Back to Journals » International Journal of Nanomedicine » Volume 14
Improved smallest peptides based on positive charge increase of the γ-core motif from PνD1 and their mechanism of action against Candida species
Authors Mello ÉO , Taveira GB, de Oliveira Carvalho A , Gomes VM
Received 28 September 2018
Accepted for publication 15 December 2018
Published 9 January 2019 Volume 2019:14 Pages 407—420
DOI https://doi.org/10.2147/IJN.S187957
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Thomas Webster
Érica de Oliveira Mello, Gabriel Bonan Taveira, André de Oliveira Carvalho, Valdirene Moreira Gomes
Laboratório de Fisiologia e Bioquímica de Microrganismos, Centro de Biociências e Biotecnologia, Universidade Estadualdo Norte Fluminense Darcy Ribeiro, Campos dos Goytacazes, Rio de Janeiro, Brazil
Background: Plant defensins have a hallmark γ-core motif (GXCX3-9C) that is related to their antimicrobial properties. The aim of this work was to design synthetic peptides based on the region corresponding to the PνD1 defensin γ-core that are the smallest amino acid sequences that bear the strongest biological activity.
Methods: We made rational substitutions of negatively charged amino acid residues with positively charged ones, and the reduction in length in the selected PνD1 γ-core sequence to verify whether the increased net positive charges and shortened length are related to the increase in antifungal activity. Herein, we opted to evaluate the action mechanism of γ33-41PνD1++ peptide due to its significant inhibitory effect on tested yeasts. In addition, it is the smallest construct comprising only nine amino acid residues, giving it a better possibility to be a prototype for designing a new antifungal drug, with lower costs to the pharmaceutical industry while still maintaining the strongest antimicrobial properties.
Results: The γ33-41PνD1++ peptide caused the most toxic effects in the yeast Candida buinensis, leading to membrane permeabilization, viability loss, endogenous reactive oxygen species increase, the activation of metacaspase, and the loss of mitochondrial functionality, suggesting that this peptide triggers cell death via apoptosis.
Conclusion: We observed that the antifungal activity of PνD1 is not strictly localized in the structural domain, which comprises the γ-core region and that the increase in the net positive charge is directly related to the increase in antifungal activity.
Keywords: defensin, antimicrobial peptide, cationic, mechanism of action
Introduction
Plant defensins are peptides identified in different plant tissues and organs, and several studies demonstrated the important role they played in the innate immune system of plants.1,2 The participation of plant defensins in the defense responses is supported by reports of their strong antimicrobial activity in vitro, not only against plant pathogenic fungi, but also against human ones;3,4 the gene expression pattern in response to pathogen infection,5 along with the demonstration that transformed plants engineering to constitutively express defensin genes, is resistant to fungal diseases.6
Plant defensins present a diversity of primary structures coded in 45–54 amino acid residues. The amino acid composition of plant defensins make them highly basic, giving the molecules a net positive charge at neutral pH. Several amino acid residues are conserved among the plant defensin primary structures, such as the strictly conserved eight cysteine residues.7,8 The eight cysteine residues bond to each other in specific pairs, forming four disulfide bridges, which are responsible for the stabilization of their three-dimensional structure,7,8 allowing this family of plant peptides to resist harsh environmental conditions, such as protease degradation and pH and temperature extremes.9 Despite the variation in the primary structure, a completely opposite effect is observed at the tertiary structure level, which is well defined and conserved, generally consisting of one α-helix and three antiparallel β sheets.10 The diversity of plant defensins at their primary structure level is responsible for their different biological activities in vitro.7,11,12 Among the activities already described, the best characterized is their growth inhibition of a large variety of filamentous fungi and yeasts, including those pathogenic to humans.3,7,13–17
The mechanism of action involved in the fungal growth inhibition by plant defensins has been investigated by several authors, and the evidence demonstrates several steps involving an extracellular mechanism acting on the cell wall and/or plasma membrane, further acting on intracellular targets.7,9,15,18 In addition to this whole membrane permeabilization process involving plant defensins, some defensins have been shown to induce apoptosis or programmed cell death in susceptible yeast and fungal species.16,17
In regard to the plant defensin structures, studies have demonstrated that two disulfide bridges, those formed between the Cys21 and Cys25 located in the α-helix and the Cys45 and Cys47 located in the last β-sheet, form a structural arrangement named the cysteine-stabilized αβ motif (CSαβ motif), characteristic of peptides with antimicrobial activity.7,19,20 In addition to the CSαβ motif, plant defensins also have another conserved framework region characterized, called the γ-core, which has the conserved sequence (GXCX3-9C, where X may be any amino acid residue).1,21 Some studies have shown that the region responsible for the biological activity of plant defensins resides in the γ-core region. De Samblanx et al22 showed that the substitution or removal of the amino acid residues located in this region, which encompasses β2 and β3 sheets, reduced the biological activity of RsAFP2 (a defensin from Raphanus sativus). Sagaram et al23 demonstrated that the main determinants of the antifungal activity of MsDef1 and MtDef4 (defensins from Medicago sativa and Medicago truncatula, respectively) reside in their γ-core motifs. In another study, it was shown that the interaction of Psd1 defensin from Pisum sativum with the membrane is mediated, in part, by the amino acid residues that compose the γ-core region.1
Knowing that the region comprising the plant defensin γ-core is related to the antimicrobial properties of these peptides, the principal aim of this work was to design synthetic peptides based on the region corresponding to the PνD1 defensin γ-core that are the smallest amino acid sequences that bear the strongest biological activity. PνD1 is an isolated defensin from Phaseolus vulgaris seeds with antifungal properties,14,15,18 anti-Leishmania activity,24 and anticancer activity.25 Additionally, knowing that the charge, length, and hydrophobicity influence the activity of antimicrobial peptides,26 we made rational substitutions of negatively charged amino acid residues with positively charged ones, as well as the reduction in length in the selected PνD1 defensin γ-core sequence to verify whether the increased net positive charge and the shortened length are related to the increase in antifungal activity. In addition, some tests were carried out to better understand the mechanism of action of the peptides derived from the γ-core region of PνD1 defensin.
