Back to Journals » International Journal of Nanomedicine » Volume 15
Filter-Membrane-Based Ultrafiltration Coupled with Surface-Enhanced Raman Spectroscopy for Potential Differentiation of Benign and Malignant Thyroid Tumors from Blood Plasma
Authors Liang X , Miao X, Xiao W, Ye Q , Wang S, Lin J, Li C, Huang Z
Received 7 October 2019
Accepted for publication 1 February 2020
Published 1 April 2020 Volume 2020:15 Pages 2303—2314
DOI https://doi.org/10.2147/IJN.S233663
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Thomas Webster
Xiaozhou Liang,1 Xuchao Miao,1 Weijin Xiao,2 Qin Ye,3 Sisi Wang,4 Juqiang Lin,1 Chao Li,2,5,6 Zufang Huang1
1Fujian Normal University, Ministry of Education, Key Laboratory of Optoelectronic Science and Technology for Medicine, Fujian Provincial Key Laboratory for Photonics Technology, Fuzhou, People’s Republic of China; 2Department of Pathology, School of Basic Medical Sciences, Fujian Medical University, Fuzhou, People’s Republic of China; 3Department of Head and Neck Surgery, Fujian Cancer Hospital, Fujian Medical University Cancer Hospital, Fuzhou, People’s Republic of China; 4Department of General Surgery, Fujian Medical University Union Hospital, Fuzhou, People’s Republic of China; 5Department of Pathology, Fujian Cancer Hospital, Fujian Medical University Cancer Hospital, Fuzhou, People’s Republic of China; 6Fujian Provincial Key Laboratory of Translational Cancer Medicine, Fuzhou, People’s Republic of China
Correspondence: Zufang Huang Tel +86 591 834 65373
Email [email protected]
Objective: The objective of this study is to evaluate the performance and feasibility of surface-enhanced Raman spectroscopy coupled with a filter membrane and advanced multivariate data analysis on identifying and differentiating benign and malignant thyroid tumors from blood plasma.
Patients and Methods: We proposed a membrane filter SERS technology for the differentiation between benign thyroid tumor and thyroid cancer. That is to say, by using filter membranes with optimal pore size, the blood plasma samples from thyroid tumor patients were pretreated with the macromolecular proteins being filtered out prior to SERS measurement. The SERS spectra of blood plasma ultrafiltrate obtained using filter membranes from 102 patients with thyroid tumors (70 thyroid cancers and 32 benign thyroid tumors) were then analyzed and compared. Two multivariate statistical analyses, principal component analysis-linear discriminate analysis (PCA-LDA) and Lasso-partial least squares-discriminant analysis (Lasso-PLS-DA), were performed on the SERS spectral data after background subtraction and normalization, as well as the first derivative processing, to analyze and compare the differential diagnosis of benign thyroid tumors and thyroid cancer.
Results: SERS measurements were performed in blood plasma acquired from a total of 102 thyroid tumor patients (benign thyroid tumor N=32; thyroid cancer N=70). By using filter membranes, the macromolecular proteins in blood plasma were effectively filtered out to yield high-quality SERS spectra. 84.3% discrimination accuracy between benign and malignant thyroid tumor was achieved using PCA-LDA method, while Lasso-PLS-DA yields a discrimination accuracy of 90.2%.
Conclusion: Our results demonstrate that SERS spectroscopy, coupled with ultrafiltration and multivariate analysis has the potential of providing a non-invasive, rapid, and objective detection and differentiation of benign and malignant thyroid tumors.
Keywords: surface-enhanced Raman spectroscopy, thyroid tumor, filter membrane, silver nanoparticles, blood plasma
Introduction
The thyroid, which produces thyroid hormone, is one of the most critical and reactive endocrine organs, and playing a vital role in regulating body function and maintaining the proper environment, as well as promoting homeostasis, which controls the production of heat and energy.1,2 In 2018, close to 567,000 cases of thyroid cancer were reported worldwide, ranking in ninth place for incidence,3 and the overall incidence rate of thyroid tumor has increased by 6% per year, a fourth most common cancer in women.4
Traditional diagnostic modalities such as thyroid isotope scanning, Computerized Tomography (CT), Magnetic Resonance Imaging (MRI) scanning, b-mode ultrasound examination, and other approaches provide specific auxiliary diagnostic information.5,6 However, CT and MRI have poor diagnostic sensitivity and specificity for thyroid nodules due to the lack of biological information, and are challenging to detect small lymph nodes in the neck. Fine-needle aspiration biopsy (FNAB) provides pathological information but is invasive and subject to inter-observer discrepancy.7 Moreover, the differentiation of benign and malignant thyroid tumors is difficult to achieve via FNAB given the limited biopsy accuracy and cytological similarities between benign and malignant tumors. Therefore, to provide a real-time monitoring of thyroid cancer from diagnosis to post-treatment, a non-invasive repeatable biopsy that provides molecular information is urgently needed, especially those rapid, more accurate, and objective methods that can complement existing clinical settings to greatly improve diagnostic sensitivity, specificity and differentiation.
