Back to Journals » Cancer Management and Research » Volume 12
hsa_circRNA_101237: A Novel Diagnostic and Prognostic Biomarker and Potential Therapeutic Target for Multiple Myeloma
Authors Liu X, Tang H, Liu J, Wang X
Received 4 December 2019
Accepted for publication 21 January 2020
Published 20 March 2020 Volume 2020:12 Pages 2109—2118
DOI https://doi.org/10.2147/CMAR.S241089
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Bilikere Dwarakanath
Xiao Liu,1,* Hao Tang,1,* Jing Liu,1 Xiang Wang2
1The Third Xiangya Hospital of Central South University, Changsha 410013, People’s Republic of China; 2Department of Hematology, First Affiliated Hospital of Anhui Medical University, Hefei 230022, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Xiang Wang
Department of Hematology, First Affiliated Hospital of Anhui Medical University, Hefei 230022, People’s Republic of China
Email [email protected]
Background: It has been demonstrated that circular RNA (circRNA) plays a crucial role in the occurrence and development of tumors, but the diagnostic and predictive value of most circRNAs in tumor patients remains unclear, especially for multiple myeloma (MM).
Methods: High-throughput circRNA microarray-based sequencing was used to identify the differentially expressed circRNAs in MM. qRT-PCR was then employed to detect hsa_circRNA_101237 expression levels in the bone marrow tissues from 143 MM patients (65 first-episode treatment-naive patients and 78 patients with recurrent/refractory disease), MM cells and bortezomib-resistant MM cell lines. Whether hsa_circRNA_101237 can be used as a potential biomarker and therapeutic target for MM was investigated.
Results: The average expressions of hsa_circRNA_101237 in the bone marrow tissues from MM patients (especially those with recurrent/refractory disease), MM cells and bortezomib-resistant MM cell lines were increased significantly (P< 0.01). hsa_circRNA_101237 was overexpressed in patients positive for 13q14 deletion, 1q21 amplification, P53 deletion, and t(4,14) and t(14,16). hsa_circRNA_101237 was closely related to prognosis of the patients, and its high expression was associated with shorter OS and PFS. In addition, those overexpressing hsa_circRNA_101237 were less responsive to bortezomib treatment. Bioinformatic analysis indicated that hsa_circRNA_101237 interacted with 11 miRNAs and 10 candidate mRNAs. This finding may shed new light on the subsequent studies on the working mechanism and functions.
Conclusion: It was first reported that hsa_circRNA_101237 was significantly upregulated in MM. It was indicated that hsa_circRNA_101237 may be a novel biomarker for MM, and it plays a significant role in the occurrence and development of MM.
Keywords: circRNAs, MM, biomarker, diagnosis, prognosis
Background
Multiple myeloma (MM) is a plasma cell malignancy in which monoclonal plasma cells proliferate in bone marrow, resulting in an overabundance of monoclonal paraprotein (M protein), destruction of bone, and displacement of other hematopoietic cell lines.1 It is one of the most common hematological malignancies in clinic, accounting for about 10% of all hematological malignancies. The occurrence and development of MM are closely related to genetic variations, including chromosome translocation, gene mutations and cytogenetic abnormalities.2 It has been reported that abnormalities of important chromosomes and genes may lead to the progression of MM and independently predict the prognosis of patients with MM.3 According to one study, t (4;14), t (14; 16) and t(14:20) chromosomal abnormalities along with KRAS, NRAS, FAM46C, DIS3, BRAF, TRAF3 and TP53 gene mutations are associated with poor prognosis of MM patients.4,5 In spite of advance in MM treatment, about 20% of the patients relapse or die within 2 years after the diagnosis.6 Therefore, early diagnosis and effective treatment are highly important for improving the prognosis of MM patients. Individualized treatment modalities can be adopted based on the levels of specific biomarkers in MM patients, so as to optimize the efficacy and reduce the toxic and side effects.7 Identifying new biomarkers for MM facilitates the diagnosis and treatment of MM and also improves prognosis of the patients.
