Back to Journals » Clinical Epidemiology » Volume 10
How well can familial hypercholesterolemia be identified in an electronic health record database?
Authors Mues KE, Bogdanov AN, Monda KL, Yedigarova L, Liede A, Kallenbach L
Received 11 June 2018
Accepted for publication 14 September 2018
Published 15 November 2018 Volume 2018:10 Pages 1667—1677
DOI https://doi.org/10.2147/CLEP.S176853
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Vera Ehrenstein
Katherine E Mues,1 Alina N Bogdanov,2 Keri L Monda,1 Larisa Yedigarova,3 Alexander Liede,1 Lee Kallenbach2
1Center for Observational Research, Amgen Inc., Thousand Oaks, CA, USA; 2Practice Fusion, San Francisco, CA, USA; 3US Medical Affairs, Amgen Inc., Thousand Oaks, CA, USA
Background: Familial hypercholesterolemia (FH) is a condition characterized by high cholesterol levels and increased risk for coronary heart disease (CHD) that often goes undiagnosed. The Dutch Lipid Network Criteria (DLNC) are used to identify FH in clinical settings via physical examination, personal and family history of CHD, in addition to the presence of deleterious mutations of the LDLR, ApoB, and PCSK9 genes. Agreement between clinical and genetic diagnosis of FH varies. While an ICD diagnosis code was not available for coding FH until 2016, Systematized Nomenclature of Medicine (SNOMED) clinical concept codes, including genetic diagnoses, for FH have been utilized in electronic health records (EHRs).
Objective: To evaluate the concordance of identifying FH via SNOMED and ICD-10 CM codes vs the DLNC in an EHR database.
Methods: Using the Practice Fusion EHR database, the sensitivity, specificity, positive predictive value (PPV), and negative predictive value were calculated comparing an FH cohort identified via SNOMED and ICD-10 CM codes to one identified via the DLNC.
Results: Among 907,616 patients with hypercholesterolemia, 2,180 were identified as FH via SNOMED code (zero were identified via ICD-10 CM), 259 had a DLNC score 6–8 (probable FH), and 45 had a DLNC score >8 (definite FH). Compared to DLNC score >8, the sensitivity, specificity, and PPV of the FH SNOMED code were 84.4%, 99.4%, and 6.4%, respectively. Compared to DLNC score ≥6, the sensitivity was 36.8% and the specificity was 99.5% with a PPV of 18.7%.
Conclusion: Compared to the clinical criteria for FH, identification of FH patients via SNOMED diagnosis codes had high sensitivity and specificity, but low PPV. The discordance of these two techniques in identifying FH patients speaks to the challenges in identifying FH patients in large electronic databases such as administrative claims and EHR.
Keywords: familial hypercholesterolemia, Dutch Lipid Network Criteria, electronic health record, SNOMED
Introduction
Familial hypercholesterolemia (FH) is a genetic disorder characterized by elevated cholesterol levels, specifically, high levels of low-density lipoprotein cholesterol (LDL-C) often resulting in premature coronary heart disease (CHD).1 The most recent prevalence estimates of heterozygous FH in the USA suggest between 1 in 212 and 1 in 250 individuals may be affected.2,3 Historically, the prevalence of homozygous FH has been estimated to be 1 in 1,000,000, whereas new research has shown the prevalence may range from 1 in 160,000 to 1 in 300,000 individuals.1,4–7 Nordestgaard et al estimated that there are between 14 and 34 million individuals living with FH worldwide.1 Given the underutilization of genetic testing and the lack of awareness of FH in the clinical community, even the most current prevalence estimates may under-represent true prevalence of this condition in the general population. Thus, many individuals may not be getting the appropriate therapies to control their LDL-C.