Materials and methods
Microorganisms
The yeast species Candida albicans (CE022) and Candida buinensis (3982) were maintained in Sabouraud agar (1% peptone, 2% glucose, and 1.7% agar; Merck). The yeast cells were provided by Laboratório de Fisiologia e Bioquímica de Microrganismos, Centro de Biociências e Biotecnologia, Universidade Estadualdo Norte Fluminense Darcy Ribeiro, Campos dos Goytacazes, Rio de Janeiro, Brazil.
Analysis of amino acid sequence of PνD1 defensin
The amino acid sequence of the PνD1 defensin from P. vulgaris was obtained as previously described by Games et al.14 In brief, the first 21 amino acid residues of PνD1 were obtained by Edman degradation of the purified peptide. Based on this 21 amino acid sequence, a degenerated primer was designed and used in an RT-PCR together with an oligo-dT primer. The amplicon was cloned into the pTZ57R/T vector (InsTAclone™ PCR Cloning Kit, Fermentas) and then submitted to sequencing by Big Dye1 Terminator v3.1 Kit (Applied Biosystems) in an ABI 3110 Genetic Analyzer (Applied Biosystems).
Synthetic peptide design
By comparison analysis of the primary structure of PνD1 defensin with other defensin primary structures7 and by superimposing onto these sequences the sequence of the γ-core, as defined by Yount and Yeaman,27 we identified and selected the region corresponding to the PνD1 γ-core. This region encompasses the γ-core itself and amino acid residues that are part of the flanking β2 and β3 sheets. Based on these comparative analyses, two peptides with 15 amino acid residues and two peptides with 9 amino acid residues each were then chemically synthesized by the GenOne Soluções em Biotecnologia (Rio de Janeiro, Brazil). The purity level of the synthetic peptides was >95% as determined by high-performance liquid chromatography (4.6×250 mm PLRP-S 100A), using a solution A (0.1% trifluoroacetic acid in 100% acetonitrile) and a solution B (0.1% trifluoroacetic acid in 100% water) with the following gradient: 0%–15% of solution A from 0 to 25 minutes; 40%–100% of solution A from 25 to 30 minutes. The Cys (C) residues of these peptides were replaced by Ala (A) residues. Additionally, two Asp (D) residues were substituted by two Arg (R) residues in two peptides. All these peptides were resuspended in ultrapure water. Their molecular masses and isoelectric points were determined by the Expasy Compute pI/Mw tool.28–30
Effect of synthetic peptides on yeast growth
Yeast cell inocula from C. albicans and C. buinensis were removed from stock tubes containing Sabouraud agar and transferred to Petri dishes containing Sabouraud agar. Cells were grown at 30°C for 2 days. After this period, each cell aliquot was added to 10 mL sterile culture medium (Sabouraud broth, 1% peptone, 2% glucose, Merck). Yeast cells were quantified in a Neubauer chamber (LaborOptik). Initially, the C. albicans and C. buinensis yeast cells (1×104 cells mL−1) were incubated in 100 μL of Sabouraud broth containing the selected synthetic peptides at increasing concentrations from 18.35 to 293.6 μM.
The assay was performed on cell culture microplates (96 wells; Nunc) at 30°C for 24 hours. Optical readings at 620 nm (EZ Read 400, Biochrom) were collected 24 hours after the start of assay. Control cells were grown in the absence of synthetic peptides. The assay was done three times according to a methodology adapted from Broekaert et al.31 The yeast growth inhibition data were evaluated by the one-way ANOVA, and the differences of the mean at P<0.05 were considered significant. All statistical analyses were performed using the GraphPad Prism software (version 6.0 for Windows).
Cell viability
To verify whether the growth inhibition of C. buinensis cells was caused by the fungicidal or fungistatic effect of the smallest active peptide, the control cells (without the peptide) were washed and diluted 1,000-fold. A 100 μL aliquot of the dilution was spread with a Drigalski spatula on the surface of a Petri dish containing Sabouraud agar and grown at 30°C for 48 hours. At the end of this period, the colony forming units were determined, and Petri dishes were photographed.32 The same procedure was repeated with the yeasts treated with 36.7 and 73.4 μM of the smallest and the strongest active peptide for 24 hours. The experiments were performed in triplicate, and the results are shown assuming that the control represents 100% cell viability. These data were evaluated by one-way ANOVA, and the differences of the mean at P<0.05 were considered significant. All statistical analyses were performed using the GraphPad Prism software (version 6.0 for Windows).
Effect of a synthetic peptide on plasma membrane permeabilization
The plasma membrane permeabilization of C. buinensis cells treated with 25 μM of the smallest and the strongest active peptide was evaluated using Sytox green fluorescent dye (Invitrogen), according to the methodology described by Thevissen et al.33 Sytox green is a dye that has high affinity for nucleic acids and penetrates the cell only when its membrane is compromised, emitting a strong green fluorescent.