Circular RNA (circRNA), an emerging class of non-coding RNA, has a covalently closed loop structure in which the 3′ and 5′ ends are linked in a non-collinear way by a process termed “back-splicing”.8 After its first discovery in 1976 in RNA viruses,9 circRNA was later reported in eukaryotes in 1979.10 Due to the recent progress in high-throughput sequencing technology, circRNAs are found in large quantities in eukaryotes,11 especially in the brains of mammals.12 According to some reports, circRNAs are considered relevant to human neurodegenerative disease.13 During pre-RNA splicing, the 5′ and 3′ ends of exon(s) can be covalently ligated to form circRNAs through back-splicing. circRNAs are highly expressed in human cells and not easily degraded by the exonuclease, and therefore, they exist more stably in humans compared with linear RNAs, with a highly conserved sequence. circRNAs are featured by specificity to tissue/cell types and high stability, which make them suitable as biomarkers.14 Since circRNAs are stably and specifically expressed in different diseases and tissues, such as breast cancer, lung cancer, colorectal cancer, liver cancer, gastric cancer and non-small cell lung cancer, they can be used as novel biomarkers and therapeutic targets for cancers.15,16 In the present study, circRNA expressions were detected in the bone marrow tissues from patients with MM and iron deficiency anemia (IDA), and the clinical value of the candidate circRNAs was analyzed. It was found that hsa_circRNA_101237 may be a potential diagnostic biomarker for MM.
Materials and Methods
Clinical Data
From January 2012 to January 2019, 143 MM patients (65 first-episode treatment-naive patients and 78 patients with recurrent/refractory disease) and 23 IDA patients treated at the Hematology Department of the Third Xiangya Hospital of Central South University were recruited. They were divided into the experimental group (i.e., MM group) and control group, respectively. Bone marrow samples (cryopreserved at −80°C) were collected from both groups. The clinical data (including age, gender, MM classification, staging and cytogenetic abnormalities) and laboratory indicators were also collected from the experimental group.
Cell Culture
All cell lines, including THP-1 human monocytic cell line, MM cell lines MM.1S and H929, were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). The bortezomib concentration was gradually increased to obtain the bortezomib-resistant MM cell lines MM.1S/BTZ and H929/BTZ. All cells were cultured in the 1640 medium (HyClone, Logan, UT, USA) and incubated in a 5% CO2 incubator at 37°C. The log-phase cells were harvested.
RNA Extraction
Plasma cells in the bone marrow samples from MM and IDA patients were enriched by magnetic-activated cell sorting (MACS) using CD138 magnetic l’ bead. Total RNA was extracted from the enriched plasma cells using the TRIzol reagent kit (Invitrogen, USA) and preserved at −80°C. The RNA concentration and purity were detected by using a Nanodrop ND-1000 Spectrophotometer and agarose gel electrophoresis, respectively.
circRNA Microarray-Based Sequencing
Linear RNAs in the total extracted RNA were degraded using RNase R (EPicentre, USA). The residual circRNA was amplified and fluorescent cRNA was obtained by transcription. The labeled circRNA were hybridized to the Arraystar Human circRNA Array V2 (8x15K, Arraystar). The microarray was scanned with Agilent Scanner G2505C. Image analysis was conducted using Agilent Feature Extraction software (version 11.0.1.1). circRNA data were analyzed using the GeneSPring 13.0 software (Agilent). Differentially expressed circRNAs were defined as those with fold change>2.0 and P<0.05.
Verification by Quantitative RT-PCR
RNA was reversely transcribed into cDHA by SuPerScriPt Reverse TranscriPtase (Invitrogen, USA). Then circRNA expressions were detected by real-time qRT-PCR (Arraystar) on the ViiA 7 Real-time PCR System (APPlied Biosystems). Each reaction system was 10μL, consisting of 5u 2×MasterMix, 1μL/PCR specific primers F/R, and 2μL cDNA. The reaction system was added to the well-plate for PCR reaction. β-actin was used as internal control for qPCR, and the relative expression level of circRNA was reflected by the ΔCt method. All primer sequences in this paper were designed by primer 5 and synthesized by Shanghai Biotech. The primer sequences are shown in Table 1.
