Back to Journals » OncoTargets and Therapy » Volume 11
High CEP55 expression is associated with poor prognosis in non-small-cell lung cancer
Authors Jiang C, Zhang Y, Li Y, Lu J, Huang Q, Xu R , Feng Y, Yan S
Received 16 February 2018
Accepted for publication 2 May 2018
Published 17 August 2018 Volume 2018:11 Pages 4979—4990
DOI https://doi.org/10.2147/OTT.S165750
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr XuYu Yang
Chao Jiang,1 Yu Zhang,2 Yong Li,2 Jiabin Lu,2 Qitao Huang,2 Rui Xu,3 Yanfeng Feng,2 Shumei Yan2
1Department of Oncology, The People’s Hospital of Baoan District, Shenzhen, Guangdong, People’s Republic of China; 2Department of Pathology, State Key Laboratory of Oncology in South China, Collaborative Innovation Center for Cancer Medicine, Sun Yat-Sen University Cancer Center, Guangzhou, Guangdong, People’s Republic of China; 3Department of Medical Oncology, Affiliated Tumor Hospital of Guangzhou Medical College, Guangzhou, People’s Republic of China
Objectives: Lung cancer is the most common and lethal malignancy worldwide. CEP55 was found to be overexpressed in multiple types of cancer. However, the expression pattern of CEP55 and its clinical significance in non-small-cell lung carcinoma (NSCLC) have not been investigated by immunohistochemistry.
Materials and methods: In this study, we analyzed 203 primary NSCLC specimens from Sun Yat-Sen University Cancer Center to investigate the clinical role of CEP55 in lung cancer. Tissue microarray was successfully generated for immunohistochemical evaluation. The correlation between CEP55 expression and clinical characteristics and survival was analyzed statistically. The predictive effect of CEP55 and APOBEC3B (AP3B) coexpression in lung cancer patients’ prognosis was evaluated.
Results: We found that the CEP55 expression was commonly elevated in NSCLC tissues and overexpression of CEP55 was correlated with unfavorable prognosis in the patients with NSCLC. Furthermore, the combination of CEP55 and AP3B expression was significantly predictive of clinical outcome in all NSCLC patients.
Conclusion: CEP55 may act as a useful and novel prognostic biomarker for NSCLC. Further studies into the mechanism of CEP55 are warranted.
Keywords: CEP55, non-small-cell lung cancer, immunohistochemistry, prognosis
Introduction
Lung cancer is the most lethal type of cancer in the United States and worldwide. In the United States, it is predicted that 222,500 new lung cancer patients will be diagnosed, and 155,870 people will die of lung cancer in 2018.1 The death rate from lung cancer is 25%, the highest among all types of cancer. Smoking and being exposed to certain toxins are currently known direct reasons of lung cancer; however, many lung cancer cases occur without clear reasons. The recent discovery of several key driver gene mutations, including EGFR, k-ras, and ALK, opened a new era of understanding lung cancer and prompted the development of targeted therapies.2 However, most lung cancers have no symptoms at initial stages and are advanced when diagnosed, resulting in poor prognosis. Many biomarkers critical for the diagnosis and treatment of lung cancer are emerging.
CEP55, also known as c10orf3 and FLJ10540, is encoded by the CEP55 gene. It is widely expressed in different types of tissue, especially in proliferative tissues. It locates to the centrosomes at interphase and forms homodimers to assist cytokinesis.3 Recently, CEP55 was found to be involved in the regulations of PI3K/AKT pathway and cancer cell stemness.4–6 Clinically, CEP55 has been found overexpressed in multiple types of cancer, such as breast, prostate, renal, and thyroid cancers.7–10 It has been included as a gene highly associated with prognosis in several patented mRNA microarrays used to screen for cancer diagnosis.11,12 However, the association of its protein level with prognosis of lung cancer patients has never been determined before. In this study, we used immunohistochemistry (IHC) and online datasets to determine the expression level of CEP55 in lung cancer patients and investigated its correlation with clinical parameters and prognosis.
