Back to Journals » Infection and Drug Resistance » Volume 14
Functional Insights of MraZ on the Pathogenicity of Staphylococcus aureus
Authors Wang B, Duan J, Jin Y, Zhan Q, Xu Y, Zhao H, Wang X, Rao L, Guo Y, Yu F
Received 5 August 2021
Accepted for publication 8 October 2021
Published 2 November 2021 Volume 2021:14 Pages 4539—4551
DOI https://doi.org/10.2147/IDR.S332777
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Bingjie Wang,1,* Jingjing Duan,2,* Ye Jin,3,* Qing Zhan,4 Yanlei Xu,4 Huilin Zhao,1 Xinyi Wang,1 Lulin Rao,3 Yinjuan Guo,1 Fangyou Yu1,5
1Department of Clinical Laboratory, Shanghai Pulmonary Hospital, School of Medicine, Tongji University, Shanghai, People’s Republic of China; 2Department of Clinical Laboratory, Renmin Hospital, Hubei University of Medicine, Hubei, People’s Republic of China; 3Department of Laboratory Medicine, The First Affiliated Hospital of Wenzhou Medical University, Wenzhou, People’s Republic of China; 4Jiangxi Provincial Key Laboratory of Preventive Medicine, School of Public Health, Nanchang University, Nanchang, People’s Republic of China; 5Shanghai Key Laboratory of Tuberculosis, Shanghai Pulmonary Hospital, Tongji University School of Medicine, Shanghai, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Fangyou Yu; Yinjuan Guo Email [email protected]; [email protected]
Introduction: In recent years, multidrug-resistant methicillin-resistant Staphylococcus aureus has become increasingly prevalent, which raised a huge challenge to antibiotic treatment of infectious diseases. The anti-virulence strategy targeting virulent factors is a promising novel therapy for S. aureus infection. The virulence mechanism of S. aureus was needed to explore deeply to develop more targets and improve the effectiveness of anti-virulence strategies.
Results: In this study, we found mraZ was highly conserved in S. aureus, and its production is homologous with the MraZ of Escherichia coli, a transcriptional regulator involved in the growth and cell division of E. coli. To investigate the function of mraZ in S. aureus, we constructed a MW2 mraZ deletion mutant and its complementary mutant for virulence comparison. Although no remarkable influence on the growth, the mraZ deletion mutant led to significantly reduced resistance to human neutrophils and decreased virulence in Galleria mellonella model as well as mouse skin and soft tissue infection models, indicating its essential contribution to virulence and immune evasion to support the pathogenicity of S. aureus infection. RNA-Seq and quantitative RT-qPCR revealed that MraZ is a multi-functional regulator; it involves in diverse biological processes and can up-regulate the expression of various virulence genes by agr and sarA.
Conclusion: mraZ plays vital roles in the pathyogenicity of S. aureus via regulating many virulence genes. It may be an attractive target for anti-virulence therapy of S. aureus.
Keywords: Staphylococcus aureus, virulence, mraZ, agr, sarA
Introduction
Staphylococcus aureus is a common opportunistic pathogen that usually colonizes in human skin and nasal mucosa, which can cause a variety of infections, ranging from mild skin lesion to severe invasive diseases such as endocarditis, osteomyelitis, pneumonia, and even life-threatening septic shock.1–3 Moreover, with the emergence of multidrug-resistant methicillin-resistant S. aureus (MRSA) strains in recent years, S. aureus infections rise a serious concern in public health. A report from Centers for Disease Control and Prevention from USA showed that the mortality rate caused by MRSA infection was 2.88 per 100 000 population per year, which leads to approximately 100 trillion dollars in damage to the global economy.4 It is obvious that the problem of antibiotic resistance in bacteria has reached the crisis stage.
A key characteristic in pathogenicity of S. aureus that promote its infection is the production of a large array of virulence factors, such as hemolysins, pore-forming leukocidins, enterotoxin, and plasma-coagulase,5–8 which allow them to resist clearance by the host, to invade and gain access to deeper tissues, and to damage host cells. Currently, most treatments of infectious diseases in clinical are based on the ability of antibacterial agents to kill pathogens quickly. However, the harmful effects of bacterial virulence factors are almost inevitable Furthermore, treatment of infections caused by MRSA strains has become more difficult because of the emergence of multidrug-resistant strains. As conventional antibiotics become less valid, the anti-virulence strategies are drawing increasing attention. One of the advantages of the anti-virulence strategies is that it can specifically block the virulence expression of bacteria without killing or inhibiting the growth of bacteria pathogens, and avoid the selective pressure from antibiotics treatment, which will be less likely lead to the emergence of drug resistance.9,10 Therefore, the investigation of more unidentified virulence factors of Staphylococcus aureus and its pathogenic mechanism can provide more directions for the anti-virulence strategies of Staphylococcus aureus.