 |
Table 1 The Primer of three circRNAs Choose to Validate by qRT-PCR |
Statistical Analysis
All statistical analyses were performed using SPSS 20.0 software. Data were expressed as means±standard deviation. Two groups’ means were compared by t-test. ANOVA was performed to compare the means between multiple groups. The reduction rate of M protein was compared by chi-square test. Receiver operating characteristic curve (ROC) was drawn to determine the diagnostic value of circRNA. When AUC was 0.5, it was considered that circRNA had no diagnostic value. The Kaplan-Meier (K-M) survival curve was plotted and Log rank test was used to analyze whether there was significant difference in the survival rate between patients with high and low circRNA expressions. P<0.05 indicated significant difference.
Results
Selection and Validation of circRNAs
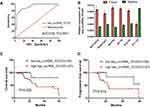
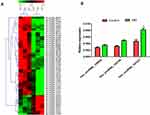
According to the results of high-throughput circRNA microarray-based sequencing in 3 MM patients and 3 IDA patients, 147 circRNAs were differentially expressed (fold change>2). Figure 1A shows the differentially expressed circRNAs, including 40 upregulated and 10 downregulated circRNAs. From them 3 most differentially expressed circRNAs were chosen (Table 2) for qRT-PCR (20 samples for MM and IDA patients, respectively). The results are shown in Figure 1B. It can be seen that hsa_circRNA_101237 was upregulated most significantly and its expression was significantly higher than that in the control group.
 |
Table 2 The Basic Information of three circRNAs Choose to Validate by qRT-PCR |
hsa_circRNA_101237 Overexpression in MM Patients and MM Cell Lines
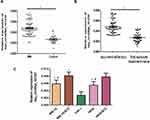
In order to further verify the results of qRT-PCR, hsa_circRNA_101237 expression levels were determined in the bone marrow tissues of 120 MM patients (Figure 2A). The results were consistent with our previous results of qRT-PCR verification, and the hsa_circRNA_101237 expression was significantly higher than that in the control group. This indicated that hsa_circRNA_101237 was stably overexpressed in MM patients and might serve as a biomarker for MM. In the next step, 143 MM patients were divided into first-episode treatment-naive group (n=65) and recurrent/refractory group (n=78). The hsa_circRNA_101237 expressions were further detected in the bone marrow tissues of the two groups (Figure 2B). The results showed that the hsa_circRNA_101237 expression in the recurrent/refractory patients was significantly upregulated compared with the first-episode treatment-naive patients, indicating that hsa_circRNA_101237 may be closely associated with the development of MM. The hsa_circRNA_101237 expression levels were also determined in the THP-1 cells and MM.1S, H929, MM.1S/BTZ and H929/BTZ cells (Figure 2C). It was found that the hsa_circRNA_101237 expression was the highest in the resistant cell lines MM.1S/BTZ and H929/BTZ, followed by the MM cell lines MM.1S and H929, and it was the lowest in the THP-1 cells, with significant differences. The above facts suggested that the hsa_circRNA_101237 upregulation may be one mechanism of bortezomib resistance in MM patients.
Correlation Between the hsa_circRNA_101237 Expression and Clinical Symptoms of MM Patients
In order to better understand the clinical value of hsa_circRNA_101237, we analyzed the correlations between the expression level of hsa_circRNA_ 101237 and the pathological features and laboratory indicators of MM patients, as shown in Table 3. It can be seen that the hsa_circRNA_101237 expression in IgG and IgA MM increased (P<0.001). In MM patients with cytogenetic abnormalities and bone destruction which were of R-ISS stage III, hsa_circRNA_101237 was overexpressed (P<0.05). In addition, hsa_circRNA_101237 was significantly upregulated in MM patients with overexpression of β2-microglobulin and lactate dehydrogenase (P<0.05).
