Back to Journals » Infection and Drug Resistance » Volume 13
Genotypic and Phenotypic-Based Assessment of Antibiotic Resistance and Profile of Staphylococcal Cassette Chromosome mec in the Methicillin-Resistant Staphylococcus aureus Recovered from Raw Milk
Authors Rahi A, Kazemeini H, Jafariaskari S, Seif A, Hosseini S, Safarpoor Dehkordi F
Received 2 September 2019
Accepted for publication 4 December 2019
Published 30 January 2020 Volume 2020:13 Pages 273—283
DOI https://doi.org/10.2147/IDR.S229499
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Azar Rahi, 1 Hamidreza Kazemeini, 2 Sedigheh Jafariaskari, 3 Ali Seif, 4 Sahar Hosseini, 5 Farhad Safarpoor Dehkordi 6
1Department of Microbiology, Kazerun Branch, Islamic Azad University, Kazerun, Iran; 2Department of Food Hygiene, Faculty of Veterinary Medicine, Amol University of Special Modern Technologies, Amol, Iran; 3Department of Parasitology, Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran; 4Doctor Veterinary Medicine, Faculty of Veterinary Medicine, Shahrekord Branch, Islamic Azad University, Shahrekord, Iran; 5Master of Food Science and Technology, Faculty of Agriculture and Food Sciences, Shahrekord Branch, Islamic Azad University, Shahrekord, Iran; 6Halal Research Center of IRI, FDA, Tehran, Iran
Correspondence: Hamidreza Kazemeini
Department of Food Hygiene, Faculty of Veterinary Medicine, Amol University of Special Modern Technologies, Amol, Iran
Email [email protected]
Background: Multidrug resistant methicillin-resistant Staphylococcus aureus (MRSA) bacteria are determined to be one of the chief causes of foodborne diseases around the world.
Purpose: This research was done to assess the genotypic and phenotypic profiles of antibiotic resistance and distribution of Staphylococcus cassette chromosome mec (SCCmec) types amongst the MRSA bacteria recovered from raw milk.
Methods: Five-hundred and ninety raw milk samples were collected and examined. MRSA bacteria were recognized using susceptibility evaluation toward oxacillin and cefoxitin disks. Profile of antibiotic resistance genes and SCCmec types were determined using the PCR. Antibiotic resistance pattern of isolates was examined using the disk diffusion.
Results: Thirty-nine out of 590 raw milk samples (6.61%) were positive for S. aureus. Twenty-eight out of 39 (71.79%) bacteria were defined as MRSA bacteria. Raw buffalo (80%) milk samples had the maximum incidence of MRSA, while raw camel (33.33%) had the minimum. MRSA bacteria harbored the maximum incidence of resistance toward penicillin (100%), tetracycline (100%), erythromycin (82.14%), gentamicin (78.57%) and trimethoprim-sulfamethoxazole (78.57%). Incidence of resistance toward more than eight classes of antibiotic agents was 28.57%. The most frequently distinguished antibiotic resistance markers were blaZ (100%), tetK (85.71%), dfrA1 (71.42%), aacA-D (67.85%), ermA (50%) and gyrA (42.85%). SCCmec IVa (29.62%), V (25%), III (14.81%) and IVb (11.11%) were the most frequently distinguished types.
Conclusion: Raw milk of dairy animals maybe sources of multidrug resistant MRSA which pose a hygienic threat concerning the consumption of raw milk in Iran. Nevertheless, further investigations are necessary to understand supplementary epidemiological features of MRSA in raw milk.
