Back to Journals » OncoTargets and Therapy » Volume 13
Identification and Validation of STAT6 as a Prognostic and Predictive Biomarker in Acute Myeloid Leukemia
Authors Liu W, Zhu F, Yan J , Liu Y, Chen C, Zhang K, Zhao X, Chen J
Received 18 July 2020
Accepted for publication 5 October 2020
Published 2 November 2020 Volume 2020:13 Pages 11165—11176
DOI https://doi.org/10.2147/OTT.S272757
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Leo Jen-Liang Su
Wei Liu,1,2 Feiyue Zhu,3 Jiazhuo Yan,4 Yi Liu,1 Cong Chen,1 Kaixuan Zhang,1 Xielan Zhao,1 Jingyuan Chen5
1Department of Hematology, Xiangya Hospital, Central South University, Changsha, Hunan 410008, People’s Republic of China; 2Department of Hematology and Hematopoietic Cell Transplantation, City of Hope National Medical Center, Duarte, CA 91010, USA; 3Department of Hematology, Loudi Central Hospital, Loudi, Hunan 417000, People’s Republic of China; 4Department of Gynaecological Radiotherapy, The Affiliated Tumor Hospital of Harbin Medical University, Harbin, Heilongjiang 150081, People’s Republic of China; 5Research Institute of Blood Lipid and Atherosclerosis, Central South University, Changsha, Hunan 410011, People’s Republic of China
Correspondence: Jingyuan Chen
Research Institute of Blood Lipid and Atherosclerosis, Central South University, No. 139 Middle Renmin Road, Furong District, Changsha, Hunan 410011, People’s Republic of China
Email [email protected]
Background: Acute myeloid leukemia (AML) is one of the most common hematological diseases in adults. The overall survival rate remains unsatisfactory. It is urgent to identify potential prognostic biomarkers and develop new molecular therapeutic strategies for AML. Signal transducer and activator of transcription (STAT) is a family of genes that encode intracellular transcription factors. STATs are associated with leukemogenesis, cellular transformation, and cell cycle in AML.
Methods: We used sequencing data and clinical data from The Cancer Genome Atlas (TCGA) and ONCOMINE to identify expression difference, gene variability and correlation as well as prognostic effects of STAT genes in AML patients. Then, we verified the expression difference of STAT6 between healthy control and AML patients and its prognostic impact in Gene Expression Omnibus (GEO) database and our own recruited cohort.
Results: The mRNA level of STAT6 was increased in AML patients among TCGA, GEO and ONCOMINE public datasets and was found to be an independent risk factor of overall survival in all AML patients and patients who only received chemotherapy by multivariate analysis. In our study, STAT6 mRNA level was markedly up-regulated in AML patients (n=105) compared to healthy donor (n=39) (P=0.0435) as a validated cohort. Patients that only received chemotherapy in high STAT6 group showed significantly lower overall survival (OS) (P=0.0055).
Conclusion: STAT6 expression was increased in AML patients. STAT6 was found to be an adverse prognosis factor in AML patients, especially those who only received chemotherapy treatments.
Keywords: acute myeloid leukemia, STAT, prognosis, TCGA, GEO
Introduction
Acute myeloid leukemia (AML) is characterized by dysfunction and abnormal clone of malignant hematopoietic stem cells.1 It is one of the most common hematological diseases in adults.2 Although many AML patients were able to benefit from stereotypical induction chemotherapy and allogeneic or autologous hematopoietic stem cell transplantation (HSCT), the overall survival rate of AML patients remains unsatisfactory. Moreover, patients present different chemotherapy response and prognosis to those therapies and the potential mechanisms are still under investigation. Recently, the development of molecule-targeting drugs has prolonged survival and improved clinical outcome of AML patients.3 Advances in epigenetic mutation studies of DNA methyltransferase 3 alpha (DNMT3A), fms related receptor tyrosine kinase 3 (FLT3), tet methylcytosine dioxygenase 2 (TET2), argininosuccinate lyase 1 (ASL1) have profoundly changed our understanding of the mechanisms of AML development. However, with the complexity in the pathogenesis of AML, the leukemogenesis is still not clearly understood. It is urgent to identify potential prognostic biomarkers and develop new molecular therapeutic strategies for AML.
