Back to Journals » OncoTargets and Therapy » Volume 13
FAM110B Inhibits Non-Small Cell Lung Cancer Cell Proliferation and Invasion Through Inactivating Wnt/β-Catenin Signaling
Authors Xie M, Cai L, Li J, Zhao J, Guo Y, Hou Z, Zhang X , Tian H, Li A, Miao Y
Received 28 January 2020
Accepted for publication 4 May 2020
Published 19 May 2020 Volume 2020:13 Pages 4373—4384
DOI https://doi.org/10.2147/OTT.S247491
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Federico Perche
Menghua Xie,1 Lin Cai,1 Jingduo Li,1 Jing Zhao,1 Yingxue Guo,1 Zaiyu Hou,1 Xiupeng Zhang,1 Hua Tian,2 Ailin Li,2 Yuan Miao1
1Department of Pathology, The College of Basic Medical Sciences and The First Affiliated Hospital of China Medical University, Shenyang, People’s Republic of China; 2Department of Radiotherapy, The First Affiliated Hospital of China Medical University, Shenyang, People’s Republic of China
Correspondence: Yuan Miao Tel +86 248 328 2248
Email [email protected]
Purpose: FAM110B is a member of the FAM110 family (family with sequence similarity 110), which is a component of the centrosome associated proteins. Previous studies have shown that FAM110B may be involved in the process of cell cycle and may play a role in carcinogenesis and tumor progression. Using an online database, we found that FAM110B may predict favorable prognosis in non-small cell lung cancer (NSCLC). Therefore, the role of FAM110B playing in NSCLC needs to be further investigated.
Patients and Methods: Online databases and immunohistochemistry were used to predict the expression and prognostic value of FAM110B in NSCLC samples. Immunofluorescence staining was used to investigate the subcellular distribution of FAM110B. Western blot, MTT, colony formation, and matrigel invasion assay were used to detect the expression and the effect of FAM110B on mediating proliferation and invasion in NSCLC cell lines.
Results: In this study, immunohistochemistry results showed that FAM110B expression significantly correlated with early TNM staging (P=0.020) and negative regional lymph node metastasis (P=0.006). Kaplan–Meier survival analysis found that the median survival time of patients with positive FAM110B expression (56.181± 2.348 months) was significantly longer than those with negative FAM110B expression (47.701± 2.997 months, P=0.024). Moreover, overexpression of FAM110B inhibited the proliferation and invasion of A549, H1299, and LK2 cell lines. Conversely, FAM110B RNAi exerted opposite effects in the above cell lines. Furthermore, FAM110B overexpression downregulated the active β‐catenin, phosphorylation of GSK-3β (p-GSK-3β), cyclin B1, cyclin D1, MMP2, and MMP7, and upregulated the phosphorylation of β-catenin (p-β-catenin) in A549 and H1299 cells. Besides, the FAM110B-induced depressions of p-GSK-3β and active β-catenin were reversed after being treated with Wnt/β-catenin inhibitor, XAV-939.
Conclusion: In summary, our results demonstrated that the overexpression of FAM110B restricts the proliferation and invasion of NSCLC cells by inhibiting Wnt/β-catenin signaling. Our study reveals the antitumor function of FAM110B in NSCLC and indicates that FAM110B is a potential therapeutic target.
