Back to Journals » Cancer Management and Research » Volume 13
LncRNA SBF2-AS1 Promotes Diffuse Large B-Cell Lymphoma Growth by Regulating FGFR2 via Sponging miR-494-3p
Received 30 September 2020
Accepted for publication 9 December 2020
Published 22 January 2021 Volume 2021:13 Pages 571—578
DOI https://doi.org/10.2147/CMAR.S284258
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Yong Teng
Dong-Wei Fu, Ai-Chun Liu
Department of Hematology, Harbin Medical University Cancer Hospital, Harbin, Heilongjiang 150081, People’s Republic of China
Correspondence: Ai-Chun Liu
Department of Hematology, Harbin Medical University, Cancer Hospital, Harbin, Heilongjiang 150081, People’s Republic of China
Email [email protected]
Purpose: Currently, there is no efficient and feasible method for diffuse large B-cell lymphoma (DLBCL) in clinical practice, and the main reason is the unclear pathogenesis of DLBCL, which leads to a high fatality rate of DLBCL.
Methods: Therefore, it is meaningful to explore the molecular mechanism of DLBCL and find a targeted therapeutic approach from the molecular level.
Results: Long non-coding RNA (lncRNA) SBF2-AS1 was highly expressed in DLBCL tissues and cell lines. Silencing of SBF2-AS1 inhibited the viability and growth of OCI-LY-3 cells. Furthermore, SBF2-AS1 acted as a sponge of miR-494-3p and inhibited its expression. And miR-494-3p directly targeted FGFR2. Functionally, forced expression of miR-494-3p or knockdown of FGFR2 removed the promoted effects of lncRNA SBF2-AS1 on DLBCL development. In vivo tumorigenesis experiments indicated SBF2-AS1 accelerated tumor growth via miR-494-3p/FGFR2 axis.
Conclusion: Our study revealed that SBF2-AS1 promoted the growth of DLBCL, which were mediated by miR-494-3p/FGFR2 axis.
Keywords: diffuse large B-cell lymphoma, SBF2-AS1, miR-494-3p, FGFR2, tumorigenesis
Introduction
Lymphoma is a malignant tumor originating from the lymphatic hematopoietic system.1 It mainly presents as painless lymphadenopathy, hepatosplenomegaly, and systemic symptoms such as fever and emaciation. Lymphoma includes two types: non-Hodgkin’s lymphoma (NHL) and Hodgkin’s lymphoma (HL).2 NHL is mainly divided into B cell, T cell and NK cell lymphomas.3 The incidence of diffuse large B-cell lymphoma (DLBCL) accounts for about 40%.4 Currently, the clinical treatment for DLBCL patients is limited. The main reason is the unclear pathogenesis of DLBCL, which leads to a high fatality rate of DLBCL.5 Therefore, it is of great clinical significance to explore the molecular mechanism of DLBCL and find a targeted therapeutic approach from the molecular level.
LncRNA is a class of RNA molecules with a length of more than 200 nucleotides. It was first discovered and reported by Nakajima et al in mice by large-scale sequencing in 2002.6 Because lncRNA has no clear open reading framework, it is not involved in or encodes very few proteins, making it considered to be the “noise” of gene transcription at the time of discovery. LncRNA underwent splicing, polyadenylation and 5ʹ capping, thus having a similar structure to mRNA.7 In recent years, lncRNA has been found to be abnormally expressed in various cancers, such as hepatocellular carcinoma (lncRNA-HEIH),8 breast adenocarcinoma (lncRNA-HOTAIR),9 and prostate cancer (PlncRNA-1).10 It has been proved that many lncRNAs took part in the tumor process and are expected to become tumor markers.11 The role of lncRNA in DLBCL has also been reported. Peng et al found that lncRNA LUNAR1 increased in DLBCL cell lines and tissues.12 The expression of transcription factor E2F1 and cyclin D1 was down-regulated and the expression of p21 was increased after the silencing of LUNAR1, thus inhibiting cell proliferation. Evidence has indicated that lncRNA SBF2-AS1 participated in multiple cancers, including gastric cancer,13 breast cancer,14 and pancreatic cancer.15 Nevertheless, whether SBF2-AS1 is involved in DLBCL is unclear.
