Back to Journals » International Journal of Nanomedicine » Volume 12
Highly sensitive protein detection via covalently linked aptamer to MoS2 and exonuclease-assisted amplification strategy
Authors Gao L , Li Q, Deng Z, Brady B, Xia N, Zhou Y , Shi H
Received 5 July 2017
Accepted for publication 25 September 2017
Published 25 October 2017 Volume 2017:12 Pages 7847—7853
DOI https://doi.org/10.2147/IJN.S145585
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Linlin Sun
Li Gao,1 Qin Li,1 Zebin Deng,1 Brendan Brady,2 Ni Xia,1 Yang Zhou,1 Haixia Shi3
1Institute of Life Sciences, Jiangsu University, Zhenjiang, 2Department of Physics, University of Victoria, Victoria, BC, Canada, 3Department of Physical Education, Dalian Jiaotong University, Dalian, People’s Republic of China
Abstract: Molybdenum disulfide (MoS2) has shown highly attractive superiority as a platform for sensing. However, DNA physisorption on the surface of MoS2 was susceptible to nonspecific probe displacement and false-positive signals. To solve these problems, we have developed a novel MoS2–aptamer nanosheet biosensor for detecting thrombin using a covalently linked aptamer to the MoS2 nanosheet. Ten percent Tween 80 was used to prevent thrombin from nonspecific binding and to rapidly form thiol-DNA/gold nanoparticle (AuNP) conjugates. Furthermore, an MoS2 and exonuclease coassisted signal amplification strategy was developed to improve the detection limit for thrombin. We used the hybridization of the aptamer molecules and the matched strand with a 5' terminal thiol to immobilize the aptamer molecules on the surface of AuNPs in AuNPs@MoS2 nanocomposites. Exonuclease digested the single-strand aptamer and released the thrombin, which was then detected in the next recycle. With the coassisted amplification strategy, a 6 fM detection limit was achieved, showing that this method has higher sensitivity than most reported methods for thrombin detection. The results presented in this work show that this method of covalently attaching the aptamer and using the coassisted amplification is a promising technique for the detection of protein in medical diagnostics.
Keywords: Molybdenum disulfide, aptamer, thrombin, protein detection, high sensitivity
Introduction
Two-dimensional layered structures of molybdenum disulfide (MoS2) with unique characteristics analogous to graphene have recently attracted much attention for energy harvesting and nanoelectronic applications because of their extraordinary thermal conductivity, robustness, and unusual optical and energy harvesting properties.1,2 In contrast to graphene, MoS2 nanosheets hold great promise as novel nanomaterials for biomedical applications as they can be synthesized on a large scale and can be directly dispersed in aqueous solution without the aid of surfactants. MoS2 nanosheets have been shown to adsorb single-stranded DNA by the van der Waals force between nucleobases and the basal plane of MoS2 nanosheets,2 acting as an efficient dye quencher in aqueous solution for detection. DNA, protein, and metal ions have been detected by using simple DNA absorption on the surface of MoS2.3–5 However, the physisorption of fluorescently labeled DNA was susceptible to nonspecific probe displacement and the false-positive signals.6 MoS2 can stabilize metal nanoparticles and form composites which extend its functionalities as a novel catalytic, magnetic, and optoelectronic nanomaterial. It has additionally been reported that MoS2 has great potential in biological molecules detection.7 Highly sensitive detection for protein has been a key target for sensors. In this study, thrombin was used as an example in protein. Thrombin is a serine protease in blood that is involved in some physiological and pathological processes in inflammation, wound cicatrization, blood solidification, and platelet activation.8,9 Thrombin concentrations vary considerably with different coagulation defects. Furthermore, it is related to the development of a variety of diseases and is thus considered as a marker related to various diseases.10,11 Therefore, thrombin detection with high sensitivity is important in early diagnosis and clinical practice. In this work, a covalent linkage between the aptamer and the surface of MoS2 was used to solve the problems of nonspecific probe displacement and the appearance of a false-positive signals. In addition, Tween 80 was used to prevent the nonspecific binding of thrombin on the MoS2 surface. To the best of our knowledge, this was the first time that Tween 80 was combined with a covalent linking aptamer to MoS2 for overcoming these problems. Furthermore, an MoS2 and exonuclease (Exo) coassisted signal amplification strategy was developed in order to improve the detection limit for this protein based on this sensor.