Signal transducer and activator of transcription (STAT) is a family of genes that encode intracellular transcription factors in regulating cell proliferation, differentiation, and cell cycle progression in cancer. The STAT family includes seven members: STAT1, STAT2, STAT3, STAT4, STAT5A, STAT5B, STAT6.4 STATs play an essential role in chromatin structure and crosstalk with various oncogenic pathways in inflammation and cancer.5 Several studies demonstrated that higher STAT protein expression improved carcinogenesis and influenced prognosis of patients. Prognostic and predictive roles of STAT gene family were also observed in glioblastoma, gastric cancer as well as lung cancer.6–9 There were also researchers demonstrating that STATs participated in the pathogenesis of AML, and inhibition of STAT induced pathways was found to enhance leukemogenesis. Enhanced activations of STATs were also shown to be related with disease treatments.10,11
Improvements in sequencing methods provided us with a more comprehensive understanding of the biology of AML. The gene expression patterns in bioinformatic datasets help provide information on expression differences, related clinical characteristics and functions and clinical outcomes related to these genes. In our study, we aimed to explore the mRNA expressions of STATs on the clinical characteristics and prognostic significance of AML patients. We examined the relationship between expression profiles of STATs and clinical outcomes in AML patients not only from bioinformatic datasets such as The Cancer Genome Atlas (TCGA), Gene Expression Omnibus (GEO) but also from AML patients we recruited.
Methods
Public Database
The gene expression database of ONCOMINE (https://www.oncomine.org/), GEPIA (http://gepia.cancer-pku.cn/), GEO (https://www.ncbi.nlm.nih.gov/geo/), Xena (https://xenabrowser.net/) and TCGA (https://www.cancer.gov/tcga) are publicly accessible. To have an overview (Figure 1) of the mRNA expressions of STATs in leukemia in the ONCOMINE database, we defined the selection thresholds as fold-change = 1.5, p-value = 0.001. For a further detailed analysis of STAT in AML in different ONCOMINE databases, the screening threshold was identified as P <0.05 (Supplemental Table 1). For STAT family genes, the analysis of the comparison between cancer and normal sets was performed. STATs expression in AML patients and normal was analyzed using GEPIA, consisting of the AML patients’ data from TCGA and the normal data from GTEXs. The TCGA Acute Myeloid Leukemia database was screened for adult AML patients with complete corresponding clinical features, STAT 1–6 expression data and excluded the overall survival time of zero from the Xena database. A total of 132 patients who met the criteria were included in the study; among them, 72 received chemotherapy treatment only, and 60 patients received auto/allo-HSCT. The major clinical features used in the diagnosis of AML patients are presented in Table 2. In addition, the gene expression datasets GSE13159 and GSE9476 of AML were also included from the Gene Expression Omnibus (GEO) database.
 |
Table 1 Unicox and Multicox Regression Analysis for Variablesfor Total Survival, Survival After Chemotherapy and HSCT in AML Among TCGA Cohort |
 |
Table 2 Clinic-Pathologic Characteristics in AML Patients of Different STAT6 Expression Among TCGA Cohort |
Patients and Ethics
A cohort of 105 AML patients and 39 healthy donors were enrolled between March 2016 to June 2019 in this study. The FAB of AML patients was classified according to the 2016 World Health Organization (WHO) criteria. The corresponding clinical features of AML patients are presented in Supplemental Table 4. The overall survival was defined as the time from diagnosis to death or the follow-up was terminated on November 15th, 2019. This study was conducted by the Declaration of Helsinki. It was approved by the ethics committee of Xiangya Hospital, Central South University.
RT-qPCR
Bone marrow mononuclear cells (BMMNCs) were separated using Ficoll-Hypaque (GE Healthcare, United States). Total RNA was extracted from BMMNCs with Trizol reagent (Life Technologies, United States). Reverse transcription to cDNA was performed using PrimeScript Kit (TaKaRa, Japan). Real-time PCR using Cham Q Universal SYBR Green Master Mix (Vazyme, China) was completed on the ViiATM7 RT-PCR system (Applied Biosystems, USA). The primers used for STAT6 expression were: forward: 5ʹ-GTTCCGCCACTTGCCAATG-3ʹ, reverse: 5ʹ- TGGATCTCCCCTACTCGGTG-3ʹ. Relative STAT6 expression mRNA levels were calculated by 2−∆∆CT and were normalized to internal control (β-Actin).