Keywords: FAM110B, NSCLC, Wnt/β-catenin signaling, proliferation, invasion
Introduction
The centrosome is primarily responsible for cell cycle progression and microtubule organization.1,2 The centrosome plays a critical role in cell shape maintenance, cell polarity, and cell motility. It also acts as a hub for multiple signaling pathways.3,4 The structure of centrosome is comprised of centrioles and pericentriolar matrix.1,4,5 The function of centrosome is tightly controlled by the proteins residing in the pericentriolar matrix which are thus called centrosome associated proteins.1,6,7 Recent studies indicated that the centrosome and its associated proteins were seen as potential anti-cancer drug targets.8–10
FAM110 family proteins (family with sequence similarity 110) belong to the centrosome associated proteins which were firstly identified in 2007.11 There are 3 family members of FAM110 proteins (FAM110A, FAM110B, and FAM110C) that localize in centrosomes and assist in nucleating and organizing microtubules. Among these FAM110 family members, FAM110B was the putative founder of the whole family. To the best of our knowledge, the published literature of FAM110B is limited. Although the previous study has demonstrated that overexpression of FAM110B resulted in G1 phase arrest in HEK293T cells, the function of FAM110B in human cancers, especially the relationship of FAM110B expression with clinicopathological features of patients with non-small cell lung cancer (NSCLC) is largely unknown.11
β-catenin is a multifunctional protein that regulates tissue organization, development, and bipolar spindle formation.12,13 Besides localizes to cell-cell contacts, nucleus, and cytosol, β-catenin is also found at centrosomes.13,14 Previous studies have demonstrated that β-catenin was a component of centrosome.13,14 Overexpression of β-catenin, which is involved in the regulation of centrosomes, leads to the disorder of centrosomes and the loss of the anchoring of cortical microtubules.15 β-catenin is regulated by its upstream Wnt signaling molecules, which control the nuclear translocation of β-catenin by regulating its phosphorylation status. The dephosphorylating form of β-catenin is seen as the active-β-catenin and contributes to its aberrant nuclear overexpression.16 Previous studies revealed that aberrant β-catenin expression was an independent prognostic marker of NSCLC.17 As a member of centrosome associated proteins, we hypothesize that FAM110B may play a role in regulating the expression and phosphorylation of β-catenin and thereby impact tumor progression of NSCLC.
To clarify our hypothesis, we evaluated the expression and subcellular distribution of FAM110B in both tissue samples and cell lines of NSCLC, including their clinicopathological correlations. Meanwhile, we also explored the impacts of FAM110B on cell proliferation and invasion after FAM110B overexpression or FAM110B RNAi. In conclusion, our data suggested that FAM110B inhibited the development of NSCLC by suppressing the activation of Wnt/β-catenin signaling pathway.
Patients and Methods
GEPIA Dataset Analysis
Gene Expression Profiling Interactive Analysis (GEPIA) is an interactive web application that analyzes the RNA sequencing expression data based on 9736 tumors and 8587 normal samples from the Cancer Genome Atlas (TCGA) and Genotype-tissue Expression dataset projects. It provides pivotal interactive and customizable functions such as tumor/normal differential expression analysis, profiling according to cancer types or pathological stages, patient survival analysis, similar gene detection, correlation analysis, and dimensionality reduction analysis. The analysis was performed in line with the protocol provided by the website. Protocol s.io: http://gepia.cancer-pku.cn.
The Kaplan–Meier Plotter Analysis
Kaplan–Meier Plotter contains gene expression data and survival information of 2190 clinical OC patients, which is an online database including microarray gene expression data and survival information provided by Gene Expression Omnibus, TCGA, and Cancer Biomedical information Grid. In this research, the Kaplan–Meier Plotter was utilized to evaluate the prognostic value of the mRNA expression of FAM110B. Protocol s.io: https://kmplot.com.
Patients and Specimens
This research has been approved by the local institutional review committee of China Medical University. Each participant signed informed consent and agreed to store their information in the hospital databases for their specimens to be utilized for the research. Tissue samples from 183 patients were obtained from the patients who underwent complete surgical resection from 2010 to 2012 at the First Affiliated Hospital of China Medical University. The clinicopathological features of these patients were summarized in Table 1. The survival of each patient was calculated from the day of surgery to the date of death or the end of follow-up. Histological diagnosis and grading were evaluated according to the 2015 World Health Organization (WHO) classification of lung tumors.18 Tumor staging was referred to the eighth edition of the American Joint Committee on Cancer (AJCC) lung cancer staging system.19
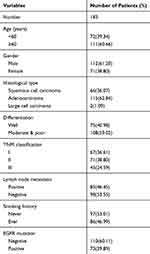 |
Table 1 The Clinicopathological Features of Patients in This Study |
Immunohistochemistry
The streptavidin-peroxidase method using a diaminobenzidine kit (MaiXin, Shenzhen, China) was used for immunohistochemical staining. The slides were then incubated with the primary antibody FAM110B (1:100, Sigma, St.Louis, MO; HPA008318) at 4°C overnight. Sections were incubated with phosphate-buffered saline (PBS) as a negative control rather than the primary antibody. A semi-quantitative was used to score FAM110B immunostaining by evaluating the scale of the representative tumor region, the intensity, and cell percentage. Each slide selected five views randomly and no less than 100 cancer cells were evaluated per view. The intensity of the FAM110B score was indicated as 0 (no staining), 1 (weak staining), 2 (moderate staining), and 3 (strong staining). Staining percentage was scored as 0 (0%), 1 (1–25%), 2 (26–50%), 3 (51–75%), and 4 (76–100%). The final score of 0–16 was multiplied by each score. FAM110B status was divided into negative expression or positive/overexpression expression according to (score < 4) or (score ≥ 4). All tumor sections were randomly analyzed by two independent investigators.