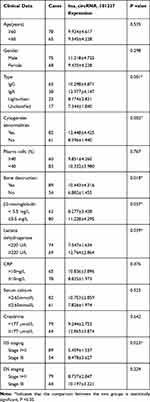 |
Table 3 A Correlation Analysis Between the Clinical Data of 143 MM Patients and hsa_circRNA_101237 Expressions |
hsa_circRNA_101237 Could Be Used as a Diagnostic and Prognostic Indicator for MM
In order to determine the diagnostic value of hsa_circRNA_101237 for MM patients, the ROC curve was drawn for MM. The results are shown in Figure 3A. hsa_circRNA_101237 had a high diagnostic accuracy for MM, and its AUC was 0.92 (P<0.0001). This indicated that hsa_circRNA_101237 may be a specific and sensitive diagnostic biomarker for MM. The prognosis of MM patients can be influenced by cytogenetic abnormalities. The expressions of hsa_circRNA_101237 in MM patients harboring different chromosomal or genetic variations were analyzed, and the results are shown in Figure 3B. It was found that the hsa_circRNA_101237 expression was significantly increased in patients positive for 13q14 deletion, 1q21 amplification, P53 deletion and t(4,14) and t(14,16) gene mutations (P<0.05), but it was decreased significantly in those positive for t(11,14) gene mutations (P<0.05). The correlation between hsa_circRNA_101237 expressions and prognosis of MM patients was further analyzed through the survival curve. Survival analysis indicated that the overall survival (OS) and progression-free survival (PFS) were shortened significantly in those with upregulated hsa_circRNA_101237 (Figure 3C and D, P<0.05).
miRNA and mRNA Target Prediction of hsa_circRNA_101237
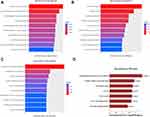
Competitive inhibition of miRNA by the sponging effect is the most important working mechanism of circRNA. In order to explore the possible working mechanism of hsa_circRNA_101237, TargetScan and miRanda were used to predict the miRNA and mRNA targets that might interact with hsa_circRNA_101237. As shown in Figure 4, 11 differentially expressed miRNA and 10 candidate mRNAs interacted with hsa_circRNA_101237. Later, GO (Gene ontology) enrichment analysis was performed to investigate the possible roles of hsa_circRNA_101237 in the cellular activities, including biological processes (Figure 5A), cell components (Figure 5B) and cell functions (Figure 5C). Figure 5D shows the possible signal pathways related to hsa_circRNA_101237 revealed by the KEGG (Kyoto Encyclopedia of Genes and Genomes) analysis, including the PI3K-Akt signaling pathway, chemokine signaling pathway, cell cycle-related pathway, and the signaling pathway related to the interactions between cytokines and their receptors.
 |
Figure 4 Functional analysis of circRNA. hsa_circRNA_101237‐miRNA‐mRNA coexpression network established using hsa_circcircRNA_101237. |
hsa_circRNA_101237 Overexpression Influenced the Treatment Response of MM Patients
One hundred forty-three patients were enrolled for efficacy evaluation, including 79 patients in the bortezomib-containing treatment group (BD group) and 64 patients in the non-bortezomib-containing treatment group (TT group). After 4 cycles of bortezomib-containing treatment, patients with a low expression of hsa_circRNA_101237 showed a higher M protein reduction rate compared to those receiving non-bortezomib-containing treatment (78.1% vs. 32.5%, p<0.001); for patients with an overexpression of hsa_circRNA_101237, after four cycles of bortezomib-containing treatment, the M protein reduction rate was lower compared to those receiving non-bortezomib-containing treatment (38.3% vs. 61.4%, p<0.05) (Figure 6). It was indicated that the hsa_circRNA_101237 expression may influence the responsiveness of MM patients to bortezomib treatment.
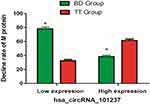 |
Figure 6 Influence of hsa_circRNA_101237 expressions on the responsiveness to bortezomib-containing treatment. *P<0.05, compared with the TT group. |
Discussion
MM is a hematological malignancy featured by abnormal proliferation of plasma cells in the bone marrow,17 which may lead to a variety of symptoms, including hypercalcemia, anemia, renal failure, lytic bone lesions, infection, neurologic symptoms and amyloidosis. MM is more common in the middle-aged and elderly males,18 and its incidence has been increasing gradually as China enters the aging society. MM is a highly heterogeneous disease, and the patients’ survival varies considerably with clinical manifestations, chromosomal and genetic variations. So far, there are no treatment modalities to cure such a high malignant disease as MM. Developing novel early diagnostic, prognostic and therapeutic targets is crucial for MM treatment.