Materials and methods
Chemicals and materials
The aptamer for thrombin was purchased from Sangon Biotechnology Co., Ltd. (Shanghai, People’s Republic of China) with high-performance liquid chromatography purification and mass spectrometry confirmation. The complementary sequence of thrombin aptamer modified with thiol used in this study is as follows: 5′-thiol–AGTCACCCCAACCTGCCCTACCACGGACT-3′. The underlined sequences can have a hairpin-like structure.
Thrombin-binding aptamer was as follows: 5′-AAAA GTCCGTG GTAGGGCA GGTTGGGGTGA CT-FAM-3′. The underlined sequences can be easily recognized by RecJf Exo.
Human α-thrombin with purity more than 95%, was obtained from Haematologic Technologies Inc (Essex, VT, USA). Exo was purchased from New England Biolabs (Beijing) Ltd (Beijing, People’s Republic of China). Gold nanoparticles (AuNPs) with an average diameter of 13±2 nm along with Tween 80 and other proteins were purchased from Sigma-Aldrich Chemical Co. (Shanghai, People’s Republic of China). All solutions that were used in the experiments were prepared with Milli-Q water (18.2 MΩ·cm). Human blood serum samples were obtained from Affiliated Hospital of Jiangsu University.
Preparation of AuNPs@MoS2 nanocomposites
MoS2 nanosheets were prepared by chemical exfoliation according to the method reported by Eda et al.12 Atomic force microscopy was carried out, revealing that the thickness of MoS2 nanosheet was ~1 nm (Figure S1) and confirming that a single-layer MoS2 nanosheet was obtained.13 AuNPs can be selectively formed on the edge sites or defective sites of MoS2 nanosheets by noncovalent bonding.14 Then, 1 mL of AuNPs was added into 1 mL of MoS2 and ultrasonically dispersed. Ten milliliters of water was slowly added after reacting for 24 h. After the colloids were collected by centrifugation at a speed of 10,000 rpm for 10 min, the precipitate was washed with water three times and ultrasonically dispersed in washing buffer (10 mM Tris-HCl, pH 7.4).15
Instrumentation
AuNPs@MoS2 nanocomposites were characterized by transmission electron microscopy. A fluorescence spectrometer (F-4600; Hitachi Co. Ltd., Tokyo, Japan) with an xenon lamp excitation source was employed to record fluorescence spectra. The excitation was set at 490 nm and the emission was monitored at 518 nm.
Fluorescence spectra measurements
One milliliter of Milli-Q water (18.2 MΩ·cm) containing 10 nM aptamer was added to 2 mL of Milli-Q water (18.2 MΩ·cm) containing 40 μg/mL MoS2. Then, different concentrations of thrombin were added and incubated for 30 min for detection of sensitivity. DNA/AuNPs conjugates were rapidly formed in 0.01 M phosphate-buffered saline (0.1 M NaCl, 10 mM phosphate, pH 7.4) at 50°C.16 Exo digestion was carried out at 37°C in buffer (50 mM NaCl, 10 mM Tris-HCl, 10 mM MgCl2, 1 mM dithiothreitol, pH 7.9). The fluorescence measurements were carried out using a Cary Eclipse spectrophotometer. The optical path length of the quartz fluorescence cell was 1.0 cm. The emission spectra were recorded in the wavelength range of 510–600 nm with excitation at 480 nm. The curves were plotted using the fluorescence intensity at 520 nm. The fitting of the experimental data was accomplished with the software Origin 8.0.
Selectivity assays
In order to ensure the selectivity of the method toward thrombin, other proteins (immunoglobulin G, lysozyme, and bovine serum albumin) were added into the Milli-Q water (18.2 MΩ·cm) containing 10 nM aptamer with 40 μg/mL of the MoS2. The concentration of each protein was 0.001 nM. The samples were incubated for 30 min at room temperature for further measurements using the spectrophotometer, and the conditions followed were the same as that for thrombin.