Immunofluorescence Staining
After being fixed with 4% paraformaldehyde for 15 minutes, cells were incubated with 0.1% Triton X-100 for 15 minutes and then sealed with goat serum at room temperature for 2 hours. The cells were incubated with primary antibodies FAM110B (1:100, Sigma, St. Louis, MO; HPA008318) overnight at 4°C, and then incubated with secondary antibody for 2 hours at 37°C. DAPI was used to stain the nuclei and a fluorescence microscope (Olympus, Tokyo, Japan) was used to observe samples.
Cell Culture and Transfection
The human normal bronchial epithelial (HBE), and the A549, PC9 cancer cell lines were obtained from the American Type Culture Collection (ATCC; Manassas, VA). The H1299 and H460 cancer cell lines were obtained from the Shanghai Cell Bank (Shanghai, China). The LK2 cell line was a gift from Dr. Hiroshi Kijima (Department of Pathology and Bioscience, Hirosaki University Graduate School of Medicine, Hirosaki, Japan). All the cell lines were cultured in RPMI 1640 (Invitrogen, Carlsbad, CA, USA) or DMEM (Invitrogen) containing 10% calf serum at 37°C and 5% CO2. The pCMV6-ddk-myc empty vector, pCMV6-ddk-myc-FAM110B plasmid (RC206875), and the negative control (NC)-siRNA (SR30004) were all purchased from Origene (Rockville, MD, USA). The FAM110B-siRNA (sc-77299) was purchased from Santa (Santa Cruz Biotechnology, Inc. CA, USA) which composed of a pool of two target-specific 19–25 nt siRNAs. Lipofectamine 3000 transfection reagent (Invitrogen) was used for cell transfections. After transfection, cells were incubated for 48 hours and analyzed using Western blot. For specific inhibition of Wnt/β-catenin signaling, a Wnt/β-catenin signaling inhibitor, XAV-939, was incorporated into the medium of the cultured cells at a concentration of 30nM for 24 hours at 37°C (S1180, Selleck Chemicals, Houston, TX, USA).
Western Blot
Lysis buffer was used to extract the proteins from cells and the Bradford method20 was used to quantify all the proteins. 10% SDS-PAGE was used to separate the total protein samples of fifty μg and transferred the samples onto polyvinylidene fluoride membranes (PVDF; Millipore, Billerica, MA, USA). The following primary antibodies were used to incubate membranes at 4°C overnight: MYC-tag (2276S), β-catenin (S33/S37/Thr41; 8814S), p-β-catenin (S33/S37/ST41; 9561S), GSK-3β (12456S), p-GSK-3β (S9; 5558S), Cyclin B1 (4135S), Cyclin D1 (2978S), MMP2 (40994S), MMP7 (3801S) (1:1000; Cell Signaling Technology Danvers, MA, USA); FAM110B (1:1000, Sigma); GAPDH (1:1000, Sigma, St.Louis, MO; USA). Washed membranes were incubated with peroxidase-conjugated anti-mouse or anti-rabbit IgG (Santa Cruz Biotechnology) for 2 hours at 37 °C. Electrochemiluminescence (Pierce, Rockford, IL, USA) was used to visualize bound proteins and a bio-imaging system (DNR Bio-Imaging Systems, Jerusalem, Israel) was used to detect.