The mraZ gene is highly conserved in many bacterial species. In some Gram-positive bacteria, mraZ is associated with the cell division and cell wall biosynthesis, which plays a vital role in the growth of bacteria.11–13 A previous study demonstrated that the overproduction of Escherichia coli MraZ could inhibit cell division and was lethal in rich medium at high induction levels and in minimal medium at low induction levels.11 Notably, the MraZ from E. coli and S. aureus shares 36% sequence identity. However, limited studies have evaluated the role of mraZ gene in S. aureus. The biological functions of the mraZ gene in S. aureus still require further investigation.
In this study, to better understand the function of MraZ in S. aureus, we constructed the MW2 mraZ deletion mutant and its complementary mutant, and investigated the effects of MraZ on the bacterial growth, the virulence and immune evasion capacity. We found that MraZ is a multi-functional regulator, especially plays a crucial part in S. aureus virulence and immune evasion by regulating the expression of extensive virulence genes.
Materials and Methods
Bacterial Strains and Culture Conditions
The bacterial strains and plasmids used in this study are listed in Table 1. Given MW2 is a typical community-acquired MRSA isolate with enhanced virulence, and its genetic background has been well established, it was used for the construction of mraZ knockout and complementation. E. coli DH5α and E. coli DC10B was used for staphylococcal cloning host. E. coli strains were cultured in Luria-Bertani (LB) medium, and Staphylococcus strain were grown at tryptic soy broth (TSB) medium. All strains were shaken in an incubator at 37 °C, ~220 rpm (adjusted appropriately). The medium was supplemented with an appropriate antibiotic (ampicillin 100 mg/L, chloramphenicol 10 mg/L and anhydrous tetracycline 50 μg/L) as needed.
 |
Table 1 The Bacterial Strains and Plasmids Used in the Present Study |
Construction of mraZ Knockout Mutant and Complemented Strain
A knockout mutant of mraZ was constructed using the temperature-sensitive plasmid pKOR1 based on the homologous recombination.14 Briefly, the upstream and downstream fragments of mraZ were amplified, linked by the T4 enzyme, and ligated to the vector pKOR1, resulting in recombinant pKOR1-ΔmraZ. This recombinant plasmid was consecutively transferred into Escherichia coli DH5α for amplification, into DC10B for bypass the S. aureus restriction barriers, and then electrotransfered into MW2. Under the dual pressure of high temperature and antibiotics, ΔmraZ was generated by the homologous recombination of the upstream and downstream homologous arm in the pKOR1-ΔmraZ with the MW2 genome, as previously described.14 The deleted mutant ΔmraZ was verified by PCR with DNA sequencing and quantitative reverse transcription-PCR (qRT-PCR).
To further verify whether the biological alteration of the deletion mutant was caused by the absence of mraZ, we constructed the complementary mutant strain ΔmraZ-C using the shuttle plasmid pRB473. Briefly, the marZ with its promoter regions was amplified and inserted into pRB473. The pRB473-mraZ was transferred into ΔmraZ by electroporation. The primers used above are listed in Table 2.
 |
Table 2 Primers Used for Construction of Δmraz and Δmraz-C |
RNA Extraction and qRT-PCR
Total RNA of S. aureus was extracted using RNA extraction kit (Tiangen Biotech, Beijing, China) following the manufacturer’s instructions. Briefly, overnight cultures were diluted 1:200 into fresh TSB and incubated at 37°C. After 9 h, the prepared lysostaphin and lysozyme solution were mixed with bacterium suspension for disrupt the cell walls, and then RNA extracted was followed the kit instructions. A PureLink RNA Mini Kit (Invitrogen, Carlsbad, CA, USA) was used to get purified RNA, and then cDNA was obtained using a PrimeScript RT reagent kit (TaKaRa, Japan). The housekeeping gene gyrB was used as an internal reference gene, and relative quantification of Expression levels were calculated by the 2−ΔΔCt method.15 The primer pairs used in qRT-PCR are listed in Table 3.
 |
Table 3 Primers Used in qRT-PCR |
Growth Assay
S. aureus strains were grown to stationary phase (12h) and then diluted (1:200) in TSB medium. All the cultures were incubated at 37°C with shaking at 220rpm. The OD600 value was measured hourly for 24h for bacterial growth curves. Meanwhile, the cells were plated in serial dilutions on TSB agar at 1h intervals for 24h, and the colony-forming units (CFU) were counted for viable bacterial count at indicated time points. The assay was performed in duplicate.