Materials and methods
Patient selection
This study was approved by the medical ethics committee of Sun Yat-Sen University Cancer Center. A total of 203 primary lung cancer specimens were obtained from primary surgery from October 2000 to April 2007. The selective criteria are as previously described.13 To avoid confusion in data analysis, neoadjuvant chemotherapy patients were excluded. The histologic grade and clinical stage of the tumors were defined as previously described.13 The final survival status of patients was confirmed in December 2013. Patients whose tissues and medical records were used for this study had provided written informed consent.
IHC staining
IHC staining was performed to detect CEP55 expression according to the protocol previously described,13 with minor changes. Tissue sections were deparaffinized with dimethylbenzene and then rehydrated through 100%, 95%, 90%, 80%, and 70% ethanol for 3 minutes each. After the PBS wash step, the slides were boiled for 15 minutes in citric acid in a microwave oven, followed by endogenous peroxidase blocking with 3% hydrogen peroxide. To retrieve antigen, the slides were then boiled in tris(hydroxymethyl) aminomethane-EDTA buffer (pH 8.0) in a microwave for 30 minutes. Nonspecific antigen binding was inhibited by incubation in 10% normal goat serum for 20 minutes. CEP55 antibody (1:1,000 dilution, Abgent, Rabbit Ig, San Diego, CA, USA) was incubated with slides overnight at 4°C in a moist chamber. For negative controls, slides were incubated with normal goat serum instead of the primary antibody. After the overnight incubation, the slides were sequentially incubated with biotinylated goat anti-rabbit antibody, streptavidin–peroxidase conjugate, and 3,3′-diaminobenzidine. Normal alveolar and bronchial epithelial mucosa were utilized as negative controls.
Immunoreactivity score (IRS) assessment
Two independent pathologists (Shanshan Lyu and Shumei Yan) who were blinded to the clinicopathological information performed the IRS assessment for CEP55 expression. The scoring criteria are as described previously.13 The staining results were scored based on the 2 criteria: 1) the percentage of positive tumor cells in the tissue: 0 (0%), 1 (1%–10%), 2 (11%–50%), 3 (51%–75%), and 4 (76%–100%) and 2) the intensity of staining: 0, absent; 1, weak; 2, moderate and 3, strong. IRS was calculated by multiplying (1) and (2) (range from 0 to 12). The specimens were rescored if the difference between the two pathologists was >3.
Statistical analysis
The CEP55 IRS cutoff value was determined by the median of the staining results of 203 specimens. The correlation between CEP55 expression and clinicopathological features was analyzed by Pearson’s χ2 test. Disease-free survival (DFS) was defined as the time from surgery to regional relapse or distant metastasis. Overall survival (OS) was defined as the time from surgery to death. DFS and OS curves in different subgroups were constructed by the Kaplan–Meier method. To test significant differences between 2 survival curves, we used HRs and their 95% CIs of covariates in the analyses of DFS or OS by Cox proportional hazards model. Multivariate analysis was performed for all of the parameters that were significant in the univariate analysis. Receiver operating characteristic (ROC) curve analysis was used to assess the predictive value of clinicopathologic features.
Data from previous publication were also utilized in this study.13 Statistical analysis was performed using SPSS software (standard version 16.0, SPSS Inc., Chicago, IL, USA). A difference was considered statistically significance if the 2-sided probability value was less than 0.05.