3-(4,5-Dimethylthiazol-2-Yl)-2,5-Diphenyltetrazolium Bromide Assay (MTT)
After 24 hours transfection, cells were cultured in 96-well plates in medium containing 10% fetal bovine serum. Cell activity was determined by MTT assay for 5 consecutive days. In short, each well was added 20 µL of 5 mg/mL MTT (thiazolyl blue) solution and incubated at 37°C for 4 hours. The medium was then removed, and the resultant MTT was solubilized in 150 µL of DMSO. A spectrophotometer was used to measure the absorbance at a wavelength of 490 nm.
Colony Formation Assay
The cells were transferred into three 6-cm cell culture dishes (800 cells per dish) and then incubated for 14 days. Washing plates with PBS and staining with Giemsa. No less than 50 cell colonies were counted in each dish. Each experiment was in triplicate.
Matrigel Invasion Assay
Coated with 20 µL Matrigel with a dilution rate of 1:6 (BD Bioscience), a 24-well transwell chamber with an 8-µm pore size (Costar, Cambridge, MA) was required for matrigel invasion assay. RPMI 1640 (Invitrogen) with 10% serum was added in the lower chamber. Cells were trypsinized in serum-free medium at a density of 2 × 105 cells/wells, transferred to the upper Matrigel chamber after 48-hours transfection, and then incubated for 18 hours. The uninvaded cells were removed with a cotton tip and the cells passing through the filter were fixed with 4% paraformaldehyde and then stained with hematoxylin.
Statistical Analysis
All data were analyzed by the software SPSS version 20. Pearson’s Chi-square test was used to assess the correlations between FAM110B expression and clinicopathological factors. Kaplan–Meier survival analysis was used to evaluate the prognostic value of FAM110B and compared by the Log rank test. The Western blot and the invasive assay results were analyzed using the Mann–Whitney U-test. P < 0.05 was considered statistically significant.
Results
FAM110B Expression Correlated with Early TNM Staging and the Favorable Prognosis of NSCLC Patients
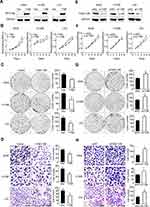
We evaluated the differential expression of FAM110B from the GEPIA online database. The results of the analysis revealed that the mRNA level of FAM110B in non-cancerous lung tissues was higher than in adenocarcinoma and squamous cell lung carcinomas (Figure 1A). Subsequent analysis from the KMplotter database showed that FAM110B predicted favorable prognosis in patients with NSCLC (Figure 1B).
Using immunohistochemistry, we investigated the expression of FAM110B in 183 NSCLC tissue samples. Our data showed that FAM110B was positively expressed in 48.6% (89/183) cases and the subcellular distribution of FAM110B was in the cytoplasm. FAM110B tended to present moderate or strong intensity in normal bronchial epitheliums (Figure 1C), whereas showed dim or negative expression in cancer cells (Figure 1D–H). Statistical analysis indicated that FAM110B expression significantly correlated with early TNM staging (P=0.020) and negative regional lymph node metastasis (P=0.006; Table 2). FAM110B expression was not significantly correlated with age, gender, histological differentiation, and histological type, smoking history, and EGFR mutation (P>0.05). Subsequent Kaplan–Meier survival analysis found that the median survival time of patients with positive FAM110B expression (56.181±2.348 months) was significantly longer than those with negative FAM110B expression (47.701±2.997 months; P=0.024; Figure 1I).
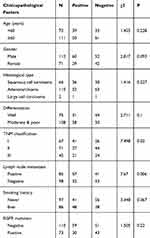 |
Table 2 The Association Between FAM110B Expression and the Clinicopathological Features |
Subsequent Western blotting assay revealed that the protein level of FAM110B was significantly lower in NSCLC tissues than that in the paired adjacent non-cancerous lung epithelium (0.733±0.120 vs 1.243±0.075; P<0.05; Figure 1J). In NSCLC cell lines, as well as in the HBE cell line, we found 4 of 5 NSCLC cell lines showed lower FAM110B expression than HBE cell line (Figure 1K). Also, in A549, H1299, and LK2 cell lines, immunofluorescence results found that the subcellular distribution of FAM110B was also localized in the cytoplasm (Figure 1L).
