Back to Journals » Infection and Drug Resistance » Volume 14
Fluorescence in situ Hybridization (FISH) in the Microbiological Diagnostic of Deep Sternal Wound Infection (DSWI)
Authors Spindler N, Moter A , Wiessner A, Gradistanac T, Borger M, Rodloff AC, Langer S, Kikhney J
Received 8 March 2021
Accepted for publication 4 June 2021
Published 21 June 2021 Volume 2021:14 Pages 2309—2319
DOI https://doi.org/10.2147/IDR.S310139
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Héctor Mora-Montes
Nick Spindler,1 Annette Moter,2,3 Alexandra Wiessner,2,3 Tanja Gradistanac,4 Michael Borger,5 Arne C Rodloff,6 Stefan Langer,1 Judith Kikhney2,3
1Department of Orthopedic Surgery, Traumatology and Plastic Surgery, University Hospital Leipzig, Leipzig, Germany; 2Charité – Universitätsmedizin Berlin, corporate member of Freie Universität Berlin, Humboldt-Universität zu Berlin, and Berlin Institute of Health Institute of Microbiology, Infectious Diseases and Immunology, Biofilmcenter, Berlin, Germany; 3MoKi Analytics GmbH, Berlin, Germany; 4Department of Pathology, University Clinic Leipzig, Leipzig, Germany; 5Department of Cardiac Surgery, Leipzig Heart Center, University of Leipzig, Leipzig, Germany; 6Institute of Microbiology and Epidemiology of Infectious Diseases, University Hospital Leipzig, Leipzig, Germany
Correspondence: Nick Spindler
Department of Orthopedic Surgery, Traumatology and Plastic Surgery, University Hospital Leipzig, Leipzig, Germany
Tel +49-341 - 9717140
Fax +49-341 - 9717139
Email [email protected]
Purpose: Postoperative mediastinitis after cardiac surgery is still a devastating complication. Insufficient microbiological specimens obtained by superficial swabbing may only detect bacteria on the surface, but pathogens that are localized in the deep tissue may be missed. The aim of this study was to analyze deep sternal wound infection (DSWI) samples by conventional microbiological procedures and fluorescence in situ hybridization (FISH) in order to discuss a diagnostic benefit of the culture-independent methods and to map spatial organization of pathogens and microbial biofilms in the wounds.
Methods: Samples from 12 patients were collected and analyzed using classic microbiological culture and FISH in combination with molecular nucleic acid amplification techniques (FISHseq). Frequency of and the time to occurrence of a DSWI was recorded, previous operative interventions, complications, as well as individual risk factors and the microbiologic results were documented.
Results: Tissue samples were taken from 12 patients suffering from DSWI. Classical microbiological culture resulted in the growth of microorganisms in the specimens of five patients (42%), including bacteria and in one case Candida. FISHseq gave additional diagnostic information in five cases (41%) and confirmed culture results in seven cases (59%).
Conclusion: Microbial biofilms are not always present in DSWI wounds, but microorganisms are distributed in a “patchy” pattern in the tissue. Therefore, a deep excision of the wound has to be performed to control the infection. We recommend to analyze at least two wound samples from different locations by culture and in difficult to interpret cases, additional molecular biological analysis by FISHseq.