Functional Analysis
The analyses of genetic alterations frequency, co-expression, and interaction network of STAT genes were performed with an online tool cBioPortal (https://www.cbioportal.org/). Analysis of gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways of STATs and neighboring genes showed enrichment using the online tool Metascape (https://metascape.org/).
Statistical Analyses
All statistical analyses were completed using SPSS 22.0 and GraphPad Prism 8.0. Pearson Chi-square analysis or Fisher exact test was used for the comparison of categorical variables, whereas Mann–Whitney’s U-test was used for the comparison of continuous variables. The prognostic effect of STAT6 expression was analyzed through Kaplan-Meier analysis using the Log rank test. Univariate and multivariate proportional hazard regression analysis was performed using Cox regression. The P-value <0.05 (two-tailed) in all statistical analyses was defined as statistically significant.
Results
Expression Levels of STATs in AML Patients
Seven STAT family members have been identified in human cancers. We compared the mRNA levels of STATs in cancers with those in normal tissue samples by using ONCOMINE databases (Figure 1 and Supplemental Table 1). ONCOMINE analysis showed that, for STAT1, STAT2 and STAT5A, they increase in some datasets and decrease in other datasets. For STAT4, STAT5B and STAT6, more datasets showed a decreased STAT in leukemia patients. For detailed analysis in ONCOMINE datasets, we further searched the expression of STATs in AML patients. In the Andersson Leukemia dataset, STAT2, STAT5A and STAT5B were increased by 1.339, 1.295 and 1.428 folds, respectively, in AML patients compared to healthy people (P = 0.018, 0.01 and 0.001, respectively). In the Valk Leukemia dataset, the increase was seen in STAT2, STAT3 and STAT6 and the P values compared with normal bone marrow were 0.015, 0.027 and 7.09E-4. Moreover, STAT2 and STAT6 were also significantly higher in the Haferlach Leukemia dataset. STAT2 increased by 1.171 folds while STAT6 increased by 1.265 folds, and the P-values compared with control were 8.31E-4 and 1.85E-7 accordingly (Supplemental Table 1). To make a more detailed comparison in AML, we further analyzed the AML patients from the TCGA database. We found mRNA expression levels of STAT2, STAT4 and STAT6 were significantly higher in AML than in normal tissues by using GEPIA (Figure 2).
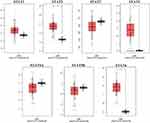 |
Figure 2 Expression of STAT genes in AML patients compared with healthy control. |
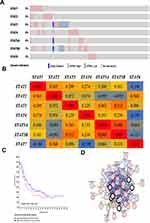
Genetic Alterations and Correlations of STATs Genes in AML
The percentages of genetic alterations in STAT family members for leukemia varied from 3 to 9% for individual genes based on the TCGA Provisional dataset (STAT1, 5%; STAT2, 5%; STAT3, 9%; STAT4, 6%; STAT5A, 4%; STAT5B, 7%; STAT6, 3%; Figure 3A). Pearson correlation analysis was conducted using expression data (RNA Seq V2 RSEM) of STAT family members collected from the cBioPortal online tool for Leukemia (TCGA, Provisional). The results indicated a significant positive correlation among STAT5B and STAT5A, STAT5B and STAT3, except these, only mild to moderate correlation between others (Figure 3B). Kaplan–Meier plot and Log rank test indicated no significant difference in overall survival (OS) between the cases with alterations in one of the query genes and those without (P = 0.220, Figure 3C). Next, we constructed the network for STATs and the 50 most frequently altered neighboring genes by using cBioPortal (Figure 3D and Supplemental material).