Undoubtedly, tumors are initially caused due to biochemical changes in the cells or tissues.8 Along with the tumor development, abnormal expression of oncogenes in tissues or cells will directly fluctuate with variations of disease-related biochemical substances. Moreover, specific antigens and secretions closely related to the disease are more or less presented in the abnormal tissues and body fluids of patients.9 Raman spectroscopy provides molecular fingerprint information of tested samples with the advantages of being performed in a non-destructive and label-free manner, and thus has been widely used for characterization and analysis of tumor tissues and single cells.10 Moreover, the surface-enhanced Raman scattering (SERS) can tremendously overcome the shortcomings of the conventional Raman signal by providing highly sensitive and specific characteristic Raman fingerprints of the analytes being measured, thereby significantly improving detection sensitivity (up to trace level).11 SERS has now been employed for label-free detection of biomarkers in various body fluids such as blood plasma, urine, and saliva.12–14 For tumors of different types, grades and stages, the biochemical composition of the patient’s blood plasma and the relative expression of disease-related biomarkers may be different, which may eventually result in unique SERS spectral signatures. In other words, differences in fingerprint characteristics in the SERS spectrum can provide an objective basis for the potential differential diagnosis of cancer.15 In our previous studies, we have employed the SERS technique to distinguish benign and malignant thyroid tissue samples coupled with principal component analysis-linear discriminate analysis (PCA-LDA).16 The development and application of non-invasive diagnostic tools based on blood plasma genotyping (also known as “liquid biopsy”) is beneficial in reducing the need for tissue biopsy, thus bringing new insights into cancer diagnosis and prognosis. We hypothesize that label-free SERS measurements would provide an unbiased snapshot of the totality of all biomolecular species (rather than focusing on a single entity) in the blood plasma specimen and, when combined with multivariate analysis, would capture latent biological differences that are encoded in the vibrational fingerprints.
Herein, we proposed a filter-based label-free SERS method for optimization of the SERS signal from blood plasma samples of thyroid tumor patients, followed by in-depth multivariate data analysis to accurately and objectively identify and distinguish benign and malignant thyroid tumors. By using filter membrane with proper cut-off weights, the macromolecular proteins in the blood plasma can be effectively filtered to obtain high-quality SERS spectra. Unlike the drying strategies involved in conventional SERS detection, SERS detection of blood plasma was performed in a liquid environment to ensure reliable and objective detection of proteins and analytes under physiological conditions. Furthermore, SERS spectroscopy of blood plasma samples combined with advanced multivariate data analysis, such as PCA-LDA and Lasso-partial least squares-discriminant analysis (Lasso-PLS-DA), was evaluated for the differential diagnosis of benign and malignant thyroid tumors. As a proof-of-concept, our results show that the combination of filter-based SERS spectra and multivariate analysis can provide and highlight potential pathological diagnostic information that reflects biochemical differences and changes at the molecular level. The integration of molecular level detection with routine pathological diagnosis is expected to pave the way for accurate classification and pathological diagnosis of thyroid tumor.
Materials and Methods
Chemicals
All chemicals were purchased from Sigma-Aldrich, unless otherwise stated. All aqueous solutions were prepared with deionized water (resistivity of 18.2 MΩ.cm) purified with a Milli-Q reagent grade water system (Merck Millipore, Billerica, MA, USA).
Patients and Samples Collection
The blood plasma samples from 102 thyroid tumor patients were provided by Fujian Provincial Cancer Hospital. All 32 benign thyroid tumor patients were histologically confirmed as 30 nodular goiter and 2 follicular adenoma, with an average age of 50±12; while all 70 thyroid cancer patients were classified as 67 papillary carcinoma and 3 follicular carcinoma, with an average age of 46±14. All participants provided written informed consent for enrollment in this study, ethics approval was obtained from Fujian Provincial Cancer Hospital Ethics Committee (Approval No. SQ2016-034-01), and the study followed the guidelines of Declaration of Helsinki. After 12 hrs of overnight fasting, 2 mL of blood samples from each patient was first collected into a tube containing an anticoagulant (EDTA) between 7:00 and 8:00 am. Blood plasma was then obtained after centrifugation at 3000 rpm for 10 min and stored at −80°C until further SERS analysis.
AgNPs Synthesis and Characterization
Silver nanoparticles (AgNPs) were prepared using the method reported by Leopold et al’s.17 Briefly, 4.5 mL of 0.1 M NaOH was mixed with 5 mL of 60 mM HONH3Cl, followed by the addition of 90 mL of 1.1 mM AgNO3 rapidly. The mixture was vigorously stirred until the solution turned milky gray. The freshly prepared AgNPs was then centrifuged at a speed of 10,000 rpm for 10 min. TEM image of AgNPs was characterized by a transmission electron microscope (Hitachi, H-7650) at an accelerating voltage of 80kV, and the absorption spectra were collected with a UV–Vis spectrometer (PerkinElmer, Lambda 950) at room temperature.