Genetic variants leading to FH are present in the LDLR, APOB, or PCSK9 genes, accounting for >90%, 5%, and 1% of FH cases, respectively.5,8–11 However, FH is most frequently diagnosed clinically based on elevated levels of LDL-C, premature CHD, family history of cardiovascular disease (CVD) and/or high LDL-C, and physical signs of elevated cholesterol.4 Although a positive genetic test for FH is conclusive evidence of the disease, FH cannot be ruled out in the result of a negative genetic test. In fact, ~20% of clinically evident FH patients do not appear to have a mutation identified in the FH-associated genes.12 A recent study by Khera et al found that ~2% of individuals free of coronary artery disease (CAD) with LDL-C ≥190 mg/dL (identified as FH patients based solely on LDL-C) tested positive for a mutation in one of the FH-associated genes.13 The same study also found the risk of CAD was higher among individuals with a genetic mutation for FH, even within LDL-C strata, suggesting that cumulative lifelong exposure to elevated LDL-C is associated with a higher CAD risk.
Concordance between the genetic and clinical diagnosis of FH varies widely due to inconsistencies in how FH is clinically diagnosed as well as lack of full understanding of the genetic basis of the disease.13–16 There are several existing clinical criteria used to diagnosis FH: 1) the Dutch Lipid Network Criteria (DLNC); 2) the Simon Broome criteria; and 3) the Make Early Diagnosis to Prevent Early Death (MEDPED) criteria.1,5,15,17 These three sets of criteria are similar in their diagnostic parameters and are generally viewed as interchangeable by clinicians. The DLNC is perhaps the most widely recognized score and uses five criteria (both clinical and genetic) to identify FH: family history of CVD, clinical history of premature CHD, physical examination for xanthomas or corneal arcus, very high LDL-C on repeated measurements, and a causative mutation detected by genetic testing.
Until the recent introduction of the ICD, 10th Revision (ICD-10 CM-CM) code for FH in October 2016 (ICD-10 CM E78.01), an ICD diagnosis code for FH did not exist in the USA, making it difficult to identify individuals with FH within large healthcare databases. However, Systematized Nomenclature of Medicine (SNOMED) clinical concept codes for FH do exist. SNOMED is a comprehensive and precise clinical coding system that was developed and supported by the International Health Terminology Standards Organization and the US National Library of Medicine as a standard for electronic exchange of clinical health information.18 The Office of the National Coordinator for Health Information Technology has approved SNOMED as a vocabulary code set to ensure consistent clinical terminology between systems.19 As such, SNOMED codes are often implemented and mapped to other coding systems within EHR software applications. The SNOMED system includes a code for FH, as well as specific codes for heterozygous and homozygous FH, and FH associated with a mutation in a specific gene (LDLR, APOB, and PCSK9).
The recent introduction of the ICD-10 CM code for FH and the increased utilization of SNOMED codes provide an opportunity to more easily identify FH patients in EHR systems. However, it is unclear whether physicians are utilizing these codes while doing a clinical work-up for FH through criteria such as the DLNC. For example, providers may enter in clinical components of the DLNC without explicitly recording that the patient has FH; alternatively, providers may record that the patient has FH without explicitly recording the clinical components that comprise it. Thus, our objective was to evaluate the concordance of FH diagnosis codes with DLNC clinical criteria for FH within an EHR database.
Methods
Patient population
We used data from the Practice Fusion EHR database that consists of patient-level data from their proprietary cloud-based ambulatory EHR platform, currently in use at over 30,000 practices in all 50 US states. A majority of Practice Fusion practices are single provider or small group practices. Data are available for over 38 million unique patients starting in 2010, of which over 16 million are currently active on the platform. Data are updated daily and are made available for analysis in a Health Information Privacy Protection Act-compliant de-identified research database. Substantial data validation and quality checks are performed as part of this process. Amgen licensed the Practice Fusion EHR data for analytic use. Institutional review board approval was not necessary for this study, as we did not intervene on patients whose data are included in the Practice Fusion EHR.
SNOMED codes were introduced into the Practice Fusion EHR platform in 2013. When entering a diagnosis into a structured field, Practice Fusion EHR users search for clinically meaningful terms related to a disease condition, for example, “familial hypercholesterolemia”. From there, a drop-down list of matching diagnosis codes appears. A physician will then choose the code that best captures the patient’s diagnosis, which is then automatically mapped to all other corresponding diagnosis codes. For example, a physician may type in “familial hypercholesterolemia” into the record. A drop-down list of diagnosis codes appears from which a physician chooses the ICD-10 CM code for FH. It is automatically mapped to the corresponding SNOMED and IC-9 codes (Figure S1). As opposed to claims-based coding, the recording of diagnosis in an EHR is not related to billing and reimbursement.