After the growth inhibition assay, C. buinensis yeast cells grown in the absence (control) and in the presence of the smallest and the strongest active peptide were incubated with Sytox green at a final concentration of 0.2 μM (dissolved in dimethyl sulfoxide), according to instructions provided by the manufacturer. A positive control with 300 mM ethanol was done. After 15 minutes incubation at 25°C with constant agitation at 500 rpm, the cells were observed under an optical microscope (Axioplan.A2, Zeiss) coupled to an AxioCAM MRc5 (Zeiss) camera and the images were analyzed by the Axiovision software version 4.0 (Zeiss). The microscope was equipped with a set of fluorescent filters for fluorescein detection (excitation wavelength between 450 and 490 nm and emission of 500 nm). The results represent triplicate experiments.
Determining the induction of intracellular oxygen reactive species
To evaluate whether the mechanism of action of the smallest and the strongest active peptide involves the induction of oxidative stress, the fluorescent probe 2′,7′-dichlorofluorescein diacetate (H2DCFDA) was used according to the methodology described by Mello et al.15 This dye enters the cell passively, is deacetylated by intracellular esterases and after being oxidized by reactive oxygen species (ROS), becomes fluorescent. After 24 hours incubation with 25 μM of the smallest and the strongest active peptide, 50 μL of C. buinensis cells grown in the absence (control) and in the presence of the peptide were incubated with 20 μM of the H2DCFDA probe (Calbiochem), for 2 hours at 25°C with constant agitation at 500 rpm. A positive control was done with 300 mM hydrogen peroxide. After this period, the cells were observed under an optical microscope (Axioplan. A2, Zeiss) coupled to an AxioCAM MRc5 (Zeiss) camera, and the images were analyzed by the Axiovision software version 4.0 (Zeiss). The microscope was equipped with a set of fluorescent filters for fluorescein detection (excitation wavelength between 450 and 490 nm and emission of 500 nm). The results represent triplicate experiments.
Analysis of mitochondrial functionality
Mitochondrial functionality was assessed by fluorescent dye Rhodamine 123 (Sigma). Rhodamine 123 is a cationic fluorescent dye that has a high affinity to the electrical potential of membranes; thus, it marks active mitochondria in living cells resulting in a bright red fluorescence. In contrast, the treated samples present a weak or absent fluorescent signal. This test was done as described in the section “Effect of synthetic peptides on yeast growth” with the following differences: after incubation with 25 μM of the smallest and the strongest active peptide for 12 and 24 hours, the C. buinensis yeast cells were resuspended, incubated with 10 μg mL−1 of Rhodamine 123 with constant agitation at 500 rpm for 2 hours and protected from light and then analyzed by differential interference contrast (DIC) under an optical microscope equipped with a fluorescence filter (with an excitation wavelength of 506 nm and emission wavelength of 530 nm). Control cells (in the absence of the peptide) had the same treatment as cells treated with the peptide and a positive control with cells treated with 300 mM ethanol for 30 minutes before the dye addition.
Detection of caspase activity induced by a synthetic active peptide
The detection of caspase activity was performed using the CaspACE FITC-VAD-FMK marker (Promega), as described by the manufacturer. The FITC-VAD-FMK marker is an analog of the Z-VAD-FMK caspase inhibitor (carbobenzoxy-valyl-alanyl-aspartyl-[O-methyl]-fluoromethylketone) that enters the cell and irreversibly binds to activated caspases. This test was done as described in the section “Effect of synthetic peptides on yeast growth” with the following differences: after incubation with 25 μM of the smallest and the strongest active peptide for 24 hours, the C. buinensis cells were resuspended, washed once in 500 μL PBS (10 mM NaH2PO4, 0.15 M NaCl) with pH 7.4 and resuspended in 50 μL of the staining solution (supplied by the kit) containing 50 μM FITC-VAD-FMK marker. After incubation for 20 minutes at 30°C with constant agitation at 500 rpm, the cells were again washed in 500 μL PBS and resuspended in 20 μL PBS. Negative control (in the absence of the peptide) and positive control cells (incubated with 300 mM acetic acid) had the same treatment as cells treated with the peptide. The cells were analyzed by DIC on the optical microscope equipped with a fluorescence filter for fluorescein detection (excitation wavelength 450–490 nm and emission wavelength 500 nm).
Results
Synthetic peptide design
After the analysis of the primary structure of PνD1, previously obtained by Games et al,14 and other defensin peptides,7 the region corresponding to the PνD1 γ-core was selected and is highlighted in blue (Figure 1A). Based on the PνD1 γ-core sequence, together with parts of the β2 and β3 sheets (from Arg31 to Lys45), an amino acid stretch giving rise to the peptide of 15 residues with the following amino acid sequence was chosen: RSGRARDDFRAWATK, which was called γ31-45PνD1 (Figure 1A). Another peptide, based on the γ31-45PνD1 design, had its net positive charge increased with the replacement of the two Asp residues, at positions 37 and 38, by two Arg residues, giving rise to the following amino acid sequence: RSGRARRRFRAWATK, which was called γ31-45PνD1++ (Figure 1A). Two other peptides of nine residues each, smaller than γ31-45PνD1 and γ31-45PνD1++, and had the sequences comprising Gly33 to Cys41 (PνD1 γ-core itself) with the amino acid sequences GRARDDFRA and GRARRRFRA, which were called γ33-41PνD1 and γ33-41PνD1++, respectively, were also synthesized (Figure 1A). All the synthesized peptides had their Cys residues replaced by Ala residues (Figure 1A). These substitutions were made to prevent the formation of disulfide bridges. Similarly, Schaaper et al34 replaced the Cys residues by α-aminobutyric acid, to avoid the formation of such undesired bridges. The biochemical characteristics of synthesized peptides are shown in Figure 1B.