Keywords: biofilm, FISH, mediastinitis, microorganisms, DSWI
Introduction
Postoperative mediastinitis after cardiac surgery is still a devastating complication. The incidence ranges between 0.5% and 4%, with a mortality rate of up to 50%.1–4 It regularly requires multiple revisions, prolonged use of antibiotics and leads to impaired perfusion of the sternal region and thus to extended hospitalization.5
The treatment consists of a multistage procedure consisting in debridement, conditioning of the sternal wound most commonly by negative pressure wound therapy (NPWT) and reconstruction of the wound using a pedicled or free myocutaneous flap.6–9 However, concomitant antibiotic therapy is necessary for successful treatment. The identification of the causing pathogens is required for a targeted antibiotic therapy, which has a superior outcome to broad range antibiotic regimen.10 However, conventional microbiological culture techniques are not always successful in the identification of the microorganisms present in a sample, in particular when microbial biofilms are involved.11 In addition, the samples to be examined in deep sternal wound infections (DSWI) are often obtained from the surface of the sternal wound using a wound. Hereby, only superficial microorganisms of the wound can be obtained. Pathogens that are localized in the depth of the tissue are missed using this type of sample collection and may in consequence result in persisting wound healing disorders. In addition, microbiological culture results of superficial wound samples are often negative and do not differentiate between contamination or infecting colonization of the wound.12
Therefore, we addressed the question whether an analyzing technique that allows the culture-independent identification of both superficial, on the surface lying microorganisms, and of those located in the deep tissue might provide additional information. Fluorescence in situ hybridization (FISH) in combination with molecular nucleic acid amplification techniques (FISHseq) is a culture-independent method for identification of microbial pathogens within the tissue context.13–15 As a microscopic technique, FISH has the unique potential to visualize, localize, and identify bacteria within the tissue as well as on the sample surface and to stage the formation of the microorganisms (planktonic, microcolonies or biofilms).12,16
The aim of this study was to analyze DSWI samples by both FISHseq and conventional microbiological procedures using scaling techniques in order to discuss a diagnostic benefit of the culture-independent methods and to map the presence of microbial biofilms in the wounds.
Materials and Methods
Study Population
The retrospective single center study protocol was approved by the Institutional Ethics Committee. The study has been registered at the Deutsches Register Klinischer Studien/German register for clinical studies by the number DRKS00015106.
From October 2014 to April 2015, microbiological samples from 12 patients suffering from DSWI were collected and analyzed using classic microbiological culture and FISHseq. All patients met the criteria of the Centers for Disease Control and Prevention (CDC) for DSWI.17 They all had undergone open-heart surgery at the Leipzig Heart Center. Sample material was collected during the revision surgery when debridement of the sternal wound was performed.
Classification of DSWI
Patients suffering from DSWI were classified according to the criteria by Pairolero and Arnold.18 This classification categorizes DSWI based on the duration of signs of infection and clinical findings. Wounds correspond to type I if an infection develops within a few days until two weeks after cardiac intervention, with or without sternal instability in the absence of mediastinal suppuration. Type II infections occur until the eighth week and usually involve cellulitis, purulent wound drainage, and obvious communication with the sternum and mediastinum. Commonly, it is associated with sternal osteomyelitis. Type III represents late infections with an onset longer than two months after the initial cardiac surgery and typically involve a chronically draining sinus tract into the sternum and costochondral.
Surgical Management and Sampling
Twenty-four hours prior to surgical revision, the pre-existing antibiotic therapy was stopped. During surgery, the patient was placed on his/her back, the arms spread to the sides.
Disinfection of the deep wound was omitted; the surrounding skin disinfection was administered as standard technique.
Prior to the debridement, microbiological samples of the deep wound were taken. For the classic microbiological culture analysis, tissue samples were collected from the deep wound edges as well as the mediastinum using a wound swab.
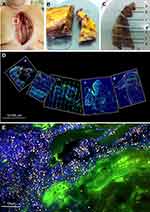
For the FISHseq analysis, samples of the wound edge (1.5 × 1.5 cm) were taken (Figure 1A). The three-dimensional bloc consisted of the three layers skin, subcutaneous fat and pectoralis muscle (Figure 1B). After excision of the sample, the tissue orientation was marked using sutures and the sample placed intra-operatively into the FISH fixation solution (MoKi Analytics, Berlin, Germany). On average, three samples were taken per wound. The entire sampling workflow is summarized in Figure 1.
Microbiological Culture
The specimens submitted to microbiological analysis were deep soft tissue samples from the sternal region sampled at the revision surgery with a wound swab. The samples were placed intra-operatively into a nutrient broth and immediately transported to the microbiological laboratory. Liquid and agar cultures were incubated for at least 48 h at 37°C in aerobic and anaerobic conditions. After 24 h, the first evaluation took place, after 48 h the final evaluation.