Results
Patient characteristics
Fifty-four females and 149 males, aged from 30 to 79 years (median 60.0 years), were included in this study. Among them, 122 were smokers and 81 were nonsmokers. The specific clinicopathological characteristics of the 203 patients are listed in Table 1.
  | Table 1 Correlation between CEP55 expression and clinicopathological variables of lung cancer cases |
Expression of CEP55 in non-small-cell lung carcinoma (NSCLC) and associations with clinicopathological characteristics
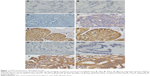
In this study, the median CEP55 IRS value was <8.0. IRS scores less than this value were considered low expression, and CEP55 IRS scores, greater than this value, were considered high expression. CEP55 staining of NSCLC tissue and normal alveolar and bronchial epithelial mucosa revealed that immunoreactivity primarily rests in the cytoplasm of tumor cells (Figure 1). Low expression of CEP55 was observed in 37.4% (76/203) of the NSCLC samples, whereas 62.6% (127/203) of NSCLC samples had high expression of CEP55 (Table 1). The expression of CEP55 correlated closely with nodal status (P=0.004). No other statistical associations were observed between CEP55 expression and age, gender, smoking, tumor diameters, histology, histology grade, visceral pleural invasion, tumor status, TNM staging, and adjuvant chemotherapy (P=0.464, P=0.623, P=0.237, P=0.662, P=0.880, P=0.334, P=0.640, P=0.996, P=0.129, and P=0.110, respectively).
CEP55 expression and patient prognosis
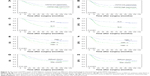
All patients participated for the duration of this study. About 102 (50.24%) patients were deceased, and 101 patients were still alive at the end of the duration of this study. The 5-year OS rates of the whole cohort were 53.2% (108/203). OS and DFS curves for the whole cohort are shown in Figure 2. Importantly, patients with high expression of CEP55 had a shorter OS and DFS compared with those with low expression (OS: mean of 73.04 versus 89.88 months, P=0.003; DFS: mean of 63.77 versus 84.73 months, P=0.005, Figure 2, Table 2). CEP55 can differentiate between tumors in N or TNM staging and between those which have or have not been treated, with adjuvant chemotherapy (Figure 2). High CEP55 expression was significantly associated with poor prognosis in NSCLC patients who are aged ≤60, females, nonsmokers, patients with tumor diameter >3.5 cm, G2/3, and patients who had not received without adjuvant chemotherapy (Figure 3, Table 2, DFS: P=0.006, P<0.001, P=0.027, P=0.006, P=0.014, and P=0.022; OS: P=0.005, P<0.001, P=0.018, P=0.004, P=0.008, and P=0.007).
Furthermore, univariate analysis using Cox’s proportional hazard model showed that the following parameters correlated significantly with OS: tumor diameter, histological grade, tumor status, nodal status, TNM staging, AP3B expression, CEP55 expression, and adjuvant chemotherapy (Table 3). CEP55 and AP3B expression and clinicopathologic parameters that were significant in univariate analysis were further analyzed in multivariate analysis. The result demonstrated that nodal status, AP3B expression, and CEP55 expression were independent prognostic factors that affected NSCLC patients’ OS (Table 3, P=0.050, P=0.003, P=0.036, respectively). High expression of CEP55 was a predictor for poor prognosis (HR, 1.617; 95% CI, 1.033–2.532; Table 3). High expression of AP3B was also a predictor for poor prognosis of NSCLC patients (HR, 1.893; 95% CI, 1.245–2.877; Table 3).
  | Table 3 Univariate and multivariate analyses of OS of lung cancer patients |
Combined increased expression of CEP55 and AP3B is correlated with poor prognosis in NSCLC
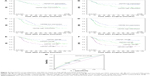
APOBEC3B, AP3B, is a newly defined DNA cytosine deaminase. Cytosine deamination resulting in cytosine to thymine (C-to-T) transition mutations, which is catalyzed by the APOBEC family, is a major source of DNA mutation in many different tumors. We retrieved data from 203 patients analyzed for a previous study13 to analyze whether CEP55 and AP3B coexpression affects NSCLC prognosis. As shown in Figure 4, when all NSCLC patients were stratified by AP3B expression, we found that the prognosis of patients with low CEP55 had significantly higher DFS and OS (P<0.05, Figure 4A and B). However, in patients with high AP3P expression, patients with low expression of CEP55 had significantly higher OS (P<0.05, Figure 4C), but showed no difference in DFS (P=0.065, Figure 4D).