Analysis of Pigments Production
To evaluate pigment production, bacteria were incubated into 3mL TSB medium at 37°C with shaking. After 24h, the cultures were centrifuged at 8000 rpm for 10 min, and pigment production was assessed visually by the color of centrifugated precipitation.
Hemolytic Capacity
The hemolytic capacity of S. aureus isolates was evaluated based on the zone of clearing around colonies plated on sheep blood agar plates. Briefly, S. aureus strains were cultivated to stationary phase (12h), the cells were collected, washed twice with PBS, and adjusted to OD600=1; then 10ul each culture suspension was spotted on sheep blood agar plates. The zones of hemolysis (diameter) were measured after 24h of 37°C incubation. The experiments were conducted in triplicate.
Neutrophil Killing Assays
Neutrophils with >95% purity and viability were isolated from the venous blood of healthy volunteers using the Human Neutrophil Isolation Kit (Haoyang, Tianjin, China). In brief, S. aureus isolates were grown to mid-logarithmic growth phase, collected, washed, and resuspended in PBS. After the opsonization of 5% normal human serum, 1.0×105 CFU bacterial suspension were mixed with 1.0×106 neutrophils in RPMI 1640 medium at 37°C with gentle rotation. Samples were taken in 1h, 2h, and 3h for CFU enumeration. 0.25%TritonX-100 was added to each sample to ensure lysis of phagosomes before diluting and plating on TSB agar. The experiments were done in triplicate.
Galleria mellonella Infection Model
As previously described, larvae of G. mellonella weighing 250–350mg (Tianjin huiyude Biotech, Tianjin, China) were stored in the dark at 10°C and used within a week of delivery. Overnight cultures of S. aureus were collected, washed, and resuspended in PBS. G. mellonella was injected with 3.0×106 CFU/larvae, via the last right proleg using Hamilton syringe. Each experimental group contained 10 larvae. Larvae with injected sterile PBS and uninjected larvae as control. The treated larvae were placed in clean, sterile plastic petri dish and incubated at 37°C. The survival rate was monitored every 12h for 3 days. The results were analyzed by GraphPad Prism 6 software. The experiment was repeated three times with similar results.
Mouse Model of Skin Abscess
The mouse skin infection model was performed as described previously. Female BALB/C nude mice aged 4 to 6 weeks old and weighing about 22g were used for the experiments. In brief, all mice were administered with sterile feed, pure water one week before the time of use to adapt to the experimental environment. S. aureus strains were grown to mid-logarithmic phase as described above. The mice were injected with 100μL PBS containing 1.0×107 CFU S. aureus, and mice injected with 100μL PBS as control. Each experimental group contained 10 mice. Abscess formations were observed and recorded daily for 7 days. The length (L) and width (W) were measured to calculate the size of the abscesses follow the formula: A =π (L×W)/2. All animals were sacrificed 7 days post infection.
Cytokine Expression in Bacteremia Mouse Model
BALB/C nude mice were immunodeficient animals with reduced cellular immunity and humoral immune function, which might affect the production of inflammatory cytokines. Therefore, BALB/C female mice with complete immunity capacity aged 4 to 6 weeks old were used for the bacteremia model and were tested for the expression levels of cytokine. Briefly, mice were randomly divided into four groups, and each group contains 10 mice. The mice were infected by intraperitoneal injection with 100ul PBS containing 1.0×107 CFU S. aureus, and mice injected with 100μL PBS as control. They were raised under the same conditions for 3L days, and then 7 mice were randomly selected for further analysis (exclude the death mice). Blood samples were obtained, and the level of cytokines in the sera of each mouse was detected by ELISA method.
RNA-Sequencing and Data Analysis
S. aureus strains were inoculated for 9h in TSB medium at 37°C, and total RNA was extracted as above. Qualitative RNA samples were analyzed by RNA-seq (transcriptome sequencing) using an Illumina HiSeq X platform and the pe150 (150 bp double-stranded assay) strategy. Differential expression analysis was performed by DEG-seq software on the RNA sequencing reads. Gene was considered significantly differentially regulated with log2 (fold change) | > 1 and P< 0.005.
Statistical Analysis
Statistical comparisons were performed using SPSS (version 16.0) and GraphPad Prism 8. Two groups comparison was analyzed using unpaired t-test. Three or more groups comparison was analyzed using one-way or two-way ANOVA. The survival curve was analyzed by the Log rank test (Mantel-Cox). The threshold for statistical significance adopted in all analyzes was P<0.05.