Growth inhibition assay
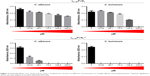
To analyze the effect of the synthetic peptides on the growth of yeasts, C. albicans and C. buinensis, γ31-45PνD1, γ31-45PνD1++, γ33-41PνD1, and γ33-41PνD1++ peptides were used at increasing concentrations from 18.35 to 293.6 μM (Figures 2 and 3).
With the γ31-45PνD1 peptide, we observed that all used concentrations inhibited the growth of yeast C. albicans, reaching 40% inhibition at the concentration of 293.6 μM. The yeast C. buinensis had its growth inhibited starting from the 73.4 μM concentration, and it was possible to observe the 100% inhibition, when the peptide concentration was 293.6 μM (Figure 2). When the positive net charge of this peptide was increased by the synthesis of γ31-45PνD1++, the peptide concentration of 18.35 μM inhibited 60% growth of the yeast C. albicans, and starting from the concentration of 73.4 μM, the yeast growth was completely inhibited (Figure 2). For yeast C. buinensis, the lowest used peptide concentration of 18.35 μM completely inhibited yeast growth (Figure 2).
When we used the smallest peptides, we observed that γ33-41PνD1 was not able to inhibit the growth of C. albicans at any concentrations tested (Figure 3). For C. buinensis, this peptide inhibits growth by 15% and 17% at the two highest used concentrations, respectively (Figure 3). However, the peptide γ33-41PνD1++ that had its positive net charge increased showed a greater inhibition potential, being able to inhibit the growth of C. albicans at all concentrations tested and reaching 63% inhibition at the concentration of 293.6 μM (Figure 3). For C. buinensis, this peptide had an even greater inhibitory potential, being able to inhibit 100% of its growth starting from the concentration of 36.7 μM (Figure 3).
Based on the results from Figures 2 and 3, the designed smallest peptide that had the strongest antimicrobial activity was γ33-41PνD1++ when incubated with C. buinensis. Because of this outcome, this peptide and this yeast species were chosen for the next experiments.
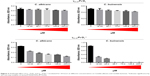
Cell viability
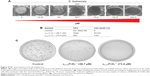
Based on the results obtained in the growth inhibition assay, we found that the minimal inhibitory concentration (MIC) of γ33-41PνD1++ peptide was 36.7 μM for C. buinensis cells (Figure 4A). The γ33-41PνD1++ peptide at 36.7 and 73.4 μM induced the viability loss in C. buinensis cells. The cells treated with 36.7 μM of this peptide had an 84% loss of viability and the cells treated with 73.4 μM had a 100% loss of viability, showing that the effect of this peptide is fungicidal to C. buinensis (Figure 4B and C). Starting from these experiments, all following assays were performed with yeast C. buinensis with the γ33-41PνD1++ at 25 μM, because at this concentration, below the determined MIC, we were able to obtain reasonable amounts of cells for visualization by optical microscopy.
Effect of γ33-41PνD1++ on plasma membrane permeabilization
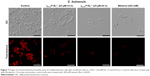
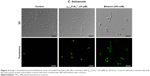
The ability of the γ33-41PνD1++ to permeabilize the C. buinensis plasma membrane was analyzed using Sytox green. The analysis by fluorescence microscopy revealed that C. buinensis cells were labeled by the dye, when treated with γ33-41PνD1++ at 25 μM; thus, these data suggest that γ33-41PνD1++ acts on the plasma membrane of C. buinensis, structurally compromising it and allowing its permeabilization for the labeling dye. In the control (in the absence of peptide), no fluorescence was observed and in the positive control (300 mM ethanol), fluorescent cells were observed (Figure 5).
Reactive oxygen species assay detection
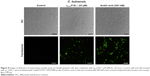
After the growth inhibition assay, C. buinensis cells were incubated with the H2DCFDA probe for the detection of endogenous ROS production. When incubated in the presence of 25 μM of γ33-41PνD1++, the cells showed ROS labeling (Figure 6), suggesting that a γ33-41PνD1++-induced increase in oxidative stress may underlie the growth inhibitory effect on this yeast. In the control (in the absence of peptide), no fluorescence was observed and in the positive control (300 mM hydrogen peroxide), fluorescent cells were observed (Figure 6).
Analysis of mitochondrial functionality
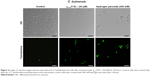
In C. buinensis cells treated with γ33−41PνD1++ at 25 μM for 12 hours is observed a diminished mitochondrial activity as indicated by the weaker red fluorescent signal of Rhodamine 123 (Figure 7) in comparison to the untreated cells, and after additional 24 hours in the presence of γ33−41PνD1++ it is also observed a shrinkage and condensation of cell cytoplasm. A similar weak fluorescent signal was observed for the 300 mM ethanol-treated samples (positive control). Control cells with functional mitochondria evinced a strong signal of Rhodamine 123 fluorescence (Figure 7). This result indicated that after treatment with γ33-41PνD1++, the C. buinensis cells lost their electrical mitochondrial membrane potential, causing the dysfunction of their mitochondria.
Detection of caspase activity induced by γ33-41PνD1++
The detection of caspase activity was performed using the CaspACE FITC-VAD-FMK in situ marker. The C. buinensis yeast cells grown in the absence (control) and in the presence of 25 μM of γ33-41PνD1++ were subsequently incubated with the FITC-VAD-FMK marker. It can be observed that the cell treatment with γ33-41PνD1++ resulted in the activation of metacaspases, making it possible to observe a strong labeling of these cells. Positive control cells, treated with 300 mM acetic acid, a known apoptosis inducer in yeast,35 also showed labeling. However, the untreated cells (negative control) did not show metacaspase activity (Figure 8). These data suggest a programmed cell death that occurs by an apoptotic pathway.