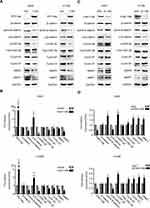
FAM110B Inhibits Proliferation and Invasion in NSCLC Cells
We overexpressed exogenous FAM110B by transfecting FAM110B specific cDNA plasmid into A549, H1299, and LK2 cell lines (Figure 2A). The results of MTT and colony formation assay revealed that overexpression of FAM110B remarkably inhibited the proliferation in the above cell lines. (Figure 2B and C) Similarly, using a transwell matrigel assay, we also observed that the overexpression of FAM110B prominently decreased the invasiveness (Figure 2D). Conversely, knockdown of endogenous FAM110B significantly enhanced the proliferation (Figure 2E–G) and invasiveness (Figure 2H) of A549, H1299, and LK2 cell lines.
FAM110B Prevented the Activation of Wnt/β-Catenin Signaling Pathway
To elucidate the underlying mechanism of FAM110B-induced inhibition of proliferation and invasion of lung cancer cells, we investigated the impact of FAM110B on the expression and phosphorylation of key proteins in Wnt signaling pathway. In A549 and H1299 cells, Western blotting results revealed that FAM110B overexpression resulted in reduced phosphorylation of GSK-3β (p-GSK-3β), active β-catenin (dephosphorylated β-catenin), Cyclin B1, Cyclin D1, MMP2 and MMP7, and the expression of p-β-catenin (phosphorylated β-catenin) was upregulated, while the protein levels of total β-catenin and GSK-3β showed no remarkable changes (Figure 3A and B). On the contrary, when we downregulated the expression of endogenous FAM110B, the levels of p-GSK-3β, active-β-catenin, Cyclin B1, Cyclin D1, MMP2, and MMP7 were correspondingly upregulated, and the protein level of total β-catenin and GSK-3β still showed no visible changes (Figure 3C and D).
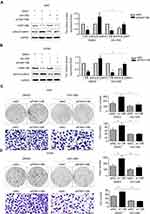
Wnt/β-Catenin Inhibitor XAV-939 Can Significantly Reverse the Effect of FAM110B on NSCLC
To further confirm whether FAM110B suppresses the development of NSCLC cells through Wnt/β-catenin signaling, we incorporated XAV-939, a specific Wnt/β-catenin inhibitor, into the medium of A549 and H1299 cells. Western blotting results showed that XAV-939 significantly attenuated the effect of FAM110B knockdown on the elevation of active-β-catenin protein levels in both A549 and H1299 cells (Figure 4A and B). Moreover, the effects of FAM110B RNAi on facilitating the proliferation and invasion of A549 and H1299 cells were also weakened by XAV-939 incorporation in the above-mentioned cell lines (Figure 4C and D).
Discussion
In this study, we identified that FAM110B localized in the cytoplasm of NSCLC cells which suppressed the proliferation and invasion through inhibiting the activation of Wnt/β-catenin signaling pathway. Moreover, NSCLC patients with positive FAM110B expression significantly correlated with favorable clinical outcomes.
