Back to Journals » OncoTargets and Therapy » Volume 13
Identification of Long Non-Coding RNA SNHG Family as Promising Prognostic Biomarkers in Acute Myeloid Leukemia
Received 3 June 2020
Accepted for publication 3 August 2020
Published 24 August 2020 Volume 2020:13 Pages 8441—8450
DOI https://doi.org/10.2147/OTT.S265853
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Leo Jen-Liang Su
Jian Shi,1,* Weifeng Ding,2,* Hong Lu3
1Enzymology Laboratory, Affiliated Hospital of Nantong University, Nantong 226001, Jiangsu, People’s Republic of China; 2Department of Laboratory Medicine, Affiliated Hospital of Nantong University, Nantong 226001, Jiangsu, People’s Republic of China; 3Eye Institute, Department of Ophthalmology, Affiliated Hospital of Nantong University, Nantong 226001, Jiangsu, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Hong Lu
Eye Institute, Department of Ophthalmology, Affiliated Hospital of Nantong University, 20 Xisi Road, Nantong 226001, Jiangsu, People’s Republic of China
Email [email protected]
Background: Small nucleolar RNA host gene (SNHG) family members are newly recognized lncRNAs, which have been revealed to be oncogenes in several cancers. However, little studies investigated the expression and clinical implications of SNHGs in AML.
Methods: Herein, we systemically determined the prognostic role of the expression of SNHG family members in acute myeloid leukemia (AML).
Results: Among the expression of all SNHG family members, we identified that only SNHG7 and SNHG12 expression were found to have prognostic effects on overall survival (OS) and leukemia-free survival (LFS) in AML by Cox regression univariate analysis. Furthermore, Kaplan–Meier analysis showed that SNHG7 higher-expressed cases had markedly longer OS and LFS time than SNHG7 lower-expressed cases, whereas SNHG12 higher-expressed cases had markedly shorter OS and LFS time than SNHG12 lower-expressed cases. Interestingly, SNHG7 and SNHG12 expression were also associated with several prognosis-related clinical/molecular features such as white blood cell counts, FAB/cytogenetic classifications, IDH1 mutation, RUNX1 mutation, and NPM1 mutation. Despite the associations, Cox regression multivariate analysis confirmed the independent prognostic impact of SNHG7 and SNHG12 expression in AML. Notably, we further validated that both SNHG7 and SNHG12 expression was significantly increased in newly diagnosed AML patients.
Conclusion: Our findings demonstrated that SNHG7 and SNHG12 expression act as independent prognostic indicators in AML.
Keywords: LncRNA, SNHG, expression, prognosis, AML
Introduction
Acute myeloid leukemia (AML), the most common adult leukemia, is a highly cytogenetically and molecularly heterogeneous blood cancer.1 Cytogenetic abnormalities and molecular alterations play key roles in the processes of AML occurrence and development such as cell self-renewal, apoptosis, proliferation, and differentiation.2 These pathological changes eventually lead to hematopoietic failure and adverse prognosis of AML patients.3 Although numerous strategies, such as chemotherapy, hematopoietic stem cell transplantation (HSCT), and immunotherapy, have been applied to treat AML, the prognosis of this disease is still poor.3 Consequently, it is urgent to identify new prognostic/predictive biomarkers and therapeutic targets for AML.
Over the last decade, non-coding RNAs account for 90% of human genome which do not codify for proteins but play a role in the regulation of functions have been shown to have multiple applications in the diagnosis, prognosis and therapeutic approach of various types of human cancers, including AML.4,5 Non-coding RNAs can be classified into subtypes based on molecular size including microRNAs which defined as 19–25 nt in length and long non-coding RNAs (lncRNAs) which usually contain more than 200 nt in length.6 So far, a large number of lncRNAs, such as H19, HOTAIR, UCA1, CASC15, MEG3, PANDAR, CCDC26, and NEAT1, have been explored in AML.7,8 Small nucleolar RNA host gene (SNHG) family members (SNHGs) including SNHG1, SNHG2/GAS5, SNHG3, SNHG4, SNHG5, SNHG6, SNHG7, SNHG8, SNHG9, SNHG10, SNHG11, SNHG12, SNHG13/DANCR, SNHG15, SNHG17, SNHG20 and SNHG28, are newly recognized lncRNAs, which have been revealed to be oncogenes in several cancers.9 Also, several members of SNHG family including SNHG1, SNHG3, and SNHG5 have been found to be dysregulated and play a crucial role in leukemogenesis, and also have prognostic value in AML.10–14 Since little studies investigated the expression and clinical implications of SNHGs in AML, we systemically determined the prognostic role of SNHGs expression in patients with AML.