Thus, the present study was designed to clarify SBF2-AS1 function in DLBCL and further explore underlying mechanisms.
Materials and Methods
Clinical Samples
The samples needed for the study were from the tissues of 50 DLBCL patients treated in Harbin Medical University Cancer Hospital. Tumor involvement was studied by taking tumor tissue and surrounding normal tissue, respectively. All the patients supported our research and signed informed consent and the Ethics Committee of Harbin Medical University Cancer Hospital.
Cell Culture and Transfection
The commonly used OCI-LY-3, OCI-LY-7, OCI-LY-10, SU-DHL-4 and SU-DHL −6 cell lines of DLBCL were purchased from ATCC cell center of the United States. The cells were cultured in DMEM medium with 10% fetal bovine serum and 1% dual-antibody solution in an incubator at 37°C and 5% CO2. The culture medium was changed every day and passed every 3 days. 500μM miRNA or 2 μg plasmid or 2 μg siRNA was transfected into cells, which was mediated by LipofectamineTM 2000 (Invitrogen, Carlsbad, CA, USA). And plasmid or miRNA or small interfering RNA (si-RNA) were constructed and purchased from by Ribobio company (Guangzhou, China).
RNA Isolation and qRT-PCR
We used trizol method to extract RNA, and RNA concentration and purity were determined using NanoDrop 2000 (Thermo Scientific, USA). RNA is used as transcription template to reverse transcribe into cDNA. Then SYBR Premix Ex TaqII was selected for RT PCR reaction. The expression value of the normal group was set as 1, and the relative expression of the experimental group was expressed as 2−ΔΔCT. GAPDH was used as internal control.
Protein Isolation and Western Blot
The tissue or treated cells were lysed with lysis buffer for 30 minutes. After centrifugation, the supernatant was separated and placed in a 0.5mL centrifuge tube. BCA method was used to determine the concentration of each sample, and protein loading treatment and quantitative protein samples were used. The samples were electrophoretic with polyacrylamide gel and then transferred to PVDF membrane. The PVDF membrane carrying protein was sealed with 1%BSA for 2 hours and incubated with primary antibody at 4°C overnight. The second antibody was incubated the next day, and the amount of protein samples on the PVDF membrane was detected by chemical radiography.
CCK8 Assay
100 μ L cell suspension was prepared in 96-well plate. The plates were pre-cultured in incubator for 24 hours (37°C, 5% CO2). 10 μ l different concentrations of substances were added to the culture plate. The plates were incubated in the incubator for 12 h. Add 10 μ l of CCK8 solution to each well (be careful not to generate bubbles in the hole, they will affect the OD reading). The plates were incubated in incubators for 1–4 hours. The absorbance at 450 nm was determined by enzyme-linked immunosorbent assay.
Edu Assay
Cells at logarithmic growth stage were taken and seeded in 96-well plates with 4×103~1×105 cells per well. The cells were cultured to the normal growth stage. Transfection was performed according to experimental requirements. An appropriate amount of 50 μM Edu culture medium (Ribobio, Guangzhou) was prepared by diluting Edu solution 1000:1. The cells were incubated at 20 μM Edu for 12 h. The cells were then fixed with 4% paraformaldehyde, stained with DAPI for 5 minutes, and cell proliferation was recorded by taking photos with a microscope.
Wound Healing Assay
The cells were cultured in a 6-well culture plate, and after 24 h of growth at a density, they should achieve a monolayer fusion of about 70–80%. Gently and slowly scrape a single layer with the tip of a new 200 μL pipette through the center of the hole. After scraping, wash the hole gently with medium twice to remove the isolated cells. The cells were cultured in a fresh serum-free medium. Cell migration was recorded at 0 h, 24 h and 48 hours after culture.
Transwell Assay
The matrix glue and basic medium 1640 were fully mixed according to 1:3. A mixture of 50 μL matrix glue and basal medium 1640 was added to the bottom of the chamber. The culture plates with small Chambers were placed in a 5% CO2 incubator for 30 minutes. Single-cell suspension was prepared and the cell concentration was adjusted to lx105/mL. In the 24-well plate, a small chamber with and without coated matrix glue was set, and a complete medium containing 10% serum was added. Cell suspension of 200 μL was slowly added into a small chamber and cultured at 37°C and 5% CO2 for 24 hours. The cells in the small chamber were removed with a wet cotton swab and fixed immediately with formaldehyde for 5 minutes. After that, the small chamber was taken out and dried. Crystal violet was used for dyeing for 20min. Then, the chamber was rinsed with water and dried. The number of transmembrane cells was observed and counted under the microscope.