Keywords: methicillin-resistant Staphylococcus aureus, raw milk, antibiotic resistance mechanisms, SCCmec types
Introduction
Milk of animal species contains assortment of imperative dietary supplements including proteins, carbohydrate, fats, minerals and vitamins with boost advantageous effects for human life.1 Therefore, their regular daily consumption has been extensively suggested. However, there is evidence that raw milk of animal species might contain different types of threatening foodborne pathogens.2–5
Most cases of foodborne outbreaks are associated with the consumption of food contaminated with foodborne bacterial pathogens,6–16 especially Staphylococcus aureus (S. aureus).17–20 S. aureus is a bacterium of the Firmicutes family originating from the human nose and skin. S. aureus is considered one of the chief causes of hospital and community-acquired infections and foodborne diseases recognized by weakness, vomiting, nausea, abdominal cramps and toxic shock syndrome.17–20
Foodborne S. aureus bacteria are typically associated with boost prevalence of antibiotic resistance.17–20 Today, methicillin-resistant S. aureus (MRSA) has developed a significant issue in both health care units and the community.17–20 Recognized data described that approximately 70% of S. aureus bacteria recovered from the health care units and the community were simultaneously resistant toward penicillins and cephalosporins.17–21 They are responsible for about 100,000 morbidity with near to 20% mortality per year in the United States.21 Higher pathogenicity of MRSA bacteria,17–21 their inclusive levels of resistance toward numerous kinds of antibiotic agents, especially penicillins, aminoglycosides, macrolides, tetracyclines and fluoroquinolones17–21 and their foodborne aspects17–20 have amplified the clinical and microbial importance of MRSA in popularly consumed foodstuffs, particularly milk. Furthermore, foodstuffs containing MRSA bacteria are considered as imperative reservoirs of antibiotic resistance genes.17–21 Boost incidence of the genes encode resistance toward penicillins (blaZ), aminoglycosides (aacA-D), tetracyclines (tetK and tetM), macrolides (ermA, ermB, msrA, msrB and mefA), fluoroquinolones (gyrA and grlA), lincosamides (linA), folate inhibitors (dfrA1), phenicols (cfr), and ansamycins (rpoB) is one of the chief ways for occurrence of severe antibiotic resistance.17–20
The mecA gene is another imperative antibiotic resistance marker responsible for resistance toward methicillin. It is associated with a 21- to 67-kb molecular element named staphylococcal chromosomal cassette mec (SCCmec)22 characterized by mec and the ccr genetic markers. SCCmec elements are characteristically divided into 11 different types based on to the positioning ccr and mec genes.22 SCCmec IV is additionally divided to IVa, IVb, IVc and IVd alleles.22 A mobile genetic element, SCCmec, plays an important role in staphylococci pathogenesis and occurrence of resistance toward penicillins.22
MRSA bacteria have rarely been examined in raw milk to evaluate microbial security, sanitation circumstances through milking, and storage periods. Thus, the existing survey was done to investigate the incidence rate, antimicrobial resistance properties and distribution of SCCmec types of the MRSA bacteria recovered from raw bovine, ovine, caprine, buffalo, and camel milk samples in Iran.
Materials and Methods
Samples
A total of 590 raw milk samples including bovine (n=130), ovine (n=120), caprine (n=120), camel (n=110), and buffalo (n=110) were randomly collected during a one-year period (2016 to 2017) from the shopping centers of different parts of Iran. None of the milk samples were not packed. All samples were stored in a refrigerator. Samples of raw milk were distributed by milk carrying specific trucks to shopping centers. A total of 50 mL were collected from each raw milk sample using a sterile laboratory tubes. Samples were proximately transferred to laboratory using cool bags. All milk samples presented usual physical properties such as odor, consolidation and color.
Isolation and Identification of S. aureus
Twenty-five grams of each of the collected samples were blended with 225 mL of buffered peptone water (EMD Millipore, Billerica, MA, USA). At that time, solutions were homogenized using Stomacher (Interscience, Saint-Nom, France). At that point, 5 mL of the achieved solution was transferred into 50 mL trypticase soy broth (TSB; EMD Millipore) supplemented with 10% NaCl and 1% sodium pyruvate and incubated for 18 h at 35°C. At that moment, a loopful of the culture was transferred into Baird-Parker agar supplemented with egg yolk tellurite emulsion (EMD Millipore) and incubated at 37°C for about 24 h. Black shiny colonies enclosed with significant zones were identified using biochemical tests as introduced before.23
Identification of Methicillin-Resistant S. aureus Bacteria
Antibiotic susceptibility tests were applied for this purpose. Susceptibility of S. aureus isolates were tested against cefoxitin (30 µg) and oxacillin (1 µg) antibiotic disks. Experiment was completed by the instructions of the Clinical and Laboratory Standards Institute (CLSI).24 Confirmation of MRSA isolates were additionally performed using the PCR-based detection of mecA gene.23
Antibiotic Susceptibility Test of MRSA Bacteria
Phenotypic pattern of antibiotic resistance of MRSA bacteria was investigated using the disk diffusion method on the Mueller–Hinton agar (EMD Millipore). Principles of CLSI were applied for this purpose.25 Diverse kinds of antibiotic agents including aminoglycosides (amikacin (30 µg/disk) and gentamicin (10 µg/disk)), fluoroquinolones (levofloxacin (5 µg/disk) and ciprofloxacin (5 µg/disk)), lincosamides (clindamycin (2 µg/disk)), macrolides (erythromycin (15 µg/disk) and azithromycin (15 µg/disk)), penicillins (penicillin (10 µg/disk), tetracyclines (doxycycline (30 µg/disk) and tetracycline (30 µg/disk)), phenicols (chloramphenicol (30 µg/disk)), folate pathway inhibitors (trimethoprim-sulfamethoxazole (25 µg/disk)) and ansamycins (rifampin (5 µg/disk)) were applied for this goal (Oxoid, UK). Method was completed using the protocol labeled beforehand.23,25
PCR-Based Amplification of Antibiotic Resistance Genes and SCCmec Types in MRSA Bacteria
Table 1 reveals the set of primers and PCR circumstances applied for detection of genotyping pattern of antibiotic resistance and SCCmec types.26–33 A programmable DNA thermo-cycler (Eppendorf Mastercycler 5330, Eppendorf-Nethel-Hinz GmbH, Hamburg, Germany) was applied for this goal.