circRNA is a special class of endogenous noncoding RNAs widely present in the organisms and mainly composed of exons and (or) introns. circRNAs regulate the expressions of target genes primarily through the sponging effect of miRNA,19 and play an important role in disease development and progression, especially cancers. The use of second-generation sequencing technology and microarray analysis has resulted in the identification of more and more differentially expressed circRNAs, which are stably expressed in many tumor cells and can be used as a novel marker for tumors. For example, Chen et al20 reported the use of hsa_circ_0000190 as a biomarker for gastric cancer, the sensitivity and specificity of which even surpassed those of carcinoembryonic antigen (CEA) and CA19-9. Yao et al21 proposed that circ RNA_100876 was closely associated with non-small cell lung cancer and that its overexpression predicted shorter survival. Gao et al22 and Tian et al23 identified circRNA1656 and hsa_circ_0004585 as the biomarkers for ovarian cancer and colorectal cancer, respectively, by using microarray analysis. Besides, other studies have indicated that tumor cells contain thousands of circRNA transcripts, which account for the majority of all transcripts. This fact implies the potential of circRNAs as novel diagnostic biomarkers and therapeutic targets for cancers.24 The present study showed that hsa_circRNA_101237 was overexpressed in the bone marrow tissues of MM patients and might serve as a novel diagnostic and prognostic biomarker.
By high-throughput circRNA microarray-based sequencing, the differentially expressed circRNAs in the bone marrow of MM patients were identified, from which the three most significantly differentially expressed circRNAs were selected for qRT-PCR and verification. The results showed that hsa_circRNA_101237 was significantly upregulated. Our study was the first attempt to demonstrate that hsa_circRNA_101237 was upregulated in MM patients, and the diagnostic and prognostic value of hsa_circRNA_101237 in MM patients was investigated. Combining with the clinical data, it was found that the hsa_circRNA_101237 expression was closely associated with MM typing, bone destruction, chromosomal and genetic variations. It can be inferred that hsa_circRNA_101237 was clinically valuable. Chromosomal or genetic variations may affect the prognosis of MM patients. In order to investigate whether hsa_circRNA_101237 influences the prognosis of MM patients, the hsa_circRNA_101237 expressions in patients with different chromosomal or genetic variations were further analyzed. The results showed that hsa_circRNA_101237 was upregulated in patients positive for 13q14 deletion, 1q21 amplification, P53 deletion, and t(4,14) and t(14,16) gene mutations, while it was contrary for t(11,14) gene mutations. It has been long confirmed that the genes mentioned above are closely associated with the prognosis of MM patients. For example, Jung et al25 showed that del (13q) was the only important genetic factor influencing OS of the first-episode MM patient. Grzasko et al26 believed that compared with 13q14 deletion alone, 13q14 deletion with 1q21 amplification could significantly decrease OS of the patients. Studies by Oh et al27 and Shin et al28 confirmed that the poor prognosis of MM patients was associated with t(4,14) gene mutations. In addition, it was also found that the hsa_circRNA_101237 expression in recurrent/refractory MM patients was higher than that in the first-episode treatment-naive patients. It was indicated that hsa_circRNA_101237 was closely associated with the progression of MM.
Bortezomib is a novel inhibitor targeting protease, and its combination with conventional chemotherapeutic agents can increase the overall response of newly diagnosed MM (NDMM), improving the patients’ life quality and prolonging OS.29 Bortezomib has also achieved a good efficacy in refractory/recurrent MM (RRMM).30 However, the long-term use of bortezomib may finally result in the resistance of MM cells. To better understand the resistance mechanism, we analyzed the correlation between hsa_circRNA_101237 and bortezomib resistance. Cell experiments showed that the hsa_circRNA_101237 was increased significantly in bortezomib-resistant cell lines and that hsa_circRNA_101237 overexpression was associated with a poor response to bortezomib-containing chemotherapy in MM patients. Moreover, the M protein reduction rate decreased after treatment. The results above suggested that hsa_circRNA_101237 was possibly involved in bortezomib resistance in MM patients.