Luciferase Assay
The HEK293 cells were cultured and inoculated in 24-well plates for 10–24 hours (80% confluence). Lipo200 was used to transfect WT-SBF2-AS1/Mut-SBF2-AS1 or WT-FGFR2/Mut-FGFR2 reporter gene plasmid into the cells. 72 hours after the Dual – Luciferase Reporter assay system detection to extract the protein. After adding miR-494-3p mimic or miR-NC, luciferase activity was detected again. Calculate the relative fluorescence intensity.
In vivo Tumor Growth Assay
SU-DHL-6 cells of each group were prepared for inoculation for subculture for 15 generations, and the concentration was adjusted to 5x107/0.1mL/site and then divided into different packs. The cell suspension is blown away. Eighteen BALB/C female nude mice (4–5 weeks, weight around 20g) were selected and grouped and numbered. Each nude mouse was weighed. The right armpit was disinfected with 75% alcohol, and 0.1mL cell suspension was injected at each site, about LCM was injected, to reduce leakage and contamination. After inoculation, the mice were observed daily. After 4 weeks, the nude mice were collected and killed by excessive carbon dioxide, the tumors were removed, and the tumors were photographed, weighed and recorded after all the surrounding connective tissue was removed. Tumor volume (mm3): V (Mm3) = S2 (Mm2) × L (Mm)/2. All animal experiments were performed in compliance with institutional guidelines, the National Institutes of Health guide for the care and use of laboratory animals (NIH Publications No. 8023, revised 1978), and had been approved by the Institutional Animal Care and Use Committee of Harbin Medical University Cancer Hospital. (Protocol Registry Number: A-2018-014).
Statistical Analysis
We used SPSS17.0 and Graphpad Prism 7.0 software for statistical analysis. The measurement data (measurement data) were expressed by mean ±standard deviation. t-test or variance analysis used to analyze the statistics of two groups. The Chi-square test was for counting data.
Results
Upregulation of lncRNA SBF2-AS1 in DLBCL Patients
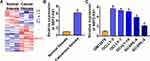
We collected 50 patients diagnosed DLBCL in our hospital, and microassay was used to determine the differentially expressed lncRNAs in DLBCL tissues comparing with normal adjacent tissues, which showed an upregulation of lncRNA SBF2-AS1 in DLBCL tissues (Figure 1A). Then, qRT-PCR analysis confirmed that lncRNA SBF2-AS1 was elevated in DLBCL cancer tissues comparing with normal tissues (Figure 1B). As well, SBF2-AS1 level was detected in DLBCL cell lines, which also showed an increase of SBF2-AS1 in DLBCL cell lines (OCI-LY-3, OCI-LY-7, OCI-LY-10, SU-DHL-4, SU-DHL-6) comparing with normal human B-lymphocytes GM12878 (Figure 1C) (Figure S1).
Deletion of SBF2-AS1 Suppressed DLBCL Cells Growth
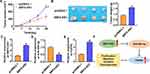
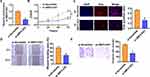
Because SBF2-AS1 had the highest expression in OCI-LY-3 cells according to Figure 1C, we then inhibited SBF2-AS1 expression in OCI-LY-3 cells using SBF2-AS1 siRNA, and si-SBF2-AS1 reduced SBF2-AS1 levels (Figure 2A). Next, we assessed si-SBF2-AS1 function. CCK-8 assay was performed to test cell viability, and data indicated that si-SBF2-AS1 inhibited OCI-LY-3 cell viability (Figure 2B). And Edu results showed that deletion of SBF2-AS1 reduced Edu positive cell numbers (Figure 2C). The wound-healing assay suggested that si-SBF2-AS1 suppressed cell migrate rate (Figure 2D). And transwell assay exhibited that si-SBF2-AS1 reduced invasive ability of OCI-LY-3 cells (Figure 2E).