Preparation of Blood Plasma Ultrafiltrate and SERS Measurement
In preparation of blood plasma ultrafiltrate, Millipore Amicon Ultra 0.5 mL centrifugal filter devices (Merck, Germany), of cut-off weights of 10 kDa, 30 kDa, 50 kDa, and 100 kDa were employed to reduce the interference of large amounts of high-abundance proteins,18 thus retaining and concentrating low-molecular-weight proteins with high diagnostic value from each blood plasma sample.19 20 µL of blood plasma ultrafiltrate mixed with identical volume of AgNPs were prepared and an aliquot of 5 µL of each sample was placed on a clean aluminum block. SERS spectra were acquired using a confocal Raman system (Renishaw) with an excitation laser of 785 nm. A 20× long working distance (NA = 0.25) objective was used to focus the laser and to collect the scattered light from sample surface. Prior to large-scale experiments, detection parameters such as integration time (30 s) as well as laser power (2 mW) were optimized to yield high performance. Raman spectra were collected over a range of 400 to 1800 cm−1 using the WiRE 3.4 software, which covers the fingerprint region of the blood plasma samples. Each sample was represented with the average spectrum calculated from triplicate, random measurements of the sample droplet.
SERS Spectral Pre-Processing and Multivariate Data Analysis
The Raman instrument was wavenumber calibrated using the wafer 520 cm−1 band. Raman spectra recorded were intensity normalized to compensate for spectral response due to effects caused by laser power variations at the sample, and restricted to the fingerprint wavenumber region (400–1800 cm−1) for analysis.
PCA was applied to analyze the variance of spectral data, which reduces the dimensionality of the spectral dataset to a few dimensions characteristic of the maximum variance in the dataset.20 This transformation converts a set of spectral recordings into a set of values of linearly uncorrelated variables that form an orthogonal basis set. Similar to our previous multivariate statistical analysis,21 PC scores with lowest p-values (PC1, PC3, and PC17) based on the independent T-test were firstly selected. Linear discriminant analysis (LDA) was then used to achieve the diagnostic line (decision boundary, which separated the data-points into regions that were actually the classes they belong to); where the discriminant function coefficients were calculated using the three PCs from PCA with significant differences and the known classification as inputs. Finally, the leave-one-out and cross-validation method were then performed to avoid overoptimistic results and assess their validity. After the PCA-LDA analysis, posterior probabilities are computed using SPSS software, which represent the probabilities of being classified as benign thyroid tumor and thyroid cancer patients based on the corresponding blood plasma SERS spectra.
Least absolute shrinkage and selection operator (Lasso) is essentially a compression estimate, basically based on the form we are familiar with representing multiple linear regression. In general, the advantage of Lasso is that it drives the parameters to zero, eliminating the characteristics of regression.22 Therefore, Lasso automatically selects more relevant features and discards other features during the iteration. This advantage also leads to a reasonable noise suppression effect of the Lasso. A sparse model with a small non-zero coefficient can be obtained using the Lasso model. After obtaining useful spectral variables with Lasso, we noticed that some spectral regions have more useful variables than others. To avoid noise interference with some variables’ uncertainty on the prediction results, we chose an integrated SERS spectral range containing one or more useful spectral variables to establish a further model. PLS-DA is then employed to classify the benign and malignant thyroid tumor based on the spectral regions selected above.
PCA-LDA and Lasso-PLS-DA23 were both used to analyze and compare the differential diagnosis of benign thyroid tumors and thyroid cancer, and to avoid overoptimistic results, they were then both validated using leave-one-out cross-validation. The receiver operating characteristic (ROC) curve was calculated from the posterior probabilities derived from the above analysis and represents the diagnostic performance of the model and the corresponding area under the curve (AUC) was calculated (see Figure S1 for the flow chart of multivariable model steps).
The SPSS 19.0 software package (SPSS Inc., Chicago, Illinois) was used for independent T-test, PCA, LDA, and ROC analysis. Homemade program in Matlab (Matlab R2013b, Mathworks, USA) was employed to conduct the Lasso-PLS-DA analysis. Origin 2017 software (OriginLab, USA) was employed to create the scatter plots as well as the Raman spectra figures.