Results and discussion
Design strategy for thrombin detection using exonuclease coassisted amplification strategy
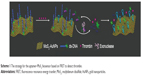
The strategy for the aptamer–MoS2 biosensor based on fluorescence resonance energy transfer (FRET) to detect thrombin is shown in Scheme 1. To avoid nonspecific displacement of the aptamer probe by nontarget molecules, a covalent linkage was established between the carboxyfluorescein (FAM) dual-labeled thrombin aptamer with its complementary sequence modified with a thiol group and the AuNPs on MoS2. The hybridization of the aptamer molecules and the matched strand with 5′ terminal thiol immobilized the aptamer molecules specifically on the surface of AuNPs. AuNPs@MoS2 nanocomposites quenched the fluorescence of FAM, which was labeled on the 3′ end of the aptamer, when the aptamer was immobilized on the surface of AuNPs. Binding of thrombin to the aptamer induced conformational changes of the aptamer, causing the fluorophore to leave the surface of MoS2.17 In this case, the matched strand forms a hairpin-shaped structure on the surface of MoS2 and is not recognized by Exo. RecJf Exo degraded the aptamer in the aptamer–thrombin complex in the direction 5′→3′, causing thrombin to be released into the solution for target recycling.16 Tween 80, a nonionic surfactant that strongly interacts with MoS2 through its hydrocarbon,18,19 was used to prevent the protein from nonspecific binding to improve the detection limit.
Sensitivity of the detection for thrombin using a covalent linking aptamer-based sensor
If the MoS2 concentration was very low, the signal-to-noise ratio was also very low. Furthermore, high MoS2 concentration would quench the fluorescence of the cleaved short FAM–DNA fragments. Therefore, the amount of MoS2 should be optimized. As shown in Figure S2, 60 μg/mL MoS2 was taken as the optimized concentration for aptamer physisorption for detection of thrombin. As shown in Figure S3, 30 min was chosen as the incubation time for further measurements at room temperature. As shown in Figure S4, the fluorescence responses of the aptamer–MoS2 nanosheet significantly increased with increasing the concentrations of thrombin from 0 to 60 nM. A linear correlation (R2=0.97) (Figure S4B) existed between the value of F/F0-1, where F0 and F were the values of fluorescence intensities without and with thrombin, respectively, and the concentration of thrombin over the range 0.0005–20 nM is also shown in this figure. This detection limit using the physisorption of fluorescently labeled aptamer was 0.122 pM (Figure S4). The prepared MoS2 nanosheets had a large surface area, which can load more AuNPs. This was followed by transmission electron microscopy, which revealed the presence of the AuNPs@MoS2 nanocomposites. As shown in Figure 1, AuNPs were dispersed on the surface of MoS2.
  | Figure 1 TEM image of AuNPs dispersed on the surface of MoS2. |
Sixty micrograms per milliliter AuNPs@MoS2 nanocomposites was also taken as the optimized concentration for covalent aptamer for detection of thrombin (Figure S5). The detection limit of this assay was calculated as 0.106 pM on the basis of the 3σ/slope (Figure S6) by using the covalent linking aptamer to MoS2-based fluorescent assay. However, the covalent aptamer was less susceptible to nonspecific probe displacement and false-positive signals. Zhao and Wang11 have also reported on the detection of thrombin using AuNPs with fluorescence-labeled aptamers. The detection limit was 0.14 nM; higher than 0.106 pM. This showed that AuNPs@MoS2 nanocomposites can improve the detection limit compared to AuNPs only.20
Measurement of thrombin using a covalent linking aptamer with Tween 80-based sensors
Tween 80 effectively stabilized AuNPs in salt solutions by forming a protective layer on the surfaces of AuNPs. DNA/AuNPs conjugates can be rapidly formed in 0.01 M PBS (0.1 M NaCl, 10 mM phosphate, pH 7.4) at 50°C followed by a ligand exchange between Tween 80 and thiol-DNA.18 The aptamer modified with thiol was immobilized on the surface of AuNPs in 2.5 h using 10% (v/v) Tween 80 compared to more than 12 h without using Tween 80. Tween 80 was also reported as a blocking agent that prevents the absorbability of nonspecific binding materials by interacting with the AuNPs@MoS2 nanocomposites through its hydrocarbon.18–20 In this study, it improved the sensitivity by preventing nonspecific adsorption of thrombin on the surface of MoS2. Figure S7A illustrates the fluorescence emission spectra of MoS2-aptamer with varying concentrations of thrombin. Figure S7B shows the values of F/F0-1, for the assay with the concentration of thrombin. The inset reveals a linear correlation (R2=0.99) between the value of [F/F0-1] and the concentration of thrombin over the range 0.0004–20 nM. The detection limit was improved to 0.03 pM on the basis of the 3σ/slope (Figure S7B), which was better than the 0.106 pM limit without Tween 80.