Ethics Statement
All procedures involving human participants were performed in accordance with the ethical standards of First Affiliated Hospital of Wenzhou Medical University. All volunteers gave written informed consent in accordance with the Declaration of Helsinki prior to donating blood. All animal assays were carried out in accordance with The Regulation on the Management of Laboratory Animals for the welfare of the laboratory animals, and the protocol was approved by the Institutional Animal Care and Ethics Committee at First Affiliated Hospital of Wenzhou Medical University.
Results
Construction of mraZ Deletion Mutant Strains
In the genome of S. aureus MW2 (GeneBank accession number: GCF_000011265.1), mraZ is a 432bp gene encoding a 143 amino acid protein with contained SpoVT-AbrB DNA-binding domains, and it is located downstream of the putative cysteine ligase encoding gene bshC and upstream of the 16S rRNA methyltransferases encoding gene rsmH.
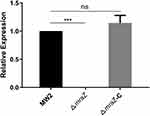
The temperature-sensitive shuttle plasmid pKOR1 was used to generate the ΔmraZ mutant in the MW2 strain. The complementary strain of ΔmraZ mutant named ΔmraZ-C was constructed using the vectors pRB473. All mutants were verified by PCR with sequencing and qRT–PCR. As shown in Figure 1, the ΔmraZ mutant and ΔmraZ-C were successfully constructed.
MraZ Has No Effect on the Growth and Pigment Production, While Enhances the Hemolysis Capacity of S. aureus
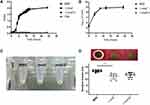
In order to investigate the potential impact of the mraZ deletion on S. aureus growth rate, the growth curves and the viable bacterial counts of strains MW2, ΔmraZ and ΔmraZ-C were determined. Compared with MW2 and ΔmraZ-C, no substantial differences in the growth curve or the viable bacterial count were found in ΔmraZ, indicating that the absence of mraZ gene did not affect the growth of S. aureus (Figure 2A and B). We also found that there is no difference in pigment production between ΔmraZ and MW2 or ΔmraZ-C (Figure 2C). We further examined the hemolysis ability of the bacteria, one of the most important virulence phenotypes of S. aureus, and found that the hemolytic capability of ΔmraZ was significantly reduced (Figure 2D).
MraZ Makes a Significant Contribution to Virulence of S. aureus in the G. mellonella Infection Model
To investigate whether the absence of mraZ can lead to less virulence, we first compared the survival rates of the wide-type strain MW2, ΔmraZ deletion mutant and ΔmraZ-C in G. mellonella infection model. As shown in Figure 3, the survival rate of G. mellonella infected with ΔmraZ deletion mutant was significantly higher than the wide-type strain, while the virulence of ΔmraZ-C was restored to the wild-type level.
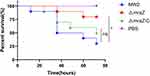 |
Figure 3 Galleria mellonella infection model. The survival rates of the G. mellonella after infection with MW2, ΔmraZ and ΔmraZ-C were expressed by the Kaplan-Meier survival plot. *p<0.05; ns p≥0.05. |
MraZ of S. aureus Promotes the Formation of Skin Abscesses in Mouse
To further verify the role of mraZ in the invasiveness and acute virulence potential of S. aureus, we established mouse models of skin abscess infection against MW2, ΔmraZ and ΔmraZ-C and compared their ability to cause pathological lesions or abscesses. As show in Figure 4, all the mice infected with these three stains were found skin lesion; however, the abscess area of the ΔmraZ was significantly smaller than the MW2, indicating that MraZ plays a considerable role in the process of abscess formation.
MraZ of S. aureus Helps Evade the Elimination of Human Neutrophils
To determine the role of mraZ in S. aureus immune evasion, we evaluated the survival rates of the wide-type strain MW2, ΔmraZ and ΔmraZ-C in human neutrophils. As shown in Figure 5, the survival rate of ΔmraZ was lower than the parent strain after 1 h of incubation (p < 0.05). After incubated for 2h, a significant reduction of the survival rate of ΔmraZ deletion mutant was found compared with the wide-type strain MW2. It indicated that the presence of mraZ gene could help resist clearance by the host innate immune system, thereby likely contributing to the promotion of virulence.