FISH and Nucleic Acid Amplification Techniques (FISHseq)
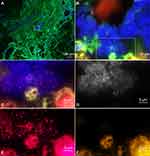
For FISHseq, fixed samples were processed as described previously.12 Briefly, the tissue was embedded in methacrylate, sectioned and submitted to FISH. In a first step, FISH was carried out using the pan-bacterial probe EUB338 combined with a nonsense probe NONEUB338 to exclude unspecific probe binding and 4′,6-diamidino-2-phenylindole (DAPI) for nucleic acid detection. The green channel was left without FISH probe to control tissue background and autofluorescence. In case of positive FISH signals with the EUB338 probe, hybridization with genus- and species-specific FISH probes was carried out in a second step. Epifluorescence microscopy was carried out using a Zeiss AxioImagerZ2 and the ZEN software (Carl Zeiss, Jena, Germany).
Consecutive sections of the same tissue block as for FISH were used for DNA preparation followed by broad-range bacterial polymerase chain reaction (PCR) using 16S rRNA-gene directed primers and sequencing.19 Sequences were analyzed using a commercial 16S rRNA database (SmartGene, Lausanne, Switzerland).
Data Collection
Hospital records and microbiology laboratory records were reviewed retrospectively for every patient. The frequency of and the time to occurrence of a DSWI was recorded, as well as baseline data, previous cardiac operative interventions, complications, individual risk factors, and length of hospital stay. The microbiological results included the identification of pathogens as well as the associated antimicrobial sensitivity tests performed by minimum inhibitory concentration measurement.
Statistical Analysis
All data were evaluated descriptively using the computer-based data analysis program SPSS. Quantitative variables were described using the mean (± SD) for normally distributed data.
Results
Epidemiologic Characteristics
Tissue samples were taken from 12 patients suffering from DSWI, 7 (58%) male and 5 (42%) female. The mean age at the time of revision was 66.8 ± 11.7 years (range: 47–84 years).
In our population, 83% of the patients had arterial hypertonia, 42% had diabetes mellitus and 58% were obese. As shown in Table 1, our population represents a characteristic risk profile for DSWI.
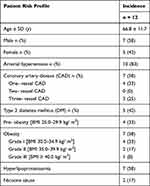 |
Table 1 Patient Risk Profile |
Heart surgery took place electively in nine cases (76%), urgently in one cases (8%), and for emergency reasons in two cases (16%). The preoperative European System for Cardiac Operative Risk Evaluation (EuroSCORE) was 9.8 (range: 0.9–46.6). Bypass surgery alone was performed in combination with a valvular reconstruction or replacement in seven patients, one patient was operated on a congenital heart defect. In one patient, an aortic bow replacement including replacement of the aortic valve was performed, one patient received a minimal invasive and two patients an open valve replacement.
Time between primary cardiac operation and diagnosis of the DSWI varied strongly within the study group (range: 19–245 days). However, no patient showed an early type I infection according to Pairolero and Arnold. Six patients (50%) showed a type II, and six patients (50%) a late infection (Type III). Average time to infection was 74 days after cardiac surgery.
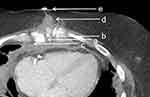
The computed tomography scans of all patients showed a non-union of the sternum. In 8 cases, the cerclages were torn out of the bone; in 4 cases, the osteosynthesis was only loosened (Figure 2). In all cases, a mediastinal abscess with a wide fistula extending to the skin was present.
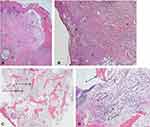
The pathologic evaluation showed an up to high-grade chronic and ulcero-phlegmonous inflammation of the resected soft tissue; In the higher magnification the characteristic superficial ulceration of the overlying keratinized stratified squamous epithelium (dermis) with associated chronic and acute inflammatory cell infiltrate such as dermal fibrosis becomes visible. The resected bone and cartilage presented necrotic fragments surrounded by prominent fibrosis of the medullary canal and a chronic and acute inflammatory cell infiltrate containing lymphocytes and neutrophil granulocytes (Figure 3).
Microbiological Culture and FISHseq Results
Classical microbiological culture resulted in the growth of microorganisms in the specimens of five patients (42%), including bacteria and in one case Candida (Table 2).