Optimization of experiment conditions for exonuclease coassisted amplification strategy
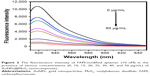
As shown in Figure 2, the fluorescence intensity of aptamer decreased as the concentration of AuNPs@MoS2 nanocomposites increased. When the concentration of AuNPs@MoS2 nanocomposites was increased to 40 μg/mL, 90% of the original fluorescence intensity was quenched. Therefore, 40 μg/mL AuNPs@MoS2 nanocomposites was chosen as the optimal concentration. The different enzyme cleavage times were also measured in the presence of 0.03 U μL-1 Exo at 37°C in buffer (50 mM NaCl, 10 mM Tris-HCl, 10 mM MgCl2, 1 mM dithiothreitol, pH 7.9).21 As shown in Figure S8, in the presence of Exo, the fluorescence intensity increased rapidly and then plateaued due to the complete digestion of DNA by Exo at different digestion times in the presence of 0.03 U μL-1 Exo. Considering the results in Figure S3, 30 min was chosen as the incubation time and detection time for next recycle.
Measurement of thrombin using exonuclease coassisted amplification strategy
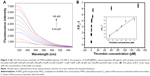
Figure 3A illustrates the fluorescence emission spectra of MoS2-aptamer with various concentrations of thrombin. The corresponding values of [F/F0-1] for the assay are shown in Figure 3B. The inset in Figure 3B shows the linear correlation (R2=0.98) between the value of [F/F0-1] and the concentration of thrombin from 0.04 to 140 pM.
The detection limit was improved to 6 fM on the basis of the 3σ/slope (Figure 3B), which compared well with most existing aptasensors for thrombin. It was lower than those obtained from other reported assays (Table 1), such as surface plasmon resonance (50 nM),22 graphene oxide-aptamer sensor (4.8 pM),18 paper analytical device (16 nM),23 and quartz crystal microbalance (0.1nM),24 graphene nanocomposites (0.01 nM),25 and electrochemical sandwich assay (1.5 pM).26 It was also lower than that obtained from similar mechanisms using other sensors, such as graphene oxide sensor (0.9 pM),16 strip biosensor (4.9 pM),27 and AuNP sensor (0.31 nM).28 In addition to the low detection limit, this method is also simple, label-free, and inexpensive. Therefore, this sensor is a promising technique for protein detection.
  | Table 1 Comparison of different methods for thrombin detection |
Selectivity of the detection for thrombin using exonuclease coassisted amplification strategy
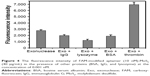
Selectivity is a critical factor to assess the performance of the proposed sensor. Thrombin aptamers are artificial single-strand oligonucleic acid sequences that bind to thrombin with high affinity and specificity and are selected in vitro with a combinatorial method by systematic evolution of ligands by exponential enrichment.28 Therefore, the detection of nonspecific proteins such as bovine serum albumin, IgG, and lysozyme were used as controls to test the sensor’s selectivity recognition to thrombin in this experiment. The response signals of the different proteins were much smaller than that of thrombin. The response to thrombin was two-fold higher in comparison to the other proteins measured at the same concentration (Figure 4). These results implied that this sensor offered high selectivity toward thrombin. If proteins were structurally more similar to thrombin, they could possibly interfere with false-positive results and the baseline FRET signal. Therefore, they cannot be used for checking false-positives and the baseline FRET signal.