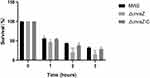 |
Figure 5 Survival rates of S. aureus MW2, ΔmraZ and ΔmraZ-C after incubated with human neutrophils. *p<0.05; **p<0.01; ***p<0.001. |
The mraZ Deletion Mutant Has Lower Pro-Inflammatory Cytokine Expression Levels in Mouse Bacteremia Model
Cytokine play major roles in the innate immune system by the interaction of pro-inflammatory cytokines such as TNF-α and IL-6 and anti-inflammatory cytokines such as IL-10 and IL-4. Therefore, we injected the bacteria into abdominal cavity of mice and determined the cytokines levels of MW2, ΔmraZ and ΔmraZ-C to evaluate the role of mraZ in the inflammatory response in S. aureus infection. As shown in Figure 6, the levels of TNF-α and IL-6 in ΔmraZ were both significantly lower than the other two strains (p < 0.01), while IL-4 and IL-10 have no significant difference in these strains, suggesting that the presence of mraZ can stimulate the production of pro-inflammatory cytokines and promote the inflammatory response to S. aureus.
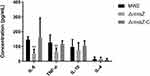 |
Figure 6 Comparison of Cytokine expression levels in mice after infection with MW2, ΔmraZ and ΔmraZ-C, respectively. **p<0.01. |
Transcriptome Comparison of MW2 and Δmraz
With the involvement of the marZ gene in virulence and immune evasion of S. aureus, we speculated MraZ might involve in regulating the expression of various genes. Therefore, we performed RNA-Seq analysis on MW2 and MW2ΔmraZ. Compared with the wild strain of MW2, there were 335 genes in the ΔmraZ mutant strain with differential expression ≥2-fold, of which 187 were up-regulated and 148 were down-regulated (Figure 7A). Furthermore, The Go enrichment analysis revealed that the differentially expressed genes (DEGs) are related to multiple functions, mainly the decomposition and anabolism of multiple substances, DNA replication and repair, gene expression regulation, ribosome synthesis, material transport, cell division and infection (Figure 7B). Therefore, MraZ can act as a multi-functional regulator and affect the expression of various genes.
In the mraZ deletion mutant, the transcription levels of the two-component system agr was down-regulated 8.51-fold (23.09-fold). The expression of dlt, an operon mediates d-alanine incorporation into teichoic acid and a determinant of defensin resistance that is essential for resistance to human neutrophil killing and virulence in mice,16 was down-regulated 2.58-fold (21.37-fold) in MW2ΔmraZ. In addition, the transcription of clfA encoding fibrinogen-binding MSCRAMM clumping factors A was significantly down-regulated 2.16-fold (21.11 fold), which might associated with the decreased virulence of S. aureus17 (Table 4). Although the expression of Staphylokinase encoding gene sak, superantigen-like protein encoding gene ssl13 were up-regulated, the transcription of the beta-class phenol-soluble modulin (PSM) encoding gene psmβ, alpha family PSM encoding gene psmɑ 3 and hld, Alpha-hemolysin encoding gene hly/hla, and enterotoxin type C encoding gene sec4 were down-regulated 61.82-fold (25.95 fold), 50.21-fold (25.65 fold), 33.13-fold (25.05 fold), 2.13-fold (21.09 fold) and 3.12-fold (21.64 fold), respectively (Table 4). These data suggested that the decreased virulence and compromised immune phenotype resulted from the absence of mraZ was associated with the down-regulated expression of agr, dlt, clfA, psmɑ, psmβ, hly/hla, hld and sec4.
 |
Table 4 Gene Expression Changes Associated with Virulence or Immune Evasion |
The Transcription Levels of Well-Established Virulence Factors and Regulatory Genes in the MW2, Δmraz and Δmraz-C
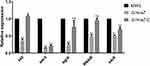
According to the absence of mraZ contributed to the virulence and immune evasion phenotype, we next measured the expression levels of several well-established virulence factors and regulator genes in detail in the MW2, ΔmraZ and ΔmraZ-C by qRT-PCR. As shown in Figure 8, the expression of hld and sec4 were significantly down-regulated in ΔmraZ strain (p<0.05); hemolysin and enterotoxin encoded by these two genes were the topical exotoxins contributed to the virulence of S. arueus, conforming that the mraZ affected the expression of virulence factors such as hemolysin or enterotoxin in S. aureus. The agr system consists of two adjacent transcripts RNA II and RNA III. RNA III can drive expression of most agr dependent virulence factors, such as the Alpha-hemolysin and delta-hemolysin, and agrA can promote the transcription of PSM genes.18 In agreement with the transcriptome data, the expression level of the agrA and RNAIII in ΔmraZ were significantly lower than the MW2 strain (p < 0.05). Furthermore, we also investigated the expression of the major S. aureus virulence regulatory locus sarA under the same conditions. We found that the expression level of sarA in ΔmraZ also significantly decreased (p<0.05). Additionally, except sec4, the expression of most above measured virulence genes and regulatory genes in the MW2ΔmraZ-C were approximately recovered to the wild-type strain MW2 (Figure 8). Therefore, it is likely that the mraZ can activate the expression of the virulence factors by up regulating several key regulatory genes such as agr and sarA.