 |
Table 1 Target Genes, Oligonucleotide Primers and PCR Conditions Used for Detection of Antibiotic Resistance Genes and SCCmec Types Amongst MRSA Bacteria Recovered from Raw Milk |
Statistical Analysis
SPSS 21.0 statistical software (IBM Corporation, Armonk, NY, USA) was applied for arithmetical analysis of data. Significant relations between data achieved from different groups and parameters were analyzed using the chi-squared test and Fisher’s exact two-tailed tests. P value <0.05 was determined as arithmetical significant level.
Results
Incidence of S. aureus and MRSA Bacteria
Table 2 signifies the distribution of S. aureus and MRSA bacteria in diverse kinds of raw milk samples. Thirty-nine out of 590 raw milk samples (6.61%) were positive for S. aureus. Raw buffalo (9.09%) and bovine (8.46%) milk samples had the maximum incidence of S. aureus, while raw camel (2.72%) milk samples had the minimum. Twenty-eight out of 39 (71.79%) bacteria were defined as MRSA bacteria. Raw buffalo (80%) and ovine (77.77%) milk samples had the maximum prevalence of MRSA bacteria, while raw camel (33.33%) milk samples had the minimum. Arithmetical important difference was seen for the prevalence of MRSA bacteria between buffalo and camel (P <0.05) and bovine and camel (P <0.05) raw milk samples.
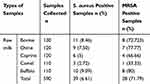 |
Table 2 Total Prevalence of S. aureus and MRSA Bacteria in Different Types of Raw Milk |
Antibiotic Resistance Pattern of MRSA Bacteria
Table 3 signifies the phenotypic pattern of antibiotic resistance of MRSA bacteria recovered from diverse kinds of raw milk samples. MRSA bacteria harbored the maximum incidence of resistance toward penicillin (100%), tetracycline (100%), erythromycin (82.14%), gentamicin (78.57%), trimethoprim-sulfamethoxazole (78.57%), and doxycycline (71.42%) antibiotic agents. MRSA bacteria exhibited lower incidence of resistance toward rifampin (14.28%), amikacin (17.85%), chloramphenicol (28.57%), azithromycin (32.14%), and levofloxacin (32.14%) antibiotic agents.
 |
Table 3 Antibiotic Resistance Pattern of the MRSA Bacteria Recovered from Raw Milk Samples |
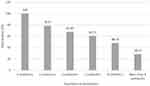
Prevalence of Multidrug Resistant MRSA Bacteria
Figure 1 signifies the incidence of resistance toward multiple groups of antibiotics. We found that all of the MRSA bacteria recovered from diverse kinds of raw milk samples had at least resistance toward four diverse classes of antibiotic agents, though incidence of resistance toward more than eight groups of antibiotics was 28.57%.
Distribution of Antibiotic Resistance Genes
Table 4 signifies the genotypic pattern of antibiotic resistance amongst the MRSA bacteria recovered from diverse kinds of raw milk samples. The most generally identified antibiotic resistance genes were blaZ (100%), tetK (85.71%), dfrA1 (71.42%) and aacA-D (67.85%). Incidence of ermA and gyrA antibiotic resistance genes were 50% and 42.85%, respectively. Incidence of msrB (10.71%), rpoB (10.71%), ermB (25%), cfr (25%), grlA (28.57%) and linA (28.57%) were lower than other identified resistance genes. Arithmetical important difference was seen between the incidence of ermA and ermB (P <0.05), msrA and msrB (P <0.05), tetK and tetM (P <0.05), and gyrA and grlA (P <0.05) antibiotic resistance genes.
 |
Table 4 Distribution of Antibiotic Resistance Genes Amongst the MRSA Bacteria Recovered from Raw Milk |
Distribution of SCCmec Types
Table 5 signifies the incidence of SCCmec types amongst the MRSA bacteria recovered from diverse kinds of raw milk samples. SCCmec IVa (29.62%), V (25%), III (14.81%) and IVb (11.11%) were the most routinely identified kinds amongst the MRSA bacteria. Incidence of SCCmec IVd (3.70%) and II (3.70%) was low. Arithmetical important difference was seen between the incidence of SCCmec types IVa and II (P <0.05), IVa and IVd (P <0.05), V and II (P <0.05) and V and IVd (P <0.05).