Discussion
Plant defensins are peptides whose activity and mechanisms of action are primarily described against a wide variety of filamentous fungi and yeasts; however, their mechanism of action has not yet been fully elucidated.3,7 The defensins have a structure composed of an α-helix and two β-sheets connected by four disulfide bridges. Yount and Yeaman27 showed that the peptides stabilized by disulfide bridges possess amino acid residues that are important for antimicrobial activity in the γ-core region that encompasses β2 to β3 sheets. Many studies have already shown that the activity of plant defensins is related to this structural domain.22,23,36
In the previous work performed by our group, the PνD1defensin isolated from P. vulgaris seeds inhibited growth of various yeasts.14,15,18 In this study, we designed synthetic peptides based on the PνD1 γ-core. Initially, we designed two peptides of 15 residues that comprised the region from Arg31 to Lys45, called γ31-45PνD1 and γ31-45PνD1++, the latter having its positive net charge increased (Figure 1). Some studies have identified charge, length, and hydrophobicity as the parameters that influence the activity of antimicrobial peptides.26,27 Regarding the length, it has already been reported that larger peptides that cover the entire β2 and β3 sheets, where the γ-core region is inserted, have higher activity.34 Sagaram et al23,37also demonstrated that synthesis of peptides extending beyond the γ-core region improves antifungal activity and that the synthetic peptide (GMA-4C) derived from the γ-core region also contained the C-terminus of MtDef4 defensin with cationic and hydrophobic amino acids that were important for its antifungal activity. Regarding the charge, Lacerda et al10 demonstrated that the positively charged amino acids located in the γ-core were essential for the antifungal activity of peptides, since the substitution of neutral residues by positively charged amino acid residues in the γ-core region increased their inhibitory activity against pathogenic fungi. The mechanism behind this interconnection between positive charge increasing and antimicrobial inhibition increasing is based mainly in the opposite charge attraction.38 Fungal cells have negatively charged structures in their cell walls, such as phosphomanolipids,39 that may serve as primary anchor sites for the positively charged peptides. Then the peptides became able to interact with the membrane. The interaction with the membrane is possible because of the amphipathic character of antimicrobial peptides which turning them able to interact with the negative charge of phospholipids hear groups and also with the hydrophobic core of the membrane.11,33 This interaction explains the ability of antimicrobial peptide to cause membrane permeabilization. Similarly, the His and Arg residues present in the γ-core were important for the oligomerization and antifungal activity of the peptide derived from MtDef5 defensin against the filamentous fungi Neurospora crassa and Fusarium graminearum.36 Thus, we designed the peptide γ31-45PνD1, which had additional amino acids of the flanking β2 and β3 sheets, beside the amino acids that compose the γ-core and increased the charge of γ31-45PνD1++ by replacing two Asp37-38 by two Arg. When we tested the antifungal activity of γ31-45PνD1, it inhibited the growth of C. albicans at all concentrations tested. For C. buinensis, we observed inhibition only starting from the concentration of 73.4 μM, with MIC being 293.6 μM concentration. When we increased the net positive charge of γ31-45PνD1++, it exhibited higher antifungal activity. All tested concentrations of this peptide inhibited the C. albicans growth, with MIC being 73.4 μM. For yeast C. buinensis, MIC was determined at the lowest concentration used, at 18.35 μM (Figure 2).
Although some previous works have shown that the peptide length influenced its antifungal activity, if a smaller molecule obtained from the native sequence of a peptide still had biological activity, it would be more commercially interesting due to the low cost of its synthesis and could be seen as a promising molecule for the control of fungal diseases.34 To find an even smaller molecule with antifungal potential that could be used as a novel drug, we designed two peptides of nine amino acid residues each that comprised the γ-core region from Gly33 to Cys41, called γ33-41PνD1 and γ33-41PνD1++, where the latter also had its positive net charge increased as discussed above (Figure 1). Our choice of the smallest sequence with the strongest biological activity is based on requirements of desired characteristics for the pharmaceutical industry to alleviate problems like toxicity, drug stability, specificity, and less immunogenicity.40 For these reasons, we chose to work with the γ33-41PνD1++ peptide.
When we tested the activity of these peptides, the γ33-41PνD1 peptide at different concentrations did not inhibit the growth of C. albicans cells, whereas for yeast C. buinensis, it showed low activity at high concentrations (Figure 3). These results are consistent with the works of Schaaper et al34 and Sagaram et al37 in the sense that the smallest peptides, only encompassing the γ-core, as in our study, did not bear strong biological activity, at least for C. albicans and C. buinensis. Additionally, as in another example of Sagaram et al,23 the γ-core alone may or may not present biological activity similar to the entire defensin. These authors tested the activity of two synthetic peptides called GMA1 and GMA4 derived, respectively, from the γ-core of the defensins MsDef1 and MtDef4, against the filamentous fungus F. graminearum. The peptide GMA1, even at the concentration of 96 μM, had no antifungal activity. In contrast, the peptide GMA4 exhibited antifungal activity at 6 μM, and when used at 12 μM, was able to completely inhibit the conidial germination of the tested fungus. These results make it evident that only the γ-core region of the MtDef4 defensin is sufficient for antifungal activity; however, the γ-core of the MsDef1 defensin by itself is not sufficient for antifungal activity, similar to the γ-core (γ33-41PνD1) of the PνD1 defensin.