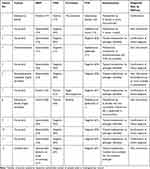 |
Table 2 Microbiological Culture and FISH/PCR Results |
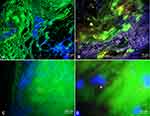
Microscopic analysis using the nucleic acid stain DAPI revealed bacteria in three cases and questionable bacterial structures in nine patients. All three DAPI-positive cases also showed a FISH-signal with 16S rRNA-directed probes indicating a high ribosome content, importantly including one culture-negative case. The formation of the bacteria in these three cases ranged from single microorganisms to biofilms (Figure 4).
In one culture-positive case, FISHseq analysis confirmed the culture results (Enterococcus faecalis), however four other cases, remained negative/questionable (Enterobacteriaceae, Stenotrophomonas maltophilia and Candida). In one case, where culture detected E. faecalis, FISHseq found instead massive amounts of Staphylococcus epidermidis biofilms (Figure 5).
Taken together, FISHseq gave additional diagnostic information in five cases (41%), confirmed culture results in seven cases (59%).
On average, in this study, three samples were taken per wound to be investigated by FISHseq. Out of the three cases with positive microscopy results, microorganisms were seen in 1/7 samples, 1/3 samples and 6/6 samples.
Discussion
The study revealed that FISHseq confirms culture results, and, furthermore, gives additional diagnostic information including therapy-relevant findings that was missed by culture. Therefore, FISHseq diagnostic supports the interpretation of culture and clinical findings and can identify possible revelation of a pathogen change.
Due to the complex situation of DSWI, a multidisciplinary team is needed for optimal patient management consisting of cardiothoracic and plastic surgeons, as well as a clinical microbiologist specialized in infectious diseases. Diagnosis and optimal therapy of DSWI require, beside radical surgical debridement and plastic reconstruction, knowledge of the infectious agent as well as quantity and localization of the infectious foci for a specific antibiotic therapy. However, debridement is done often too superficial and remaining microorganisms in the deep tissue regions of the wound support persistence of the infection.
Though the classic microbiological analysis of wound specimens often gives essential information about the identity and resistance pattern of bacteria present, it lacks information about their spatial formation (planktonic single bacteria, microcolonies or biofilm-forming bacteria), their localization within the tissue of the wound as well as their state of activity. Since the FISH probes bind to rRNA present primarily in metabolically active bacteria, it is highly likely that the FISH-positive bacteria were viable at the time of sampling. DAPI-stained microorganisms not visualized by FISH probably correspond to dead or metabolically inactive bacteria.20,21 In addition, culture is not able to discriminate between infection and contamination with skin flora upon sampling. DSWIs regularly show a wide variety of different bacteria supporting the persistence of a wound healing disorder.
Fluorescence in situ hybridization (FISH) in combination with 16S RNA-directed PCR and sequencing (FISHseq) is a molecular method that has proven useful for the detection and identification of bacteria in clinical samples, in particular in biofilm-associated diseases and culture-negative infections.22,23 As a microscopic technique, FISH visualizes an infection landscape of the tissue sample showing where specific microbial species are found, and in which state they are. This is important because resistant bacteria may persist in the deep tissue of the sternal wound. Especially artificial material, as wire cerclage, is an excellent carrier for different bacteria and hereby may support this chronic infection. However, FISH is clearly not a stand-alone technique and is ideally combined with other diagnostic methods.
Here, we found that FISHseq provided new diagnostic information in five out of 12 cases (41%), including therapy-relevant findings like the identification of Staphylococcus aureus that was missed by culture, revelation of a pathogen change (successful therapy of E. faecalis leaving S. epidermidis to thrive in the wound, Figure 5), and support of the interpretation of culture or clinical findings. In addition, FISHseq identified in one case the typical skin flora member S. epidermidis that often is suspected to be a sampling contamination, as the causative pathogen of the mediastinitis.