Materials and Methods
Patients
A total of 173 AML patients were obtained for SNHGs expression data from The Cancer Genome Atlas (TCGA) databases.15 Clinical and molecular characteristics of these patients including age, gender, white blood cell (WBC) counts, peripheral blood (PB) blasts, bone marrow (BM) blasts, French-American-British (FAB) subtypes, karyotypes, and the frequencies of AML-associated genetic mutations were obtained. Treatments of these patients were induction chemotherapy together with chemotherapy and HSCT as consolidation treatment as reported.15
Another cohort of 50 AML patients and 25 healthy volunteers from the Affiliated Hospital of Nantong University was also enrolled in the study. The study was approved by the Institutional Review Board of the Affiliated Hospital of Nantong University, and all participants provided informed consents.
Samples Preparation, RNA Isolation, and Reverse Transcription
Peripheral blood (PB) specimens were collected from 25 controls and 50 AML patients at diagnosis time. PB nucleated cells were obtained after using red blood cell lysis buffer (Solarbio, Beijing, China). Total RNA was extracted from PB nucleated cells using Trizol reagent (Invitrogen, Carlsbad, CA, USA). Reverse transcription was performed to synthesize cDNA using PrimeScript™ RT reagent Kit (TaKaRa, Tokyo, Japan). The program of reverse transcription was performed according to the manufacturer’s instructions.
RT-qPCR
Real-time quantitative PCR (RT-qPCR) was conducted to detect SNHG7, SNHG12 and GAPDH transcript using TB Green Premix Ex Taq™ II (TaKaRa, Tokyo, Japan). The primers used for SNHG7 were 5ʹ-GTGACTTCGCCTGTGATGGA-3ʹ (forward) and 5ʹ-TGCTGCCTGGCTTTGGTT-3ʹ (reverse). The primers used for SNHG12 were 5ʹ-AGATGGTGGTGAATGTGGC-3ʹ (forward), and 5ʹ-AGTCTTGATGGGACCGTTTT-3ʹ (reverse). The primers used for GAPDH were 5ʹ- AATCCCATCACCATCTTCCAG-3ʹ (forward) and 5ʹ-GAGCCCCAGCCTTCTCCAT-3ʹ (reverse). Housekeeping gene GAPDH was detected as the reference gene. Relative SNHG7 and SNHG12 transcript level was calculated based on 2-∆∆CT method.
Statistical Analysis
Mann–Whitney’s U-test and Pearson Chi-square/Fisher exact test were used for the comparison of continuous variables and categorical variables, respectively. The effect of SNHG7 and SNHG12 expression on leukemia-free survival (LFS) and overall survival (OS) analyzed through Cox regression analysis and Kaplan-Meier analysis. The two-tailed P value <0.05 in all statistical analyses was defined as statistically significant.
Results
Identification of Prognosis-Related SNHGs Expression in AML
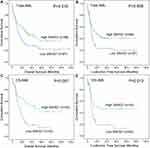
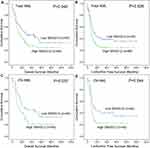
In order to evaluate the prognostic significance of SNHGs expression in AML, we extracted the expression data of SNHGs (SNHG1, SNHG2/GAS5, SNHG3, SNHG4, SNHG5, SNHG6, SNHG7, SNHG8, SNHG9, SNHG10, SNHG11, SNHG12, SNHG13/DANCR, SNHG15, SNHG17, SNHG20, and SNHG28) in AML from the TCGA databases. Prognostic significance of SNHGs expression was analyzed between two groups (lower and higher) divided by the median level of each SNHG member mRNA, respectively. By Cox regression univariate analysis, only SNHG7 and SNHG12 expression were found to have prognostic effects on OS and LFS among both total AML and cytogenetically normal AML (CN-AML) patients (Table 1). Furthermore, among both total AML and CN-AML, Kaplan-Meier analysis also showed that SNHG7 higher-expressed cases had markedly longer OS and LFS time than SNHG7 lower-expressed cases (Figure 1), whereas SNHG12 higher-expressed cases had markedly shorter OS and LFS time than SNHG12 lower-expressed cases (Figure 2).