Results and Discussion
Optimization of SERS Signals
Representative UV–Vis spectrum of AgNPs solution was shown in Figure S2, where the maximum peak locating at 416 nm indicates the characteristic localized surface plasmon resonance (LSPR) peak of AgNPs. The inset shows the representative TEM image of AgNPs, with an average particle size of 35±4 nm. Rather than using the original concentration, for the following experiment, the AgNPs colloid was centrifuged and adjusted to 20 times the initial concentration for better performance.24 785 nm laser excitation was purposely chosen for Raman spectroscopy, since it offers an excellent balance between Raman efficiency, sample fluorescence, and heat absorption.25 Although the maximum LSPR peak is far from our excitation wavelength, components in the blood plasma (such as proteins, analytes, and salts) can contribute to the analyte–colloid and colloid–colloid interactions, which finally cause nanoparticles to aggregate, resulting in a red-shift of the LSPR.26 This phenomenon can be easily confirmed by plotting the absorption spectra with different volume ratios of blood plasma and concentrated AgNPs (see Figure S3). To investigate the effect of volume ratios between concentrated AgNPs and blood plasma to SERS effects, various volume ratios between them were prepared to achieve the optimum SERS enhancement and the most reliable SERS spectral results. It was discovered that when the concentrated AgNPs was mixed with the blood plasma sample at a volume ratio of 1:1, the SERS spectrum relatively shows the best signal strength enhancement over the entire wavenumber range and the greatest signal-to-noise ratio (SNR) (see Figure S4). The reasonable explanation is that low volume ratio of blood plasma to AgNPs will not generate sufficient SERS enhancement; however, excessive AgNPs in the mixture tend to form aggregates and cause precipitation, thus result in lower SERS enhancement, which finally leads to unstable and irreproducible SERS performance. In addition, we found that the most intense SERS spectrum occurs when the LSPR is blue-shifted from the 785 nm excitation, which agrees well with the recent studies by Turzhitsky et al27,28 that the maximum SERS signal is recorded when the excitation wavelength is red-shifted to the LSPR, due to the competitive relationship between SERS enhancement and extinction in nanoparticle colloids and the resulting balance of incident light and scattered Raman signals.
Blood plasma contains a large amount of substances that can be used for disease diagnosis. Although Raman spectroscopy was deemed to provide detailed molecularly specific fingerprints of compounds in blood plasma, fluorophores with molecular weights greater than 10 kDa27 are considered to be the main source of interference when using Raman to detect potential diagnostic biomarkers such as low-abundance proteins and small molecules. By applying ultrafiltration via filter-membranes with proper cut-off weights, it is expected to effectively filter out macromolecules that cause fluorescent and Raman backgrounds rather than useful diagnostic signals,29 which is also confirmed by our results (see Figure S5). To further comprehensively evaluate the SERS detection of blood plasma ultrafiltrate, we compared the SERS enhancement of blood plasma ultrafiltrate after treatment with various membrane pore sizes. Our experimental results showed that the most intense SERS spectral signals and the corresponding high signal-to-noise ratio SERS spectrum were obtained (see Figure S6) by mixing the concentrated AgNPs with the blood plasma filtrated using the 50 kDa membrane filter.
SERS Spectral Features of the Blood Plasma Samples
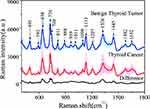
Figure 1 shows the average SERS spectra of blood plasma obtained from benign thyroid tumor samples (blue line with green-shaded area) as well as thyroid cancerous samples (red line with pink-shaded area), and their difference spectrum (bottom curve). From the average SERS spectra, a series of prominent peaks can be observed, ie, 495 cm−1 (C-OH3 torsion of the methoxy group), 638 cm−1 (C-S stretch and C-C twisted protein tyrosine), 726 cm−1 (adenine, C-S (protein)), 740 cm−1 (C-S stretch (one of the three thiocyanate peaks)), 1133 cm−1(palmitic acid, fatty acid), 1207 cm−1 (mainly tyrosine), 1326 cm−1 (nucleic acids and phosphates), 1445 cm−1 (methyl (CH3) and sub-methyl (CH2)), indicating a similar Raman profile which shares the major Raman peaks between these two groups, while the major spectral differences occur at peak intensities due to the difference in relative concentration of relevant biochemical components. To facilitate better visualization and highlight the spectral differences between these two groups, difference spectrum was also calculated, which features that the main difference lies in the relative intensity of the bands at 495 cm−1, 592 cm−1, 638 cm−1, 726 cm−1, 740 cm−1, 1326 cm−1, and 1445 cm−1. For example, at 592 cm−1 and 1326 cm−1, the C-H bond vibration in nucleic acids and phospholipids was observed, especially the peak intensity of 1326 cm−1 in thyroid tumor was significantly higher than that in the cancer group. The peak intensity of 1445 cm−1 in thyroid tumor was clearly higher than those from thyroid cancer, which was mainly attributed to the different bending degrees between methyl (CH3) and sub-methyl (CH2).30 Our blood plasma SERS assay is performed in solution rather than air-dried conditions, thereby effectively avoiding inconsistencies in Raman signals caused by spatial heterogeneity. In addition, as the brownian motion of the molecules during the measurement process essentially aids in achieving average of the SERS signal throughout the sample, thus ensuring the stabilization of the natural functional structure of biomolecules and protein molecules in blood plasma as well as the accomplishment of reproducible and reliable SERS detection.31 The tentative band assignments based on the SERS spectra of reference compounds with biological origin32 are listed in Table 1.