SBF2-AS1 Suppressed miR-494-3p Expression
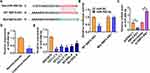
To explore the molecular mechanism of SBF2-AS1 in DLBCL, we used miRanda database and found a potential binding between SBF2-AS1 and miR-494-3p (Figure 3A). Then luciferase assay showed miR-494-3p inhibited activity of WT SBF2-AS1 not mut SBF2-AS1 in HEK293 cells (Figure 3B). And overexpression of SBF2-AS1 inhibited miR-494-3p level, while silencing of SBF2-AS1 promoted miR-494-3p level (Figure 3C). Furthermore, miR-494-3p decreased in DLBCL tissues and cells (Figure 3D and E).
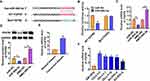
MiR-494-3p Directly Targeted FGFR2
Through Targetscan we found bases pairing of miR-494-3p and FGFR2 (Figure 4A). Followed luciferase analysis suggested miR-494-3p directly inhibited FGFR2 expression (Figure 4B). Furthermore, miR-494-3p suppressed FGFR2 mRNA and protein expression, but AMO-494-3p increased FGFR2 level in DLBCL cells (Figure 4C and D). What’s more, there was a dramatic enrichment of FGFR2 in DLBCL tissues and cells (Figure 4E and F).
SBF2-AS1 Promoted DLBCL Cells Growth by Modulating miR-494-3p/FGFR2
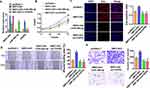
Because SBF2-AS1 had the lowest expression in SU-DHL-6 cells according to Figure 1C, we then forced expression of SBF2-AS1 with miR-494-3p mimics or si-FGFR2 in SU-DHL-6 cells (Figure 5A). SBF2-AS1 promoted cell proliferation, migration and invasion in SU-DHL-6 cells (Figure 5B–E). However, miR-494-3p or si-FGFR2 removed the promoting role of SBF2-AS1 on DLBCL cells (Figure 5B–E).
SBF2-AS1 Promoted in DLBCL Tumorigenesis
We implemented tumor formation in nude mice. SU-DHL-6 cells were subcutaneously injected into nude mice, and then, lentivirus packaged SBF2-AS1 was injected through tail vein. SBF2-AS1 enhanced tumor volume (Figure 6A), and increased tumor weight (Figure 6B). In addition, isolated tumor tissues had a higher SBF2-AS1 level after injection (Figure 6C). Moreover, injection of SBF2-AS1reduced the mRNA level of miR-494-3p, and induced FGFR2 expression (Figure 6D and E).
Discussion
DLBCL is an extremely complex and invasive disease.16 Although rituximab combined with CHOP regimen (R-CHOP) is effective in the treatment of DLBCL, a third of patients will have a recurrence/refractory condition.17 Due to its high heterogeneity, DLBCL can be divided into many subtypes, and different histological types have different responses to treatment.18 The present study firstly identified an increase of lncRNA SBF2-AS1 in DLBCL tissues. And functional experiments showed that silencing SBF2-AS1 inhibited DLBCL cell growth. Then, we found that SBF2-AS1 sponged miR-494-3p, and miR-494-3p targeted FGFR2 using database and luciferase assay. Moreover, SBF2-AS1 accelerated DLBCL cell growth and tumorigenesis via miR-494-3p/FGFR2 axis (Figure 6F).
In recent years, it has been reported that lncRNA is closely related to tumors, and acts as prognostic indicator or therapeutic target.19 A previous study has shown that lncRNA AGPG was upregulated in multiple cancers, which related to poor prognosis.20 In addition, lncRNA contributes to the development and activation of B cell.21 Therefore, from the perspective of lncRNA, it has become a research direction for researchers to find its influence on the prognosis of DLBCL patients.22 Interestingly, we found an obvious elevation of lncRNA SBF2-AS1 in DLBCL tissues and cell lines. And our finding suggested that SBF2-AS1 might be involved in DLBCL development, but the underlying mechanism needs to be further explored.
Considering abnormal expression is the basis of abnormal function, we speculated that SBF2-AS1 might contribute to the process of DLBCL. Then, we silenced SBF2-AS1 expression in OCI-LY-3 cells that had a higher SBF2-AS1 level. The results showed that si-SBF2-AS1 inhibited cell viability, proliferation, migration and invasion. And our data were similar with the previous study.14 These data revealed that SBF2-AS1 regulated the viability and growth of DLBCL cells, and SBF2-AS1 might be a biomarker and target of treatment of DLBCL patients.