Analytical application of exonuclease coassisted amplification strategy
To assess the analytical reliability and application of the constructed aptasensor, a series of human serum samples were prepared by first centrifuging at 3,000 rpm for 15 min followed by the addition of various concentrations of thrombin into the human serum. Affiliated Hospital of Jiangsu University provided human blood serum samples. Each sample was analyzed three times. The detection results for the prepared samples are presented in Table 2. The recovery (between 83.51% and 103.02%) and relative standard deviation (between 3.76% and 7.01%) were acceptable, which indicated that this sensor had promising sensing abilities in real samples.
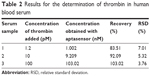  | Table 2 Results for the determination of thrombin in human blood serum |
Conclusion
This was the first example of a covalent aptamer for protein (thrombin) detection based on the MoS2-aptamer system. The previous assays involved only physisorbed aptamer. In this work, a covalent aptamer for protein detection based on MoS2-aptamer system was prepared. The same sequence was also used to prepare physisorbed probes for comparison. Both types of sensors had similar sensitivity. However, the covalent sensor was much more resistant to nonspecific probe displacement and false-positive signals. Furthermore, 10% Tween 80 was used to shorten the reaction time between AuNPs and aptamer and improved the detection limit by preventing the thrombin from being absorbed on the surface of MoS2. Exo digested the single-strand aptamer and released the thrombin, which was then detected for next recycle. With the coassisted amplification strategy, the detection limit was 6 fM. This method is not restricted to protein detection. By covalently attaching other types of aptamers, DNA, RNA, and the detection of other multiple analytes may be achieved.
Acknowledgments
This work is supported by the Jiangsu University (No 13JDG005), Jiangsu University Graduate Project (Y15A171), Natural Science Foundation of Liaoning Province (201601268), the Universities Natural Science Research Project of Jiangsu Province (No 15KJB180001), and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Disclosure
The authors report no conflicts of interest in this work.
References
Ge J, Ou EC, Yu RQ, Chu X. A novel aptameric nanobiosensor based on the self-assembled DNA–MoS2 nanosheet architecture for biomolecule detection. J Mater Chem B. 2013;2:625–628. | ||
Zhu CF, Zeng ZY, Li H, Li F, Fan CH, Zhang H. Single-layer MoS2-based nanoprobes for homogeneous detection of biomolecules. J Am Chem Soc. 2013;135:5998–6001. | ||
Li BL, Zou HL, Lu L, et al. Size-dependent optical absorption of layered MoS2 and DNA oligonucleotides induced dispersion behavior for label-Free detection of single-nucleotide polymorphism. Adv Funct Mater. 2015;25:3541–3550. | ||
Mao K, Wu Z, Chen Y, Zhou X, Shen A, Hu J. A novel biosensor based on single-layer MoS2 nanosheets for detection of Ag+. Talanta. 2015;132:658–663. | ||
Xiang X, Shi JB, Huang FH, Zheng MM, Deng QC, Xu JQ. MoS2 nanosheet-based fluorescent biosensor for protein detection via terminal protection of small-molecule-linked DNA and exonuclease III-aided DNA recycling amplification. Biosens Bioelectron. 2015;74:227–232. | ||
Wang KM, Tang ZW, Yang CY, et al. Molecular engineering of DNA: molecular beacons. Angew Chem Int Ed Engl. 2009;48:856–870. | ||
Su S, Zhang C, Yuwen LH, et al. Creating SERS hot spots on MoS2 nanosheets with in situ grown gold nanoparticles. ACS Appl Mater Interfaces. 2014;6:18735–18741. | ||
Kongsuphol P, Arya SK, Wong CC, Polla LJ, Park MK. Coiled-coil peptide based sensor for ultra-sensitive thrombin detection. Biosens Bioelectron. 2014;55:26–31. | ||