The published literature of FAM110B was limited to date. However, in the preceding studies, the role of FAM110B playing in malignant tumors was controversial. Based on genome-wide DNA and RNA data, Vainio et al suggested that FAM110B may have a potential oncogenic role in castration-resistant prostate cancer.21 Based on the data from online-database, Xi et al identified that FAM110B was associated with favorable survival time of pancreatic adenocarcinoma patients.22 In the present study, we found that FAM110B may act as a tumor suppressor in human NSCLC cells. We identified that FAM110B expression was lower in both NSCLC tissue samples and cell lines using immunohistochemistry and Western blot. Our results are consistent with the data from online-database that FAM110B expression tends to be downregulated in NSCLC. Moreover, statistical analysis results in online databases, and our cohort indicated that FAM110B expression significantly correlated with early TNM staging, negative regional lymph node metastasis, and favorable prognosis in NSCLC patients. The survival analysis of the present study is consistent with the previous study in which they identified FAM110B expression was associated with favorable clinical outcome in pancreatic adenocarcinoma cohorts. Recent studies have shown that cigarette smoking is a major preventable risk factor for lung cancer23 and may alter metabolic pathways and increase the risk of lung cancer development.24 Smoke-induced mechanisms are recognized to act on cell growth and tumor invasiveness by deregulating of oncogenes or anti-oncogenes23 and high smoking exposition is associated with adverse prognostic factors for lung cancer such as HIF.25 The exposure of lung epithelial cells to cigarette smoke could stimulate aberrant EGFR phosphorylation.24 It has been shown that the EGFR mutations are common in lung cancer and nonsmokers have a higher frequency of EGFR mutations than smokers in EGFR mutated patients.24 In this study, our statistical analysis indicated that FAM110B expression had no connection with smoking history and EGFR mutation. The previous study and ours suggested that FAM110B may be a useful marker in predicting survival in several human malignant tumors, especially in NSCLC.
Previous studies demonstrated that centrosome was comprised of two centrioles embedded in pericentriolar materials. The functions of centrosome were tightly regulated by the centrosome associated proteins which localized in centrioles and pericentriolar materials.26–28 FAM110 protein family was identified as centrosome associated proteins in 2007. However, there were only limited studies focusing on the roles of FAM110 protein family playing in carcinogenesis, especially on FAM110B. In the present study, by transfecting with FAM110B plasmid or FAM110B siRNA, we found that FAM110B overexpression suppressed the proliferation and invasion, and decreased the expression of Cyclin B1, Cyclin D1, MMP2, and MMP7, while FAM110B knockdown exerted opposite effect, which indicated that FAM110B played a tumor-suppressive role in the development of NSCLC. Besides, our results showed that overexpression of FAM110B reduced the phosphorylation of GSK-3β and the expression of active β-catenin, which resulted in the inactivation of Wnt/β-catenin signaling pathway. Available data indicate that Wnt signaling substantially impacts NSCLC tumorigenesis, prognosis, and resistance to therapy, with loss of Wnt signaling inhibitors by promoter hypermethylation or other mechanisms appearing to be particularly important. Wnt inhibitors may restore the sensitivity of Wnt signaling.29 In this study, Wnt/β-catenin inhibitor XAV-939 significantly attenuated the effect of FAM110B knockdown on the elevation of active-β-catenin protein levels in both A549 and H1299 cells. And the effects of FAM110B RNAi on facilitating the proliferation and invasion were also weakened by XAV-939 incorporation in the above-mentioned cell lines. Multiple reports demonstrated that key Wnt/β-catenin signaling components localized in the centrosome.30–33 Both β-catenin and the components of the β-catenin degradation complex are localized at the centrosome, and the stabilization of β-catenin was critical to centrosome splitting.31,32 Furthermore, in addition to the well-known effect of β-catenin stabilization, abnormal activation of Wnt signaling also caused centrosome splitting, which eventually led to centrosome aberrations, a hallmark of cancer cells.13,31 The significance of β-catenin accumulation for centrosome disjunction inspired a new model in which β-catenin has been attributed to a central role.31 Reportedly, FAM110B silencing decreased the expression of β-catenin in prostate cancer cells.21 NSCLC is the leading cause of cancer death and the pathogenesis of NSCLC is complex.34 The high proliferation rate of cancer cells is usually along with altered vascularization25 and the pathogenesis of NSCLC is related to a variety of protein markers, such as the well- known EGFR, HIF, and PD-L, which may significantly influence prognosis, grading, and staging.24,25,34 In this study, we identified FAM110B as a novel component of Wnt/β-catenin signaling in regulating the development of NSCLC. Therefore, our study suggested that FAM110B/GSK-3β/Wnt/β-catenin signaling axis may play a crucial role in regulating the malignant behavior of lung cancer cells. Certain limitations of our study should be noted. Our results showed that the FAM110B overexpression inhibited Wnt/β-catenin signaling, thereby restricting the proliferation and invasion of lung cancer cells. However, as the regulation of FAM110B in lung cancer remains unclear, further investigations are required to clarify the specific regulatory mechanisms.