To determine whether coexpression of CEP55 and AP3B influence survival, patients were divided into 4 groups: 1 (CEP55−/AP3B−), 2 (CEP55+/AP3B−), 3 (CEP55−/AP3B+), and 4 (CEP55+/AP3B+). Group 1 patients had the best survival rate, and group 4 patients had the worst survival rate (mean OS: 99.82 vs 58.12 months; mean DFS: 97.41 vs 49.42 months). Group 2 and group 3 had moderate survivals rates.
ROC curves were plotted to evaluate the patients’ survival status to assess prognostic values of CEP55, AP3B, or combined expression in NSCLC. ROC curve analysis showed promising predictive significance of CEP55 combined with AP3B expression in all NSCLC patients (area under the curve =0.665, P<0.001, Figure 4G). ROC curves of CEP55 and AP3B showed area under the curve of 0.600 and 0.616, which is also predictively significant (P=0.013 and P=0.004, respectively, Figure 4G).
Discussion
In this study, we showed that high expression of CEP55 is predictive of worse prognosis in lung cancer patients. Nodal status, AP3B expression, and CEP55 expression were independent prognostic factors that affected prognosis. Patients with low CEP55 and AP3B expression had the best survival rate, and patients with high CEP55 and AP3B expression had the worse survival rate. Previously, CEP55 had been shown to be a useful biomarker in breast, prostate, renal, and thyroid cancers.7–10 However, its predictive value in lung cancer had not been reported.
CEP55 is a 55 kD centrosome- and midbody-associated protein. It contains 464 amino and was initially discovered as a key regulator of cytokinesis.14,15 Cytokinesis is a short and final step of the cell cycle during which the cytoplasmic division of a cell occurs and two daughter cells form. CEP55 localizes on the centrosome throughout the cell cycle and is recruited to the mitotic spindle and midbody during mitosis to recruit downstream machinery to it, which is critical for the correct assembly of the midbody. Consistently, CEP55 depletion in cells led to an increase in multinucleated cells and cells arrested at the midbody stage, which demonstrated the critical role of CEP55 in cytokinesis.16,17
Apart from the cell cycle, CEP55 and the midbody are also associated with cell stemness. Stem cells and cancer cells accumulate more midbodies by evading autophagy, while differentiated cells possess high autophagic activity and do not accumulate midbodies.6 Interestingly, our clinical analysis showed some similar results. We found that patients with higher histologic grade and TNM stages had a high ratio of positive CEP55 staining compared to those with well-differentiated tumors and lower TNM stages. Additionally, patients with lymph node metastasis demonstrated a higher ratio of positive CEP55 staining than those without lymph node metastasis. These clinical observations suggest that high expression of CEP55 correlates with cancer cell stemness, which is consistent with previous studies.
Another possible explanation of the high ratio of CEP55 expression in more advanced stage lung cancer patients is due to its activation of PI3K/AKT pathway, a major prosurvival pathway in cancer cells. CEP55 has been reported to bind to PIK3CA and stabilize it, leading to increased S473 phosphorylation of AKT in lung cancer.18 In addition, the CEP55/PI3K/AKT pathway leads to increased migration and invasion of lung cancer cells.19 Thus, the activation of AKT could be a critical oncogenic factor of these advanced stage lung cancer patients.