 |
Table 1 Tentative Major Raman Peak Assignments in the Obtained SERS Spectra of Blood Plasma |
Multivariate Data Analysis for Differential Diagnosis
The complexity of biomolecular components in blood plasma can lead to a significant overlap of vibrational modes observed in Raman data. As mentioned above, SERS spectra from benign thyroid tumor and thyroid cancer shared a high similarity in spectral profiles. In order to effectively reveal the spectral differences and evaluate the ability of SERS to distinguish between benign and malignant thyroid tumor samples, two multivariate analysis methods, PCA-LDA and Lasso-PLS-DA, were adopted in the experiment. After dimensionality reduction by PCA, the PC scores and loadings were obtained.33 However, not all PCs are meaningful. Some are due to noise, and some may have insignificant meanings. Subsequently, an independent T-test was performed to screen for the most diagnostically significant PCs (PC1, PC3 and PC17, accounting for a variance of 47.5%, 8.5%, and 0.5%, respectively), where p-values for PC3 and PC17 are smaller than 0.05, except that p-value for PC1 is slightly larger than 0.05, thereby enhancing the reliability of SERS analysis.
Figure 2 shows the scatter plot of PCA in the benign thyroid tumor group and the thyroid cancer group, where the dotted lines indicate the best separation of the two groups. It can be seen that different combinations of PC components with significant p-value differences can yield different discriminant classification effects.34 According to the calculation results, the corresponding sensitivities were 84.3%, 64.3%, and 61.4%, respectively; the specificities were 43.8%, 43.8%, and 53.1%, respectively (as shown in Table 2). The results indicate that different PC combinations provide different differential accuracies between benign thyroid tumor and thyroid cancer samples. PCA here does not yield the best classification results for our SERS data, although it offers a great way to summarize high dimension datasets into subsets of variables.
 |
Figure 3 Scatter plots of posterior probability of blood plasma SERS spectra belonging to benign thyroid tumor and thyroid cancer, based on (A) PC1 and PC3, (B) PC1 and PC17, and (C) PC3 and PC17. |
 |
Table 2 Differentiation of Benign Thyroid Tumor and Thyroid Cancer Group Based on Different PC Combinations |
To improve the diagnostic results of blood plasma SERS spectra, three PC components (PC1, PC3, and PC17) with the lowest P-values were introduced into the LDA model in three different combinations for differential classification (as shown in Figure 3). LDA arranges discriminant functions by maximizing the variances in the SERS data between different sample groups while minimizing the variances between the samples of the same group.35 After the loading of diagnostically significant PCs into the LDA model, leave-one-out cross-validation on the limited Raman data sheet was implemented to avoid biased diagnostics. The threshold on the posterior probability for the decision boundary is 0.5.36 The diagnostic classification results (sensitivity, specificity, and diagnostic accuracy) based on PCA-LDA analysis of blood plasma SERS spectra for separation of thyroid cancer and benign thyroid tumor subjects were: 90.0% (63/70), 43.8% (14/32) and 75.5% (77/102) for the PC1 vs PC3 group; 65.7% (46/70), 59.4% (19/32) and 63.7% (65/102) for PC1 vs PC17; 71.4% (50/70), 56.3% (18/32) and 66.7% (68/102) for PC3 vs PC17 (as shown in Table 3).
 |
Table 3 The Diagnostic Classification Results of PCA-LDA in Blood Plasma SERS Spectra of Thyroid Cancer and Thyroid Tumors |
To assess the prediction accuracy of the PCA-LDA-based diagnostic algorithm, we plotted the receiver operating characteristic curve (ROC) (as shown in Figure 4), in which the ROC was used to assess the performance of the discrimination of PCA-LDA for benign thyroid tumor and thyroid cancer. The ROC curve represents how the true positive rate (Sensitivity) changes with the false-positive rate (1-Specificity) for different cut-offs. The area under the curve (AUC) value indicates the ability of the algorithm to distinguish these two categories. For the discrimination of benign thyroid tumor and thyroid cancer based on three different PC combinations, the calculated AUC results are: 0.675, 0.613, and 0.627, respectively, indicating that PCA-LDA based diagnostic algorithms yield in average an acceptable discrimination result for benign thyroid tumor and thyroid cancer patients.37
Although the signal-to-noise ratio of the spectrum signal may be reduced in some cases, the use of derivative spectroscopy can reduce or eliminate the spectral overlap caused by the relatively wideband components; then highlight the subtle differences in the two spectra, and ultimately improve the diagnosis sensitivity. So far, the derivative spectroscopy has been commonly used in the interpretation and exploitation of data obtained from near-infrared and ultraviolet spectroscopy.38 However, there are few studies on the optimization and improvement of the application of derived analytical methods to Raman data.39
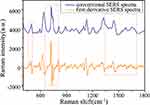
Figure 5 compares the original SERS spectra and the corresponding first derivative spectra of benign thyroid tumors in the range of 400–1800 cm−1. The red-dotted frame shows three representative peak regions such as 450–500 cm−1, 650–800 cm−1, 1350–1700 cm−1. It is evident that the first-derivative spectrum not only remarkably highlights the shoulder and overlapping Raman bands hidden in the original SERS spectrum, but also shows richer Raman peaks and significantly increase the resolution of the SERS spectrum, which might potentially provide high-quality data for further discrimination analysis input.40
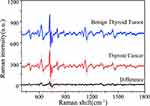
The mean first-order derivative and difference SERS spectrum for benign thyroid tumor (32, blue curve) and thyroid cancer (70, red curve) are compared in Figure 6, while the black curve indicates the corresponding difference spectrum. Compared with the conventional filter SERS spectrum, the first-order derivative filter SERS spectrum has a significant number of negative and positive peaks. For instance, in the range of 750–800 cm−1 (within the red-dotted frame), the two positive peaks (726 cm−1, 740 cm−1) in the difference spectrum of the conventional blood plasma ultrafilter SERS spectrum (see Figure 1) is now turned into two positive peaks (721 cm−1, 738 cm−1) and two negative peaks (729 cm−1, 748 cm−1) after the first-order derivative transformation.
