Tomenson M, Cervigne NK, et al. Programmed cell death 4 loss increases tumor cell invasion and is regulated by miR-21 in oral squamous cell carcinoma. Mol Cancer. 2010;9:238. doi:10.1186/1476-4598-9-254
16. Liu T, Chen G, Sun D, et al. Exosomes containing miR-21 transfer the characteristic of cisplatin resistance by targeting PTEN and PDCD4 in oral squamous cell carcinoma. Acta Biochim Biophys Sin (Shanghai). 2017;49(9):808–816. doi:10.1093/abbs/gmx078
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.