 |
Table 5 Distribution of SCCmec Types Amongst the MRSA Bacteria Recovered from Raw Milk |
Discussion
Prior to the 1990s, the majority of MRSA bacteria were hospital-associated (HA-MRSA) strains. Then, community-associated MRSA (CA-MRSA) prompted to occur infections outside the health-care and/or hospital environments. Recorded surveys revealed the occurrence of livestock-associated MRSA (LA-MRSA) in animals and/or livestock fields. The extensive developments in LA-MRSA and CA-MRSA have elevated the query as to whether MRSA is certainly a foodborne microbe.34 Furthermore, surveys on MRSA are interesting due to their considerable prevalence in diverse kinds of foodstuffs.34
Findings of the existing investigation revealed that the contamination rate of milk samples was 4.74% (28/590). Incidence of the MRSA in raw milk samples in our survey was lower than those of Italy (20%)35 and Turkey (17%),36 while it was higher than those of England (2.30%)37 and Germany (2.30%).38 Investigations conducted in the US along with other countries, including North America, Canada, Africa, Asia and Europe, have recovered MRSA mostly from dissimilar kinds of food and dairy samples.34,39 Some dairy animals are the main sources of MRSA bacteria. The possibility of primary presence of MRSA bacteria in raw milk samples due to the occurrence of sub-clinical mastitis in dairy animals and thus their transmission to raw milk, the opportunity of transmission of multidrug resistant MRSA from the milking halls, and also infected staff into the raw milk are the most important probable reasons for presence of MRSA bacteria.
Irregular and unauthorized prescription of antibiotics are the probable reasons for high prevalence of antibiotic resistance in the current survey. Additionally, boost incidence of antibiotic resistance was attended with boost incidence of specific antibiotic resistance genes. Furthermore, our findings showed that some of the MRSA bacteria exhibited higher incidence of resistance toward antibiotics used for human beings which can indirectly show their anthropogenic source. Conversely, some others exhibited higher incidence of resistance toward antibiotics used for animals which can circuitously demonstrate their animal origins. This conclusion was comparable with those of Hasanpour Dehkordi et al17 and Safarpoor Dehkordi et al20 which were both conducted on Iranian food samples. Comparable resistance of MRSA recovered from dissimilar kinds of foodstuffs and clinical specimens have also been determined toward aminoglycosides,19,20,40–43 cephems,19,20,40–42 penicillins,19,20,40–42 macrolides,19,20,40–42 tetracyclines,19,20,40,41 fluoroquinolones,19,20,40–43 lincosamides,19,20,40–42 folate inhibitors,19,20,40–43 phenicols19,20,40,41 and ansamycins19,20,40,41 antibiotic agents. Fowoyo and Ogunbanwo44 revealed that the S. aureus bacteria recovered from ready-to-eat foodstuffs exhibited the boost incidence of resistance toward trimethoprim–sulfamethoxazole (74.90%), ampicillin (86.70%), cefotaxime (3.50%), amoxicillin–clavulanic acid (52.50%), ciprofloxacin (23.90%), oxacillin (35.70%), gentamicin (11.40%), erythromycin (15.70%), and ofloxacin (7.10%) which was relatively similar to our findings. Boost incidence of resistance toward chloramphenicol (28.57%) maybe due to its unlawful and unselective prescription especially in veterinary medicine. Akanbi et al45 reported that blaZ, mecA, rpoB, ermB and tetM were the most generally identified antibiotic resistance genes amongst the S. aureus bacteria recovered from food samples in South Africa which was relatively similar to our findings. Similar to our findings, high distribution of mecA, gyrA, grlA and cfr was also described in the S. aureus bacteria recovered from chicken meat in Egypt.46 Another Iranian investigation47 showed that oxacillin, gentamicin, penicillin, tetracycline and erythromycin resistant S. aureus bacteria recovered from milk and dairy products carried considerable incidence of blaZ, aacA-aphD, mecA, tetK and tetM, ermB, ermA, ermT, ermC, msrB and msrA antibiotic resistance markers likewise to our survey.