Functional Enrichment Analysis of STATs in AML Patients
The functions of STAT family members and their co-activated genes were predicted by analyzing GO and KEGG in Metascape. The top 20 GO enrichment items were classified into 2 functional groups: biological process group (16 items) and molecular function group (4 items) (Figure 4A and B and Supplemental Table 2). STATs and their neighboring genes were enriched in cytokine-mediated signaling pathway and cytokine production, peptidyl-tyrosine phosphorylation, cell differentiation, such as myeloid cell leukocyte and myeloid leukocyte differentiation, positive regulation of cell migration, transmembrane receptor protein tyrosine kinase signaling pathway, positive regulation of kinase activity and so on. The functions for these genes were mainly enriched in GTPase activity, protein kinase activity, protein phosphatase binding and MAP kinase activity.
The top 12 KEGG pathways for the STAT family members and their neighboring genes are shown in Figure 4C and D and Supplemental Table 3. Among these pathways, pathways in cancer and Ras cell signaling, Cytokine-cytokine receptor interaction, Natural killer cell mediated cytotoxicity and Endocytosis were found to be related to multiple tumor development and were involved in leukemia tumorigenesis and pathogenesis.
To better understand the relationship between STAT family members and leukemia, a Metascape protein–protein interaction enrichment analysis was performed. The protein–protein interaction network and MCODE components identified in the gene lists are shown in Figure 4E and F.
Prognostic Value of STAT6 in AML Patients
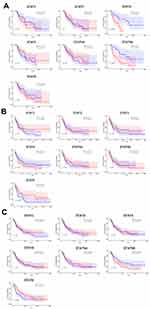
To investigate the prognostic value of STATs expression in AML patients, we further studied the role of STATs genes in the survival of AML patients from the TCGA database (divided AML patients into two groups according to the median level of STATs). Based on the Kaplan-Meier analysis, the STATs showed no correlation with the overall survival rate among total AML patients and patients with auto/allo-HSCT. However, in patients who have only received chemotherapy, higher expression of STAT6 showed a poor prognosis (Figure 5A-C). Furthermore, we evaluated the prognostic effect of STATs expression in AML patients by using Cox regression analysis; in both unicox and multicox analysis, STAT6 was found as an independent predictor for overall survival in all AML patients and AML patients only received chemotherapy (Table 1).
Clinical Features of STAT6 in AML Patients
To explore the clinical features in AML patients, the comparison of clinical/laboratory characteristics of AML patients between two groups (STAT6high and STAT6low) was summarized in Table 2. There were no significant differences between STAT6high and STAT6low groups in sex, PM blasts and BM blasts (P > 0.05). However, STAT6high cases had significantly higher white blood cells (WBC) and older age (P = 0.002 and 0.018). Additionally, significant differences in distributions of French-American-British (FAB) classifications and cytogenetics were observed. STAT6 high expression was markedly negatively correlated with FAB-M3 (P = 0.005) and t (15;17) cytogenesis (P=0.009). No significant differences were found between STAT6 expression among the gene mutations.
Verification of STAT6 in Expression and Prognosis Value in Validated AML Patients
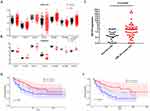
We analyzed data from GEO datasets, GSE13159 and GSE9476, to verify the expression of STAT6 in AML patients (Figure 6A and B). In both datasets, STAT6 showed an increased expression in AML compared to healthy control. We next determined the expression and prognostic impact of STAT6 in recruited AML patients. Patients’ characteristics are shown in Supplemental Table 4. Follow-up data were available in 105 patients. Firstly, we observed that STAT6 was overexpressed in AML patients compared to healthy donors (Figure 6C). By Kaplan-Meier analysis, only patients who received chemotherapy treatment and have a lower expression of STAT6 showed higher over-all survival (Figure 6E). Cox regression analysis also identified that STAT6 expression could be an independent factor of OS in AML patients who received chemotherapy treatment (Supplemental Table 5). Although not significant, there was also a higher survival tendency in the overall survival of STAT6low group in all AML patients (Figure 6D).