Since not all of the useful variables were distributed in each SERS bands, and some bands had more useful variables than others, therefore, to achieve better classification, removal of irrelevant and correlated data are required. Meanwhile, weak peaks may make large contributions to discrimination. To address these limitations, three spectral regions (981–972, 1520–1511, and 1572–1562 cm−1) containing the most critical spectral variables from the SERS spectrum were selected for further analysis using Lasso-PLS-DA. In addition, the minimum angle regression is used as the Lasso model parameter, because it is more intuitive to relate to useful features than other parameters.
The Lasso algorithm coupled with leave-one-out cross-validation (LOOCV) was used to extract the SERS spectral features closely associated with different cancer pathologies.41 In LOOCV, one sample (ie, one spectrum) was taken out from all these 102 samples, and the rest of blood spectra were then used for reconstruction by the Lasso algorithm for classification of the selected spectrum. This process is repeated until all blood plasma samples that are retained are classified.
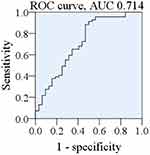
To assess the prediction accuracy of the Lasso-PLS-DA based diagnostic algorithm, the ROC was plotted (Figure 7), with the AUC of being 0.714.
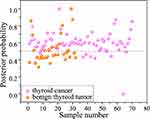
Compared with the PCA-LDA results, the sensitivity was significantly increased by 90.0% (63/70), and the specificity was 59.4% (19/32). The diagnostic accuracy of posterior probability scatter plot for separating the benign thyroid tumor and thyroid cancer is 80.4% (Figure 8). In this regard, the Lasso-PLS-DA algorithm provides better results for differentiating benign thyroid tumor and thyroid cancer by establishing a classification model based on significant Raman characteristics.
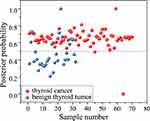
Based on the capability of the derivative spectroscopy to extract fine spectral features, we distinguished the SERS spectra of benign and malignant thyroid tumors by converting their mathematical spectral curves into first-order derivatives, expecting to enhance and highlight the sensitivity of the fine spectral features. Figure 9 shows the posterior probabilities of blood plasma belonging to benign thyroid tumor and thyroid cancer, together with Lasso-PLS-DA algorithms with LOOCV. Specifically, SERS technique together with Lasso-PLS-DA and LOOCV modeling provides a diagnostic accuracy of 90.2% (sensitivity of 97.1%; specificity of 75.0%) for differentiation between benign thyroid tumor and thyroid cancer, which is superior to the results calculated by the other methods described above.
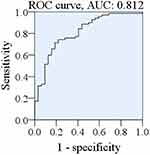
Similarly, we compared the first-order derivative of the normalized SERS spectral data based on Lasso-PLS-DA analysis. The receiver operating characteristic curve (ROC) is plotted (Figure 10), showing an AUC of 0.812.
Conclusions
In summary, we demonstrated the successful combination of filter membrane with SERS spectroscopy to obtain the SERS detection and effective optimization of blood plasma SERS signals from patients with benign thyroid tumor and thyroid cancer under physiological environment. Moreover, coupled with advanced multivariate analysis such as Lasso-PLS-DA and first-derivative analysis on the SERS data, a better differential diagnosis between benign thyroid tumor and thyroid cancer was obtained, with a diagnostic accuracy of 90.2%. This study demonstrates the potential of employing label-free SERS method as a practical and reliable technique to complement the existing clinical methods for the differentiation of benign and malignant thyroid tumors.