Assess the distribution of SCCmec types is a practical method to find presence of HA-MRSA and CA-MRSA bacteria. Findings of epidemiological investigations revealed that presence of SCCmec types I, II and III indirectly showed occurrence of HA-MRSA bacteria, while presence of IV and V types represented the occurrence of CA-MRSA bacteria.48,49 Our findings showed that all of the SCCmec types had diverse distribution in the MRSA bacteria recovered from raw milk samples which may have assumed the presence of both HA and CA-MRSA bacteria. Moreover, SCCmec types IVa (29.62%) and V (25%) had the highest distribution amongst all studied elements. This finding may assume that most of the MRSA bacteria were probably originated from milk of infected animals. In keeping with this, SCCmec type III had also considerable prevalence (14.81%) which may assume that some of the MRSA bacteria had hospital or health-care origin and were probably transmitted from the contaminated workers of the milking halls. Johnson50 reported similar results for the boost incidence of SCCmec IV in retail meat samples. In a survey which was carried out by Vossenkuhl et al51 most of MRSA bacteria recovered from turkey meat samples carried SCCmec V (58.10–71.90%) and IVa (19–27.0%). Type III (0–1.2%) was detected periodically which was comparable to our findings. Zhang et al52 reported a the high prevalence of SCCmec III in their food samples. Boost incidence of SCCmec types IVa and V in food samples with animal origin has also been reported previously.38,53,54
Conclusions
By and large, we recognized boost incidence of S. aureus and MRSA bacteria in bovine, camel, caprine, ovine, and buffalo milk samples on top of boost incidence of genotypic and phenotypic profiles of antibiotic resistance and SCCmec types. The existing survey is the first report of the genotypic evaluation of antibiotic resistance and SCCmec typing of the MRSA bacteria in raw buffalo and camel milk samples. High prevalence of MRSA bacteria and substantial incidence of resistance toward erythromycin, penicillin, gentamicin, tetracycline, trimethoprim-sulfamethoxazole and doxycycline antibiotic agents and blaZ, tetK, dfrA1, aacA-D ermA and gyrA antibiotic resistant genes may pose a possible menace regarding the consumption of raw milk samples in Iran. Presence of multidrug resistant MRSA bacteria may show indiscriminate and unauthorized prescription of antibiotic agents in Iranian dairy animal farms. Most of MRSA bacteria harbored SCCmec types IV and V which may have assumed their possible community-acquired origins. However, some of the MRSA bacteria harbored SCCmec types I, II, and III which may assume their possible health care or hospital origins. Incidence of resistance toward human-based and also animal-based antibiotics can indirectly show the origin of MRSA bacteria. Ample boiling of raw milk beforehand consumption and prevention from cross-contamination can diminish the risk of virulent and resistant MRSA bacteria. However, supplementary surveys are necessary to comprehend more advanced epidemiological features of the MRSA bacteria in raw milk of dairy animal species.
Ethical Criteria
The contemporary survey was accepted by the ethical research committee of the Department of Food Hygiene, Faculty of Veterinary Medicine, Amol University of Special Modern Technologies, Amol, Iran.
Acknowledgments
Authors would like to thank Dr. Behsan Hemmatinezhad and Dr. Manoochehr Moumeni Sharaki for their clinical supports. Authors would also thank from Dr. Moshtaba Masoodimanesh for his imperative support in laboratory examinations.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Watson RR, Collier RJ, Preedy VR. Nutrients in Dairy and Their Implications for Health and Disease. Elsevier Academic Press; 2017.
2. Momtaz H, Safarpoor Dehkordi F, Taktaz T, Rezvani A, Yarali S. Shiga toxin-producing Escherichia coli isolated from bovine mastitic milk: serogroups, virulence factors, and antibiotic resistance properties. Sci World J. 2012;2012. doi:10.1100/2012/618709
3. Ranjbar R, Dehkordi FS, Shahreza MHS, Rahimi E. Prevalence, identification of virulence factors, O-serogroups and antibiotic resistance properties of Shiga-toxin producing Escherichia coli bacteria isolated from raw milk and traditional dairy products. Antimicrob Resist Infect Control. 2018;7(1):53. doi:10.1186/s13756-018-0345-x
4. Safarpoor Dehkordi F, Barati S, Momtaz H, Hosseini Ahari SN, Nejat Dehkordi S. Comparison of shedding, and antibiotic resistance properties of listeria monocytogenes isolated from milk, feces, urine, and vaginal secretion of bovine, ovine, caprine, buffalo, and camel species in Iran. Jundishapur J Microbiol. 2013;6(3):284–294. doi:10.5812/jjm.6616
5. Safarpoor Dehkordi F, Valizadeh Y, Birgani T, Dehkordi K. Prevalence study of brucella melitensis and brucella abortus in cow’s milk using dot enzyme linked immuno sorbent assay and duplex polymerase chain reaction. J Pure Appl Microbiol. 2014;8:1065–1069.