Discussion
AML is a malignant hematological disease and is associated with many different cytogenetic abnormalities with genetic alterations detected at diagnosis. It is the leading cause of leukemia-associated deaths worldwide each year. Although significant progress was made in the last two decades, etiologies, pathogenesis and biomarker of the disease are not yet clarified and are still worth researching. Meanwhile, for patients who received standard therapy, effects and prognosis vary. The STAT gene family contains seven different members, STAT1 to STAT6, and is a group of proteins located mainly in nucleus and modulates target genes for cell proliferation, differentiation and apoptosis. The function of STAT genes has been validated in several cancer types.4,12,13 STAT-related transcription factors were found constitutively activated in peripheral blood cells from AML patients.14
Furthermore, the JAK/STAT pathway is heavily involved in AML, and blocking phosphorylation of JAK-STAT could inhibit proliferation of AML cells in vitro and in vivo.15–17 Among different STAT proteins, it was confirmed that STAT1 is activated in AML while inhibiting its activation decreases leukemia cell proliferation.18,19 STAT3 and STAT5 are hyperactivated in the progression and nosogenesis of myeloid leukemia. STAT3 and STAT5 usually collaborate with upstream oncogenic drivers such as FLT3-ITD, BCL-ABL and JAK2.20 Inhibition of STAT3 was found to have potent anti-leukemia activity, and blocking the expression of STAT5 could inhibit proliferation and enhance apoptosis of AML cells.21,22 STAT6 induced by interleukin 4 (IL-4) also has an anti-leukemia effect in primitive AML cells.23
Although basic to translational research have found a potent role for STATs in the disease pathogenesis and disease inhibition, the role of STATs as biomarkers for disease diagnosis and prognosis is still worth investigating. Our research identified an expression difference in healthy control and AML patients and prognosis differences in patients with high or low expression of STATs in TCGA and ONCOMINE database. Herein, we selected STAT gene expression to analyze and investigate whether the change of expression of different STATs was related to clinical parameters as well as prognosis.
STAT6 is a member of the STAT gene family and was found to play different roles according to cancer type.12,24–26 In AML, STAT6 was increased in AML blasts and participated in the pathogenesis of AML.23,27 Through analysis of expression differences and prognosis values, we demonstrated that expression of STAT6 was increased in AML patients. Although it failed to achieve a significant role in predicting OS, STAT6 high expression was related to poor prognosis in patients who only received chemotherapy. Thereafter, we verified these results with the GEO databases and our cohort. Overexpression of STAT6 was found in both GEO database GSE13159 and GSE9476. We also recruited 105 AML patients, who had undergone chemotherapy or HSCT. STAT6 increased in these patients while a poor prognosis with higher STAT6 expression in patients only received chemotherapies was also verified. In accordance with our finding, STAT6 level and activation were confirmed to have a prognostic value in patients who underwent chemotherapy in several cancer types, such as primary CNS lymphoma, B cell lymphoma and colon cancer,28–30 and may relate to the patients’ immune function.31
It is well known that STAT signaling can be activated by diverse cytokines that are involved in many biological processes, such as immunity, cell division, cell death, and cancer development. From our GO and KEGG analyses, we demonstrated that STATs are enriched in cell differentiation, cell migration and pathways in cancer, which are all risk factors in AML progression. In the AML research field, STAT6 is mainly triggered by IL-4, SHP-1 and JAK. In these signaling pathways, STAT6 usually get triggered and phosphorylated.24,32,33 Our research was the first to find and validate the expression differences of STATs in AML patients and the prognostic value of the STAT6 gene in AML patients only-received chemotherapy treatment. These results may lead to further investigations of the gene function of STAT6. Whether STAT6 could be a prognostic factor or some important target, more evidence should be proved.
Abbreviations
AML, Acute myeloid leukemia; STAT, signal transducer and activator of transcription; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; GEPIA, Gene Expression Profiling Interactive Analysis; TCGA, The Cancer Genome Atlas; HSCT, hematopoietic stem cell transplantation; OS, Overall Survival; GeneMANIA, Gene Multiple Association Network Integration Algorithm; PPI, Protein–protein interaction.
Data Sharing Statement
The data used to support the findings of this study are available from the corresponding author upon request.
Ethics Approval and Consent to Participate
This study was approved by the Ethics Committee of Xiangya Hospital, Central South University in China, and all subjects gave written informed consent.
Acknowledgments
We thank Dr. Jiang Xie, Dr. Lige Xiao, Dr. Guanteng Yang, Dr. Yan Huang for collecting the clinical data.