Acknowledgment
The project was supported by the National Natural Science Foundation of China (Nos. 11874006, 61775037), the Natural Science Foundation of Fujian Province, China (No. 2018J01786, 2019J01513), Fujian Provincial Health Technology Project (No. 2017-CX-10), and Program for Changjiang Scholars and Innovative Research Team in University (IRT1115).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Dittrich R, Beckmann MW, Oppelt PG, et al. Thyroid hormone receptors and reproduction. J Reprod Immunol. 2011;90(1):58–66. doi:10.1016/j.jri.2011.02.009
2. Tasneem A. The incidence of hyperprolactinaemia and associated hypothyroidism- local experience from Lahore Centre for Nuclear Medicine. Mayo Hospital Lahore, Government College for Boys Gulberg Lahore. PJNM. 2011;1:49–55.
3. Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492.
4. Davies L, Welch HG. Current thyroid cancer trends in the United States. JAMA Otolaryngol Head Neck Surg. 2014;140(4):317–322. doi:10.1001/jamaoto.2014.1
5. LG T. Diagnostic ultrasound in clinical thyroid investigation. J Clin Endocrinol Metab. 1971;36:709716.
6. Gharib H, Papini E. Thyroid nodules: clinical importance, assessment, and treatment. Endocrinol Metab Clin North Am. 2007;36(3):707–735. doi:10.1016/j.ecl.2007.04.009
7. Castro MR, Hossein Gharib MD. Thyroid fine-needle aspiration biopsy-progress, practice, and Pitfalls. AACE. 2003;9(2):128–136.
8. Kolanjiappan K, Ramachandran CR, Manoharan S. Biochemical changes in tumor tissues of oral cancer patients. Clin Biochem. 2003;36(1):61–65. doi:10.1016/S0009-9120(02)00421-6
9. Sarandakou A, Protonotariou E, Rizos D. Tumor markers in biological fluids associated with pregnancy. Crit Rev Clin Lab Sci. 2007;44(2):151–178. doi:10.1080/10408360601003143
10. Wachsmann-Hogiu S, Weeks T, Huser T. Chemical analysis in vivo and in vitro by Raman spectroscopy–from single cells to humans. Curr Opin Biotechnol. 2009;20(1):63–73. doi:10.1016/j.copbio.2009.02.006
11. Botti S, Almaviva S, Cantarini L, et al. Trace level detection and identification of nitro-based explosives by surface-enhanced Raman spectroscopy. J Raman Spectrosc. 2013;44(3):463–468. doi:10.1002/jrs.4203
12. Hunter R, Sohi AN, Khatoon Z, et al. Optofluidic label-free SERS platform for rapid bacteria detection in serum. Sens Actuat B Chem. 2019;300:126907. doi:10.1016/j.snb.2019.126907
13. Joseph MM, Narayanan N, Nair JB. Exploring the margins of SERS in practical domain- An emerging diagnostic modality for modern biomedical applications. Biomaterials. 2018;181:140–181. doi:10.1016/j.biomaterials.2018.07.045
14. Zheng X-S, Jahn IJ, Weber K, et al. Label-free SERS in biological and biomedical applications: recent progress, current challenges and opportunities. Spectrochim Acta Part A Mol Biomol Spectrosc. 2018;197:56–77. doi:10.1016/j.saa.2018.01.063
15. McAughtrie S, Faulds K, Graham D. Surface enhanced Raman spectroscopy (SERS): potential applications for disease detection and treatment. J Photochem Photobiol C Photochem Rev. 2014;21:40–53. doi:10.1016/j.jphotochemrev.2014.09.002
16. Zuanfang L, Chao L, Li C, Lin D, et al. Surface-enhanced Raman spectroscopy for differentiation between benign and malignant thyroid tissues. Laser Phys Lett. 2014;11:045602.
17. Leopold N, Lendl B. A new method for fast preparation of highly surface-enhanced Raman scattering (SERS) active silver colloids at room temperature by reduction of silver nitrate with hydroxylamine hydrochloride. J Phys Chem b. 2003;107(24):5723–5727. doi:10.1021/jp027460u
18. Wisniewski JR, Zougman A, Nagaraj N, et al. Universal sample preparation method for proteome analysis. Nat Methods. 2009;6(5):359–362. doi:10.1038/nmeth.1322
19. Bonnier F, Baker MJ, Byrne HJ. Vibrational spectroscopic analysis of body fluids: avoiding molecular contamination using centrifugal filtration. Anal Methods. 2014;6(14):5155. doi:10.1039/c4ay00891j
20. Svante Wold KE, Geladi P. Principal component analysis. Chemom Intell Lab Syst. 1987;2(13):37–52. doi:10.1016/0169-7439(87)80084-9
21. Feng S, Chen R, Lin J, et al. Nasopharyngeal cancer detection based on blood plasma surface-enhanced Raman spectroscopy and multivariate analysis. Biosens Bioelectron. 2010;25(11):2414–2419. doi:10.1016/j.bios.2010.03.033
22. Kukreja SL, Löfberg J, Brenner MJ. A least absolute shrinkage and selection operator (LASSO) for nonlinear system identification. Elsevier. 2006;39(1):814–819.