Competitive inhibition of miRNA as the molecular sponge is the most important working mechanism of circRNA. It has been found that circRNAs adsorb specific miRNAs to influence the expression of mRNA, thereby performing their biological functions.31,32 Our bioinformatic analysis identified 11 miRNAs and 10 candidate mRNAs which might interact with hsa_circRNA_101237. It was also found that hsa_circRNA_101237 might be involved in various cellular activities and signaling pathways, which further contributes to the development and occurrence of MM. These results provide a novel clue to further investigate the pathogenic and chemotherapeutic resistance mechanism of hsa_circRNA_101237 in MM.
In this study, hsa_circRNA_101237 overexpression in MM patients was found and verified by microarray-based sequencing, which was associated with development, prognosis and responsiveness of MM to bortezomib treatment. Thus hsa_circRNA_101237 may be used as a novel diagnostic and prognostic biomarker and potential therapeutic target for MM. However, more efforts should be made to develop novel biomarkers for MM and improve the diagnosis, treatment and prognosis of MM.
Ethics Approval and Informed Consent
This study was approved by the Ethics Committee of The Third Xiangya Hospital, Central South University. Informed consent was obtained from each subject in accordance with the Declaration of Helsinki.
Author Contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Funding
This work was supported by The New Xiangya Talent Project of the Third Xiangya Hospital, Central South University, China (No. 20170307).
Disclosure
The authors report no declarations of interest in this work.
References
1. Rajkumar SV, Kumar S. Multiple myeloma: diagnosis and treatment. Mayo Clin Proc. 2016;91(1):101–119. doi:10.1016/j.mayocp.2015.11.007
2. Shah V, Sherborne AL, Walker BA, et al. Prediction of outcome in newly diagnosed myeloma: a meta-analysis of the molecular profiles of 1905 trial patients. Leukemia. 2018;32(1):102–110. doi:10.1038/leu.2017.179
3. Chan NC, Chan NP. Recurrent cytogenetic abnormalities in multiple myeloma. Methods Mol Biol. 2017;1541:295–302.
4. Lakshman A, Alhaj Moustafa M, Rajkumar SV, et al. Natural history of t(11;14) multiple myeloma. Leukemia. 2018;32(1):131–138. doi:10.1038/leu.2017.204
5. Walker BA, Boyle EM, Wardell CP, et al. Mutational spectrum, copy number changes, and outcome: results of a sequencing study of patients with newly diagnosed myeloma. J Clin Oncol. 2015;33(33):3911–3920. doi:10.1200/JCO.2014.59.1503
6. Lonial S, Boise LH, Kaufman J. How I treat high-risk myeloma. Blood. 2015;126:1536–1543. doi:10.1182/blood-2015-06-653261
7. Chim CS, Kumar SK, Orlowski RZ, et al. Management of relapsed and refractory multiple myeloma: novel agents, antibodies, immunotherapies and beyond. Leukemia. 2018;32(2):252–262. doi:10.1038/leu.2017.329
8. Lasda E, Parker R. Circular RNAs: diversity of form and function. RNA. 2014;20:1829–1842. doi:10.1261/rna.047126.114
9. Sanger HL, Klotz G, Riesner D, et al. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A. 1976;73:3852–3856. doi:10.1073/pnas.73.11.3852
10. Hsu MT, Coca-Prados M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature. 1979;280:339–340. doi:10.1038/280339a0
11. Xia S, Feng J, Chen K, et al. CSCD: a database for cancer-specific circular RNAs. Nucleic Acids Res. 2018;46(D1):D925–D929. doi:10.1093/nar/gkx863
12. You X, Vlatkovic I, Babic A, et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci. 2015;18:603–610. doi:10.1038/nn.3975
13. Kumar L, Haque S, Haque R, et al. Circular RNAs: the emerging class of non-coding RNAs and their potential role in human neurodegenerative diseases. Mol Neurobiol. 2017;54(9):7224–7234. doi:10.1007/s12035-016-0213-8
14. Kristensen LS, Hansen TB, Veno MT, et al. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene. 2018;37(5):555–565. doi:10.1038/onc.2017.361
15. Meng S, Zhou H, Feng Z, et al. CircRNA: functions and properties of a novel potential biomarker for cancer. Mol Cancer. 2017;16(1):94. doi:10.1186/s12943-017-0663-2
16. Yang Z, Xie L, Han L, et al. Circular RNAs: regulators of cancer-related signaling pathways and potential diagnostic biomarkers for human cancers. Theranostics. 2017;7:3106–3117. doi:10.7150/thno.19016
17. Manier S, Salem KZ, Park J, et al. Genomic complexity of multiple myeloma and its clinical implications. Nat Rev Clin Oncol. 2017;14(2):100–113. doi:10.1038/nrclinonc.2016.122