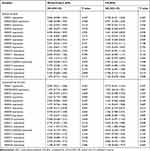 |
Table 1 Cox Regression Univariate Analysis of Variables for Overall Survival and Leukemia-Free Survival in AML Patients |
Validation of SNHG7/12 Overexpression in Newly Diagnosed AML
In order to explore the expression pattern of SNHG7 and SNHG12 in AML, we further examined SNHG7 and SNHG12 mRNA in newly diagnosed AML patients. By RT-qPCR results, both SNHG7 and SNHG12 expression were significantly increased in newly diagnosed AML as compared with normal controls (Figure 3).
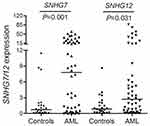 |
Figure 3 SNHG7/12 expression in AML. SNHG7/12 transcript level in controls and AML patients, which was detected by RT-qPCR. |
Clinical Implications of SNHG7/12 Expression in AML
Due to the prognostic effect of SNHG7 and SNHG12 expression in AML, we further analyzed the associations of SNHG7/12 expression with clinical/biological features of AML patients. As presented in Table 2, patients with higher expression of SNHG7 presented lower WBCs and higher percentage of PB blasts than those with lower expression of SNHG7 patients. Moreover, significant difference was observed between two groups among the distributions of FAB classifications (Table 2). Higher expression of SNHG7 was frequently occurred in FAB-M1/M2 and less frequently happened in FAB-M4/5 (Table 2). Although no significant difference was observed between two groups among the distributions of cytogenetic classifications, higher expression of SNHG7 was closely associated −7/del(7) subtype (Table 2).
 |
Table 2 Correlation of SNHG7/SNHG12 Expression with Clinic-Pathologic Characteristics in AML |
Regarding SNHG12, patients with higher expression of SNHG12 presented higher percentage of PB blasts than those with lower expression of SNHG12 patients (Table 2). Moreover, significant difference was observed between two groups among the distributions of cytogenetic classifications (Table 2). Higher expression of SNHG12 was less frequently occurred in inv(16) and other subtypes (Table 2). Although no significant difference was observed between two groups among the distributions of FAB classifications, higher expression of SNHG12 was frequently occurred in FAB-M1 and less frequently happened in FAB-M4 (Table 2).
SNHG7/12 Expression Associated with Gene Mutations in AML
We also observed the associations of SNHG7/12 expression with AML-associated gene mutations. Higher SNHG7 expression was associated with FLT3 and NPM1 wild type as well as IDH1 mutation (Table 2). In addition, higher SNHG12 expression was associated with RUNX1 wild type (Table 2). In order to confirm the significant correlations of SNHG7/12 expression with these gene mutations, we also compared the SNHG7/12 expression with and without these gene mutations. As presented in Figure 4, patients with IDH1 and RUNX1 mutations showed significantly higher SNHG7 expression (P=0.001 and 0.037, respectively), whereas cases with NPM1 mutation showed markedly higher SNHG12 expression (P=0.014).