23. Chen G, Lin X, Lin D, et al. Identification of different tumor states in nasopharyngeal cancer using surface-enhanced Raman spectroscopy combined with Lasso-PLS-DA algorithm. RSC Adv. 2016;6(10):7760–7764. doi:10.1039/C5RA24438B
24. Shi Y, Liu W, Chen C. Two-step centrifugation method for subpicomolar surface-enhanced raman scattering detection. Anal Chem. 2016;88(9):5009–5015. doi:10.1021/acs.analchem.6b01194
25. Synytsya A, Judexova M, Hoskovec D, et al. Raman spectroscopy at different excitation wavelengths (1064, 785 and 532 nm) as a tool for diagnosis of colon cancer. J Raman Spectrosc. 2014;45(10):903–911. doi:10.1002/jrs.4581
26. Wang X, Li Y, Wang H, et al. Gold nanorod-based localized surface plasmon resonance biosensor for sensitive detection of hepatitis B virus in buffer, blood serum and blood plasma. Biosens Bioelectron. 2010;26(2):404–410. doi:10.1016/j.bios.2010.07.121
27. Rohleder D, Kiefer W, Petrich W. Quantitative analysis of serum and serum ultrafiltrate by means of Raman spectroscopy. Analyst. 2004;129(10):906–911. doi:10.1039/b408927h
28. Li M, Kang JW, Dasari RR, et al. Shedding light on the extinction-enhancement duality in gold nanostar-enhanced Raman spectroscopy. Angew Chem Int Ed Engl. 2014;53(51):14115–14119. doi:10.1002/anie.v53.51
29. Harper RG, Workman SR, Schuetzner S, et al. Low-molecular-weight human serum proteome using ultrafiltration, isoelectric focusing, and mass spectrometry. Electrophoresis. 2004;25(9):1299–1306. doi:10.1002/(ISSN)1522-2683
30. Harris AT, Garg M, Yang XB, et al. Raman spectroscopy and advanced mathematical modelling in the discrimination of human thyroid cell lines. Head Neck Oncol. 2009;1(1):38. doi:10.1186/1758-3284-1-38
31. Spedding FH, Stamm RF. The raman spectra of the sugars in the solid state and in solution I. The raman spectra of α‐ and β‐d‐glucose. J Chem Phys. 1942;10(3):176–183. doi:10.1063/1.1723681
32. Mosier-Boss PA, Lieberman SH. Detection of volatile organic compounds using surface enhanced Raman spectroscopy substrates mounted on a thermoelectric cooler. Anal Chim Acta. 2003;488(1):15–23. doi:10.1016/S0003-2670(03)00676-7
33. Tenenbaum JB, de Silva V, Langford JC. A global geometric framework for nonlinear dimensionality reduction. Science. 2000;290(5500):2319–2323. doi:10.1126/science.290.5500.2319
34. Li MX, Yeung JMY, Cherny SS, et al. Evaluating the effective numbers of independent tests and significant p-value thresholds in commercial genotyping arrays and public imputation reference datasets. Hum Genet. 2012;131(5):747–756. doi:10.1007/s00439-011-1118-2
35. Guo Y, Hastie T, Tibshirani R. Regularized linear discriminant analysis and its application in microarrays. Biostatistics. 2007;8(1):86–100. doi:10.1093/biostatistics/kxj035
36. Westerhuis JA, Hoefsloot HCJ, Smit S, et al. Assessment of PLSDA cross validation. Metabolomics. 2008;4(1):81–89. doi:10.1007/s11306-007-0099-6
37. Pencina MJ, D’ Agostino RB, D’ Agostino RB, et al. Evaluating the added predictive ability of a new marker: from area under the ROC curve to reclassification and beyond. Stat Med. 2008;27(2):157–172. discussion 20712. doi:10.1002/(ISSN)1097-0258.
38. Larouche N, Stansfield BL. Classifying nanostructured carbons using graphitic indices derived from Raman spectra. Carbon. 2010;48(3):620–629. doi:10.1016/j.carbon.2009.10.002
39. Notingher I, Jell G, Notingher PL, et al. Multivariate analysis of Raman spectra for in vitro non-invasive studies of living cells. J Mol Struct. 2005;744747:179–185. doi:10.1016/j.molstruc.2004.12.046
40. Tan SC, Khor E. The degree of deacetylation of chitosan- advocating the first derivative UV-spectrophotometry method of determination. Talanta. 1998;45(4):713–719. doi:10.1016/S0039-9140(97)00288-9
41. Cawley GC, Talbot NL. Fast exact leave-one-out cross-validation of sparse least-squares support vector machines. Neural Networks. 2004;17(10):1. doi:10.1016/j.neunet.2004.07.002
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.