Conclusion
In summary, our results demonstrate that the overexpression of FAM110B restricts the proliferation and invasion of NSCLC cells by inhibiting Wnt/β-catenin signaling. These findings highlight the importance of the FAM110B/Wnt/β-catenin signaling pathway in NSCLC progression. Our study reveals the antitumor function of FAM110B in NSCLC and indicates FAM110B to be a potential therapeutic target.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 81472805 to Yuan Miao) and the Natural Science Foundation of Liaoning Province (No. 2019-MS-382 to Yuan Miao).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Schatten H. The mammalian centrosome and its functional significance. Histochem Cell Biol. 2008;129(6):667–686. doi:10.1007/s00418-008-0427-6
2. Chavali PL, Putz M, Gergely F. Small organelle, big responsibility: the role of centrosomes in development and disease. Philos Trans R Soc Lond B Biol Sci. 2014;369(1650):20130468. doi:10.1098/rstb.2013.0468
3. Azimzadeh J, Bornens M. Structure and duplication of the centrosome. J Cell Sci. 2007;120(13):2139–2142. doi:10.1242/jcs.005231
4. Nigg EA, Raff JW. Centrioles, centrosomes, and cilia in health and disease. Cell. 2009;139(4):663–678. doi:10.1016/j.cell.2009.10.036
5. Gonczy P. Towards a molecular architecture of centriole assembly. Nat Rev Mol Cell Biol. 2012;13(7):425–435. doi:10.1038/nrm3373
6. Andersen JS, Wilkinson CJ, Mayor T, Mortensen P, Nigg EA, Mann M. Proteomic characterization of the human centrosome by protein correlation profiling. Nature. 2003;426(6966):570–574. doi:10.1038/nature02166
7. Archinti M, Lacasa C, Teixido-Travesa N, Luders J. SPICE–a previously uncharacterized protein required for centriole duplication and mitotic chromosome congression. J Cell Sci. 2010;123(18):3039–3046. doi:10.1242/jcs.069963
8. Larsen CP, Austyn JM, Morris PJ. The role of graft-derived dendritic leukocytes in the rejection of vascularized organ allografts. Recent findings on the migration and function of dendritic leukocytes after transplantation. Ann Surg. 1990;212(3):308–317. doi:10.1097/00000658-199009000-00009
9. Pihan GA. Centrosome dysfunction contributes to chromosome instability, chromoanagenesis, and genome reprograming in cancer. Front Oncol. 2013;3:277. doi:10.3389/fonc.2013.00277
10. Godinho SA, Pellman D. Causes and consequences of centrosome abnormalities in cancer. Philos Trans R Soc Lond B Biol Sci. 2014;369(1650):1650. doi:10.1098/rstb.2013.0467
11. Hauge H, Patzke S, Aasheim CH. Characterization of the FAM110 gene family. Genomics. 2007;90(1):14–27. doi:10.1016/j.ygeno.2007.03.002
12. Nelson WJ, Nusse R. Convergence of Wnt, beta-catenin, and cadherin pathways. Science. 2004;303(5663):1483–1487. doi:10.1126/science.1094291
13. Kaplan DD, Meigs TE, Kelly P, Casey PJ. Identification of a role for beta-catenin in the establishment of a bipolar mitotic spindle. J Biol Chem. 2004;279(12):10829–10832. doi:10.1074/jbc.C400035200
14. Bahmanyar S, Kaplan DD, Deluca JG, et al. beta-Catenin is a Nek2 substrate involved in centrosome separation. Genes Dev. 2008;22(1):91–105. doi:10.1101/gad.1596308
15. Ligon LA, Karki S, Tokito M, Holzbaur EL. Dynein binds to beta-catenin and may tether microtubules at adherens junctions. Nat Cell Biol. 2001;3(10):913–917. doi:10.1038/ncb1001-913
16. MacDonald BT, Tamai K, He X. Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell. 2009;17(1):9–26. doi:10.1016/j.devcel.2009.06.016
17. Xu X, Sun PL, Li JZ, Jheon S, Lee CT, JH C. Aberrant Wnt1/β-catenin expression is an independent poor prognostic marker of non-small cell lung cancer after surgery. J Thorac Oncol. 2011;6(4):716–724. doi:10.1097/JTO.0b013e31820c5189
18. Travis WD, Brambilla E, Burke AP, Marx A, Nicholson AG, eds. WHO Classification of Tumours of the Lung, Pleura, Thymus, and Heart. WHO Classification of Tumours.