On the other hand, when we tested the antifungal activity of γ33-41PνD1++, with the positive net charge increased by replacing two Asp37-38 residues by two Arg residues, we observed that it inhibited the growth of C. albicans and C. buinensis, with the MIC for C. buinensis of 36.7 μM. Previous studies showed that higher the net positive charge, the higher will be the antifungal activity of peptides, and the positively charged amino acids, such as Arg and Lys, were very important for antifungal activity. The synthetic peptide MBGo1 derived from RsAFP2 defensin, which had its net positive charge increased due to the substitution of some amino acid residues by Arg residues, showed higher activity against the filamentous fungus Fusarium culmorum.34 Sagaram et al23 showed that MtDef4 defensin had five basic amino acids in its γ-core, and MsDef1 defensin had two basic amino acids and two acidic amino acids. After an exchange of the γ-core region between these two defensins, MsDef1 had a two fold higher antifungal activity, thus showing the importance of positively charged amino acid residues for the higher antifungal activity of this peptide.
In the growth inhibition assays, γ33-41PνD1++ peptide showed higher activity against C. buinensis cells, and this inhibitory effect on yeast growth was fungicidal (Figure 4). Taveira et al41,42 found that another cationic peptide isolated from Capsicum annuum fruits, belonging to thionin family, called CaThi, had a fungicidal effect against six species of Candida genus and Fusarium solani. It is worth emphasizing that the fungicidal characteristic is very important for the development of new therapeutic drugs, since substances that present fungistatic effect can contribute to the development of resistant microorganisms.43,44
Considering the results obtained, we began to investigate the mechanism of action of γ33-41PνD1++, responsible for the inhibition of C. buinensis growth. One characteristic of the cationic antimicrobial peptides is their ability to permeabilize the plasma membrane of microorganisms.15,36 Initially, we used Sytox green, a dye that has high affinity for nucleic acids and penetrates cells only when their plasma membrane is compromised. We observed that the C. buinensis cells, when treated with the γ33-41PνD1++ peptide, were marked by the dye (Figure 5). Sagaram et al23 showed that MsDef1γ4 peptide was also able to permeabilize the membrane; however, the tested fungus was F. graminearum. In another work, Islam et al36 found that the cationic amino acid residues, present in the γ-core of MtDef5 defensin, were responsible for its antifungal activity and that at micromolar concentrations, MtDef5 caused the membrane permeability of the filamentous fungi F. graminearum and N. crassa. These same authors also observed that there was an increase in the production of intracellular ROS in F. graminearum and N. crassa hyphae after these cells were treated with MtDef5.
ROS have been considered as primary regulators of cell death and are linked to many crucial apoptotic pathways in yeasts.45 Studies have shown that an increase of ROS in the medium can be toxic to the organisms, leading to the destruction of several cell types through apoptotic pathways.1,17 Previously, our research group showed that PνD1 induced ROS in C. albicans and Fusarium oxysporum cells.15 In this study, we observed that C. buinensis cells were labeled by the 2′,7′-dichlorofluorescein diacetate probe when they were treated with γ33-41PνD1++, thus indicating that it caused an increase in endogenous ROS production in these cells (Figure 6). Due to the increased ROS production in the presence of γ33-41PνD1++, our next objective was to verify if the apoptotic process could be taking place when the C. buinensis cells were treated with γ33-41PνD1++. For this process, we verified the presence of active metacaspases46 and the dissipation of mitochondrial membrane potential in these cells. Caspases play a central role in signaling death by apoptosis. These are described as specific cysteine-containing aspartate proteases that are among the key markers of the apoptotic pathway, including yeasts.46,47 In these two assays, it was possible to observe that γ33-41PνD1++ activated the metacaspases in C. buinensis cells and caused a collapse of mitochondrial membrane potential in these cells (Figures 7 and 8). Taveira et al42 found that CaThi caused an activation of metacaspase in F. solani, indicating that programmed cell death could be triggered by CaThi in this fungus. In another study, Soares et al17 showed that ApDef1 defensin, isolated from the seeds of Adenanthera pavonina, caused an increase in the ROS production and accumulation that led to permeabilization of the plasma membrane and, consequently, the death of Saccharomyces cerevisiae cells through a metacaspase-dependent apoptotic process. It has also been observed by Vieira et al48 that C. albicans cells lost mitochondrial functionality when treated with the Lp-Def1 defensin, isolated from Lecythis pisonis seeds.
Conclusion
In conclusion, taken together, our results suggest that the antifungal activity of PνD1 is not strictly localized in the structural domain, which comprises the γ-core region; however, we can state that the addition of amino acid residues beyond the γ-core region, which comprises parts of the β2 and β3 sheets, is important for antifungal activity. Additionally, we observed that the increase in the net positive charge is directly related to the increase in antifungal activity. In this work, we opted to evaluate the mechanism of action of the γ33-41PνD1++ peptide due to its significant inhibitory effect on tested yeast. In addition, γ33-41PνD1++ is the smallest construct comprising only nine amino acid residues, which gives it a better possibility to be the basis for a design of a new antifungal drug, with lower costs to the pharmaceutical industry, but still maintaining its antimicrobial properties. In relation to the antifungal activity of γ33-41PνD1++, it was fungicidal, and its effects on the growth inhibition of C. buinensis are related to membrane permeabilization, an endogenous increase in ROS, the loss of mitochondrial functionality and caspase activation, suggesting that γ33-41PνD1++ triggers C. buinensis cell death via apoptosis, as demonstrated by the key markers of this pathway.