We conclude that CEP55 is an independent prognostic factor in lung cancers. In the Kaplan–Meier plots, lung cancer patients with high CEP55 expression have shorter survival time than those with low CEP55 expression. We also extracted Kaplan–Meier plots from SurveExpress online database and found similar results, further supporting our conclusions. As mentioned before, overexpression of CEP55 has been reported to predict poor prognosis in ovarian carcinoma, esophageal squamous cell carcinoma, bladder, breast, and thyroid cancers. Of note, in the last few decades, CEP55 has been identified as being overexpressed in mRNA microarray expression profiles of many human cancers, including gastric, hepatocellular, lung, and bladder carcinoma.4,18,20,21 CEP55 was included in the top 70 most highly overexpressed genes in CIN 70 and in a signature in patented prognostic kits for the prediction of malignancy risk, prognosis, and therapy resistance in multiple human cancers.22 CEP55 was also part of commercial signatures from Myriad Genetics Inc. and PAM50 for diagnosis and prognostic prediction of lung and other types of cancers.23
More importantly, it has been reported that CEP55 can act as a cancer vaccine target. It is not only a tumor-associated antigen but also aberrantly expressed in tumors, making it ideal for immunotherapy vaccines.24 Previous studies have shown that CEP55 peptides were able expand and activate cytotoxic T lymphocyte clonally in vitro and in vivo.25,26 Exciting results have been shown in the treatment of chemotherapy-resistant colon cancer cells.
Our results fill in the blank in the prognostic prediction of lung cancer patients by IHC, which strengthened the clinical significance of CEP55. All these findings demonstrate that both CEP55 RNA and protein levels can act as prognostic factors in multiple cancers and might be a good treatment target in cancer immunotherapies.
Another intriguing investigation direction is about the regulation of CEP55 by driver genes in cancers. Although it has been demonstrated that FOXM1 transactivates CEP55 and PLK1 stabilizes it, few papers cover the relationship of CEP55 with key driver mutations in lung cancer, such as ALK, EGFR, k-ras, and met.12,27 As an important oncogenic factor, the activity of CEP55 must be strictly regulated in cells. Thus, it deserves more studies both clinically and mechanistically. We admit that this study has limitations. It was conducted only in one institution; further verifications from other independent investigators are needed. Additionally, key mutation information was not included in our data, so correlational analyses between CEP55 and other known mutations could not be performed.
Overall, our local and online analyses led to the conclusion that lung cancer patients with high CEP55 expression have shorter OS and DFS compared with those with low CEP55 expression. CEP55 might be associated with stemness of lung cancer cells, and it can be an independent predictor of survival for patients with lung cancer. Further studies are warranted.
Disclosure
The authors report no conflicts of interest in this work.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. | ||
Chan BA, Hughes BG. Targeted therapy for non-small cell lung cancer: current standards and the promise of the future. Transl Lung Cancer Res. 2015;4(1):36–54. | ||
Fabbro M, Zhou BB, Takahashi M, et al. Cdk1/Erk2- and Plk1-dependent phosphorylation of a centrosome protein, Cep55, is required for its recruitment to midbody and cytokinesis. Dev Cell. 2005;9(4):477–488. | ||
Chen CH, Lu PJ, Chen YC, et al. FLJ10540-elicited cell transformation is through the activation of PI3-kinase/AKT pathway. Oncogene. 2007;26(29):4272–4283. | ||
Wang G, Liu M, Wang H, et al. Centrosomal protein of 55 regulates glucose metabolism, proliferation and apoptosis of glioma cells via the Akt/mTOR signaling pathway. J Cancer. 2016;7(11):1431–1440. | ||