6. Hemmatinezhad B, Khamesipour F, Mohammadi M, Safarpoor Dehkordi F, Mashak Z. Microbiological investigation of O‐Serogroups, virulence factors and antimicrobial resistance properties of Shiga Toxin‐producing Escherichia coli isolated from Ostrich, Turkey and Quail meats. J Food Safe. 2015;35(4):491–500. doi:10.1111/jfs.12199
7. Atapoor S, Dehkordi FS, Rahimi E. Detection of Helicobacter pylori in various types of vegetables and salads. Jundishapur J Microbiol. 2014;7(5):e10013. doi:10.5812/jjm.10013
8. Safarpoor Dehkordi F, Parsaei P, Saberian S, et al. Prevalence study of theileria annulata by comparison of four diagnostic t. Bulgar J Vet Med. 2012;15:2.
9. Rahimi E, Sepehri S, Dehkordi FS, Shaygan S, Momtaz H. Prevalence of Yersinia species in traditional and commercial dairy products in Isfahan Province, Iran. Jundishapur J Microbiol. 2014;7:4. doi:10.5812/jjm.9249
10. Momtaz H, Davood Rahimian M, Safarpoor Dehkordi F. Identification and characterization of Yersinia enterocolitica isolated from raw chicken meat based on molecular and biological techniques. J Appl Poult Res. 2013;22(1):137–145. doi:10.3382/japr.2012-00549
11. Ghorbani F, Gheisari E, Dehkordi FS. Genotyping of vacA alleles of Helicobacter pylori bacteria isolated from some Iranian food items. Trop J Pharm Res. 2016;15(8):1631–1636. doi:10.4314/tjpr.v15i8.5
12. Safarpoor Dehkordi F, Khamesipour F, Momeni M. Brucella abortus and Brucella melitensis in Iranian bovine and buffalo semen samples: the first clinical trial on seasonal, senile and geographical distribution using culture, conventional and real-time polymerase chain reaction assays. Kafkas Uni Vet Fak Derg. 2014;20(6):821–828. doi:10.9775/kvfd.2014.10827
13. Safarpoor Dehkordi F, Haghighi N, Momtaz H, Rafsanjani MS, Momeni M. Conventional vs real-time PCR for detection of bovine herpes virus type 1 in aborted bovine, buffalo and camel foetuses. Bulgar J Vet Med. 2013;16(2):102−111.
14. Nejat S, Momtaz H, Yadegari M, Nejat S, Safarpour Dehkordi F, Khamesipour F. Seasonal, geographical, age and breed distributions of equine viral arteritis in Iran. Kafkas Univ Vet Fak Derg. 2015;21(1):111–116. doi:10.9775/kvfd.2014.11934
15. Rahimi E, Yazdanpour S, Dehkordi F. Detection of Toxoplasma gondii antibodies in various poultry meat samples using enzyme linked immuno sorbent assay and its confirmation by polymerase chain reaction. J Pure Appl Microbiol. 2014;8(1):421–427.
16. Ranjbar R, Masoudimanesh M, Dehkordi FS, Jonaidi-Jafari N, Rahimi E. Shiga (Vero)-toxin producing Escherichia coli isolated from the hospital foods; virulence factors, o-serogroups and antimicrobial resistance properties. Antimicrob Resist Infect Control. 2017;6(1):4. doi:10.1186/s13756-016-0163-y
17. Hasanpour Dehkordi A, Khaji L, Sakhaei Shahreza M, et al. One-year prevalence of antimicrobial susceptibility pattern of methicillin-resistant Staphylococcus aureus isolated from raw meat. Trop Biomed. 2017;34(2):396–404.
18. Madahi H, Rostami F, Rahimi E, Dehkordi FS. Prevalence of enterotoxigenic Staphylococcus aureus isolated from chicken nugget in Iran. Jundishapur J Microbiol. 2014;7:8. doi:10.5812/jjm.10237
19. Momtaz H, Dehkordi FS, Rahimi E, Asgarifar A, Momeni M. Virulence genes and antimicrobial resistance profiles of Staphylococcus aureus isolated from chicken meat in Isfahan province, Iran. J Appl Poult Res. 2013;22(4):913–921. doi:10.3382/japr.2012-00673
20. Safarpoor Dehkordi F, Gandomi H, Akhondzadeh Basti A, Misaghi A, Rahimi E. Genotypic and phenotypic characterization of antibiotic resistance of methicillin-resistant Staphylococcus aureus isolated from hospital food. Antimicrob Resist Infect Control. 2017;6(1):104. doi:10.1186/s13756-017-0257-1
21. Klevens RM, Morrison MA, Nadle J, et al. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. J Am Med Assoc. 2007;298(15):1763–1771. doi:10.1001/jama.298.15.1763
22. Liu J, Chen D, Peters BM, et al. Staphylococcal chromosomal cassettes mec (SCCmec): a mobile genetic element in methicillin-resistant Staphylococcus aureus. Microb Pathog. 2016;101:56–67. doi:10.1016/j.micpath.2016.10.028
23. Fijałkowski K, Peitler D, Karakulska J. Staphylococci isolated from ready-to-eat meat–identification, antibiotic resistance and toxin gene profile. Int J Food Microbiol. 2016;238:113–120. doi:10.1016/j.ijfoodmicro.2016.09