19. Amin MB, Edge S, Greene F, et al. eds. AJCC Cancer Staging Manual.
20. Bradford MM. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72(1–2):248–254. doi:10.1016/0003-2697(76)90527-3
21. Vainio P, Wolf M, Edgren H, et al. Integrative genomic, transcriptomic, and RNAi analysis indicates a potential oncogenic role for FAM110B in castration-resistant prostate cancer. Prostate. 2012;72(7):789–802. doi:10.1002/pros.21487
22. Xi T, Zhang G. Integrated analysis of tumor differentiation genes in pancreatic adenocarcinoma. PLoS One. 2018;13(3):e0193427. doi:10.1371/journal.pone.0193427
23. Pezzuto A, Citarella F, Croghan I, Tonini G. The effects of cigarette smoking extracts on cell cycle and tumor spread: novel evidence. Future Sci OA. 2019;5(5):FSO394. doi:10.2144/fsoa-2019-0017
24. Condoluci A, Mazzara C, Zoccoli A, Pezzuto A, Tonini G. Impact of smoking on lung cancer treatment effectiveness: a review. Future Oncol. 2016;12(18):2149–2161. doi:10.2217/fon-2015-0055
25. Pezzuto A, Perrone G, Orlando N, et al. A close relationship between HIF-1α expression and bone metastases in advanced NSCLC, a retrospective analysis. Oncotarget. 2019;10(66):7071–7079. doi:10.18632/oncotarget.27378
26. Kellogg DR, Moritz M, Alberts BM. The centrosome and cellular organization. Annu Rev Biochem. 1994;63(1):639–674. doi:10.1146/annurev.bi.63.070194.003231
27. Doxsey S. Re-evaluating centrosome function. Nat Rev Mol Cell Biol. 2001;2(9):688–698. doi:10.1038/35089575
28. Bornens M. Centrosome composition and microtubule anchoring mechanisms. Curr Opin Cell Biol. 2002;14(1):25–34. doi:10.1016/S0955-0674(01)00290-3
29. Stewart DJ. Wnt signaling pathway in non-small cell lung cancer. J Natl Cancer Inst. 2014;106(1):djt356. doi:10.1093/jnci/djt356
30. Kim SM, Choi EJ, Song KJ, et al. Axin localizes to mitotic spindles and centrosomes in mitotic cells. Exp Cell Res. 2009;315(6):943–954. doi:10.1016/j.yexcr.2009.01.013
31. Hadjihannas MV, Bruckner M, Behrens J. Conductin/axin2 and Wnt signalling regulates centrosome cohesion. EMBO Rep. 2010;11(4):317–324. doi:10.1038/embor.2010.23
32. Mbom BC, Nelson WJ, Barth A. β-catenin at the centrosome: discrete pools of β-catenin communicate during mitosis and may co-ordinate centrosome functions and cell cycle progression. Bioessays. 2013;35(9):804–809. doi:10.1002/bies.201300045
33. Bryja V, Cervenka I, Cajanek L. The connections of Wnt pathway components with cell cycle and centrosome: side effects or a hidden logic? Crit Rev Biochem Mol Biol. 2017;52(6):614–637. doi:10.1080/10409238.2017.1350135
34. Pezzuto A, Cappuzzo F, D’Arcangelo M, et al. Prognostic value of p16 protein in patients with surgically treated non-small cell lung cancer; relationship with Ki-67 and PD-L1. Anticancer Res. 2020;40(2):983–990. doi:10.21873/anticanres.14032
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.