Acknowledgments
This work was performed at the Universidade Estadual do Norte Fluminense Darcy Ribeiro. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil(CAPES) – Finance Code 001. We acknowledge the financial support from the Brazilian agencies CNPq(305766/2013-9) FAPERJ (E-26/203090/2016; E-26/202.132/2015; E-26/202.735/2016). We are grateful to Souza LCD and Kokis VM for general laboratory technical support.
Author contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
de Coninck B, Cammue BPA, Thevissen K. Modes of antifungal action and in planta functions of plant defensins and defensin-like peptides. Fungal Biol Rev. 2013;26(4):109–120. | ||
Cools TL, Vriens K, Struyfs C, et al. The antifungal plant defensin HsAFP1 is a phosphatidic acid-interacting peptide inducing membrane permeabilization. Front Microbiol. 2017;8:2295. | ||
Carvalho AO, Gomes VM. Plant defensins – prospects for the biological functions and biotechnological properties. Peptides. 2009;30(5):1007–1020. | ||
Jarczak J, Kościuczuk EM, Lisowski P, et al. Defensins: natural component of human innate immunity. Hum Immunol. 2013;74(9):1069–1079. | ||
Jha S, Chattoo BB. Expression of a plant defensin in rice confers resistance to fungal phytopathogens. Transgenic Res. 2010;19(3):373–384. | ||
Sarkar P, Jana K, Sikdar SR. Overexpression of biologically safe Rorippa indica defensin enhances aphid tolerance in Brassica juncea. Planta. 2017;246(5):1029–1044. | ||
Carvalho AO, Gomes VM. Plant defensins and defensin-like peptides – biological activities and biotechnological applications. Curr Pharm Des. 2011;17(38):4270–4293. | ||
Wong JH, Ip DC, Ng TB, Chan YS, Fang F, Pan WL. A defensin-like peptide from Phaseolus vulgaris cv. “King Pole Bean”. Food Chem. 2012;135(2):408–414. | ||
Parisi K, Shafee TMA, Quimbar P, van der Weerden NL, Bleackley MR, Anderson MA. The evolution, function and mechanisms of action for plant defensins. Semin Cell Dev Biol. Epub 2018 Feb 23. | ||
Lacerda AF, Vasconcelos EA, Pelegrini PB, Grossi de Sa MF. Antifungal defensins and their role in plant defense. Front Microbiol. 2014;5(459):1–10. | ||
Thevissen K, Ferket KK, François IE, Cammue BP. Interactions of antifungal plant defensins with fungal membrane components. Peptides. 2003;24(11):1705–1712. | ||
Hegedüs N, Marx F. Antifungal proteins: more than antimicrobials? Fungal Biol Rev. 2013;26(4):132–145. | ||
Broekaert WF, Cammue BPA, de Bolle MFC, et al. Antimicrobial peptides from plants. Crit Rev Plant Sci. 1997;16(3):297–323. | ||
Games PD, dos Santos IS, Mello EO, et al. Isolation, characterization and cloning of a cDNA encoding a new antifungal defensin from Phaseolus vulgaris L. seeds. Peptides. 2008;29(12):2090–2100. | ||
Mello EO, Ribeiro SF, Carvalho AO. The antifungal activity of PνD1, a plant seed defensin of Phaseolus vulgaris, involves plasma membrane permeabilization, inhibition of medium acidification and induction of reactive oxygen species in yeast cells. Curr Microbiol. 2011;62:1209–1217. | ||
Thevissen K, de Mello Tavares P, Xu D, et al. The plant defensin RsAFP2 induces cell wall stress, septin mislocalization and accumulation of ceramides in Candida albicans. Mol Microbiol. 2012;84(1):166–180. | ||
Soares JR, José Tenório de Melo E, da Cunha M, et al. Interaction between the plant ApDef1 defensin and Saccharomyces cerevisiae results in yeast death through a cell cycle- and caspase-dependent process occurring via uncontrolled oxidative stress. Biochim Biophys Acta Gen Subj. 2017;1861(1 Pt A):3429–3443. | ||
de O Mello É, dos Santos IS, de O Carvalho A, et al. Functional expression and activity of the recombinant antifungal defensin PνD1r from Phaseolus vulgaris L. (common bean) seeds. BMC Biochem. 2014;15(1):7. | ||
Cornet B, Bonmatin JM, Hetru C, Hoffmann JA, Ptak M, Vovelle F. Refined three-dimensional solution structure of insect defensin A. Structure. 1995;3(5):435–448. | ||
Thomma BP, Cammue BP, Thevissen K, Defensins P. Plant defensins. Planta. 2002;216(2):193–202. | ||
Muñoz A, Chu M, Marris PI, et al. Specific domains of plant defensins differentially disrupt colony initiation, cell fusion and calcium homeostasis in Neurospora crassa. Mol Microbiol. 2014;92(6):1357–1374. | ||
de Samblanx GW, Goderis IJ, Thevissen K, et al. Mutational analysis of a plant defensin from radish (Raphanus sativus L.) reveals two adjacent sites important for antifungal activity. J Biol Chem. 1997;272(2):1171–1179. | ||
Sagaram US, Pandurangi R, Kaur J, Smith TJ, Shah DM. Structure-activity determinants in antifungal plant defensins MsDef1 and MtDef4 with different modes of action against Fusarium graminearum. PLoS One. 2011;6(4):e18550. | ||