Numerous reports proved that lncRNAs act as a sponge of miRNAs and inhibited their expression and activity. LncRNA UCA1 modulated miR-182 through ceRNA mechanism, and promoted renal cancer progression.23 And lncRNA HGBC facilitated gallbladder cancer development via sponging miR-502-3p.24 To illuminate the mechanism of SBF2-AS1 in DLBCL, we searched for microRNAs that could bind with SBF2-AS1. And SBF2-AS1 can be a sponge of miR-494-3p and inhibited its expression. miR-494-3p was decreased in DLBCL tissues and cells, which proved it might negatively correlate with DLBCL development.
FGFR2, a member of fibrosis growth factor receptor (FGFR) family, is involved in a wide range of malignant diseases and nominated as oncogenic drivers.25 It has been reported that degradation of FGFR2 benefited the treatment of cholangiocarcinoma.26 In the present study, we were surprised to find FGFR2 was o potential target of miR-494-3p, and FGFR2 was increased in DLBCL tissues and cells. Followed functional experiments suggested that overexpression of miR-494-3p or knockdown of FGFR2 removed the promoted effects of lncRNA SBF2-AS1 on DLBCL development. As well, in vivo tumorigenesis experiments indicated SBF2-AS1 accelerated tumor growth via miR-494-3p/FGFR2 axis.
Conclusion
In summary, our data revealed that SBF2-AS1 promoted the progression of DLBCL, which was mediated by miR-494-3p/FGFR2 axis. And this study might provide new understanding for DLBCL mechanism, and be helpful for DLBCL treatment.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Lang JY, Ma K, Guo JX, Sun H. Oxidative stress induces B lymphocyte DNA damage and apoptosis by upregulating p66shc. Eur Rev Med Pharmacol Sci. 2018;22(4):1051–1060. doi:10.26355/eurrev_201802_14388
2. Jabagi MJ, Goncalves A, Vey N, et al. Risk of hematologic malignant neoplasms after postoperative treatment of breast cancer. Cancers (Basel). 2019;11(10):1463. doi:10.3390/cancers11101463
3. Au KM, Wang AZ, Park SI. Pretargeted delivery of PI3K/mTOR small-molecule inhibitor-loaded nanoparticles for treatment of non-Hodgkin’s lymphoma. Sci Adv. 2020;6(14):eaaz9798. doi:10.1126/sciadv.aaz9798
4. Lamy T, Damaj G, Soubeyran P, et al. R-CHOP 14 with or without radiotherapy in nonbulky limited-stage diffuse large B-cell lymphoma. Blood. 2018;131(2):174–181. doi:10.1182/blood-2017-07-793984
5. Tsuyama N, Sakata S, Baba S, et al. BCL2 expression in DLBCL: reappraisal of immunohistochemistry with new criteria for therapeutic biomarker evaluation. Blood. 2017;130(4):489–500. doi:10.1182/blood-2016-12-759621
6. Nakajima D, Okazaki N, Yamakawa H, et al. Construction of expression-ready cDNA clones for KIAA genes: manual curation of 330 KIAA cDNA clones. DNA Res. 2002;9(3):99–106. doi:10.1093/dnares/9.3.99
7. Choudhari R, Sedano MJ, Harrison AL, et al. Long noncoding RNAs in cancer: from discovery to therapeutic targets. Adv Clin Chem. 2020;95:105–147.