FISHseq found only in one case out of 12 patients a biofilm-associated infection. This finding renders the notion of biofilms as ubiquitous threats in DSWI as questionable, although the patient number investigated here is admittedly rather low. Although DSWI are not directly comparable to chronic wounds, in contrast, a meta-analysis by Malone et al described the presence of biofilms in chronic wounds as high as 78.2%.24 However, in particular typical skin flora members should not be underestimated in their potential to indeed form biofilms. The detection of biofilms is important because biofilm infections are responsible for culture-negative results and therapy recalcitrance and may require a prolonged or escalated antibiotic therapy regimen in combination with a more aggressive re-debridement of the affected tissue.11 All six samples investigated in the biofilm-case of this study were positive in the FISH-analysis as opposed to all other cases of this study, where only part of the investigated samples per wound was positive or questionable in the FISH-analysis (Table 2). This result is corroborated by findings from chronic wounds, were microscopic techniques also found a patchy distribution of microorganisms in the wounds.25,26
Limitations of the FISHseq technique are the small sample size (2 µm sections) to be analyzed in comparison to the size of the wound (Figure 1). As microorganisms are not evenly distributed in the sternal wounds, optimal diagnostic results are obtained when several samples are investigated, ideally in combination with culture. For prosthetic joint infections, at least three and optimally five or six intraoperative tissue specimens are recommended for culture before perioperative antibiotics are administered.27 This may apply to both classical microbiological culture as well as molecular biological techniques.28 However, the sampling error is higher for a microscopic analysis with a small sample size to be analyzed as compared to culture where larger tissue samples are investigated. On the other hand, in this study, we found, that microscopic evaluation of several samples per wound was able to pinpoint the sample(s) most promising for PCR and sequencing later on. A disadvantage of culture is the danger of contamination.28 Therefore, in each individual DSWI case, the attending physician has to bargain between the costs for sample analyses and optimal diagnostic outcome. However, patients that would profit from this diagnostics are usually in a highly critical state and FISHseq may provide a decisive diagnosis; the cost of FISHseq would be already justified when the stay in intensive care unit is reduced by one day only.
In our study group, DSWI occurred as a mid- to late-time infection, averaging 74 days after cardiac surgery. The prolonged duration of the incubation period in these patients most likely reflects the reduced immunological state of the patients. Such timing is representative for the fact that cardiac surgery can lead to a systemic inflammatory response syndrome followed by an anti-inflammatory reaction; and aftermath of the surgical intervention that can persist for long periods of time after the actual cardiac surgery.29 These findings correlate with our microscopic observations in the FISHseq analysis of the sternal wounds. FISHseq gives information not only about superficial spatial formation of the bacteria, but also about their penetration of the soft tissue. We found active microorganisms also in the deep layers of the wounds responsible for persisting infection. Therefore, the knowledge about the three dimensional bacterial arrangement has a direct influence on the further way of treatment. Knowing about a resident biofilm or bacteria in the depth of soft tissue would lead to re-debride the affected tissue.
Therefore, we conclude that it is not sufficient in DSWI to debride the wounds only superficially using a sharp spoon, but a complete excision of the wound has to be performed to control the infection.
Conclusion
DSWI is a devastation complication after open chest surgery in a particularly vulnerable patient group. In addition to the local suppression of the immune system by the surgical intervention, often these patients suffer from other severe underlying diseases affecting the immune status. Surgical debridement including deep tissue regions of the wound is of utmost importance to control the infection. Microbial biofilms are not always present in DSWI wounds, but microorganisms are distributed in a “patchy” pattern in the tissue. Therefore, careful decision making regarding diagnostic sampling and analyses techniques is required ideally by a multidisciplinary DSWI team.
The molecular analysis by FISHseq had a major impact on our treatment plan. The regularly taken samples support clinical decision-making in cases of persisting DSWI and/or difficult to interpret-cases. Hereby, we received important additional information on where and to which extend to perform the debridement to obtain clear wound edges.
Abbreviations
CDC, Centers for disease control and prevention; DAPI, 4′,6-diamidino-2-phenylindole; DSWI, Deep sternal wound infection; EuroSCORE, European System for Cardiac Operative Risk Evaluation; FISH, fluorescence in situ hybridization; FISHseq, FISH in combination with molecular nucleic acid amplification techniques; NPWT, negative pressure wound therapy; PCR, polymerase chain reaction; RNA, ribonucleic acid.
Data Sharing Statement
All study-related data are available from the corresponding author upon reasonable request.
Ethics Approval and Consent to Participate
All procedures complied with the ethical standards of the University of Leipzig ethics committee (021/17-ek) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The participants provided written informed consent for participation in this study; copies are available for review by the editor of this journal.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
This report was supported by the German Research Foundation and Leipzig University as part of an Open Access Publishing program.