Popovic M, Smiljanic K, Dobutovic B, Syrovets T, Simmet T, Isenovic ER. Thrombin and vascular inflammation. Mol Cell Biochem. 2012;359:301–313. | ||
Oroval M, Climent E, Coll C, et al. An aptamer-gated silica mesoporous material for thrombin detection. Chem Commun. 2013;49:5480–5482. | ||
Zhao Q, Wang X. An aptamer-capture based chromogenic assay for thrombin. Biosens Bioelectron. 2012;34:232–237. | ||
Eda G, Yamaguchi H, Voiry D, Fujita T, Chen MW, Chhowalla M. Photoluminescence from chemically exfoliated MoS2. Nano Lett. 2011;11:5111–5116. | ||
Zeng ZY, Yin ZY, Huang X, et al. Single-layer semiconducting nanosheets: high-yield preparation and device fabrication. Angew Chem Int Ed. 2011;50:11093–11097. | ||
Shi Y, Huang JK, Jin L, et al. Selective decoration of Au nanoparticles on monolayer MoS2 single crystals. Sci Rep. 2013;3:1839. | ||
Wang X, Deng WP, Shen L, Yan M, Ge SG, Yu JH. A sensitive quenched electrochemiluminescent DNA sensor based on the catalytic activity of gold nanoparticle functionalized MoS2. New J Chem. 2015;39:8100–8107. | ||
Bao T, Wen W, Zhang X, Wang S. An exonuclease-assisted amplification electrochemical aptasensor of thrombin coupling “signal on/off” strategy. Anal Chim Acta. 2015;860:70–76. | ||
Liu S, Zhang XY, Luo WX, et al. Single-molecule detection of proteins using aptamer-functionalized molecular electronic devices. Angew Chem Int Ed. 2011;50:2496–2502. | ||
Gao L, Li Q, Li RQ, et al. Highly sensitive detection for proteins using graphene oxide-aptamer based sensors. Nanoscale. 2015;7:10903–10907. | ||
Xu SM, Yuan H, Xu A, Wang J, Wu LJ. Rapid synthesis of stable and functional conjugates of DNA/gold nanoparticles mediated by Tween 80. Langmuir. 2011;27:13629–13634. | ||
Wang WJ, Chen CL, Qian MX, Zhao XS. Aptamer biosensor for protein detection using gold nanoparticles. Analyt Biochem. 2008;373:213–219. | ||
Yi HY, Xu WJ, Yuan YL, Wu YM, Chai YQ, Yuan R. A sensitive electrochemical aptasensor for thrombin detection based on exonuclease-catalyzed target recycling and enzyme-catalysis. Biosens Bioelectron. 2013;47:368–372. | ||
Bai YF, Feng F, Zhao L, et al. Aptamer/thrombin/aptamer-AuNPs sandwich enhanced surface plasmon resonance sensor for the detection of subnanomolar thrombin. Biosens Bioelectron. 2013;47:265–270. | ||
Cunningham JC, Brenes NJ, Crooks RM. Paper electrochemical device for detection of DNA and thrombin by target-induced conformational switching. Anal Chem. 2014;86:6166–6170. | ||
Yuan F, Chen HQ, Xu J, Zhang YY, Wu Y, Wang L. Aptamer-based luminescence energy transfer from near-infraredto-near-infrared upconverting nanoparticles to gold nanorods and its application for the detection of thrombin. Chem Eur J. 2014;20:2888–2894. | ||
Wang QQ, Zhou ZX, Zhai YL, et al. Label-free aptamer biosensor for thrombin detection based on functionalized graphene nanocomposites. Talanta. 2015;141:247–252. | ||
Yeh FY, Liu TY, Tseng IH, Yang CW, Lu LC, Lin CS. Gold nanoparticles conjugates-amplified aptamer immunosensing screen-printed carbon electrode strips for thrombin detection. Biosens Bioelectron. 2014;61:336–343. | ||
Qin CY, Wen W, Zhang XH, Gu HS, Wang SF. Visual detection of thrombin using a strip biosensor through aptamer-cleavage reaction with enzyme catalytic amplification. Analyst. 2015;140:7710–7717. | ||
Gao T, Ning LM, Li C, Wang HY, Li GX. A colorimetric method for protein assay via exonuclease III-assisted signal attenuation strategy and specific DNA–protein interaction. Anal Chim Acta. 2013;788:171–176. | ||
Wang L, Zhu JB, Han L, et al. Graphene-based aptamer logic gates and their application to multiplex detection. ACS Nano. 2012;6:6659–6666. | ||
Yoon J, Choi N, Ko J, Kim K, Lee S, Choo J. Highly sensitive detection of thrombin using SERS-based magnetic aptasensors. Biosens Bioelectron. 2013;47:62–67. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.