Kuo TC, Chen CT, Baron D, et al. Midbody accumulation through evasion of autophagy contributes to cellular reprogramming and tumorigenicity. Nat Cell Biol. 2011;13(10):1214–1223. | ||
Inoda S, Hirohashi Y, Torigoe T, et al. Cep55/c10orf3, a tumor antigen derived from a centrosome residing protein in breast carcinoma. J Immunother. 2009;32(5):474–485. | ||
Shiraishi T, Terada N, Zeng Y, et al. Cancer/testis antigens as potential predictors of biochemical recurrence of prostate cancer following radical prostatectomy. J Transl Med. 2011;9:153. | ||
Jones J, Otu H, Spentzos D, et al. Gene signatures of progression and metastasis in renal cell cancer. Clin Cancer Res. 2005;11(16):5730–5739. | ||
Martin KJ, Patrick DR, Bissell MJ, Fournier MV. Prognostic breast cancer signature identified from 3D culture model accurately predicts clinical outcome across independent datasets. PLoS One. 2008;3(8):e2994. | ||
Naderi A, Teschendorff AE, Barbosa-Morais NL, et al. A gene-expression signature to predict survival in breast cancer across independent data sets. Oncogene. 2007;26(10):1507–1516. | ||
Waseem A, Ali M, Odell EW, Fortune F, Teh MT. Downstream targets of FOXM1: CEP55 and HELLS are cancer progression markers of head and neck squamous cell carcinoma. Oral Oncol. 2010;46(7):536–542. | ||
Yan S, He F, Gao B, et al. Increased APOBEC3B predicts worse outcomes in lung cancer: a comprehensive retrospective study. J Cancer. 2016;7(6):618–625. | ||
Martinez-Garay I, Rustom A, Gerdes HH, Kutsche K. The novel centrosomal associated protein CEP55 is present in the spindle midzone and the midbody. Genomics. 2006;87(2):243–253. | ||
Zhao WM, Seki A, Fang G. Cep55, a microtubule-bundling protein, associates with centralspindlin to control the midbody integrity and cell abscission during cytokinesis. Mol Biol Cell. 2006;17(9):3881–3896. | ||
Bastos RN, Barr FA. Plk1 negatively regulates Cep55 recruitment to the midbody to ensure orderly abscission. J Cell Biol. 2010;191(4):751–760. | ||
van der Horst A, Simmons J, Khanna KK. Cep55 stabilization is required for normal execution of cytokinesis. Cell Cycle. 2009;8(22):3742–3749. | ||
Chen CH, Lai JM, Chou TY, et al. VEGFA upregulates FLJ10540 and modulates migration and invasion of lung cancer via PI3K/AKT pathway. PLoS One. 2009;4(4):e5052. | ||
Liu L, Mei Q, Zhao J, Dai Y, Fu Q. Suppression of CEP55 reduces cell viability and induces apoptosis in human lung cancer. Oncol Rep. 2016;36(4):1939–1945. | ||
Tao J, Zhi X, Tian Y, et al. CEP55 contributes to human gastric carcinoma by regulating cell proliferation. Tumour Biol. 2014;35(5):4389–4399. | ||
Singh PK, Srivastava AK, Rath SK, Dalela D, Goel MM, Bhatt ML. Expression and clinical significance of centrosomal protein 55 (CEP55) in human urinary bladder transitional cell carcinoma. Immunobiology. 2015;220(1):103–108. | ||
Carter SL, Eklund AC, Kohane IS, Harris LN, Szallasi Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat Genet. 2006;38(9):1043–1048. | ||
Sung JC, Lee AY, Yeatman TJ. The promise of gene signatures in cancer diagnosis and prognosis. Encyclopedia of Genetics, Genomics, Proteomics and Bioinformatics. Hoboken, NJ: John Wiley & Sons, Ltd; 2004. | ||
Inoda S, Morita R, Hirohashi Y, et al. The feasibility of Cep55/c10orf3 derived peptide vaccine therapy for colorectal carcinoma. Exp Mol Pathol. 2011;90(1):55–60. | ||
Inoda S, Hirohashi Y, Torigoe T, et al. Cytotoxic T lymphocytes efficiently recognize human colon cancer stem-like cells. Am J Pathol. 2011;178(4):1805–1813. | ||
Gao XY, Wang XL. An adoptive T cell immunotherapy targeting cancer stem cells in a colon cancer model. J BUON. 2015;20(6):1456–1463. | ||
Laoukili J, Kooistra MR, Bras A, et al. FoxM1 is required for execution of the mitotic programme and chromosome stability. Nat Cell Biol. 2005;7(2):126–136. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.