18. Suzuki K, Takahashi H. The epidemiology of multiple myeloma. Nihon Rinsho. 2015;73(1):7–12.
19. Karreth FA, Tay Y, Perna D, et al. In vivo identification of tumor-suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell. 2011;147(2):382–395. doi:10.1016/j.cell.2011.09.032
20. Chen S, Li T, Zhao Q, et al. Using circular RNA hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin Chim Acta. 2017;466:167–171. doi:10.1016/j.cca.2017.01.025
21. Yao JT, Zhao SH, Liu QP, et al. Over-expression of Circ RNA_100876 in non-small cell lung cancer and its prognostic value. Pathol Res Pract. 2017;213(5):453–456. doi:10.1016/j.prp.2017.02.011
22. Gao Y, Zhang C, Liu Y, et al. Circular RNA profiling reveals circRNA1656 as a novel biomarker in high grade serous ovarian cancer. Biosci Trends. 2019;13(2):204–211. doi:10.5582/bst.2019.01021
23. Tian J, Xi X, Wang J, et al. CircRNA hsa_circ_0004585 as a potential biomarker for colorectal cancer. Cancer Manag Res. 2019;11:5413–5423. doi:10.2147/CMAR.S199436
24. Chen Z, Zhang L, Han G, et al. A meta-analysis of the diagnostic accuracy of circular RNAs in digestive system malignancy. Cell Physiol Biochem. 2018;45(3):962–972. doi:10.1159/000487291
25. Jung HA, Jang M-A, Kim K, et al. Clinical utility of a diagnostic approach to detect genetic abnormalities in multiple myeloma: a single institution experience. Ann Lab Med. 2018;38(3):196–203. doi:10.3343/alm.2018.38.3.196
26. Grzasko N, Hajek R, Hus M, et al. Chromosome 1 amplification has similar prognostic value to del(17p13) and t(4;14)(p16;q32) in multiple myeloma patients: analysis of real-life data from the Polish Myeloma Study Group. Leuk Lymphoma. 2017;58(9):1–15. doi:10.1080/10428194.2016.1272684
27. Oh S, Koo DH, Kwon M-J, et al. Chromosome 13 deletion and hypodiploidy on conventional cytogenetics are robust prognostic factors in Korean multiple myeloma patients: web-based multicenter registry study. Ann Hematol. 2014;93(8):1353–1361. doi:10.1007/s00277-014-2057-5
28. Shin SY, Eom HS, Sohn JY, et al. Prognostic implications of monosomies in patients with multiple myeloma. Clin Lymphoma Myeloma Leuk. 2017;17(3):159–164. doi:10.1016/j.clml.2016.12.001
29. San Miguel JF, Schlag R, Khuageva NK, et al. Bortezomib plus melphalan and prednisone for initial treatment of multiple myeloma. N Engl J Med. 2008;359(9):906–917. doi:10.1056/NEJMoa0801479
30. Terpos E, Katodritou E, de la Rubia J, et al. Bortezomib-based therapy for relapsed/refractory multiple myeloma in real-world medical practice. Eur J Haematol. 2018;101(4):556–565. doi:10.1111/ejh.2018.101.issue-4
31. Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333–338. doi:10.1038/nature11928
32. Hansen TB, Jensen TI, Clausen BH, et al. Natural RNA circles function as efficient micro RNA sponges. Nature. 2013;495(7441):384–388. doi:10.1038/nature11993
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.