For this analysis, four patient cohorts of interest were identified between January 1, 2013 and October 16, 2016. The hypercholesterolemia cohort was defined as the source population by the presence of at least one of the following ICD diagnosis codes for pure hypercholesterolemia: 272.0, E78.0. Within the hypercholesterolemia cohort, the FH SNOMED/ICD-10 CM cohort was defined by the presence of either a SNOMED or ICD-10 CM diagnosis code for FH (Table 1). Also within the hypercholesterolemia cohort, the FH DLNC cohort was defined via diagnoses and LDL-C lab test results that identify patients who meet the DLNC for FH. A patient receives a specific number of points for each DLN criterion that is present, which are then summed into an overall score. The score is categorized into the following groups: >8 definite FH; 6–8, probable FH; 3–5, possible FH; and <3, unlikely FH. See Table S1 for corresponding SNOMED, Logical Observation Identifiers Names and Codes, ICD-10 CM, and ICD-9 codes used to identify the DLNC.
  | Table 1 SNOMED and ICD-10 codes for familial hypercholesterolemia Abbreviation: SNOMED, Systematized Nomenclature of Medicine. |
Statistical analysis
Within each cohort, we calculated the distribution of patient demographics, number of patient visits, available follow-up time in the database, prescribed medications, LDL-C values, and other lab values (high-density lipoprotein cholesterol [HDL], triglycerides, total cholesterol, and LDL-C/HDL-C ratio). We evaluated the prescription of lipid lowering therapy (LLT) (statins, ezetimibe, and PCSK9 inhibitors) and other concomitant medications (insulin and other glucose lowering agents, as well as antihypertensives) at any point in the patient’s record. We evaluated both the highest available and most recent LDL-C value in a patient’s record. For categorical variables, we calculated the number and percent of patients within each cohort with the variable of interest; for continuous variables, we calculated mean and SD within each cohort.
In this analysis, we compared a diagnosis of FH recorded via a SNOMED or ICD-10 CM code to the DLNC clinical diagnosis of FH. The sensitivity (Se), specificity (Sp), positive predictive value (PPV), negative predictive value (NPV), and the corresponding 95% CIs of identifying FH patients via SNOMED/ICD-10 CM codes vs several cut points of the DLNC score (DLNC >8, DLNC ≥6, and DLNC ≥3) were calculated. The calculation of Se, Sp, PPV, and NPV was limited to patients meeting at least one of the DLNC components. All analyses were conducted in IBM Netezza SQL.
Results
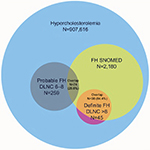
In the Practice Fusion database, 907,616 patients with hypercholesterolemia were identified between January 1, 2013 and October 16, 2016. Of these patients, 2,180 (0.24%) were identified as FH via an SNOMED code (there were no records of an ICD-10 CM code for FH), 96,392 (10.6%) had at least one criterion of the DLNC, 259 (0.03%) had a DLNC score 6–8 indicating probable FH, and 45 (0.005%) had a DLNC score >8 indicating definite FH (Table 2). Among patients identified as definite FH, 38 (84.4%) were also identified as FH via SNOMED codes, while among patients identified as probable FH, 74 (28.6%) were also identified as FH via SNOMED codes (Figure 1).