24. CLSI. Performance standards for antimicrobial susceptibility testing.
25. CLSI. Performance standards for antimicrobial susceptibility testing.
26. Lina G, Quaglia A, Reverdy M-E, Leclercq R, Vandenesch F, Etienne J. Distribution of genes encoding resistance to macrolides, lincosamides, and streptogramins among staphylococci. Antimicrob Agent Chemother. 1999;43(5):1062–1066. doi:10.1128/AAC.43.5.1062
27. Strommenger B, Kettlitz C, Werner G, Witte W. Multiplex PCR assay for simultaneous detection of nine clinically relevant antibiotic resistance genes in Staphylococcus aureus. J Clin Microbiol. 2003;41(9):4089–4094. doi:10.1128/jcm.41.9.4089-4094.2003
28. Aboshkiwa M, Rowland G, Coleman G. Nucleotide sequence of the Staphylococcus aureus RNA polymerase rpoB gene and comparison of its predicted amino acid sequence with those of other bacteria. Biochim Biophys Acta. 1995;1262(1):73–78. doi:10.1016/0167-4781(95)00054-k
29. Schmitz F-J, Jones ME, Hofmann B, et al. Characterization of grlA, grlB, gyrA, and gyrB mutations in 116 unrelated isolates of Staphylococcus aureus and effects of mutations on ciprofloxacin MIC. Antimicrob Agent Chemother. 1998;42(5):1249–1252. doi:10.1128/AAC.42.5.1249
30. Roberts MC, Sutcliffe J, Courvalin P, Jensen LB, Rood J, Seppala H. Nomenclature for macrolide and macrolide-lincosamide-streptogramin B resistance determinants. Antimicrob Agent Chemother. 1999;43(12):2823–2830. doi:10.1128/AAC.43.12.2823
31. Tang J, Zhang R, Chen J, et al. Incidence and characterization of Staphylococcus aureus bacteria isolated from food markets. Ann Microbiol. 2015;65(1):279–286. doi:10.1007/s13213-014-0859-2
32. Kehrenberg C, Schwarz S. Distribution of florfenicol resistance genes fexA and cfr among chloramphenicol-resistant Staphylococcus isolates. Antimicrob Agent Chemother. 2006;50(4):1156–1163. doi:10.1128/AAC.50.4.1156-1163.2006
33. Shittu AO, Okon K, Adesida S, et al. Antibiotic resistance and molecular epidemiology of Staphylococcus aureus in Nigeria. BMC Microbiol. 2011;11(1):92. doi:10.1186/1471-2180-11-92
34. Wendlandt S, Schwarz S, Silley P. Methicillin-resistant Staphylococcus aureus: a food-borne pathogen? Ann Rev Food Sci Technol. 2013;4:117–139. doi:10.1146/annurev-food-030212-182653
35. Riva A, Borghi E, Cirasola D, et al. Methicillin-resistant Staphylococcus aureus in raw milk: prevalence, SCC mec typing, enterotoxin characterization, and antimicrobial resistance patterns. J Food Prot. 2015;78(6):1142–1146. doi:10.4315/0362-028X.JFP-14-531
36. Paterson G, Morgan F, Harrison E, et al. Prevalence and characterization of human mecC methicillin-resistant Staphylococcus aureus isolates in England. J Antimicrob Chemother. 2013;69(4):907–910. doi:10.1093/jac/dkt462
37. Türkyılmaz S, Tekbıyık S, Oryasin E, Bozdogan B. Molecular epidemiology and antimicrobial resistance mechanisms of methicillin‐resistant Staphylococcus aureus isolated from bovine milk. Zoonoses Publ Health. 2010;57(3):197–203. doi:10.1111/j.1863-2378.2009.01257.x
38. Kreausukon K, Fetsch A, Kraushaar B, et al. Prevalence, antimicrobial resistance, and molecular characterization of methicillin-resistant Staphylococcus aureus from bulk tank milk of dairy herds. J Dairy Sci. 2012;95(8):4382–4388. doi:10.3168/jds.2011-5198
39. Sergelidis D, Angelidis A. Methicillin‐resistant Staphylococcus aureus: a controversial food‐borne pathogen. Lett Appl Microbiol. 2017;64(6):409–418. doi:10.1111/lam.12735
40. Paludi D, Vergara A, Festino AR, et al. Antimicrobial resistance pattern of methicillin-resistant Staphylococcus aureus in the food industry. J Bio Reg Homeostatic Agent. 2011;25(4):671.