do Nascimento VV, Mello Éde O, Carvalho LP, et al. PνD1 defensin, a plant antimicrobial peptide with inhibitory activity against Leishmania amazonensis. Biosci Rep. 2015;35(5):e00248. | ||
Figueira TN, Oliveira FD, Almeida I, et al. Challenging metastatic breast cancer with the natural defensin PνD1. Nanoscale. 2017;9(43):16887–16899. | ||
Schmidtchen A, Pasupuleti M, Malmsten M. Effect of hydrophobic modifications in antimicrobial peptides. Adv Colloid Interface Sci. 2014;205:265–274. | ||
Yount NY, Yeaman MR. Multidimensional signatures in antimicrobial peptides. Proc Natl Acad Sci U S A. 2004;101(19):7363–7368. | ||
Bjellqvist B, Hughes GJ, Pasquali C, et al. The focusing positions of polypeptides in immobilized pH gradients can be predicted from their amino acid sequences. Electrophoresis. 1993;14(10):1023–1031. | ||
Bjellqvist B, Basse B, Olsen E, Celis JE. Reference points for comparisons of two-dimensional maps of proteins from different human cell types defined in a pH scale where isoelectric points correlate with polypeptide compositions. Electrophoresis. 1994;15(3–4):529–539. | ||
Gasteiger E, Hoogland C, Gattiker, A, et al. Protein identification and analysis tools on the ExPASy Server. In: Walker JM, editor. The Proteomics Protocols Handbook. New York: Humana Press; 2005:571–607. | ||
Broekaert WF, Terras FRG, Cammue BPA, Vanderleyden J. An automated quantitative assay for fungal growth inhibition. FEMS Microbiol Lett. 1990;69(1–2):55–59. | ||
Vermelho AB, Pereira AF, Coelho RRR, Souto-Padrón T. Práticas de Microbiologia. Rio de Janeiro: Guanabara Koogan; 2006:239p. | ||
Thevissen K, Terras FR, Broekaert WF. Permeabilization of fungal membranes by plant defensins inhibits fungal growth. Appl Environ Microbiol. 1999;65(12):5451–5458. | ||
Schaaper WM, Posthuma GA, Plasman HH, et al. Synthetic peptides derived from the beta2-beta3 loop of Raphanus sativus antifungal protein 2 that mimic the active site. J Pept Res. 2001;57(5):409–418. | ||
Aerts AM, Carmona-Gutierrez D, Lefevre S, et al. The antifungal plant defensin RsAFP2 from radish induces apoptosis in a metacaspase independent way in Candida albicans. FEBS Lett. 2009;583(15):2513–2516. | ||
Islam KT, Velivelli SLS, Berg RH, Oakley B, Shah DM. A novel bi-domain plant defensin MtDef5 with potent broad spectrum antifungal activity binds to multiple phospholipids and forms oligomers. Biosci Rep. 2017;7:16157. | ||
Sagaram US, El-Mounadi K, Buchko GW, et al. Structural and functional studies of a phosphatidic acid-binding antifungal plant defensin MtDef4: identification of an RGFRRR motif governing fungal cell entry. PLoS One. 2013;8(12):e82485. | ||
Findlay B, Zhanel GG, Schweizer F. Cationic amphiphiles, a new generation of antimicrobials inspired by the natural antimicrobial peptide scaffold. Antimicrob Agents Chemother. 2010;54(10):4049–4058. | ||
Chaffin WL. Candida albicans cell wall proteins. Microbiol Mol Biol Rev. 2008;72(3):495–544. | ||
Ramesh S, Govender T, Kruger HG, de La Torre BG, Albericio F. Short AntiMicrobial Peptides (SAMPs) as a class of extraordinary promising therapeutic agents. J Pept Sci. 2016;22(7):438–451. | ||
Taveira GB, Carvalho AO, Rodrigues R, Trindade FG, da Cunha M, Gomes VM. Thionin-like peptide from Capsicum annuum fruits: mechanism of action and synergism with fluconazole against Candida species. BMC Microbiol. 2016;16(1):12. | ||
Taveira GB, Mello Érica O, Carvalho AO, et al. Antimicrobial activity and mechanism of action of a thionin-like peptide from Capsicum annuum fruits and combinatorial treatment with fluconazole against Fusarium solani. Biopolymers. 2017;108(3):e23008. | ||
Xie D, Yao C, Wang L, et al. An albumin-conjugated peptide exhibits potent anti-HIV activity and long in vivo half-life. Antimicrob Agents Chemother. 2010;54(1):191–196. | ||
Kołaczkowska A, Kołaczkowski M. Drug resistance mechanisms and their regulation in non-albicans Candida species. J Antimicrob Chemother. 2016;71(6):1438–1450. | ||
Carmona-Gutierrez D, Eisenberg T, Büttner S, Meisinger C, Kroemer G, Madeo F. Apoptosis in yeast: triggers, pathways, subroutines. Cell Death Differ. 2010;17(5):763–773. | ||
Madeo F, Carmona-Gutierrez D, Ring J, Büttner S, Eisenberg T, Kroemer G. Caspase-dependent and caspase-independent cell death pathways in yeast. Biochem Biophys Res Commun. 2009;382(2):227–231. | ||
Zivna L, Krocova Z, Härtlova A, et al. Activation of B cell apoptotic pathways in the course of Francisella tularensis infection. Microb Pathog. 2010;49(5):226–236. | ||
Vieira ME, Vasconcelos IM, Machado OL, Gomes VM, Carvalho AO. Isolation, characterization and mechanism of action of an antimicrobial peptide from Lecythis pisonis seeds with inhibitory activity against Candida albicans. Acta Biochim Biophys Sin. 2015;47(9):716–729. |
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.