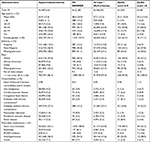
Table 2 summarizes the baseline demographic and clinical characteristics among patients in the hypercholesterolemia cohort, FH SNOMED cohort, those meeting at least 1 of the DLN criterion, those with a DLNC score 6–8, and those with a DLNC score >8. Individuals identified as FH via SNOMED code and via the DLNC were younger and a higher proportion were female than those in the source hypercholesterolemia cohort. The average number of visits in the EHR during the study period ranged from 9.5 among the FH SNOMED cohort to 21.0 among those with DLNC score >8. The average Quan Enhanced Charlson comorbidity score was highest among those with DLNC score >8 and DLNC score 6–8, followed by the hypercholesterolemia cohort, those with at least 1 DLN criterion, and the FH SNOMED cohort. Prescription of statins ranged from 48.4% among those in the FH SNOMED cohort to 68.7% among those with a DLNC score 6–8 (Table 2). Prescriptions for nonstatin LLTs were most frequent among those with a DLNC >8.
Patients who had a DLNC score 6–8 and >8 had the highest mean LDL-C values (Table 3), and ~62% of patients with a DLNC score 6–8 had an LDL-C value of at least 330 mg/dL (data not shown). The mean LDL-C value among those in the FH SNOMED cohort and those meeting at least one DLN criterion was higher than among those in the hypercholesterolemia cohort (for both most recent and highest LDL-C values) (Table 3). The mean levels of triglycerides, total cholesterol, and the LDL/HDL ratio were relatively higher among patients with DLNC score 6–8 and >8 compared to the other cohorts.
Among the full hypercholesterolemia cohort, 10.6% (N=96,392) had at least one DLN criterion compared with 27.4% (N=598) among the FH SNOMED cohort. Among hypercholesterolemia patients with at least one of the DLN criterion, LDL-C between 155 and 189 mg/dL (58.0%) was the criterion most frequently met, followed by LDL-C 190–249 mg/dL (18.3%), clinical history of premature CAD (11.2%), and clinical history of premature cerebral or peripheral vascular disease (9.1%) (Figure 2). Among FH SNOMED patients with at least one of the DLN criterion, LDL-C 190–249 mg/dL was the most frequently met (33.9%), followed by LDL-C 155–189 mg/dL (27.9%), a positive genetic test (15.6%), clinical history of premature CAD (10.9%), LDL-C 250–239 mg/dL (9.7%), and family history of premature coronary and vascular disease or LDL-C above the 95th percentile (9.4%).
Among the 96,392 patients with at least one DLN criterion, 45 had a DLNC score >8, 304 had a DLNC score ≥6, 20,048 had a DLNC score ≥3, and 598 were identified as FH via an SNOMED code. Compared to the DLNC >8, the sensitivity of the FH SNOMED code was 84.4% and the specificity was 99.4%, with a PPV of 6.4% (Table 4). Compared to a DLNC ≥6, the sensitivity was 36.8% and the specificity was 99.5% with a PPV of 18.7%.
Discussion
Compared to the clinical criteria for FH, the identification of FH patients via SNOMED diagnosis codes had high sensitivity and specificity, but low PPV. These results indicate that FH patients identified via SNOMED codes are unlikely to be identified using clinical criteria contained in their EHR, even at several cutoff levels of the DLNC. The discordance of these two techniques in identifying FH patients speaks to the challenges in identifying FH patients in large electronic databases such as administrative claims and EHR.
If we consider clinical criteria as the gold standard of an FH diagnosis, standard coding such as SNOMED not only identifies a large percentage of “true FH” patients but also identifies patients who do not meet the gold standard of FH diagnosis. This may be a true misclassification of these patients via diagnosis codes or may speak to the lack of such clinical criteria being recorded in the structured portions of the medical record. Thus, the diagnosis codes may be accurate, but the gold standard may not be recorded as easily searchable content in an EHR, making identification of FH patients difficult. These findings may also indicate that clinicians are not fully aware of the clinical symptoms of FH (such as tendon xanthoma), and thus are not looking for and noting the clinical signs within the medical record, also contributing to the under-diagnosis of the condition.