41. Sallam KI, Abd-Elghany SM, Elhadidy M, Tamura T. Molecular characterization and antimicrobial resistance profile of methicillin-resistant Staphylococcus aureus in retail chicken. J Food Prot. 2015;78(10):1879–1884. doi:10.4315/0362-028X.JFP-15-150
42. Jackson CR, Davis JA, Barrett JB. Prevalence and characterization of methicillin-resistant Staphylococcus aureus isolates from retail meat and humans in Georgia. J Clin Microbiol. 2013;51(4):1199–1207. doi:10.1128/JCM.03166-12
43. Daka D, Yihdego D. Antibiotic-resistance Staphylococcus aureus isolated from cow’s milk in the Hawassa area, South Ethiopia. Ann Clin Microbiol Antimicrob. 2012;11(1):26. doi:10.1186/1476-0711-11-26
44. Fowoyo P, Ogunbanwo S. Antimicrobial resistance in coagulase-negative staphylococci from Nigerian traditional fermented foods. Ann Clin Microbiol Antimicrob. 2017;16(1):4. doi:10.1186/s12941-017-0181-5
45. Akanbi OE, Njom HA, Fri J, Otigbu AC, Clarke AM. Antimicrobial susceptibility of Staphylococcus aureus isolated from Recreational waters and beach sand in Eastern Cape province of south Africa. Int J Environ Res Publ Health. 2017;14(9):1001. doi:10.3390/ijerph14091001
46. Osman K, Badr J, Al-Maary KS, et al. Prevalence of the antibiotic resistance genes in coagulase-positive-and negative-staphylococcus in chicken meat retailed to consumers. Front Microbiol. 2016;7:1846. doi:10.3389/fmicb.2016.01846
47. Jamali H, Paydar M, Radmehr B, Ismail S, Dadrasnia A. Prevalence and antimicrobial resistance of Staphylococcus aureus isolated from raw milk and dairy products. Food Control. 2015;54:383–388. doi:10.1016/j.foodcont.2015.02.013
48. David MZ, Cadilla A, Boyle-Vavra S, Daum RS. Replacement of HA-MRSA by CA-MRSA infections at an academic medical center in the midwestern United States, 2004–5 to 2008. PLoS One. 2014;9(4):e92760. doi:10.1371/journal.pone.0092760
49. Valsesia G, Rossi M, Bertschy S, Pfyffer GE. Emergence of SCCmec type IV and SCCmec type V methicillin-resistant Staphylococcus aureus containing the Panton-Valentine leukocidin genes in a large academic teaching hospital in central Switzerland: external invaders or persisting circulators? J Clin Microbiol. 2010;48(3):720–727. doi:10.1128/JCM.01890-09
50. Johnson AP. Methicillin-resistant Staphylococcus aureus: the European landscape. J Antimicrob Chemother. 2011;66(suppl_4):iv43–iv48. doi:10.1093/jac/dkr07
51. Vossenkuhl B, Brandt J, Fetsch A, et al. Comparison of spa types, SCCmec types and antimicrobial resistance profiles of MRSA isolated from turkeys at farm, slaughter and from retail meat indicates transmission along the production chain. PLoS One. 2014;9(5):e96308. doi:10.1371/journal.pone.0096308
52. Zhang K, McClure J-A, Elsayed S, Louie T, Conly JM. Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. J Clin Microbiol. 2005;43(10):5026–5033. doi:10.1128/JCM.43.10.5026-5033.2005
53. Argudín M, Mendoza M, González-Hevia M, Bances M, Guerra B, Rodicio M. Genotypes, exotoxin gene content and antimicrobial resistance in Staphylococcus aureus isolated from foods and food-handlers. Appl Environ Microbiol. 2012;07411–07487. doi:10.1128/AEM.07487-11
54. Bhargava K, Wang X, Donabedian S, Zervos M, da Rocha L, Zhang Y. Methicillin-resistant Staphylococcus aureus in retail meat, Detroit, Michigan, USA. Emerg Infect Dis. 2011;17(6):1135. doi:10.3201/eid/1706.101905
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