Discussion
In recent years, the anti-virulence strategies targeting multiple virulence factors of S. aureus have been proved to be an effective alternatives to antibacterial drugs in the treatment of bacterial infection; it is reported that clonal antibody against ɑ-hemolysin could serve as an effective target for monoprophylaxis or adjuvant therapy for S. aureus pneumoniae;19 Diapophytoene desaturase CrtN, a key catalytic enzyme of the golden pigment biosynthesis pathway, can also effectively block the virulence of bacteria by targeting drugs.20 However, S. aureus strains usually contain various virulence genes and display heterogeneous virulence profiles. In this study, for the first time, we revealed that MraZ highly conserved in S. aureus is a multi-functional regulator, and elucidated that it plays an essential role in the virulence and the innate host defense of S. aureus. Our research suggested a promising target for anti-virulence strategy for S. aureus infection.
Studies involving mraZ in E. coli, Corynebacterium glutamicum and other Gram-negative bacteria have described its position at the head of the division cell wall (dcw) operon (mraZ-mraW-ftsL-ftsI-murE-murF-mraY-murD-ftsW-murG-murC-ftsQ-ftsA-ftsZ-envA), which responsible for cell division and cell wall synthesis. MraZ can as regulator to control the expression of the first 11 genes of dcw operon and excessive production of MraZ inhibits cell division in E. coli.11–13 We found mraZ was highly conserved in S. aureus, while it is located downstream of the bshC gene and upstream of the rsmH gene, which is different from other reported Gram-negative bacteria. Unsurprisingly, in our study, the MraZ in S. aureus have no effect on the bacterial growth, indicating the species-specific functions of MraZ homologs in bacteria.
We found that the MraZ protein of S. aureus MW2 contains SpoVT-AbrB DNA-binding domains, which existed in some toxin-antitoxin system of bacteria.21,22 Moreover, the downstream gene rsmH was proved to be related to the virulence of S. aureus in the silkworm infection model.23 Therefore, we assumed the mraZ might related to the S. aureus pathogenicity. We first found the hemolysis capability of the mraZ deletion mutant was significantly reduced, which may weaken the ability to induce acute infections such as abscess and pneumoniae.24 We evaluated the killing ability of the wide-type strain and the mutants in the G. mellonella infection model, and we found the mraZ deletion mutant was less virulent. Given that S. aureus often cause skin and soft tissue infections (SSTIs) in human, mouse skin and soft tissue infection model was performed. Our results indicated that MraZ play an essential role in SSTIs and invasion infections. Moreover, the observed impact of mraZ on neutrophil killing revealed that MraZ contributed to immune evasion, which help explain how MraZ contribute to acute infection. Inasmuch as the success of a S. aureus infection depends on the effective evasion of host defenses and neutrophils represent the cornerstone of human innate defenses.25,26 The golden pigment of S. aureus serves as an antioxidant and is protective in killing by neutrophils, while our results indicating that the immune evasion mediated by MraZ was not associated this mechanism.26 Furthermore, our results of the cytokine TNF-α and IL-6 level from mouse bacteremia model were also verified the effect of mraZ on the immune defense. These findings indicated that MraZ might was a virulence factor of S. aureus, and play an important role on the virulence and immune evasion in S. aureus infections.
Our RNA-seq analysis suggested that MarZ was a regulator of expression of extensive genes and many of which were related to S. aureus infection and immune evasions. It is evident that the initiates and maintains of S. aureus infection involves an astounding array of virulence determinants.25 The MSCRAMM surface proteins A (clfA) that mediates attachment and invasion of tissue cells was decreased in ΔmraZ.17 The α-hemolysin is an important virulence factor in S. aureus infections; it can cause pore formation in a series of target cells and epithelial and endothelial breach to help systemic infection.24 We observed that the expression of α-hemolysin were decreased in ΔmraZ. Additionally, PSMs and δ-hemolysin were also markedly decreased in ΔmraZ; these cytolytic toxins can drive attack and eliminate immune cells such as neutrophils and promote immune evasion. Moreover, after S. aureus have been managed to ingest by neutrophils, it has reported have several mechanisms providing resistance to antimicrobial peptides (AMPs) with positively charge.1 Among them, the dlt operon can mediate the introduction of positive charge, increase the net charge of the cell surface, and inhibit neutrophil killing.16,27 Consistently, our data revealed that the deletion of mraZ can significantly reduce the expression of dlt. These findings confirmed that mraZ involved in virulence mechanisms of S. aureus and further explained that the significant effect of mraZ on virulence phenotype.