Several other studies that have evaluated the application of clinical diagnostic systems in large databases have found similar challenges in identifying FH patients. Abul-Husn et al found that when applying the DLNC to a population of individuals with severe hypercholesterolemia, only 24% of FH variant carriers were classified as either probable or definite FH, highlighting the rather low sensitivity of the DLNC when applied to an EHR database.20 In the same study, of the 4,435 individuals with LDL-C >190 mg/dL in the EHR, only 112 (2.5%) had an FH genetic variant, again indicating the low capture of genetic variants for FH and potentially the underutilization of genetic testing in this population. In a general practitioner electronic healthcare database of patients in Australia, Vickery et al found that among active patients, 0.7% had an LDL-C value ≥5 mmol/L (~190 mg/dL), but only 0.013% had a diagnosis code for FH, again indicating the low diagnosis rates of FH. Among those with an LDL-C ≥5 mmol/L, 60% had been coded for a lipid disorder, 56% had record of a statin prescription, 40% had both, and 24% had neither.21
Although there was no ICD diagnosis code for FH until October 2016, SNOMED codes offer an alternate coding system to identify patients with FH in an electronic database. The SNOMED nomenclature is the most comprehensive healthcare terminology, an international standard for the electronic exchange of clinical health information supported by the US National Library of Medicine, which has previously been incorporated into EHR-based algorithms to identify clinically relevant cardio- and cerebrovascular disease.19,22 Although practitioners may not be as familiar with SNOMED codes as other coding systems (eg, ICD-9/ICD-10 CM, HCPC), one of the strengths of the Practice Fusion data platform was the automatic mapping of physician diagnoses entered into the structured data fields into SNOMED codes. Thus, even if a physician was not familiar with the FH SNOMED codes, the Practice Fusion system automatically mapped each structured diagnosis of FH to an SNOMED code, allowing for efficient identification of FH patients within the database. Although we did use the ICD-10 CM code for identifying FH in this analysis, there were no cases identified in this manner, since the analysis was conducted through mid-October 2016, with limited (only 3 months) post-ICD-10 CM implementation. The hope is that, over time, this code will become more frequently used and awareness of FH will increase, to allow for more accurate diagnosis of FH within EHR systems.
In this analysis, only a small proportion of identified individuals with FH had evidence of a functional mutation on the LDLR, APOB, or PCSK9 gene resulting from a genetic test. These results reiterate the low frequency of and differing opinions on the incremental value of FH genetic testing, even among a population of patients who have a diagnostic code for FH, suggesting that FH remains the first and foremost clinical diagnosis. Genetic testing is expensive and may not be covered by insurance plans.23 Additionally, clinical guidelines do not recommend universal testing, but instead recommend cascade screening among first-degree relatives of an individual with FH, with universal screening limited to children <16 years with high levels of cholesterol.1,12 Finally, due to limitations in FH genetic testing, some individuals who carry a genetic mutation for FH may not be easily identifiable. This can happen for several reasons. First, a standard FH panel only covers a subset of all known variants, generally limited to those that contribute to a more severe disease phenotype. For example, although over 1,500 variants have been identified in the LDLR gene alone, not all of these are covered on a standard genetic array. Second, individuals that are compound heterozygotes (eg, two different mutations affecting two alleles) may display an FH phenotype without an easily identifiable genotype given that multiple low penetrance variants may be present.24
This study has several limitations that should be considered when interpreting the results. First, the true gold standard of an FH diagnosis is unclear. Although we compare the SNOMED codes for FH to the DLNC, and calculate sensitivity and specificity accordingly, we must interpret these findings with the understanding that clinical criteria are not the only method to diagnose FH; genetic testing may have been done and not necessarily recorded within the medical record. Despite this, patients identified as FH via the DLNC were slightly older, more female, had a higher disease burden, more statin and nonstatin LLT prescriptions, and had higher LDL-C values compared with FH patients identified via a SNOMED code. Given these findings, considering the DLNC as the gold standard in this study does not seem unreasonable. Second, the analysis was limited to only structured information captured by the EHR; FH diagnoses and results of genetic testing were not confirmed from unstructured data. We did not incorporate information captured in physician notes into the main analysis. Given that EHRs may still be relatively new to healthcare providers, it is possible that providers may prefer to record much of a patient’s information via free text. Natural language processing is a lengthy and expensive process, limiting our ability to use the unstructured information included in the EHR, and more generally, speaks to the challenge of identifying FH patients in electronic databases. Third, this analysis was limited to information captured within the Practice Fusion EHR platform, which mostly covers primary care physicians, who may be less familiar with a condition such as FH. We do not have health information collected on patients who also sought care outside of Practice Fusion’s network. Although the Practice Fusion network does capture some specialists such as cardiologists within community-based practices, it is possible that, compared to large health centers, these physicians may be less likely to diagnose and/or record a diagnosis of FH. Last, these results are not necessarily applicable to other EHR databases that may capture different types of health practitioners or may be related to an associated payer who may have specific requirements for FH diagnoses in order for appropriate treatments to be covered.