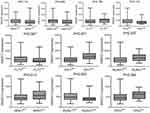 |
Figure 4 The associations of SNHG7/12 expression with gene mutations in AML. SNHG7/12 expression in AML patients with and without gene mutations. |
The Independent Prognostic Value of SNHG7/12 Expression in AML
Since SNHG7/12 expression was associated with well-known prognostic factors such as WBC and gene mutations in AML, we further performed Cox regression multivariate analysis adjusting for prognosis-related factors. As shown in Table 3, both SNHG7 and SNHG12 could act as independent prognostic factors for OS and LFS in both total AML and CN-AML.
 |
Table 3 Cox Regression Multivariate Analysis of Variables for Overall Survival and Leukemia-Free Survival in AML Patients |
Discussion
The oncogenic role of SNHGs in diverse human cancers is supported by solid scientific data, which show that they are related to stimulation of the following malignant processes: epithelial to mesenchymal transition, invasion, proliferation, cell cycle, and apoptosis evasion.9 We intended to test the SNHGs expression and determined their clinical implication in AML. In this study, we for the first time revealed clinical implications of SNHGs expression in AML. Among all members of SNHG family, we only observed that SNHG7 and SNHG12 expression have prognostic value in AML. Moreover, we also validated that both SNHG7 and SNHG12 were significantly overexpressed in newly diagnosed AML. Notably, by our study, higher SNHG7 expression was associated with favorable prognosis, whereas higher SNHG12 expression was correlated with poor prognosis in AML. These results indicated that SNHG7 and SNHG12 may play different roles in AML during occurrence and development. However, until now, no clinical or functional studies were observed regarding SNHG7 and SNHG12 in AML. In solid tumors, a variety of studies have investigated the potential role of SNHG7 in the development and progression of multiple human cancers such as bladder, breast, colorectal, esophageal, gastric, and prostate cancer, as well as osteosarcoma.16 SNHG7 was reported to promote proliferation and metastasis, while inhibiting apoptosis in these types of cancer cells.16 Moreover, high expression of SNHG7 predicts poor prognosis and poor survival for such patients.16 Also, the underlying role of SNHG12 was also determined in a number of cancers, such as breast, gastric, osteosarcoma, and glioma.17 The increased expression of SNHG12 in these cancers has been correlated with the viability, proliferation, metastasis, and invasion of tumor cells, impacting the prognosis and survival of cancer patients.17 Further functional studies are needed to investigate the underlying role of SNHG7 and SNHG12 in AML occurrence and development.
Interestingly, previous studies have shown that SNHG1 expression was up-regulated and associated with poor prognosis in AML.10 Moreover, SNHG1 promoted cell proliferation and inhibited the cell apoptosis by inhibiting miR-101 or miR-488/NUP205 axis in AML.10,11 Peng et al reported that SNHG3 elicited a growth-promoting role via sponging miR-758-3p to regulate SRGN expression in AML.12 In addition, Li et al showed that SNHG5 was increased and served as a potential prognostic biomarker in AML.13 Mechanically, SHNG5 played a crucial role in AML chemotherapy resistance by targeting the miR-32/DNAJB9 axis.14 However, we did not observe the prognostic value of SNHG1/3/5 expression in AML. The conflicting results may be attributed to the differences in ethnics and in AML subtype distribution with different phenotypes and genotypes. Due to the limitation of our clinical samples, we could not perform a validation study regarding the prognostic value of SNHG7 and SNHG12 to further confirm our results identified by TCGA data. Obviously, further studies are required to validate the results in different ethnics before SNHGs expression could be used routinely as a promising biomarker for risk stratification in AML.
Genetic alterations and epigenetic modifications are common molecular events involved in the process of leukemogenesis and interacted with each other. Evidences have shown that somatic gene mutations such as RUNX1 mutation affected transcription activation in AML.18 In our study, we further identified the association between SNHG7/12 and common gene mutations such as IDH1/2, RUNX1 and NPM1 mutations in patients with AML. However, the potential connections between SNHG7/12 expression and these gene mutations remain poorly defined. Further studies are required to determine the potential role of SNHG7 and SNHG12 overexpression during the leukemogenesis caused by IDH1/2, RUNX1 and NPM1 mutations.
Collectively, our findings demonstrated that SNHG7 and SNHG12 expression act as independent prognostic indicators in AML.
Abbreviations
AML, acute myeloid leukemia; HSCT, hematopoietic stem cell transplantation; LncRNAs, long non-coding RNAs; SNHG, small nucleolar RNA host gene; TCGA, The Cancer Genome Atlas; WBC, white blood cell; PB, peripheral blood; BM, bone marrow; FAB, French-American-British; CN-AML, cytogenetically normal AML; LFS, leukemia-free survival; OS, overall survival.