S. aureus has evolved a complex regulatory network to control virulence. It has well known that agr quorum-sensing system, some TCS systems saeRS, srrAB and arlRS and the assistant regulator sarA.18 Among them, agr can positively regulate α-hemolysin gene (hla), PSM genes (pmsɑ1- pmsɑ4, psmβ1 and psmβ2) and δ-hemolysin gene (hld);28 sarA can enhances the accumulation of many virulence factors via positive regulation of agr activity.18,29 In this study, we found that the deletion of mraZ resulted in decreasing expressions of agr and sarA, indicating that the mraZ can lead to the high expression of many virulence genes by upregulating agr, sarA. This notion is further supported by the reduced production of agr-dependent virulence factors in the mutant with reduced.
However, there are still some limitations in our study. Most of the phenotypic experiments of the complementary strain have not completely recovered to the level of its parent strain. The probable reason may be associated to the genes that only exists on the plasmid and did not restore in their chromosome. Although we confirmed that mraZ plays an important role in the virulence of S. aureus, and regulating some virulence genes and regulators, the regulatory mechanisms of mraZ in relation to virulence requires further investigated.
Conclusion
Our work demonstrates the mraZ plays vital roles in regulating many virulence genes via modulation of agr and sarA expression in S. aureus. MraZ is an attractive target for anti-virulence therapy of S. aureus.
Author Contributions
FY and YG conceived and drafted the work. BW, JD, YJ designed of the work and analyzed and interpreted of data for the work. QZ, YX, HZ, XW and LR participated in the experimental work and data analysis.FY agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All authors read and approved the final manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (81871704).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Otto M. Staphylococcus colonization of the skin and antimicrobial peptides. Expert Rev Dermatol. 2010;5(2):183–195. doi:10.1586/edm.10.6
2. Tong SY, Davis JS, Eichenberger E, Holland TL, Fowler VG. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev. 2015;28(3):603–661. doi:10.1128/CMR.00134-14
3. Tacconelli E, Tumbarello M, Cauda R. Staphylococcus aureus infections. N Engl J Med. 1998;339(27):2026–2027.
4. Centers for Disease Control and Prevention. Active bacterial core surveillance report, emerging infections program network, methicillin-resistant Staphylococcus aureus. 2014.
5. Dinges MM, Orwin PM, Schlievert PM. Exotoxins of Staphylococcus aureus. Clin Microbiol Rev. 2000;13(1):16–34, table of contents. doi:10.1128/CMR.13.1.16
6. Risley AL, Loughman A, Cywes-Bentley C, Foster TJ, Lee JC. Capsular polysaccharide masks clumping factor A-mediated adherence of Staphylococcus aureus to fibrinogen and platelets. J Infect Dis. 2007;196(6):919–927. doi:10.1086/520932
7. Thammavongsa V, Kim HK, Missiakas D, Schneewind O. Staphylococcal manipulation of host immune responses. Nat Rev Microbiol. 2015;13(9):529–543. doi:10.1038/nrmicro3521
8. Otto M. Staphylococcus aureus toxins. Curr Opin Microbiol. 2014;17:32–37. doi:10.1016/j.mib.2013.11.004
9. Dickey SW, Cheung GYC, Otto M. Different drugs for bad bugs: antivirulence strategies in the age of antibiotic resistance. Nat Rev Drug Discov. 2017;16(7):457–471. doi:10.1038/nrd.2017.23
10. Rasko DA, Sperandio V. Anti-virulence strategies to combat bacteria-mediated disease. Nat Rev Drug Discov. 2010;9(2):117–128. doi:10.1038/nrd3013
11. Eraso JM, Markillie LM, Mitchell HD, Taylor RC, Orr G, Margolin W. The highly conserved MraZ protein is a transcriptional regulator in Escherichia coli. J Bacteriol. 2014;196(11):2053–2066. doi:10.1128/JB.01370-13
12. Maeda T, Tanaka Y, Takemoto N, Hamamoto N, Inui M. RNase III mediated cleavage of the coding region of mraZ mRNA is required for efficient cell division in Corynebacterium glutamicum. Mol Microbiol. 2016;99(6):1149–1166. doi:10.1111/mmi.13295
13. Trespidi G, Scoffone VC, Barbieri G, Riccardi G, Rossi ED, Buroni S. Molecular characterization of the Burkholderia cenocepacia dcw Operon and FtsZ Interactors as new targets for novel antimicrobial design. Antibiotics. 2020;9(12):841. doi:10.3390/antibiotics9120841
14. Bae T, Schneewind O. Allelic replacement in Staphylococcus aureus with inducible counter-selection. Plasmid. 2006;55(1):58–63. doi:10.1016/j.plasmid.2005.05.005
15. Montgomery CP, Boyle-Vavra S, Roux A, Ebine K, Sonenshein AL, Daum RS. CodY deletion enhances in vivo virulence of community-associated methicillin-resistant Staphylococcus aureus clone USA300. Infect Immun. 2012;80(7):2382–2389. doi:10.1128/IAI.06172-11
16. Collins LV, Kristian SA, Weidenmaier C, et al. Staphylococcus aureus strains lacking D-alanine modifications of teichoic acids are highly susceptible to human neutrophil killing and are virulence attenuated in mice. J Infect Dis. 2002;186(2):214–219. doi:10.1086/341454
17. Farnsworth CW, Schott EM, Jensen SE, et al. Adaptive upregulation of clumping factor A (ClfA) by Staphylococcus aureus in the obese, type 2 diabetic host mediates increased virulence. Infect Immun. 2017;85(6). doi:10.1128/IAI.01005-16.