Comparing SNOMED diagnosis codes for FH to the DLNC clinical criteria, the sensitivity and specificity were high, whereas the positive predictive value was low. Thus, if considering the DLNC as the gold standard, the identification of FH patients in an EHR using a SNOMED code may result in the capture of many false positives. It is unclear if this is due to the true misclassification of FH via diagnosis codes or rather to the lack of recording of clinical symptoms of FH in the structured portions of an EHR (eg, not the clinical notes). Future work in the Practice Fusion database may include a comprehensive review of physician notes for identifying subjects with FH. Although EHRs provide an opportunity to extract physician notes with deeper clinical information, for research studies on large population-level databases, natural language processing to extract information from the notes is expensive, time-consuming, and may be limited in its accuracy. Therefore, most research studies using EHR data on a large scale will most likely only be able to use structured information to identify subjects with FH, requiring improved techniques of identifying individuals with FH via structured data. Although the recent introduction of an ICD-10 CM code for FH in the USA will mitigate some of the challenges in identifying FH subjects in EHR and claims databases, the lag in data availability and need for historic data will result in continued challenges in research. However, until FH is more fully recognized by the clinical community, the addition of an ICD-10 CM code will not completely correct the under-diagnosis of FH.
Acknowledgment
Amgen, Inc. funded this study.
Author contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
KEM, KLM, LY, and AL are employees and stockholders of Amgen Inc. ANB and LK are employees of Practice Fusion, which received funding from Amgen to conduct the study. The authors report no other conflicts of interest in this work.
References
Nordestgaard BG, Chapman MJ, Humphries SE, et al. Familial hypercholesterolaemia is underdiagnosed and undertreated in the general population: guidance for clinicians to prevent coronary heart disease: consensus statement of the European Atherosclerosis Society. Eur Heart J. 2013;34(45):3478–3490. | ||
Bucholz EM, Rodday AM, Kolor K, Khoury MJ, de Ferranti SD. Prevalence and predictors of cholesterol screening, awareness, and statin treatment among US adults with familial hypercholesterolemia or other forms of severe dyslipidemia (1999–2014). Circulation. 2018;137(21):2218–2230. | ||
de Ferranti SD, Rodday AM, Mendelson MM, Wong JB, Leslie LK, Sheldrick RC. Prevalence of Familial hypercholesterolemia in the 1999 to 2012 United States National Health and Nutrition Examination Surveys (NHANES). Circulation. 2016;133(11):1067–1072. | ||
Goldstein JK, Hobbs HH, Brown MS. Familial hypercholesterolemia. In: Scriver CR, Beaudet AL, Sly WS, Valle D, editors. The Metabolic & Molecular Bases of Inherited Diseases. 8th ed. New York: McGraw-Hill; 2001:2863–2913. | ||
Austin MA, Hutter CM, Zimmern RL, Humphries SE. Genetic causes of monogenic heterozygous familial hypercholesterolemia: a HuGE prevalence review. Am J Epidemiol. 2004;160(5):407–420. | ||
Benn M, Watts GF, Tybjaerg-Hansen A, Nordestgaard BG. Familial hypercholesterolemia in the danish general population: prevalence, coronary artery disease, and cholesterol-lowering medication. J Clin Endocrinol Metab. 2012;97(11):3956–3964. | ||
Sjouke B, Hovingh GK, Kastelein JJP, Stefanutti C. Homozygous autosomal dominant hypercholesterolaemia. Curr Opin Lipidol. 2015;26(3):200–209. | ||