Disclosure
Dr Judith Kikhney reports personal fees from and hold shares of MoKi Analytics, grants from BMBF, outside the submitted work. Professor Michael Borger reports that his hospital receives speakers’ honoraria and/or consulting fees on his behalf from Edwards Lifescience, Medtronic, Abbott, and CryoLife. Dr Annette Moter reports personal fees from and chairs of MoKi, personal fees from Moter Diagnostics, Fresenius, Chiesi, and Biomerieux, outside the submitted work. All authors declare that they have no other competing interests.
References
1. Lu JC, Grayson AD, Jha P, Srinivasan AK, Fabri BM. Risk factors for sternal wound infection and mid-term survival following coronary artery bypass surgery. Eur J Cardiothorac Surg. 2003;23(6):943–949. doi:10.1016/S1010-7940(03)00137-4
2. Mauermann WJ, Sampathkumar P, Thompson RL. Sternal wound infections. Best Pract Res Clin Anaesthesiol. 2008;22(3):423–436. doi:10.1016/j.bpa.2008.04.003
3. Salehi Omran A, Karimi A, Ahmadi SH, et al. Superficial and deep sternal wound infection after more than 9000 coronary artery bypass graft (CABG): incidence, risk factors and mortality. BMC Infect Dis. 2007;7:112. doi:10.1186/1471-2334-7-112
4. Toumpoulis IK, Anagnostopoulos CE, Derose JJ, Swistel DG. The impact of deep sternal wound infection on long-term survival after coronary artery bypass grafting. Chest. 2005;127(2):464–471. doi:10.1378/chest.127.2.464
5. Loop FD, Lytle BW, Cosgrove DM, et al. J. Maxwell Chamberlain memorial paper. Sternal wound complications after isolated coronary artery bypass grafting: early and late mortality, morbidity, and cost of care. Ann Thorac Surg. 1990;49(2):
6. Jurkiewicz MJ, Bostwick J, Hester TR, Bishop JB, Craver J. Infected median sternotomy wound. Successful treatment by muscle flaps. Ann Surg. 1980;191(6):738–744. doi:10.1097/00000658-198006000-00012
7. Klesius AA, Dzemali O, Simon A, et al. Successful treatment of deep sternal infections following open heart surgery by bilateral pectoralis major flaps. Eur J Cardiothorac Surg. 2004;25(2):218–223. doi:10.1016/j.ejcts.2003.11.019
8. Fleck TM, Koller R, Giovanoli P, et al. Primary or delayed closure for the treatment of poststernotomy wound infections? Ann Plast Surg. 2004;52(3):310–314. doi:10.1097/01.sap.0000105524.75597.e0
9. Pairolero PC, Arnold PG, Harris JB. Long-term results of pectoralis major muscle transposition for infected sternotomy wounds. Ann Surg. 1991;213(6):
10. Hranjec T, Rosenberger LH, Swenson B, et al. Aggressive versus conservative initiation of antimicrobial treatment in critically ill surgical patients with suspected intensive-care-unit-acquired infection: a quasi-experimental, before and after observational cohort study. Lancet Infect Dis. 2012;12(10):774–780. doi:10.1016/S1473-3099(12)70151-2
11. Hall-Stoodley L, Stoodley P, Kathju S, et al. Towards diagnostic guidelines for biofilm-associated infections. FEMS Immunol Med Microbiol. 2012;65(2):127–145. doi:10.1111/j.1574-695X.2012.00968.x
12. Schoenrath F, Kikhney J, Kursawe L, et al. Life on the driveline: molecular detection and fluorescence in situ hybridization-based visualization of microbial species in patients with left ventricular assist devices. J Heart Lung Transplant. 2018;37(1):163–166. doi:10.1016/j.healun.2017.09.023
13. Gescher DM, Kovacevic D, Schmiedel D, et al. Fluorescence in situ hybridisation (FISH) accelerates identification of Gram-positive cocci in positive blood cultures. Int J Antimicrob Agents. 2008;32(Suppl 1):S51–59. doi:10.1016/j.ijantimicag.2008.06.007