Author Contributions
JyC designed the study, WL performed most of the experiments, FyZ, JzY, YL, WL, CC, KxZ and XlZ collected the clinical sample and data, performed rest necessary experiments. All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Funding
There is no funding to report.
Disclosure
The authors declare that they have no competing interests.
References
1. Döhner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. N Engl J Med. 2015;373(12):1136–1152.
2. Tallman MS, Wang ES, Altman JK, et al. Acute myeloid leukemia, Version 3.2019, NCCN clinical practice guidelines in oncology. J Natl Compr Cancer Net. 2019;17(6):721–749.
3. Estey E. Acute myeloid leukemia – many diseases, many treatments. N Engl J Med. 2016;375(21):2094–2095. doi:10.1056/NEJMe1611424
4. O’Shea JJ, Holland SM, Staudt LM. JAKs and STATs in immunity, immunodeficiency, and cancer. N Engl J Med. 2013;368(2):161–170. doi:10.1056/NEJMra1202117
5. O’Shea JJ, Schwartz DM, Villarino AV, Gadina M, McInnes IB, Laurence A. The JAK-STAT pathway: impact on human disease and therapeutic intervention. Annu Rev Med. 2015;66:311–328. doi:10.1146/annurev-med-051113-024537
6. Li C, Zhou Y, Deng H, et al. Identification of therapeutic targets and prognostic biomarkers among STAT family in glioblastoma. 2020.
7. Zhang Y, Yu C. Prognostic values of signal transducers activators of transcription in gastric cancer. Biosci Rep. 2019;39:4.
8. Yang M, Chen H, Zhou L, Chen K, Su F. Expression profile and prognostic values of STAT family members in non-small cell lung cancer. Am J Transl Res. 2019;11(8):4866–4880.
9. Liu HW, Lee PM, Bamodu OA, et al. Enhanced Hsa-miR-181d/p-STAT3 and Hsa-miR-181d/p-STAT5A ratios mediate the anticancer effect of garcinol in STAT3/5A-addicted glioblastoma. Cancers. 2019;11:12. doi:10.3390/cancers11121888
10. Steensma DP, McClure R, Karp J, et al. JAK2 V617F is a rare finding in de novo acute myeloid leukemia, but STAT3 activation is common and remains unexplained. Leukemia. 2006;20(6):971–978. doi:10.1038/sj.leu.2404206
11. Zhao S, Guo J, Zhao Y, et al. Chidamide, a novel histone deacetylase inhibitor, inhibits the viability of MDS and AML cells by suppressing JAK2/STAT3 signaling. Am J Transl Res. 2016;8(7):3169.
12. Demicco EG, Harms PW, Patel RM, et al. Extensive survey of STAT6 expression in a large series of mesenchymal tumors. Am J Clin Pathol. 2015;143(5):672–682.
13. Dutta P, Zhang L, Zhang H, et al. Unphosphorylated STAT3 in heterochromatin formation and tumor suppression in lung cancer. BMC Cancer. 2020;20(1):145.
14. Gouilleux-Gruart V, Gouilleux F, Desaint C, et al. STAT-related transcription factors are constitutively activated in peripheral blood cells from acute leukemia patients. Blood. 1996;87(5):1692–1697. doi:10.1182/blood.V87.5.1692.1692
15. Choudhary C, Schwäble J, Brandts C, et al. AML-associated Flt3 kinase domain mutations show signal transduction differences compared with Flt3 ITD mutations. Blood. 2005;106(1):265–273. doi:10.1182/blood-2004-07-2942
16. Zhou J, Bi C, Janakakumara JV, et al. Enhanced activation of STAT pathways and overexpression of survivin confer resistance to FLT3 inhibitors and could be therapeutic targets in AML. Blood J Am Soc Hematol. 2009;113(17):4052–4062.
17. Faderl S, Ferrajoli A, Harris D, Van Q, Priebe W, Estrov Z. WP-1034, a novel JAK-STAT inhibitor, with proapoptotic and antileukemic activity in acute myeloid leukemia (AML). Anticancer Res. 2005;25(3B):1841–1850.
18. Liu F, Chen Y, Li G. Activation of the STAT1 protein in the peripheral blood white cells of the patients with acute myeloid leukemia. Carcinogenesis Teratogenesis Mutagenesis. 2004;16(4):214–216.