18. Jenul C, Horswill AR, Fischetti VA. Regulation of Staphylococcus aureus virulence. Microbiol Spectr. 2019;7(2). doi:10.1128/microbiolspec.GPP3-0031-2018
19. Hua L, Hilliard JJ, Shi Y, et al. Assessment of an anti-alpha-toxin monoclonal antibody for prevention and treatment of Staphylococcus aureus-induced pneumonia. Antimicrob Agents Chemother. 2014;58(2):1108–1117. doi:10.1128/AAC.02190-13
20. Wang Y, Di H, Chen F, et al. Discovery of benzocycloalkane derivatives efficiently blocking bacterial virulence for the treatment of methicillin-resistant S. aureus (MRSA) infections by targeting diapophytoene desaturase (CrtN). J Med Chem. 2016;59(10):4831–4848. doi:10.1021/acs.jmedchem.6b00122
21. Chan WT, Espinosa M, Yeo CC. Keeping the wolves at bay: antitoxins of prokaryotic type II toxin-antitoxin systems. Front Mol Biosci. 2016;3:9. doi:10.3389/fmolb.2016.00009
22. Dienemann C, Bøggild A, Winther KS, Gerdes K, Brodersen DE. Crystal structure of the VapBC toxin–antitoxin complex from Shigella flexneri reveals a hetero-octameric DNA-binding assembly. J Mol Biol. 2011;414(5):713–722. doi:10.1016/j.jmb.2011.10.024
23. Kyuma T, Kimura S, Hanada Y, Suzuki T, Sekimizu K, Kaito C. Ribosomal RNA methyltransferases contribute to Staphylococcus aureus virulence. Febs J. 2015;282(13):2570–2584. doi:10.1111/febs.13302
24. Vandenesch F, Lina G, Henry T. Staphylococcus aureus hemolysins, bi-component leukocidins, and cytolytic peptides: a redundant arsenal of membrane-damaging virulence factors? Front Cell Infect. 2012;2:12.
25. Cheung GYC, Bae JS, Otto M. Pathogenicity and virulence of Staphylococcus aureus. Virulence. 2021;12(1):547–569. doi:10.1080/21505594.2021.1878688
26. Jong NW, Kessel KP, Strijp JA. Immune evasion by Staphylococcus aureus. Microbiol Spectr. 2019;7(2):7–12.
27. Peschel A, Otto M, Jack RW, Kalbacher H, Jung G, Götz F. Inactivation of the dlt Operon inStaphylococcus aureus confers sensitivity to defensins, protegrins, and other antimicrobial peptides. J Biol Chem. 1999;274(13):8405–8410. doi:10.1074/jbc.274.13.8405
28. Cheung GY, Wang R, Khan BA, Sturdevant DE, Otto M, Payne SM. Role of the accessory gene regulator agr in community-associated methicillin-resistant Staphylococcus aureus pathogenesis. Infect Immun. 2011;79(5):1927–1935. doi:10.1128/IAI.00046-11
29. Zielinska AK, Beenken KE, Joo HS, et al. Defining the strain-dependent impact of the Staphylococcal accessory regulator (sarA) on the alpha-toxin phenotype of Staphylococcus aureus. J Bacteriol. 2011;193(12):2948–2958. doi:10.1128/JB.01517-10
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.