14. Trebesius K, Leitritz L, Adler K, Schubert S, Autenrieth IB, Heesemann J. Culture independent and rapid identification of bacterial pathogens in necrotising fasciitis and streptococcal toxic shock syndrome by fluorescence in situ hybridisation. Med Microbiol Immunol. 2000;188(4):169–175. doi:10.1007/s004300000035
15. Wagner M, Horn M, Daims H. Fluorescence in situ hybridisation for the identification and characterisation of prokaryotes. Curr Opin Microbiol. 2003;6(3):302–309. doi:10.1016/S1369-5274(03)00054-7
16. Frickmann H, Zautner AE, Moter A, et al. Fluorescence in situ hybridization (FISH) in the microbiological diagnostic routine laboratory: a review. Crit Rev Microbiol. 2017;43(3):263–293. doi:10.3109/1040841X.2016.1169990
17. Garner JS, Jarvis WR, Emori TG, Horan TC, Hughes JM. CDC definitions for nosocomial infections, 1988. Am J Infect Control. 1988;16(3):128–140. doi:10.1016/0196-6553(88)90053-3
18. Pairolero PC, Arnold PG. Management of infected median sternotomy wounds. Ann Thorac Surg. 1986;42(1):1–2. doi:10.1016/S0003-4975(10)61822-X
19. Mallmann C, Siemoneit S, Schmiedel D, et al. Fluorescence in situ hybridization to improve the diagnosis of endocarditis: a pilot study. Clin Microbiol Infect. 2010;16(6):767–773. doi:10.1111/j.1469-0691.2009.02936.x
20. Santos Ferreira I, Kikhney J, Kursawe L, et al. Encapsulation in polymeric microparticles improves daptomycin activity against mature staphylococci biofilms-a thermal and imaging study. AAPS PharmSciTech. 2018;19(4):1625–1636. doi:10.1208/s12249-018-0974-7
21. Sutrave S, Kikhney J, Schmidt J, et al. Effect of daptomycin and vancomycin on Staphylococcus epidermidis biofilms: an in vitro assessment using fluorescence in situ hybridization. PLoS One. 2019;14(8):e0221786. doi:10.1371/journal.pone.0221786
22. Moter A, Gobel UB. Fluorescence in situ hybridization (FISH) for direct visualization of microorganisms. J Microbiol Methods. 2000;41(2):85–112. doi:10.1016/S0167-7012(00)00152-4
23. Eichinger S, Kikhney J, Moter A, Wie√üner A, Eichinger WB. Fluorescence in situ hybridization for identification and visualization of microorganisms in infected heart valve tissue as addition to standard diagnostic tests improves diagnosis of endocarditis. Interact Cardiovasc Thorac Surg. 2019;29(5):678–684. doi:10.1093/icvts/ivz159
24. Malone M, Bjarnsholt T, McBain AJ, et al. The prevalence of biofilms in chronic wounds: a systematic review and meta-analysis of published data. J Wound Care. 2017;26(1):20–25. doi:10.12968/jowc.2017.26.1.20
25. Fazli M, Bjarnsholt T, Kirketerp-M√∏ller K, et al. Nonrandom distribution of Pseudomonas aeruginosa and Staphylococcus aureus in chronic wounds. J Clin Microbiol. 2009;47(12):4084. doi:10.1128/JCM.01395-09
26. Kirketerp-M√∏ller K, Jensen P, Fazli M, et al. Distribution, organization, and ecology of bacteria in chronic wounds. J Clin Microbiol. 2008;46(8):2717. doi:10.1128/JCM.00501-08
27. Mikkelsen DB, Pedersen C, HØJbjerg T, SchØNheyder HC. Culture of multiple peroperative biopsies and diagnosis of infected knee arthroplasties. APMIS. 2006;114(6):449–452. doi:10.1111/j.1600-0463.2006.apm_428.x
28. Xu Y, Larsen LH, Lorenzen J, et al. Microbiological diagnosis of device-related biofilm infections. APMIS. 2017;125(4):289–303. doi:10.1111/apm.12676
29. Robertson CM, Coopersmith CM. The systemic inflammatory response syndrome. Microbes Infect. 2006;8(5):1382–1389. doi:10.1016/j.micinf.2005.12.016
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.