Ethics Statements
All procedures performed in studies involving human participants were approved by the Ethics Committee of Affiliated Hospital of Nantong University with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all patients included in this study.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Döhner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. N Engl J Med. 2015;373(12):1136–1152. doi:10.1056/NEJMra1406184
2. Chen J, Odenike O, Rowley JD. Leukaemogenesis: more than mutant genes. Nat Rev Cancer. 2010;10(1):23–36. doi:10.1038/nrc2765
3. Döhner H, Estey E, Grimwade D, et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129(4):424–447.
4. Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011;1(5):391–407. doi:10.1158/2159-8290.CD-11-0209
5. Garitano-Trojaola A, Agirre X, Prósper F, Fortes P. Long non-coding RNAs in haematological malignancies. Int J Mol Sci. 2013;14(8):15386–15422. doi:10.3390/ijms140815386
6. Beermann J, Piccoli MT, Viereck J, Thum T. Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev. 2016;96(4):1297–1325. doi:10.1152/physrev.00041.2015
7. Zhang TJ, Zhou JD, Zhang W, et al. H19 overexpression promotes leukemogenesis and predicts unfavorable prognosis in acute myeloid leukemia. Clin Epigenetics. 2018;10:47. doi:10.1186/s13148-018-0486-z
8. Gourvest M, Brousset P, Bousquet M. Long noncoding RNAs in acute myeloid leukemia: functional characterization and clinical relevance. Cancers. 2019;11(11):1638. doi:10.3390/cancers11111638
9. Zimta -A-A, Tigu AB, Braicu C, Stefan C, Ionescu C, Berindan-Neagoe I. An emerging class of long non-coding RNA with oncogenic role arises from the snoRNA host genes. Front Oncol. 2020;10:389. doi:10.3389/fonc.2020.00389
10. Tian M, Gong W, Guo J. Long non-coding RNA SNHG1 indicates poor prognosis and facilitates disease progression in acute myeloid leukemia. Biol Open. 2019;8(10):bio046417. doi:10.1242/bio.046417
11. Bao XL, Zhang L, Song WP. LncRNA SNHG1 overexpression regulates the proliferation of acute myeloid leukemia cells through miR-488-5p/NUP205 axis. Eur Rev Med Pharmacol Sci. 2019;23(13):5896–5903. doi:10.26355/eurrev_201907_18334
12. Peng L, Zhang Y, Xin H. lncRNA SNHG3 facilitates acute myeloid leukemia cell growth via the regulation of miR-758-3p/SRGN axis. J Cell Biochem. 2020;121(2):1023–1031. doi:10.1002/jcb.29336
13. Li J, Sun CK. Long noncoding RNA SNHG5 is up-regulated and serves as a potential prognostic biomarker in acute myeloid leukemia. Eur Rev Med Pharmacol Sci. 2018;22(11):3342–3347. doi:10.26355/eurrev_201806_15154
14. Wang D, Zeng T, Lin Z, et al. Long non-coding RNA SNHG5 regulates chemotherapy resistance through the miR-32/DNAJB9 axis in acute myeloid leukemia. Biomed Pharmacother. 2019;123:109802. doi:10.1016/j.biopha.2019.109802
15. Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368(22):2059–2074. doi:10.1056/NEJMoa1301689
16. Zhou Y, Tian B, Tang J, et al. SNHG7: a novel vital oncogenic lncRNA in human cancers. Biomed Pharmacother. 2020;124:109921. doi:10.1016/j.biopha.2020.109921
17. Tamang S, Acharya V, Roy D, et al. SNHG12: an LncRNA as a potential therapeutic target and biomarker for human cancer. Front Oncol. 2019;9:901. doi:10.3389/fonc.2019.00901
18. Mujahed H, Miliara S, Neddermeyer A, et al. AML displays increased CTCF occupancy associated with aberrant gene expression and transcription factor binding. Blood. 2020;136(3):339–352. doi:10.1182/blood.2019002326
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