Brown MS, Goldstein JL, Arnold W, Perdue S. Expression of the familial hypercholesterolemia gene in heterozygotes: mechanism for a dominant disorder in man. Science. 1974;185(4145):61–63. | ||
Soria LF, Ludwig EH, Clarke HRG, Vegat GL, Grundyt SM, Mccarthy BJ. Association between a specific apolipoprotein B mutation and familial defective apolipoprotein B-100 (genetic disease/cholesterol metabolism). Genetics. 1989;86:587–591. | ||
Innerarity TL, Weisgraber KH, Arnold KS, et al. Familial defective apolipoprotein B-100: low density lipoproteins with abnormal receptor binding. Proc Natl Acad Sci U S A. 1987;84(19):6919–6923. | ||
Abifadel M, Varret M, Rabès JP, et al. Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat Genet. 2003;34(2):154–156. | ||
Goldberg AC, Hopkins PN, Toth PP, et al. Familial hypercholesterolemia: screening, diagnosis and management of pediatric and adult patients: clinical guidance from the National Lipid Association Expert Panel on Familial Hypercholesterolemia. J Clin Lipidol. 2011;5(3):133–140. | ||
Khera AV, Won HH, Peloso GM, et al. Diagnostic yield and clinical utility of sequencing familial hypercholesterolemia genes in patients with severe hypercholesterolemia. J Am Coll Cardiol. 2016;67(22):2578–2589. | ||
Civeira FInternational Panel on Management of Familial Hypercholesterolemia, International Panel on Management of Familial Hypercholesterolemia. Guidelines for the diagnosis and management of heterozygous familial hypercholesterolemia. Atherosclerosis. 2004;173(1):55–68. | ||
Williams RR, Hunt SC, Schumacher MC, et al. Diagnosing heterozygous familial hypercholesterolemia using new practical criteria validated by molecular genetics. Am J Cardiol. 1993;72(2):171–176. | ||
Palacios L, Grandoso L, Cuevas N, et al. Molecular characterization of familial hypercholesterolemia in Spain. Atherosclerosis. 2012;221(1):137–142. | ||
Haase A, Goldberg AC. Identification of people with heterozygous familial hypercholesterolemia. Curr Opin Lipidol. 2012;23(4):282–289. | ||
National Institutes of Health. U.S. National Library of Medicine. SNOMED CT. Available from: https://www.nlm.nih.gov/healthit/snomedct/. Accessed January 11, 2017. | ||
Health and Human Services Department. 2015 Edition Health Information Technology (Health IT) Certification Criteria, 2015 Edition Base Electronic Health Record (EHR) Definition, and ONC Health IT Certification Program Modifications; 2015. Available from: https://www.federalregister.gov/documents/2015/10/16/2015-25597/2015-edition-health-information-technology-health-it-certification-criteria-2015-edition-base. Accessed August 14, 2018. | ||
Abul-Husn NS, Manickam K, Jones LK, et al. Genetic identification of familial hypercholesterolemia within a single U.S. health care system. Science. 2016;354(6319):aaf7000. | ||
Vickery AW, Ryan J, Pang J, Garton-Smith J, Watts GF. Increasing the detection of familial hypercholesterolaemia using general practice electronic databases. Heart Lung Circ. 2017;26(5):450–454. | ||
Esteban S, Rodríguez Tablado M, Ricci RI, Terrasa S, Kopitowski K. A rule-based electronic phenotyping algorithm for detecting clinically relevant cardiovascular disease cases. BMC Res Notes. 2017;10(1):281. | ||
Singh S, Bittner V. Familial hypercholesterolemia-epidemiology, diagnosis, and screening. Curr Atheroscler Rep. 2015;17(2):482. | ||
Talmud PJ, Shah S, Whittall R, et al. Use of low-density lipoprotein cholesterol gene score to distinguish patients with polygenic and monogenic familial hypercholesterolaemia: a case-control study. Lancet. 2013;381(9874):1293–1301. |
Supplementary materials
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.