8. Yang F, Zhang L, Huo XS, et al. Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology. 2011;54(5):1679–1689. doi:10.1002/hep.24563
9. Abba MC, Gong T, Lu Y, et al. A molecular portrait of high-grade ductal carcinoma in situ. Cancer Res. 2015;75(18):3980–3990. doi:10.1158/0008-5472.CAN-15-0506
10. Fang Z, Xu C, Li Y, et al. A feed-forward regulatory loop between androgen receptor and PlncRNA-1 promotes prostate cancer progression. Cancer Lett. 2016;374(1):62–74. doi:10.1016/j.canlet.2016.01.033
11. Santos JMO, Peixoto da Silva S, Gil da Costa RM, Medeiros R. The emerging role of microRNAs and other non-coding RNAs in cancer cachexia. Cancers (Basel). 2020;12(4):1004. doi:10.3390/cancers12041004
12. Peng W, Feng J. Long noncoding RNA LUNAR1 associates with cell proliferation and predicts a poor prognosis in diffuse large B-cell lymphoma. Biomed Pharmacother. 2016;77:65–71. doi:10.1016/j.biopha.2015.12.001
13. Liang C, Yue C, Liang C. The long non-coding RNA SBF2-AS1 exerts oncogenic functions in gastric cancer by targeting the miR-302b-3p/E2F transcription factor 3 axis [Retraction]. Onco Targets Ther. 2020;13:2523. doi:10.2147/OTT.S254587
14. Xia W, Liu Y, Cheng T, et al. Correction to: down-regulated lncRNA SBF2-AS1 inhibits tumorigenesis and progression of breast cancer by sponging microRNA-143 and repressing RRS1. J Exp Clin Cancer Res. 2020;39(1):60. doi:10.1186/s13046-020-01563-5
15. Hua YQ, Zhu YD, Xie GQ, et al. Long non-coding SBF2-AS1 acting as a competing endogenous RNA to sponge microRNA-142-3p to participate in gemcitabine resistance in pancreatic cancer via upregulating TWF1. Aging (Albany NY). 2019;11(20):8860–8878. doi:10.18632/aging.102307
16. Arzuaga-Mendez J, Prieto-Fernandez E, Lopez-Lopez E, et al. Cell-free DNA as a biomarker in diffuse large B-cell lymphoma: a systematic review. Crit Rev Oncol Hematol. 2019;139:7–15. doi:10.1016/j.critrevonc.2019.04.013
17. Lacy SE, Barrans SL, Beer PA, et al. Targeted sequencing in DLBCL, molecular subtypes, and outcomes: a haematological malignancy research network report. Blood. 2020;135(20):1759–1771. doi:10.1182/blood.2019003535
18. Opinto G, Vegliante MC, Negri A, et al. The tumor microenvironment of DLBCL in the computational era. Front Oncol. 2020;10:351. doi:10.3389/fonc.2020.00351
19. Huang Z, Zhou JK, Peng Y, et al. The role of long noncoding RNAs in hepatocellular carcinoma. Mol Cancer. 2020;19(1):77. doi:10.1186/s12943-020-01188-4
20. Liu J, Liu ZX, Wu QN, et al. Long noncoding RNA AGPG regulates PFKFB3-mediated tumor glycolytic reprogramming. Nat Commun. 2020;11(1):1507. doi:10.1038/s41467-020-15112-3
21. Brazao TF, Johnson JS, Muller J, et al. Long noncoding RNAs in B-cell development and activation. Blood. 2016;128(7):e10–19. doi:10.1182/blood-2015-11-680843
22. Deng L, Jiang L, Tseng KF, et al. Aberrant NEAT1_1 expression may be a predictive marker of poor prognosis in diffuse large B cell lymphoma. Cancer Biomark. 2018;23(2):157–164. doi:10.3233/CBM-160221
23. Wang W, Hu W, Wang Y, et al. Long non-coding RNA UCA1 promotes malignant phenotypes of renal cancer cells by modulating the miR-182-5p/DLL4 axis as a ceRNA. Mol Cancer. 2020;19(1):18. doi:10.1186/s12943-020-1132-x
24. Hu YP, Jin YP, Wu XS, et al. LncRNA-HGBC stabilized by HuR promotes gallbladder cancer progression by regulating miR-502-3p/SET/AKT axis. Mol Cancer. 2019;18(1):167. doi:10.1186/s12943-019-1097-9
25. Hilberg F, Tontsch-Grunt U, Baum A, et al. Triple angiokinase inhibitor nintedanib directly inhibits tumor cell growth and induces tumor shrinkage via blocking oncogenic receptor tyrosine kinases. J Pharmacol Exp Ther. 2018;364(3):494–503. doi:10.1124/jpet.117.244129
26. Lamberti D, Cristinziano G, Porru M, et al. HSP90 inhibition drives degradation of FGFR2 fusion proteins: implications for treatment of cholangiocarcinoma. Hepatology. 2019;69(1):131–142. doi:10.1002/hep.30127
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.