19. Jiang L-J, Zhang -N-N, Ding F, et al. RA-inducible gene-I induction augments STAT1 activation to inhibit leukemia cell proliferation. Proc Natl Acad Sci. 2011;108(5):1897–1902.
20. Al-Jamal HA, Mat Jusoh SA, Hassan R, Johan MF. Enhancing SHP-1 expression with 5-azacytidine may inhibit STAT3 activation and confer sensitivity in lestaurtinib (CEP-701)-resistant FLT3-ITD positive acute myeloid leukemia. BMC Cancer. 2015;15:869. doi:10.1186/s12885-015-1695-x
21. Baśkiewicz‐Masiuk M, Machaliński B. The role of the STAT5 proteins in the proliferation and apoptosis of the CML and AML cells. Eur J Haematol. 2004;72(6):420–429. doi:10.1111/j.1600-0609.2004.00242.x
22. Minus MB, Liu W, Vohidov F, et al. Rhodium(II) proximity-labeling identifies a novel target site on STAT3 for inhibitors with potent anti-leukemia activity. Angewandte Chemie (International Ed in English). 2015;54(44):13085–13089. doi:10.1002/anie.201506889
23. Peña-Martínez P, Eriksson M, Ramakrishnan R, et al. Interleukin 4 induces apoptosis of acute myeloid leukemia cells in a Stat6-dependent manner. Leukemia. 2018;32(3):588–596. doi:10.1038/leu.2017.261
24. Li BH, Yang XZ, Li PD, et al. IL-4/Stat6 activities correlate with apoptosis and metastasis in colon cancer cells. Biochem Biophys Res Commun. 2008;369(2):554–560. doi:10.1016/j.bbrc.2008.02.052
25. Cui X, Zhang L, Luo J, et al. Unphosphorylated STAT6 contributes to constitutive cyclooxygenase-2 expression in human non-small cell lung cancer. Oncogene. 2007;26(29):4253–4260.
26. Binnemars-Postma K, Bansal R, Storm G, Prakash J. Targeting the Stat6 pathway in tumor-associated macrophages reduces tumor growth and metastatic niche formation in breast cancer. FASEB J. 2018;32(2):969–978. doi:10.1096/fj.201700629R
27. Munje CR, Williamson AJ, Taylor S, et al. Integrated Nuclear Proteomics and Transcriptomics Identifies New High Frequency Dysregulated Transcription Factors in AML. Washington, DC: American Society of Hematology; 2014.
28. Yang S-H, Lee KS, Kim IS, et al. Long-term survival in primary CNS lymphoma treated by high-dose methotrexate monochemotherapy: role of STAT6 activation as prognostic determinant. J Neurooncol. 2009;92(1):65. doi:10.1007/s11060-008-9736-9
29. Häberle M-T, Kelsch E, Dorsch K, Möller P, Ritz O. Knock-down of BCL6/STAT6 sensitizes primary B cell lymphoma cells for treatment with current therapeutic agents. Oncoscience. 2014;1(4):283. doi:10.18632/oncoscience.35
30. Wang C-G, Ye Y-J, Yuan J, Liu -F-F, Zhang H, Wang S. EZH2 and STAT6 expression profiles are correlated with colorectal cancer stage and prognosis. World J Gastroenterol. 2010;16(19):2421. doi:10.3748/wjg.v16.i19.2421
31. Lesterhuis WJ, Punt CJ, Hato SV, et al. Platinum-based drugs disrupt STAT6-mediated suppression of immune responses against cancer in humans and mice. J Clin Invest. 2011;121(8):3100–3108. doi:10.1172/JCI43656
32. Reich D, Kresinsky A, Müller JP, et al. SHP1 regulates a STAT6–ITGB3 axis in FLT3ITD-positive AML cells. Leukemia. 2020;34(5):1444–1449. doi:10.1038/s41375-019-0676-5
33. Irish JM, Hovland R, Krutzik PO, et al. Single cell profiling of potentiated phospho-protein networks in cancer cells. Cell. 2004;118(2):217–228. doi:10.1016/j.cell.2004.06.028
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.